Abstract
Key message
The NF-Y gene family was identified and characterized in Populus trichocarpa by genome-wide analysis. Expression of PtoNF-Y was analyzed during floral bud development in Populus tomentosa.
Abstract
Nuclear factor Y (NF-Y) is a transcription factor widely found in eukaryotes. It is composed of three subunits: NF-YA, NF-YB, and NF-YC. NF-Y has been identified as a key regulator of multiple pathways that control responses to developmental, biotic, and abiotic factors. Although the NF-Y gene has been reported in many plants, it has not been comprehensively analyzed in poplar. In this study, a total of 46 NF-Y gene family members in the P. trichocarpa genome were accessed using NF-Y amino acid sequences from Arabidopsis thaliana as a probe. In addition, bioinformatic characterization of NF-Y gene family members in P. trichocarpa, including gene structure, chromosome location, and phylogenetic relationships, was conducted. The results of chromosome distribution showed that the 46 PtNF-Y genes were distributed among 16 chromosomes of P. trichocarpa at varying frequencies. Gene-structure analysis showed that seven members have no introns. Phylogenetic analysis indicated that the NF-Y protein may have similar functions in Arabidopsis and P. trichocarpa. We used transcriptome data from different tissues to analyze the expression of the NF-Y genes in Populus tomentosa, and verified the results by qRT-PCR. The results indicated that some PtoNF-Y genes play significant roles in the floral bud development of P. tomentosa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transcription factors (TF) play vital roles in numerous biological activities by binding to specific DNA elements of their targeted genes. Nuclear factor Y (NF-Y), as a transcription factor, is composed of three subunits: NF-YA (HAP2 or CBF-B), NF-YB (HAP3 or CBF-A), and NF-YC (HAP5 or CBF-C) (Bucher and Trifonov 1988). NF-Y is widely found in fungi, animals, and plants. It specifically binds to the cis-element CCAAT-box in eukaryotic promoters (Nardini et al. 2013). Subunits NF-YB and NF-YC form dimers in the cytoplasm and then bind to the NF-YA protein and form a trimer in the nucleus (Tom et al. 2013).
In many mammals and yeasts, a single gene encodes each of the three NF-Y subunits (Li et al. 1992). For example, in mice and humans, only one NF-Y gene encodes each NF-Y subunit. However, in plants, each NF-Y subunit gene is represented by multiple orthologs (Zhao et al. 2016). For example, in Arabidopsis thaliana, the NF-YA, NF-YB, and NF-YC subunit families have 10, 13, and 13 members, respectively (Siefers et al. 2009). In rice (Oryza sativa), the NF-YA subfamily is encoded by 10 NF-YA genes, NF-YB by 11 NF-YB genes, and NF-YC by 7 NF-YC genes (Thirumurugan et al. 2008). Likewise, in soybeans (Glycine max), the number of identified members of the three subfamilies has reached 21, 32, and 15, respectively (Quach et al. 2015). In addition, the NF-Y genes of citrus (Pereira et al. 2018), walnut (Quan et al. 2018), tomato (Li et al. 2016), foxtail millet (Feng et al. 2015), sorghum (Malviya et al. 2016), and maize (Zhang et al. 2016) have previously been studied, and their numbers of members have increased in plants. This expansion of NF-Y gene number might contribute to the formation the well-established TF network and regulate plant growth and development (Calvenzani et al. 2012; Petroni et al. 2012).
Recent studies have revealed that NF-Y genes play a key role in plant response to various abiotic and biotic stresses. Overexpression of ATNF-YA5 and ATNF-YB1 in plants generally increases their growth under drought stress (Li et al. 2008). However, nfya5 knockout plants have the opposite response to overexpression of AtNF-YA5, becoming more sensitive to drought stress. In castor bean, cold stress significantly up-regulated the expression level of RcNF-YC6 (Wang et al. 2018). In addition, in soybean, the expression level of GmNF-YA3 was induced by various stress treatments (Ni et al. 2013). These studies have indicated that the NF-Y gene family participates in various regulatory processes in response to abiotic stress in plants, but with functional differences among members.
Many NF-Y genes have been reported to be involved in growth and developmental processes, such as embryogenesis (Fornari et al. 2013), seed germination (Liu et al. 2016), photoperiod or age-dependent flowering (Kumimoto et al. 2010; Wei et al. 2017), and fruit maturation (Li et al. 2016). NF-Y genes are also involved in plant physiological processes, including photomorphogenesis (Myers et al. 2016), regulation of nodulation during nitrogen fixation (Quach et al. 2015), and stress responses of the endoplasmic reticulum (ER) (Liu and Howell 2010).
