Abstract
Pigmentary mosaicism of the Ito type, also known as hypomelanosis of Ito, is a neurocutaneous syndrome considered to be predominantly caused by somatic chromosomal mosaicism. However, a few monogenic causes of pigmentary mosaicism have been recently reported. Eleven unrelated individuals with pigmentary mosaicism (mostly hypopigmented skin) were recruited for this study. Skin punch biopsies of the probands and trio-based blood samples (from probands and both biological parents) were collected, and genomic DNA was extracted and analyzed by exome sequencing. In all patients, plausible monogenic causes were detected with somatic and germline variants identified in five and six patients, respectively. Among the somatic variants, four patients had MTOR variant (36%) and another had an RHOA variant. De novo germline variants in USP9X, TFE3, and KCNQ5 were detected in two, one, and one patients, respectively. A maternally inherited PHF6 variant was detected in one patient with hyperpigmented skin. Compound heterozygous GTF3C5 variants were highlighted as strong candidates in the remaining patient. Exome sequencing, using patients’ blood and skin samples is highly recommended as the first choice for detecting causative genetic variants of pigmentary mosaicism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pigmentary mosaicism, or hypomelanosis of Ito (HI, MIM# 300337), is a rare neurocutaneous syndrome characterized by hypopigmented skin whorls along Blaschko’s lines. It is associated with nervous system abnormality and other ectodermal defects involving fingers, toes, and teeth (Kouzak et al. 2013; Sybert 1994). Hypopigmented lesions have a reduced number of melanocytes, which originate from neural crest cells (Carmignac et al. 2021; Kuster and Konig 1999; Lee et al. 1999). In some cases, it is difficult to determine whether the affected skin is hypopigmented or hyperpigmented (Nehal et al. 1996), and the term “pigmentary mosaicism” refers to patterned hypopigmentation or hyperpigmentation resulting from a clone of skin cells with altered melanin-producing ability (Kromann et al. 2018; Shaffer and Chernoff 2020). The prevalence of HI is unknown, but it is estimated to be between 1/10,000 and 1/8,500, making it one of the most predominant rare neurocutaneous syndromes (Barbel et al. 2015). It affects both sexes, with a female-to-male prevalence ratio of 2:1. It has long been considered that somatic mosaicism is involved in HI, because the differently pigmented skin areas correspond to the varying distribution of two mosaic cell lines in each individual. Up to 60% of individuals with HI present with multiple types of mosaic cytogenetic abnormalities, including numerical (e.g., trisomy 18 or 20, triploidy) and structural chromosomal aberrations (e.g., deletions, duplications, or translocations) (Kromann et al. 2018; Salas-Labadia et al. 2019; Shaffer and Chernoff 2020; Thomas et al. 1989). However, no medical treatments are currently available for such chromosomal abnormalities, even if they were detected. Thus, skin biopsies, only to confirm chromosomal abnormalities, have been less motivated.
Recently, a few monogenic causes of HI have been reported. Germline TFE3 (MIM* 314310) and somatic RHOA (MIM* 165390) variants were found in pigmentary mosaicism (Lehalle et al. 2020; Vabres et al. 2019; Yigit et al. 2020). Somatic or germline MTOR (MIM* 601231) variants have also been occasionally associated with pigmentary mosaicism (Gordo et al. 2018; Hadouiri et al. 2020; Handoko et al. 2019; Mirzaa et al. 2016) and a large cohort of MTOR-related HI has recently been reported (Carmignac et al. 2021). This study aims to further explore the monogenic causes of pigmentary mosaicism.
Materials and methods
Patients
Individuals who were clinically diagnosed with pigmentary mosaicism were recruited through the collaborative activity of the Japanese Society of Child Neurology between April 2018 and March 2021. A total of 11 patients with pigmentary mosaicism (clinically diagnosed as HI and predominant skin hypopigmentation) were enrolled (Fig. 1). Skin samples were obtained from the patients, whereas blood samples were collected from the patients and their parents. Fresh skin tissues were obtained by punch biopsies of hypopigmented skin. Genomic DNA was extracted from the skin and peripheral blood leukocytes. Informed consent was obtained from the patients’ parents. This study protocol was approved by the Institutional Review Board of the Yokohama City University Faculty of Medicine.
Flowchart of the study. DNA was extracted from blood leukocytes and skin lesion biopsies of patients with pigmentary mosaicism and analyzed by next-generation sequencing. Rare variants only in the skin (somatic variants) or in both the skin and blood (germline or somatic variants) were investigated. Candidate germline variants were confirmed by Sanger sequencing of trio-samples (from patients and their parents). Exome sequencing of 11 individuals with pigmentary mosaicism revealed four somatic MTOR variants, a somatic RHOA variant; de novo variants in USP9X (n = 2), TFE3 (n = 1), and KCNQ5 (n = 1); biallelic germline variants in GTF3C5; and one inherited germline variant in PHF6
Exome sequencing (ES)
The flowchart of the study design is shown in Fig. 1. DNA was extracted from skin biopsies using proteinase K and sodium dodecyl sulfate with RNase and phenol/chloroform. DNA extraction from peripheral blood leukocytes was performed using the QuickGene DNA Extraction Whole Blood Kit L (Kurabo Industries Ltd., Osaka, Japan). ES was performed at a standard depth (approximately, 40–70×) as previously described (Sakamoto et al. 2021). In brief, genomic DNA was captured with a SureSelect Human All Exon V6 Kit (Agilent Technologies, Santa Clara, CA, USA) and sequenced on a Illumina Novaseq 6000 Sequencing System. Reads were aligned to the human reference genome (hg19) using Novoalign v3.02.13 (http://www.novocraft.com/).