Numerous studies have reported that NF-Y genes are involved in the regulation of flowering timing and flower differentiation, implying that NF-Y genes play a particularly important role in flower development (Su et al. 2018; Wei et al. 2017). Poplar is an economically important tree that is highly valued for bioenergy and timber production purposes (Ji et al. 2013). Extensive research has focused on traditional breeding. Due to rapid growth and ease of transformation, P. trichocarpa is considered an ideal tree species for basic and applied research (Yan et al. 2012). Taking advantage of functional genomics studies of the plant NF-Y gene and current genomic data of P. trichocarpa, we attempted to analyze the NF-Y gene family of P. trichocarpa. In addition, the urban environmental pollution issues (such as causing traffic accident, fire risk, as well as spreading bacteria and virus) caused by female catkins of poplar during spring have attracted much attention by researchers. Meanwhile, the pollen in male floral buds has allergenic properties and poses potential health hazards to allergic people (An et al. 2011). Therefore, clarifying the flower development in poplar is of great importance. In this study, we identified 46 NF-Y genes from the P. trichocarpa genome and performed a relatively complete bioinformatics analysis, including conserved regions, phylogenetic relationships, gene-structure analysis, and chromosome localization. Moreover, we analyzed NF-Y genes in terms of their expression levels among five different tissues and seven different flower developmental stages using transcriptome data and validated these expression levels by qRT-PCR, which is particularly important for identifying candidate genes involved in regulation of the growth and flowering of P. tomentosa. Taken together, our results contribute to a more complete understanding of the function of the NF-Y genes in poplar.
Materials and methods
Identification of PtNF-Y gene family members
Using the classification criteria of NF-Y TFs in A. thaliana (Petroni et al. 2012), we obtained 36 AtNF-Y amino acid (AA) sequences from The Arabidopsis Information Resource (Swarbreck et al. 2008). These sequences included 10 NF-YA, 13 NF-YB, and 13 NF-YC subunits. To anchor the P. trichocarpa NF-Y family members, AA sequences of the Arabidopsis NF-Y members were used as queries to search for NF-Y family members of P. trichocarpa in Phytozomev12.1 (Goodstein et al. 2012). We selected all sequences with e value < 10−10 for subsequent analysis. In addition, SMART tool (http://smart.embl-heidelberg.de/) (Zhang et al. 2019) was utilized to identify NF-Y protein domains. We obtained the NF-Y candidate members of P. trichocarpa based on this comprehensive screening for use in subsequent analysis.
Chromosomal distribution and structure analysis of PtNF-Y
The PtNF-Y genes were mapped to the chromosomes using the MapInspect software (Zhang et al. 2019). The gene structures of PtNF-Y members were analyzed using the Gene-Structure Display Serve (http://gsds.cbi.pku.edu.cn/) (Hu et al. 2015).
NF-Y protein alignment and phylogenetic prediction
We obtained the genomic sequences, coding sequences (CDS) and protein sequences of PtNF-Y members using Phytozome v12.1. Multiple sequence alignments analysis of NF-Y proteins was performed using Clustal X2.1 (Larkin et al. 2007). Additionally, based on the alignments obtained, genetic distance matrices were obtained, and an unrooted phylogenetic tree was generated in MEGA7 using the neighbor-joining (NJ) method based on the PtNF-Y protein sequences of P. trichocarpa and Arabidopsis. Their evolutionary relationships were estimated with 1000 bootstrap replications (Kumar et al. 2016).
Plant material
Adult P. tomentosa trees were grown under natural conditions in the Beijing Forestry University greenhouse (Beijing, China). Roots, stems, and leave tissue were obtained from 6-month-old cultured plantlets. Dormant and germinating leaf buds were collected during the flower transition period in 2017. Male and female floral buds of P. tomentosa were collected from June 2017 to February 2018 at Beijing Forestry University. These flower buds covered seven stages of flower development, from flower induction to flower initiation and organ differentiation. In addition, all collected samples were immediately frozen in liquid nitrogen and then stored at − 80 °C until use.
Transcriptome sequencing and de novo assembly
RNA-seq libraries were generated using the Illumina kit according to the manufacturer’s protocol (Illumina, San Diego, CA, USA). Equal amount of RNA for each sample were pooled, and sequenced separately on the Illumina HiSeqTM 2000 platform using paired-end technology. The assembly method used was described previously (Chen et al. 2018). The expression levels of PtoNF-Ys were calculated as fragments per kilobase per million (FPKM) (Cole et al. 2010) reads values to determine differences in expression in various organs and tissues among NF-Y members, and heat maps were constructed using TBool software. Hierarchical cluster analysis was conducted to visualize the expression pattern of NF-Y genes in different P. tomentosa organs and tissues.
RNA isolation and qRT-PCR analysis
Total RNAs were isolated from each tissue at various developmental stages of P. tomentosa. We used Trizol Total RNA Extraction Kit (Promega, Madison, WI, USA) and RQ1 DNase to remove genomic DNA (Promega) and incubated the extracts at 65 °C for 10 min to inactivate the DNase. The RNA was detected by 1.0% agar gel electrophoresis and a NanoDrop 2000 spectrophotometer (IMPLEN, CA, USA). The total RNA was reverse-transcribed into first strand cDNA using the Reverse Transcription System (Promega). The resulting cDNA was diluted 1:10 with ddH2O and used as the template for qRT-PCR amplification.
For qRT-PCR, we first designed primers by reference to the CDS sequence of P. trichocarpa NF-Y to amplify this sequence. The amplified DNA fragments were then cloned into the pMD-18T vector (TaKaRa, Otsu, Japan) for sequencing compared with the P. trichocarpa CDS (Fig. S1). Then, qRT-PCR primers were designed based on the CDS sequence of P. tomentosa using primer 5.0 software. Finally, primers qRT-PCR was identified by PCR (Fig. S1). The PtoACTIN (GenBank accession: AY261523.1) (An et al. 2011; Zhang et al. 2008; Zheng et al. 2009) was selected as an internal control gene for normalization according to the 2−ΔΔCt method. All primers are listed in Table S1. Real-time quantitative PCR was performed using SYBR ® Premix Ex Taq ™ (TaKaRa) on the ABI PRISM 7500 Fast Real-time PCR System (Applied Biosystems, Foster City, CA, USA), The PCR program was as follows: 95 °C for 30 s, 40 cycles of 95 °C for 5 s and 60 °C for 20 s, and then, a final elongation step of 72 °C for 7 min. The plates were read at 0.2 °C intervals for 1 s from 70 to 95 °C to generate melting curves and thus verify the specificity of the amplified product. The expression values of these genes were analyzed using the 2 −ΔΔCt method (Livak and Schmittgen 2001). All analyses were performed with three technical and three biological replicates.
Results
Isolation of the NF-Y family in P. trichocarpa
To obtain information on the NF-Y genes of P. trichocarpa, Arabidopsis NF-Y protein sequences were used to search for NF-Y using the most recent P. trichocarpa genome assembly in Phytozome v12.1 and the ExPASy server (Table 1). A total of 46 NF-Y genes were identified in the P. trichocarpa genome: 11 PtNF-YA, 21 PtNF-YB, and 14 PtNF-YC. The 46 predicted P. trichocarpa NF-Y genes were named PtNF-YA1 to 11, PtNF-YB1 to 21 and PtNF-YC1 to 14, respectively. The bioinformatics data of all gene members were analyzed, including genomic DNA sequence length, transcript length, CDS length, number of AAs, theoretical molecular weight (MM), and theoretical isoelectric point (pI). The identified PtNF-Y genes encodes peptides ranging from 143 to 348 AA, with pI values ranging from 4.85 to 9.46, and molecular weights ranging from 12.81 to 41.42 kDa, as estimated using the ExPASy server.
Genomic structure and locations of PtNF-Y gene family members
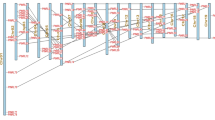
Gene-structure analysis can provide insight into the evolution of families of genes. Therefore, we determined the numbers and positions of exons and introns of the PtNF-Y genes using genomic DNA sequences in the Phytozome v12.1 database (Fig. 1). All PtNF-YA genes were separated by introns. PtNF-YA genes had four or five exons, except PtNF-YA6, which had three introns. Among the PtNF-YB genes, 11 had no introns and the others had 1–5 introns. Of the PtNF-YC genes, 10 of 14 genes were intronless, whereas PtNF-YC13 had six introns. In addition, we found that most homologs in the PtNF-Y subfamily, such as NF-YA6/A7, NF-YA4/A5, NF-YB7/B12, NF-YB6/B11, and NF-YC1/C2, had different gene structures. These results indicate that the intron/exon distribution in PtNF-Y genes is highly variable.
We used the current P. trichocarpa genome database in Phytozome v12.1 to analyze the positions of PtNF-Y genes on P. trichocarpa chromosomes. The 46 PtNF-Y genes were found to be distributed among 16 of the 19 chromosome scaffolds of P. trichocarpa (Table 1, Fig. 2).