Single nucleotide variant (SNV) and copy number variation (CNV) analysis
Germline variants in exons and at canonical splice sites (± 2 bp) were selected based on autosomal dominant and X-linked models considering de novo occurrence. Simultaneously, rare variants, assuming autosomal recessive inheritance were also considered. These variants were absent from our in-house exome database (n = 575) and the genome aggregation database (https://gnomad.broadinstitute.org/) (or at extremely low frequency if recessive). Somatic variants were extracted using MuTect2 (Cibulskis et al. 2013) and Varscan2 (Koboldt et al. 2012). We focused on variants detected by both programs. Candidate somatic variants were selected if they were covered by at least three or more reads. Known gene variants which have been previously reported in neurocutaneous syndrome-related phenotype on the Human Gene Mutation Database (HGMD) or the Catalogue of Somatic Mutations in Cancer (COSMIC) databases were prioritized among candidates with > 4% of variant allele frequencies (VAF). Candidate variants were validated by Sanger sequencing. Primers were designed using Primer3Plus, a web tool for designing primers (primer information is available on request) and PCR was performed using TaKaRa Ex Taq DNA Polymerase with 30 cycles of PCR. CNV was detected using eXome Hidden Markov Model (XHMM) and Nord’s program as described previously (Miyatake et al. 2015; Tsuchida et al. 2018; Uchiyama et al. 2021).
Results
ES analysis
ES identified plausible monogenic variants in all patients (11/11) (Fig. 1). The candidate variants information is summarized in Table 1 and Supplementary Table 1. The mean read depths of the protein coding sequences were covered with 40 reads or more. De novo somatic MTOR variants were identified in four patients [Patient 1, NM_004958.4: c.4448G>A, p.(Cys1483Tyr); Patient 2, c.5930C>T, p.(Thr1977Ile); Patient 3, c.6644C>T, p.(Ser2215Phe), Patient 4, c.7292T>C, p.(Leu2431Pro)]. Of these, three were exclusively detected in skin lesions, and one was identified in both skin and blood samples (Table 2). The somatic VAF of the MTOR mutant allele detected in skin in Patients 1, 2, 3, and 4 were 26.6% (20/70 reads), 21.3% (10/43 reads), 11.4% (6/62 reads), and 22.8% (8/35 reads), respectively. In Patient 4, the MTOR mutant read was also detected in the blood sample (5/28, 17.8%). All mosaic variants were confirmed to have signal peaks compared with wild-type variants in Sanger sequencing electropherograms (Supplementary Fig. 1). A somatic RHOA variant [NM_001664.4:c.139G>A, p.(Glu47Lys)] was found in the skin of Patient 5, with a VAF of 25.6%; this has been previously reported (Yigit et al. 2020). As for germline variants, de novo null variants in USP9X were found in two patients [Patient 7, NM_001039590.3: c.355dupT, p.(Arg121Profs*2); Patient 8, c.5834G > A, p.(Trp1945*)]. De novo TFE3 and KCNQ5 variants were identified in Patient 6 [NM_006521.6:c.560C>T, p.(Thr187Met)] and Patient 10 [NM_001160133.2:c.1477C>T, p.(Arg439Trp)], respectively. Patient 6 has been previously reported (Lehalle et al. 2020). A pathogenic PHF6 variant [NM_032458.3: c.821G>A, p.(Arg274Gln)] was maternally inherited in Patient 9, who presented hyperpigmented skin, rather than the usual hypopigmentation pattern. In Patient 11, we could neither identify de novo mosaic variants in the skin, nor any de novo germline variants. However, trio-based ES of blood DNA revealed biallelic GTF3C5 variants [NM_001286709.2: c.1160A>G; 1517G>T, p.(Lys387Arg); p.(Gly513Val)] as the candidates for this neurocutaneous phenotype.
Clinical data
Clinical photographs and images from brain imaging studies of patients are shown in Figs. 2 and 3, and clinical information is described in detail in Table 1 and in the Supplementary Information. Of the 11 patients, 9 (81.8%) exhibited intellectual disability. Patient 5 with a mosaic RHOA variant developed normally, whereas Patient 8 who had a germline USP9X variant had delayed developmental milestones (such as head control at 6 months, sitting at 11 months, walking at 17 months, and speaking a word at 2 years), but reached to the normal level of development by 6 years of age. Of the 11 patients, 7 (63.6%) had epilepsy, 6 (54.5%) exhibited brain abnormalities, 5 (45.5%) had dental anomalies, 3 (27.2%) experienced body/limb asymmetry, and another 3 (27.2%) demonstrated distinctive facial features. Macrocephaly was observed in only one case (Patient 4 with a somatic MTOR variant in blood and skin samples).
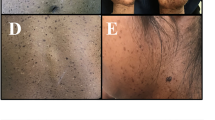
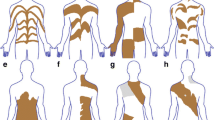
Skin lesion photographs and brain magnetic resonance imaging (MRI) studies of patients 1–4. a–f Patient 1 with a somatic MTOR somatic variant (c.4448G>A, p.Cys1483Tyr). a, c Hypopigmentation of the back and buttocks. b Axial view of T1-weighted MRI images of the brain. The brain is almost symmetrical, but the right hemisphere is slightly larger than the left. d Axial image of brain positron emission tomography (PET). The signals are significantly decreased in the right hemisphere (arrowhead). e, f Fontana–Masson stained skin biopsy of a depigmented lesion. Arrow indicates clusters of melanin-laden keratinocytes (brown areas), which are sparse in the hypopigmented skin compared to a normally pigmented skin lesion, although the number of melanocytes is not reducted (f). g Skin lesion of Patient 2. Hypopigmentation of the skin is seen only in the right side of the body. h, i T2-weighted coronal and axial images of brain MRI of Patient 2 with a somatic MTOR variant (c.5930C>T, p.Thr1977Ile). The right hemisphere is slightly larger than the left. The arrowheads show focal cortical dysplasia in bilateral occipital lobes, but predominantly in the right hemisphere. j Skin lesion of Patient 3 with a somatic MTOR variant (c.6644C>T, p.Ser2215Phe). k T2-weighted MRI brain finding of Patient 3 at day 21 after birth prior to surgery. Right hemimegalencephaly is observed in many slices. l, m Skin lesions and T2-weighted brain MRI of Patient 4 with a somatic MTOR variant (c.7292 T>C, p.Leu2431Pro) found in both skin lesion and blood leukocytes, respectively. Brain MRI of Patient 4 is normal and almost symmetrical
Skin lesion photographs and brain magnetic resonance imaging (MRI) studies of patients 5–11. a Hypopigmentation in the left leg of Patient 5 with a somatic RHOA variant. b, c Saggital and axial views of T2-weighted brain MRI, respectively. No laterality is observed. Hyperintensity with mild dilatation of ventricles and small cysts are observed. d Skin lesion of Patient 6 with a germline TFE3 variant. e, f Skin lesions of Patients 7 and 8, respectively, both with a germline USP9X variant. g, h Skin lesion and sagittal view of T2-weighted brain MRI of Patient 9 with a germline PHF6 variant. The patient’s skin showed hyperpigmentation rather than hypopigmentation. Atrophy of the bridge capsules and superior cerebellar peduncles is observed. i Skin lesion of Patient 10 with a germline KCNQ5 variant. j, k Skin lesions and sagittal view of T2-weighted brain MRI of Patient 11. Brain MRI showed polymicrogyria, loss of white matter volume, and mildly enlarged lateral ventricles
All of the four patients with somatic MTOR variants were diagnosed with epilepsy and developmental delay of varied severity after birth in addtion to skin hypopigmentation (Fig. 2 a, c, g, j, l). Patient 1 had a slightly enlarged right hemisphere on brain magnetic resonance imaging (MRI) (Fig. 2b). In a brain positron emission tomography scan, signals were significantly decreased in the right hemisphere (Fig. 2d). Patient 2 presented with unclear bilateral occipital cortexes (especially in the right) (Fig. 2i), but was able to walk and speak; therefore, his phenotype was considered relatively mild. Afebrile seizures occurred at 4 years, which were not improved with levetiracetam and lacosamide but were eventually well controlled with carbamazepine. Patient 3 had seizures 10 days after birth and exhibited severe developmental delay; thus, brain surgery was required to control refractory seizures. Brain MRI indicated that Patient 3 had apparent right hemimegalencephaly and focal cortical dysplasia (FCD) in the right occipital lobe, which was surgically corrected (Fig. 2k). Patient 4 showed macrocephaly, severe developmental delay, and epilepsy but without FCD or hemimegalencephaly on brain MRI (Fig. 2m), which is indicative of Smith–Kingsmore syndrome.