Multiple alignments analyses of the PtNF-Y genes
For each PtNF-Y subunit, multiple alignments performed using ClustalX 2.0 software revealed conserved DNA binding domains in PtNF-YA, PtNF-YB, and PtNF-YC proteins (Fig. 3). The results illustrate that PtNF-Y proteins contain evolutionarily conserved domains and more variable N- or C-terminal transcriptional regulation regions. The conserved domain of the PtNF-YA protein core contains 53 AAs, including two highly conserved domains: NF-YB/C interaction domain α1 and DNA contact domain α2. The relatively conserved linker of 21 AAs separates the alpha1 and alpha2 subdomains (Fig. 3a). The conserved core region of PtNF-YB was 86 AAs in length, which is similar to the lengths reported in other species. In addition, it also contains a central domain similar to the histidine folding domain (HFM) in the core histone H2B and plays a key role in DNA binding and protein–protein interactions (Fig. 3b). PtNF-YC subunits were also found to consist of a core histone-like sequence with a central domain about 79 AAs in length (Fig. 3c).
Phylogenetic relationships of PtNF-Y gene family
To explore the evolutionary relationships among different PtNF-Y family members, an unrooted phylogenetic tree of 46 PtNF-Y proteins was constructed (Fig. 4a). The results showed that all PtNF-Y members formed three major clusters: I (PtNF-YA), II (PtNF-YB), and III (PtNF-YC). Closely related members showed the most similar physical and chemical properties, such as MM and pI (Table 1). In addition, the result also indicated that the PtNF-YB proteins are more closely related to PtNF-YC proteins than they are to PtNF-YA proteins, implying that PtNF-YB and PtNF-YC genes may share a more recent ancestor than PtNF-YA.
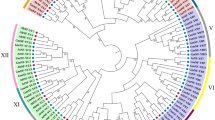
Phylogenetic analysis of P. trichocarpa NF-Y proteins. (a) Combined phylogenetic tree for 11 PtNF-YA, 21 PtNF-YB, and 14 PtNF-YC proteins constructed by the neighbor-joining method in MEGA 7.0 software with 1000 bootstrap reiterations. (b) The genes encoding different subunits formed three separate branches. Different colors indicate different subfamilies. Blue, red, and green represent NF-YA, NF-YB, and NF-YC, respectively. Proteins prefixed by the term ‘At’ and ‘Pt’ indicate NF-Y from Arabidopsis thaliana and Populus trichocarpa, respectively. Among, AtNF-YB6 and AtNF-YB9, which are marked by red triangles, represent AtL1L and AtLEC1, respectively. Numbers on branches show bootstrap values > 50%
To investigate and elucidate the phylogenetic relationships among PtNF-Y proteins and assist with functional predictions, we constructed a comprehensive phylogenetic tree using the full-length protein sequences of all the NF-Y proteins of Arabidopsis and P. trichocarpa (Fig. 4b). The phylogenetic tree shows close relationships among the proteins, all of which, except AtNF-YC10, were grouped into three subfamilies (A, B, and C). The phylogenetic tree indicated close relationships among the PtNF-Ys within each of the three subfamilies. Furthermore, in Fig. 4b, we found that PtNF-YB3, PtNF-YB5 and AtNF-YB9 (AtLEC1), AtNF-YB6 (AtL1L) were clustered in a separate subgroup. However, protein multiple alignment analysis revealed that PtNF-YB3 and PtNF-YB5 were more diverse compared to the others, due to an amino acid change from lysine (K) to aspartic acid (D) (Fig. 3, Fig S2). Therefore, based on this phylogenetic tree, we further predicted that the structures and functions of some PtNF-Y members may be similar to those of Arabidopsis.
Expression of the PtoNF-Y gene family different tissues determined by transcriptome and qRT-PCR analyses
To identify the potential functions of PtoNF-Ys in the development of P. tomentosa, we compared the relative expression levels of the PtoNF-Y genes in root, stem, leaf, leaf bud, and male and female flower buds at seven different developmental stages. The results indicated that 34 of the 46 PtoNF-Ys were expressed in all tissues, while the other 12 PtoNF-Y gene members were not detected (Fig. 5). The transcription levels of the 34 expressed PtoNF-Y genes varied among tissues. We observed four main clusters of expression among the vegetative organs (see Fig. 5, clusters I–IV). Six of the genes are clustered together (clusters I: A3, A8, B4, B14, B21, C6) have relatively high expression except in the germinating leaf bud.