Patient 5 with a mosaic RHOA variant showed skin hypopigmentation (Fig. 3a) along with abnormally high MRI signals and cysts near the right ventricle (Fig. 3b and c). Almost all patients demonstrated skin hypopigmentation (Fig. 3a, d–f, i, j) except Patient 9, who had a PHF6 variant and demonstrated skin hyperpigmentation (Fig. 3g). The PHF6 variant was inherited from the patient’s mother who also had hyperpigmented skin. The familial pedigree of Patient 9 with genetic data is shown in Supplementary Fig. 2. Regarding a PHF6 variant, there was a report demonstrating grandparental genotyping was useful for interpreting ES results (Daum et al. 2020). We confirmed that the maternal grandmother did not possess a PHF6 variant. Unfortunately, a sample derived from the maternal grandfather was unavailable for confirmation. Neither grandparent exhibited hyperpigmentation or any other symptoms; therefore, the mother’s PHF6 variant may have occurred de novo. Patient 11 was born at 35 weeks of gestation by cesarean section because of oligohydramnios. He was noted to have HI after birth (Fig. 3j), refractory epilepsy at 1 year, and severe growth and developmental delay. The family pedigree of Patient 11 with genetic data is shown in Supplementary Fig. 2.
Discussion
We identified plausible monogenic causes of pigmentary mosaicism (mostly HI) in all 11 patients examined (Fig. 1). All causal genes identified in this study are highly intolerant to variation (high pLI, LOEUF, and z scores) except for the autosomal recessive GTF3C5 variants (Table 3). The high rate of monogenic cause of pigmentary mosaicism in this study was unexpected, but may be reasonable, considering that previous genetic investigations of pigmentary mosaicism were performed using only chromosomal analysis and/or array-based comparative genomic hybridization (without ES) (Kromann et al. 2018; Salas-Labadia et al. 2019). We propose that the exome-first strategy, preferably using blood- and skin-derived DNA, should be adopted for HI because it is a neuroectodermal disorder.
De novo somatic variants were identified in almost half cases (45%, five of 11 patients). Epidermal tissue contains both keratinocytes and melanocytes. It is well known that melanocytes are neuroectoderm derived. The neural tube (differentiating to neuronal cells in the central nervous system) and neuronal crest cells (to melanocytes) are derived from the neuroectoderm (Li et al. 2020). Thus, skin hypopigmentation could accompany the brain phenotype, depending on when somatic variants arise.
Postzygotic pathogenic variants in MTOR cause FCD (MIM# 607341), hemimegalencephaly, and Smith–Kingsmore syndrome (MIM# 616638). Among the 11 patients in this study, somatic MTOR variants were the most prevalent (n = 4, 36.3%), and among the four variants, three (Patients 1–3) were exclusively observed only in skin DNA, indicating that the variants may have occurred later in the postzygotic period (after epidermal differentiation), thus, only affecting the skin cells, while Patient 4 with Smith–Kingsmore syndrome had a variant in both skin and blood, indicating earlier occurrence in the postzygotic period (before epidermal differentiation). According to a review of Gordo et al. (2018), HI was observed in 14.8% (4/27) of patients with Smith–Kingsmore syndrome. Recently, MTOR-related HI in 14 (19.7%) of 71 patients with pigmentary mosaicism was reported (Carmignac et al. 2021), indicating that MTOR is the most common cause of pigmentary mosaicism. MTOR variants are often mosaic; therefore, skin biopsy is highly recommended for a conclusive diagnosis. All MTOR variants detected in this study have been previously known, and are all registered in COSMIC database as pathogenic somatic variants in cancers. The c.4448G>A, p.(Cys1483Tyr) variant in Patient 1 has been recurrently reported and is implicated as a cause of hemimegalencephaly and FCD or Smith–Kingsmore syndrome (D’Gama et al. 2015; Gordo et al. 2018). The c.5930C>T, p. (Thr1977Ile) variant found in Patient 2 has also been frequently reported as a cause of FCD and megalencephaly with pigmentary mosaicism (Carmignac et al. 2021; Handoko et al. 2019; Mirzaa et al. 2016). Interestingly, Handoko et al. (2019) confirmed that all four patients with c.5930C>T, p.(Thr1977Ile) showed cutaneous hypopigmentation along Blaschko’s lines. These two variants [c.4448G>A, p.(Cys1483Tyr) and c.5930C>T, p.(Thr1977Ile)] have been reported as either somatic or germline variant. The c.6644C>T, p.(Ser2215Phe) variant in Patient 3 has also been indicated as a cause of FCD or hemimegalencephaly (D’Gama et al. 2017; Moller et al. 2016), while the c.7292 T>C, p.(Leu2431Pro) variant in Patient 4 has never been reported as the cause of such condition (previously reported only in cancer). Somatic MTOR variants were first described as the cause of FCD requiring brain surgery to control epilepsy (Lim et al. 2015; Nakashima et al. 2015). Three patients in our cohort with MTOR somatic variants (except patient 3) did not require brain surgery, indicating that such variants can cause a relatively mild epilepsy phenotype. The VAF of MTOR variants in skin lesions was approximately 10–30%, which is consistent with that of a previous report regardless of the disease severity (Carmignac et al. 2021). The VAF of MTOR variants in skin lesions was considerably higher than that in FCD and hemimegalencephaly lesions, and it is usually less than 10% (sometimes 1–3% or less). This is advantageous for performing ES analysis of skin-derived DNA because deep sequencing may not be essential. As for patients with MTOR variants, mTOR inhibitors such as sirolimus and everolimus are potential treatment options.
Pigmentary mosaicism caused by somatic RHOA variants is usually associated with normal neurodevelopment, although apparent brain lesions can be detected on MRI, as in Patient 5 in this cohort (Vabres et al. 2019; Yigit et al. 2020). Previous reports have included patients with clinical manifestations compatible with this unique HI, but their genetic status was not confirmed (Inoue et al. 2015; Mateus et al. 2014). In two previous studies, the genetic cause of this type of HI in 9 of 11 patients (9/11) was a recurrent somatic RHOA variant, c.139G>A, p.(Glu47Lys), and is therefore called “RHOA-related mosaic ectodermal dysplasia” (Vabres et al. 2019; Yigit et al. 2020). RHOA variants are also frequently registered in the COSMIC database.
Pathogenic germline variants were found in three X-linked genes, TFE3 (transcription factor E3), USP9X (ubiquitin-specific protease 9, MIM*300072), and PHF6 (PHD finger protein 6, MIM*300414), which are consistent with the association of HI with structural abnormalities of the X chromosome. Germline or somatic TFE3 variants cause intellectual disability with pigmentary mosaicism and storage disorder-like features in both males and females (Lehalle et al. 2020). TFE3 functions in the mechanistic target of rapamycin (mTOR) complex 1 signaling in the PIK3–AKT–mTOR pathway, which can explain the association of these variants with pigmentary mosaicism similar to MTOR variants. Lehalle et al. (2020) found pigmentary mosaicism to be present in the majority of patients with TFE3 mutations (71%, 12/17), including 40% of the males and 83% of the females. USP9X and PHF6 variants cause distinct neurodevelopmental disorders with congenital anomalies in both sexes. X-linked recessive (MIM# 300919) and dominant (MIM# 300968) intellectual developmental disorders are caused by USP9X variants, whereas Borjeson–Forssman–Lehmann syndrome (BFLS, MIM# 301900) is caused by PHF6 variants. However, pigmentary mosaicism was only seen in female patients with abnormalities in these two genes (Jolly et al. 2020; Reijnders et al. 2016; Zhang et al. 2019). The rates of skin pigment complication in females related to USP9X and PHF6 variants were 65% (11/17) (Reijnders et al. 2016) and 77% (10/13) (Zhang et al. 2019), respectively. Skin lesions in females with aberrant USP9X and PHF6 manifest as hypopigmentation and hyperpigmentation along Blaschko’s lines, respectively. USP9X plays an important role associated with mTOR in neural development both in humans and mice (Agrawal et al. 2012; Murtaza et al. 2015; Reijnders et al. 2016), which may explain how this variant causes HI. PHF6 is implicated in chromatin remodeling by interacting with the SWI/SNF complex (Zhang et al. 2019; Zweier et al. 2013), the variants of which are occasionally found in patients with Coffin–Siris syndrome (MIM# 135900) (Zweier et al. 2013). In addition, pathogenic variants in USP9X and PHF6 are also found in cancers such as pediatric leukemia (Ma et al. 2018), which is similar to the reports of MTOR and RHOA variants being frequently detected in various cancers (Kilian et al. 2021; Mirzaa and Poduri 2014; Zou et al. 2020).
Germline KCNQ5 variants cause intellectual disability or epilepsy (Lehman et al. 2017), and only a few individuals with de novo KCNQ5 variants have been reported. Skin hypopigmentation in Patient 10 with a de novo KCNQ5 variant is indeed a novel phenotypic feature, which has not previously been reported in association with KCNQ5 abnormality. Therefore, it remains uncertain whether the KCNQ5 variant is related to the skin phenotype. GTF3C5 (MIM*604890) which encodes general transcription factor IIIC subunit 5 (GTF3C5), plays a crucial role in general transcription (Crepaldi et al. 2013); however, GTF3C5 abnormalities have never been reported in human diseases. Homozygous mutant Gtf3c5tm2a(KOMP)Wtsi mice exhibited complete preweaning lethality, while heterozygous mutant mice survived (Ayadi et al. 2012). The biallelic GTF3C5 variants may potentially cause skin hypopigmentation based on the high probability score (0.99) of gene intolerance to the biallelic loss of function (Lek et al. 2016). Further studies involving patients with GTF3C5 variants are required to confirm the association of GTF3C5 with this human ectodermal disorder. It is important to study and analyze more patients harboring KCNQ5 and GTF3C5 variants and collect further evidence before establishing any conclusion.
HI is thought to be caused by somatic chromosomal structural abnormalities, but these structural abnormalities have been inconsistent, and genetic testing for diagnosis has not been actively performed until recently. Moreover, HI has sometimes been considered as a benign phenotype rather than a disease (Ruggieri and Pavone 2000; Sybert 1994). However, the high discovery rate of monogenic causes of HI prompts us to perform ES for patients with pigmentary mosaicism, preferably by collecting skin samples, as this may lead to the optimal therapeutic approaches in the future. Although using skin tissue instead of brain tissue may allow regular ES with 40–70× read coverage for reliably detecting somatic mosaic variants with > 4% of VAF in pigmentary mosaicism, higher read coverage in the NGS analysis is better to avoid missing low-prevalent (≦ 4%) variants.