Heat map of PtoNF-Y predicted expression in different tissues and organs of P. tomentosa. The abbreviations R, S, L, DLB, and GLB represent roots, stems, leaves, dormant leaf buds and germinated leaf buds. Expression potential from high to low is represented by square colors from red to green and the black is medium. FPKM value was used to create the heat map. The scale represents the relative expression level intensity of FPKM values. In this study, de novo assembly was done using pooled samples
Among the reproductive organs, we analyzed the expression patterns of male and female floral buds at seven different developmental stages by RNA-seq and qRT-PCR (Fig. 6, these genes are highly expressed in fig S3). The results showed that about 90% of the genes appear to have similar RNA-seq and qRT-PCR patterns. In addition, the result analysis found that some genes show similar expression patterns across the male and female floral bud stages. Such as, in male flower buds and female flower buds, PtoNF-YB4 and PtoNF-YB21 peak in early and late stages, respectively. However, PtoNF-YB15, PtoNF-YC2, and PtoNF-YC7 peak in the middle stages. Moreover, the patterns also showed that the same gene can exhibit contrasting peak times between male and female floral buds. Such as, in the later stages, the expression level of PtoNF-YC5 is completely opposite in male flower buds and female flower buds. The peak of PtoNF-YC14 appears in the late stage of male flower buds and the early stage of female flower buds. The above phenomenon indicated that some genes evolved different functional roles in the two floral bud types.
Expression profiles of the PtoNF-Y genes in male floral buds and female floral buds determined by qRT-PCR. (a) qRT-PCR validation of NF-Y gene expression in the male floral of P. tomentosa. (b) qRT-PCR validation of the expression levels of PtoNF-Y genes in female floral buds. The blue and orange lines represent values obtained by qRT-PCR and RNA-seq, respectively. MFB1–MFB7 represents the seven stages (from June 2017 to February 2018) of male floral bud and female floral buds development. The data represent mean ± SD. The error bars indicate the standard deviation
Discussion
In this study, based on the currently available P. trichocarpa genome sequences and the updated classification criteria of NF-Y TFs in A. thaliana (Petroni et al. 2012), 46 PtNF-Y genes were identified in P. trichocarpa (11 PtNF-YA, 21 PtNF-YB, and 14 PtNF-YC) (Table 1). Previous studies have reported 33 NF-Y members in Arabidopsis (Siefers et al. 2009), 28 in rice (Thirumurugan et al. 2008), and 33 in walnuts (Quan et al. 2018). The high number of NF-Y members in P. trichocarpa may be due to the expansion of this gene family in poplars. In contrast, soybean and tomato contain 66 and 59 NF-Y members, respectively. Compared with the numbers of NF-Y genes in these species, fewer PtNF-Y genes were identified in our study. These differences in gene numbers may be related to differences in classification criteria. The NF-Y family member classification criteria for maize and tomato include several NC2 and Dpb3/4 members (Li et al. 2016; Zhang et al. 2016).
Gene-structure alterations might lead to changes in gene or protein function. In this study, compared with the exon–intron structures of PtNF-YB and PtNF-YC, most PtNF-YA members showed a more variable and complicated exon–intron organization pattern. This result is also consistent with the NF-Y gene structures previously reported in Arabidopsis, Brassica napus, and Solanum lycopersicum. Moreover, exon–intron structure analysis provides new insight into the evolutionary relationships among genes or organisms. We analyzed the exon–intron structure of 46 PtNF-Y genes (Fig. 1), of which 26 lacked introns. However, introns are considered to be essential components of structural eukaryotic genes. Previous studies have reported that introns have many functions, such as alternative splicing and regulation of the evolution rate of genes (Lee et al. 2003; Roy and Gilbert 2006). Their deletion or alteration might lead to structural diversity and complexity, in turn affecting the evolution of the gene family.
In general, the conserved domain is located at the C-terminus of mammalian proteins; the phenomenon is more apparent in plant NF-YAs. Previous studies have shown that three histidine (H) and three arginine (R) residues are essential for DNA binding (Xing et al. 1993). The results of the current study show that these six sites were also highly conserved among all 11 PtNF-YA proteins (Fig. 3a). This finding is consistent with the previously observed residues in Arabidopsis, Oryza sativa, and Brachypodium (Cao et al. 2011; Siefers et al. 2009; Thirumurugan et al. 2008). Previous studies reported that the aspartate at D55 site was considered a critical protein interaction site of the AtNF-YB subfamily, and the Arabidopsis NF-YB subunit based on the 55th Asp in its domain divided into two classes: LEC1 and non-LEC1 (Lee et al. 2003). LEC1 has been identified to play a key role in Arabidopsis embryogenesis and the development of castor seeds (Wang et al. 2018). However, in our study, PtNF-YB3 and PtNF-YB5 changed from lysine (K) to aspartic acid (D) at this binding site. (Figure 3, Fig S2). In addition, phylogenetic relationship analysis also showed that PtNF-YB3, PtNF-YB5 and AtNF-YB9 (AtLEC1), AtNF-YB6 (AtL1L) were clustered in a separated subgroup (Fig. 4b). Therefore, we speculate that these two members are possibly orthologs of AtLEC1 and AtL1L, and propose that PtNF-YB3 and PtNF-YB5 may have functions similar to those of AtLEC1 and AtL1L.