Data availability
ES data cannot be shared to preserve the patient’s privacy, but other data are available upon reasonable requests.
References
Agrawal P, Chen YT, Schilling B, Gibson BW, Hughes RE (2012) Ubiquitin-specific peptidase 9, X-linked (USP9X) modulates activity of mammalian target of rapamycin (mTOR). J Biol Chem 287:21164–21175. https://doi.org/10.1074/jbc.M111.328021
Ayadi A, Birling MC, Bottomley J, Bussell J, Fuchs H, Fray M, Gailus-Durner V, Greenaway S, Houghton R, Karp N, Leblanc S, Lengger C, Maier H, Mallon AM, Marschall S, Melvin D, Morgan H, Pavlovic G, Ryder E, Skarnes WC, Selloum M, Ramirez-Solis R, Sorg T, Teboul L, Vasseur L, Walling A, Weaver T, Wells S, White JK, Bradley A, Adams DJ, Steel KP, Hrabe de Angelis M, Brown SD, Herault Y (2012) Mouse large-scale phenotyping initiatives: overview of the European Mouse Disease Clinic (EUMODIC) and of the Wellcome Trust Sanger Institute Mouse Genetics Project. Mamm Genome 23:600–610. https://doi.org/10.1007/s00335-012-9418-y
Barbel P, Brown S, Peterson K (2015) Identification of hypomelanosis of Ito in pediatric primary care. J Pediatr Health Care 29:551–554. https://doi.org/10.1016/j.pedhc.2015.01.002
Carmignac V, Mignot C, Blanchard E, Kuentz P, Aubriot-Lorton MH, Parker VER, Sorlin A, Fraitag S, Courcet JB, Duffourd Y, Rodriguez D, Knox RG, Polubothu S, Boland A, Olaso R, Delepine M, Darmency V, Riachi M, Quelin C, Rollier P, Goujon L, Grotto S, Capri Y, Jacquemont ML, Odent S, Amram D, Chevarin M, Vincent-Delorme C, Catteau B, Guibaud L, Arzimanoglou A, Keddar M, Sarret C, Callier P, Bessis D, Genevieve D, Deleuze JF, Thauvin C, Semple RK, Philippe C, Riviere JB, Kinsler VA, Faivre L, Vabres P, Mosaique PN (2021) Clinical spectrum of MTOR-related hypomelanosis of Ito with neurodevelopmental abnormalities. Genet Med. https://doi.org/10.1038/s41436-021-01161-6
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES, Getz G (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31:213–219. https://doi.org/10.1038/nbt.2514
Crepaldi L, Policarpi C, Coatti A, Sherlock WT, Jongbloets BC, Down TA, Riccio A (2013) Binding of TFIIIC to sine elements controls the relocation of activity-dependent neuronal genes to transcription factories. PLoS Genet 9:e1003699. https://doi.org/10.1371/journal.pgen.1003699
Daum H, Mor-Shaked H, Ta-Shma A, Shaag A, Silverstein S, Shohat M, Elpeleg O, Meiner V, Harel T (2020) Grandparental genotyping enhances exome variant interpretation. Am J Med Genet A 182:689–696. https://doi.org/10.1002/ajmg.a.61511
D’Gama AM, Geng Y, Couto JA, Martin B, Boyle EA, LaCoursiere CM, Hossain A, Hatem NE, Barry BJ, Kwiatkowski DJ, Vinters HV, Barkovich AJ, Shendure J, Mathern GW, Walsh CA, Poduri A (2015) Mammalian target of rapamycin pathway mutations cause hemimegalencephaly and focal cortical dysplasia. Ann Neurol 77:720–725. https://doi.org/10.1002/ana.24357
D’Gama AM, Woodworth MB, Hossain AA, Bizzotto S, Hatem NE, LaCoursiere CM, Najm I, Ying Z, Yang E, Barkovich AJ, Kwiatkowski DJ, Vinters HV, Madsen JR, Mathern GW, Blumcke I, Poduri A, Walsh CA (2017) Somatic mutations activating the mTOR pathway in dorsal telencephalic progenitors cause a continuum of cortical dysplasias. Cell Rep 21:3754–3766. https://doi.org/10.1016/j.celrep.2017.11.106
Gordo G, Tenorio J, Arias P, Santos-Simarro F, Garcia-Minaur S, Moreno JC, Nevado J, Vallespin E, Rodriguez-Laguna L, de Mena R, Dapia I, Palomares-Bralo M, Del Pozo A, Ibanez K, Silla JC, Barroso E, Ruiz-Perez VL, Martinez-Glez V, Lapunzina P (2018) mTOR mutations in Smith-Kingsmore syndrome: four additional patients and a review. Clin Genet 93:762–775. https://doi.org/10.1111/cge.13135