Previous studies have reported that NF-Y genes play important roles in regulating plant growth and development (Li et al. 2016; Potkar et al. 2013; Wang et al. 2018; Wei et al. 2017). In this study, the expression levels of PtoNF-Y genes (except for 12 NF-Y genes that had no expression data) were observed in five tissues and organs, indicating that PtoNF-Y is involved in regulating the growth and development of P. tomentosa (Figs. 5, 6, S3). It has been reported that AtNF-YB2 and 3 and AtNF-YC3, 4, and 9 are critical regulators of flowering time (Kumimoto et al. 2010; Wenkel et al. 2006). Here, we detected expression of their orthologs, PtoNF-YB4, PtoNF-YB14, and PtoNF-YB5-7, during the early stages of floral bud development, but the specific regulatory mechanism of this expression remains unclear. Therefore, determining how these genes regulate floral bud development and affect the growth and development of P. tomentosa are interesting questions for future research.
Comprehensive bioinformatics analysis of the NF-Y family in P. trichocarpa identified 46 genes as putative PtNF-Y genes, and the expression of PtoNF-Y genes in various P. tomentosa tissues and organs was evaluated using transcriptome data. The results suggest that the expression and transcription levels of 34 of 46 members in various tissues and organs vary among family members. The results of this study provide a basis for further studies on the functions of the PtNF-Y gene family. However, the specific functions of these genes in P. tomentosa must be further verified experimentally, and the regulatory mechanisms must also be determined in future studies.
References
An XM, Wang DM, Wang ZL, Li B, Bo WH, Cao GL, Zhang ZY (2011) Isolation of a LEAFY homolog from Populus tomentosa: expression of PtLFY in P. tomentosa floral buds and PtLFY-IR-mediated gene silencing in tobacco (Nicotiana tabacum). Plant Cell Rep 30:89–100. https://doi.org/10.1007/s00299-010-0947-0
Bucher P, Trifonov EN (1988) CCAAT box revisited: bidirectionality, location and context. J Biomol Struct Dyn 5:1231–1236. https://doi.org/10.1080/07391102.1988.10506466
Calvenzani V, Testoni B, Gusmaroli G, Lorenzo M, Gnesutta N, Petroni K, Mantovani R, Tonelli C (2012) Interactions and CCAAT-binding of Arabidopsis thaliana NF-Y subunits. PLoS One 7:e42902. https://doi.org/10.1371/journal.pone.0042902
Cao S, Kumimoto RW, Siriwardana CL, Risinger JR, Holt BF 3rd (2011) Identification and characterization of NF-Y transcription factor families in the monocot model plant Brachypodium distachyon. PLoS One 6:e21805. https://doi.org/10.1371/journal.pone.0021805
Chen Z, Rao P, Yang X, Su X, Zhao T, Gao K, Yang X, An X (2018) A global view of transcriptome dynamics during male floral bud development in Populus tomentosa. Sci Rep 8:722. https://doi.org/10.1038/s41598-017-18084-5
Cole T, Williams BA, Geo P, Ali M, Gordon K, Baren MJ, Van Salzberg SL, Wold BJ, Lior P (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Feng ZJ, He GH, Zheng WJ, Lu PP, Chen M, Gong YM, Ma YZ, Xu ZS (2015) Foxtail millet NF-Y families: genome-wide survey and evolution analyses identified two functional genes important in abiotic stresses. Front Plant Sci 6:1142. https://doi.org/10.3389/fpls.2015.01142
Fornari M, Calvenzani V, Masiero S, Tonelli C, Petroni K (2013) The Arabidopsis NF-YA3 and NF-YA8 genes are functionally redundant and are required in early embryogenesis. PLoS One 8:e82043. https://doi.org/10.1371/journal.pone.0082043
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. https://doi.org/10.1093/nar/gkr944
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Ji L, Wang J, Ye M, Li Y, Guo B, Chen Z, Li H, An X (2013) Identification and characterization of the Populus AREB/ABF subfamily. J Integr Plant Biol 55:177–186. https://doi.org/10.1111/j.1744-7909.2012.01183.x
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kumimoto RW, Zhang Y, Siefers N, Holt BF 3rd (2010) NF-YC3, NF-YC4 and NF-YC9 are required for CONSTANS-mediated, photoperiod-dependent flowering in Arabidopsis thaliana. Plant J Cell Mol Biol 63:379–391. https://doi.org/10.1111/j.1365-313X.2010.04247.x
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lee H, Fischer RL, Goldberg RB, Harada JJ (2003) Arabidopsis LEAFY COTYLEDON1 represents a functionally specialized subunit of the CCAAT binding transcription factor. Proc Natl Acad Sci USA 100:2152–2156. https://doi.org/10.1073/pnas.0437909100
Li XY, Hooft vHR, Mantovani R, Benoist C, Mathis D (1992) Intron-exon organization of the NF-Y genes. Tissue-specific splicing modifies an activation domain. J Biol Chem 267:8984
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell Online 20:2238–2251. https://doi.org/10.1105/tpc.108.059444
Li S, Li K, Ju Z, Cao D, Fu D, Zhu H, Zhu B, Luo Y (2016) Genome-wide analysis of tomato NF-Y factors and their role in fruit ripening. BMC Genomics 17:36. https://doi.org/10.1186/s12864-015-2334-2
Liu JX, Howell SH (2010) bZIP28 and NF-Y transcription factors are activated by ER stress and assemble into a transcriptional complex to regulate stress response genes in Arabidopsis. Plant Cell 22:782–796. https://doi.org/10.1105/tpc.109.072173
Liu X, Hu P, Huang M, Tang Y, Li Y, Li L, Hou X (2016) The NF-YC-RGL2 module integrates GA and ABA signalling to regulate seed germination in Arabidopsis. Nat Commun 7:12768. https://doi.org/10.1038/ncomms12768
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Malviya N, Jaiswal P, Yadav D (2016) Genome-wide characterization of nuclear factor Y (NF-Y) gene family of sorghum [Sorghum bicolor (L.) Moench]: a bioinformatics approach. Physiol Mol Biol Plants 22:33–49. https://doi.org/10.1007/s12298-016-0349-z
Myers ZA, Kumimoto RW, Siriwardana CL, Gayler KK, Risinger JR, Pezzetta D, Holt Iii BF (2016) Nuclear factor Y, SUBUNIT C (NF-YC) transcription factors are positive regulators of photomorphogenesis in Arabidopsis thaliana. PLoS Genet 12:e1006333. https://doi.org/10.1371/journal.pgen.1006333
Nardini M, Gnesutta N, Donati G, Gatta R, Forni C, Fossati A, Vonrhein C, Moras D, Romier C, Bolognesi M, Mantovani R (2013) Sequence-specific transcription factor NF-Y displays histone-like DNA binding and H2B-like ubiquitination. Cell 152:132–143. https://doi.org/10.1016/j.cell.2012.11.047
Ni Z, Hu Z, Jiang Q, Zhang H (2013) GmNFYA3, a target gene of miR169, is a positive regulator of plant tolerance to drought stress. Plant Mol Biol 82:113–129. https://doi.org/10.1007/s11103-013-0040-5
Pereira SLS, Martins CPS, Sousa AO, Camillo LR, Araujo CP, Alcantara GM, Camargo DS, Cidade LC, de Almeida AF, Costa MGC (2018) Genome-wide characterization and expression analysis of citrus nuclear factor-Y (NF-Y) transcription factors identified a novel NF-YA gene involved in drought-stress response and tolerance. PLoS One 13:e0199187. https://doi.org/10.1371/journal.pone.0199187
Petroni K, Kumimoto RW, Gnesutta N, Calvenzani V, Fornari M, Tonelli C, Holt BF 3rd, Mantovani R (2012) The promiscuous life of plant nuclear factor Y transcription factors. Plant Cell 24:4777–4792. https://doi.org/10.1105/tpc.112.105734
Potkar R, Recla J, Busov V (2013) ptr-MIR169 is a posttranscriptional repressor of PtrHAP2 during vegetative bud dormancy period of aspen (Populus tremuloides) trees. Biochem Biophys Res Commun 431:512–518. https://doi.org/10.1016/j.bbrc.2013.01.027
Quach TN, Nguyen HT, Valliyodan B, Joshi T, Xu D, Nguyen HT (2015) Genome-wide expression analysis of soybean NF-Y genes reveals potential function in development and drought response. Mol Genet Genomics MGG 290:1095–1115. https://doi.org/10.1007/s00438-014-0978-2
Quan S, Niu J, Zhou L, Xu H, Ma L, Qin Y (2018) Identification and characterization of NF-Y gene family in walnut (Juglans regia L.). BMC Plant Biol 18:255. https://doi.org/10.1186/s12870-018-1459-2
Roy SW, Gilbert W (2006) The evolution of spliceosomal introns: patterns, puzzles and progress. Nat Rev Genet 7:211–221. https://doi.org/10.1038/nrg1807
Siefers N, Dang KK, Kumimoto RW, Bynum WEt, Tayrose G, Holt BF 3rd (2009) Tissue-specific expression patterns of Arabidopsis NF-Y transcription factors suggest potential for extensive combinatorial complexity. Plant Physiol 149:625–641. https://doi.org/10.1104/pp.108.130591