Hadouiri N, Darmency V, Guibaud L, Arzimanoglou A, Sorlin A, Carmignac V, Riviere JB, Huet F, Luu M, Bardou M, Thauvin-Robinet C, Vabres P, Faivre L (2020) Compassionate use of everolimus for refractory epilepsy in a patient with MTOR mosaic mutation. Eur J Med Genet 63:104036. https://doi.org/10.1016/j.ejmg.2020.104036
Handoko M, Emrick LT, Rosenfeld JA, Wang X, Tran AA, Turner A, Belmont JW, Undiagnosed Diseases N, Lee BH, Bacino CA, Chao HT (2019) Recurrent mosaic MTOR c.5930C > T (p.Thr1977Ile) variant causing megalencephaly, asymmetric polymicrogyria, and cutaneous pigmentary mosaicism: case report and review of the literature. Am J Med Genet A 179:475–479. https://doi.org/10.1002/ajmg.a.61007
Inoue M, Fukuda M, Ishii E, Sayama K (2015) Linear leukoplakia and central nervous system lesions: a clinical clue to the diagnosis of hypomelanosis of Ito. J Pediatr 167(771):e1. https://doi.org/10.1016/j.jpeds.2015.06.024
Jolly LA, Parnell E, Gardner AE, Corbett MA, Perez-Jurado LA, Shaw M, Lesca G, Keegan C, Schneider MC, Griffin E, Maier F, Kiss C, Guerin A, Crosby K, Rosenbaum K, Tanpaiboon P, Whalen S, Keren B, McCarrier J, Basel D, Sadedin S, White SM, Delatycki MB, Kleefstra T, Kury S, Brusco A, Sukarova-Angelovska E, Trajkova S, Yoon S, Wood SA, Piper M, Penzes P, Gecz J (2020) Missense variant contribution to USP9X-female syndrome. NPJ Genom Med 5:53. https://doi.org/10.1038/s41525-020-00162-9
Kilian LS, Voran J, Frank D, Rangrez AY (2021) RhoA: a dubious molecule in cardiac pathophysiology. J Biomed Sci 28:33. https://doi.org/10.1186/s12929-021-00730-w
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, Miller CA, Mardis ER, Ding L, Wilson RK (2012) VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22:568–576. https://doi.org/10.1101/gr.129684.111
Kouzak SS, Mendes MS, Costa IM (2013) Cutaneous mosaicisms: concepts, patterns and classifications. An Bras Dermatol 88:507–517. https://doi.org/10.1590/abd1806-4841.20132015
Kromann AB, Ousager LB, Ali IKM, Aydemir N, Bygum A (2018) Pigmentary mosaicism: a review of original literature and recommendations for future handling. Orphanet J Rare Dis 13:39. https://doi.org/10.1186/s13023-018-0778-6
Kuster W, Konig A (1999) Hypomelanosis of Ito: no entity, but a cutaneous sign of mosaicism. Am J Med Genet 85:346–350. https://doi.org/10.1002/(sici)1096-8628(19990806)85:4%3c346::aid-ajmg7%3e3.0.co;2-1
Lee HS, Chun YS, Hann SK (1999) Nevus depigmentosus: clinical features and histopathologic characteristics in 67 patients. J Am Acad Dermatol 40:21–26. https://doi.org/10.1016/s0190-9622(99)70524-4
Lehalle D, Vabres P, Sorlin A, Bierhals T, Avila M, Carmignac V, Chevarin M, Torti E, Abe Y, Bartolomaeus T, Clayton-Smith J, Cogne B, Cusco I, Duplomb L, De Bont E, Duffourd Y, Duijkers F, Elpeleg O, Fattal A, Genevieve D, Guillen Sacoto MJ, Guimier A, Harris DJ, Hempel M, Isidor B, Jouan T, Kuentz P, Koshimizu E, Lichtenbelt K, Loik Ramey V, Maik M, Miyakate S, Murakami Y, Pasquier L, Pedro H, Simone L, Sondergaard-Schatz K, St-Onge J, Thevenon J, Valenzuela I, Abou Jamra R, van Gassen K, van Haelst MM, van Koningsbruggen S, Verdura E, Whelan Habela C, Zacher P, Riviere JB, Thauvin-Robinet C, Betschinger J, Faivre L (2020) De novo mutations in the X-linked TFE3 gene cause intellectual disability with pigmentary mosaicism and storage disorder-like features. J Med Genet 57:808–819. https://doi.org/10.1136/jmedgenet-2019-106508
Lehman A, Thouta S, Mancini GMS, Naidu S, van Slegtenhorst M, McWalter K, Person R, Mwenifumbo J, Salvarinova R, Study C, Study E, Guella I, McKenzie MB, Datta A, Connolly MB, Kalkhoran SM, Poburko D, Friedman JM, Farrer MJ, Demos M, Desai S, Claydon T (2017) Loss-of-function and gain-of-function mutations in KCNQ5 cause intellectual disability or epileptic encephalopathy. Am J Hum Genet 101:65–74. https://doi.org/10.1016/j.ajhg.2017.05.016