Su H, Cao Y, Ku L, Yao W, Cao Y, Ren Z, Dou D, Wang H, Ren Z, Liu H, Tian L, Zheng Y, Chen C, Chen Y (2018) Dual functions of ZmNF-YA3 in photoperiod-dependent flowering and abiotic stress responses in maize. J Exp Bot 69:5177–5189. https://doi.org/10.1093/jxb/ery299
Swarbreck D, Wilks C, Lamesch P, Berardini TZ, Garcia-Hernandez M, Foerster H, Li D, Meyer T, Muller R, Ploetz L, Radenbaugh A, Singh S, Swing V, Tissier C, Zhang P, Huala E (2008) The Arabidopsis Information Resource (TAIR): gene structure and function annotation. Nucleic Acids Res 36:D1009–D1014. https://doi.org/10.1093/nar/gkm965
Thirumurugan T, Ito Y, Kubo T, Serizawa A, Kurata N (2008) Identification, characterization and interaction of HAP family genes in rice. Mol Genet Genomics MGG 279:279–289. https://doi.org/10.1007/s00438-007-0312-3
Tom L, Stéphane DM, Pascal G, Maël B, Andreas N (2013) CCAAT-box binding transcription factors in plants: y so many? Trends Plant Sci 18:157–166
Wang Y, Xu W, Chen Z, Han B, Haque ME, Liu A (2018) Gene structure, expression pattern and interaction of nuclear factor-Y family in castor bean (Ricinus communis). Planta 247:559–572. https://doi.org/10.1007/s00425-017-2809-2
Wei Q, Ma C, Xu Y, Wang T, Chen Y, Lu J, Zhang L, Jiang CZ, Hong B, Gao J (2017) Control of chrysanthemum flowering through integration with an aging pathway. Nat Commun 8:829. https://doi.org/10.1038/s41467-017-00812-0
Wenkel S, Turck F, Singer K, Gissot L, Le Gourrierec J, Samach A, Coupland G (2006) CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 18:2971–2984. https://doi.org/10.1105/tpc.106.043299
Xing Y, Fikes JD, Guarente L (1993) Mutations in yeast HAP2/HAP3 define a hybrid CCAAT box binding domain. EMBO J 12:4647
Yan D-H, Xia X, Yin W (2012) NF-YB family genes identified in a poplar genome-wide analysis and expressed in Populus euphratica are responsive to drought stress. Plant Mol Biol Rep 31:363–370. https://doi.org/10.1007/s11105-012-0508-5
Zhang Q, Zhang ZY, Lin SZ, Zheng HQ, Lin YZ, An XM, Li Y, Li HX (2008) Characterization of resistance gene analogs with a nucleotide binding site isolated from a triploid white poplar. Plant Biol 10:310–322. https://doi.org/10.1111/j.1438-8677.2008.00029.x
Zhang Z, Li X, Zhang C, Zou H, Wu Z (2016) Isolation, structural analysis, and expression characteristics of the maize nuclear factor Y gene families. Biochem Biophys Res Commun 478:752–758. https://doi.org/10.1016/j.bbrc.2016.08.020
Zhang S, Tong Y, Li Y, Cheng ZM, Zhong Y (2019) Genome-wide identification of the HKT genes in five Rosaceae species and expression analysis of HKT genes in response to salt-stress in Fragaria vesca. Genes Genomics 41:325–336. https://doi.org/10.1007/s13258-018-0767-0
Zhao H, Wu D, Kong F, Lin K, Zhang H, Li G (2016) The Arabidopsis thaliana nuclear factor Y transcription factors. Front Plant Sci 7:2045. https://doi.org/10.3389/fpls.2016.02045
Zheng H, Lin S, Zhang Q, Lei Y, Zhang Z (2009) Functional analysis of 5′ untranslated region of a TIR-NBS-encoding gene from triploid white poplar. Mol Genet Genomics MGG 282:381–394. https://doi.org/10.1007/s00438-009-0471-5
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31570661), the National Key Program on Transgenic Research (2018ZX08020002-002-004), and the medium-long-term project of young teachers (2015ZCQ-SW-01). The English in this document has been checked by at least two professional editors, both native speakers of English. (http://www.textcheck.com/certificate/gzh38s).
Author information
Authors and Affiliations
Contributions
Juan Li, Kai Gao, and Xinmin An designed the experiments and wrote the manuscript. Juan Li, Wasif Ullah Khan, Tianyun Zhao, and Xiong Yang performed the experiments. Juan Li, Xiaoyu Yang, and Zhong Chen analyzed the data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by D.G. Peterson.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, J., Gao, K., Khan, W.U. et al. Genome-wide analysis of the poplar NF-Y gene family and its expression in floral bud development of Populus tomentosa. Trees 34, 285–296 (2020). https://doi.org/10.1007/s00468-019-01917-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-019-01917-3