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, O’Donnell-Luria AH, Ware JS, Hill AJ, Cummings BB, Tukiainen T, Birnbaum DP, Kosmicki JA, Duncan LE, Estrada K, Zhao F, Zou J, Pierce-Hoffman E, Berghout J, Cooper DN, Deflaux N, DePristo M, Do R, Flannick J, Fromer M, Gauthier L, Goldstein J, Gupta N, Howrigan D, Kiezun A, Kurki MI, Moonshine AL, Natarajan P, Orozco L, Peloso GM, Poplin R, Rivas MA, Ruano-Rubio V, Rose SA, Ruderfer DM, Shakir K, Stenson PD, Stevens C, Thomas BP, Tiao G, Tusie-Luna MT, Weisburd B, Won HH, Yu D, Altshuler DM, Ardissino D, Boehnke M, Danesh J, Donnelly S, Elosua R, Florez JC, Gabriel SB, Getz G, Glatt SJ, Hultman CM, Kathiresan S, Laakso M, McCarroll S, McCarthy MI, McGovern D, McPherson R, Neale BM, Palotie A, Purcell SM, Saleheen D, Scharf JM, Sklar P, Sullivan PF, Tuomilehto J, Tsuang MT, Watkins HC, Wilson JG, Daly MJ, MacArthur DG, Exome Aggregation C (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536:285–291. https://doi.org/10.1038/nature19057
Li M, Knapp SK, Iden S (2020) Mechanisms of melanocyte polarity and differentiation: What can we learn from other neuroectoderm-derived lineages? Curr Opin Cell Biol 67:99–108. https://doi.org/10.1016/j.ceb.2020.09.001
Lim JS, Kim WI, Kang HC, Kim SH, Park AH, Park EK, Cho YW, Kim S, Kim HM, Kim JA, Kim J, Rhee H, Kang SG, Kim HD, Kim D, Kim DS, Lee JH (2015) Brain somatic mutations in MTOR cause focal cortical dysplasia type II leading to intractable epilepsy. Nat Med 21:395–400. https://doi.org/10.1038/nm.3824
Ma X, Liu Y, Liu Y, Alexandrov LB, Edmonson MN, Gawad C, Zhou X, Li Y, Rusch MC, Easton J, Huether R, Gonzalez-Pena V, Wilkinson MR, Hermida LC, Davis S, Sioson E, Pounds S, Cao X, Ries RE, Wang Z, Chen X, Dong L, Diskin SJ, Smith MA, Guidry Auvil JM, Meltzer PS, Lau CC, Perlman EJ, Maris JM, Meshinchi S, Hunger SP, Gerhard DS, Zhang J (2018) Pan-cancer genome and transcriptome analyses of 1699 paediatric leukaemias and solid tumours. Nature 555:371–376. https://doi.org/10.1038/nature25795
Mateus AM, Silva RL, Conceicao C, Vieira JP (2014) Importance of cutaneous changes in the diagnosis of neurological diseases. Einstein (sao Paulo) 12:529–530. https://doi.org/10.1590/S1679-45082014AI2931
Mirzaa GM, Poduri A (2014) Megalencephaly and hemimegalencephaly: breakthroughs in molecular etiology. Am J Med Genet C Semin Med Genet 166C:156–172. https://doi.org/10.1002/ajmg.c.31401
Mirzaa GM, Campbell CD, Solovieff N, Goold C, Jansen LA, Menon S, Timms AE, Conti V, Biag JD, Adams C, Boyle EA, Collins S, Ishak G, Poliachik S, Girisha KM, Yeung KS, Chung BHY, Rahikkala E, Gunter SA, McDaniel SS, Macmurdo CF, Bernstein JA, Martin B, Leary R, Mahan S, Liu S, Weaver M, Doerschner M, Jhangiani S, Muzny DM, Boerwinkle E, Gibbs RA, Lupski JR, Shendure J, Saneto RP, Novotny EJ, Wilson CJ, Sellers WR, Morrissey M, Hevner RF, Ojemann JG, Guerrini R, Murphy LO, Winckler W, Dobyns WB (2016) Association of MTOR mutations with developmental brain disorders, including megalencephaly, focal cortical dysplasia, and pigmentary mosaicism. JAMA Neurol 73:836–845. https://doi.org/10.1001/jamaneurol.2016.0363
Miyatake S, Koshimizu E, Fujita A, Fukai R, Imagawa E, Ohba C, Kuki I, Nukui M, Araki A, Makita Y, Ogata T, Nakashima M, Tsurusaki Y, Miyake N, Saitsu H, Matsumoto N (2015) Detecting copy-number variations in whole-exome sequencing data using the eXome Hidden Markov Model: an “exome-first” approach. J Hum Genet 60:175–182. https://doi.org/10.1038/jhg.2014.124
Moller RS, Weckhuysen S, Chipaux M, Marsan E, Taly V, Bebin EM, Hiatt SM, Prokop JW, Bowling KM, Mei D, Conti V, de la Grange P, Ferrand-Sorbets S, Dorfmuller G, Lambrecq V, Larsen LH, Leguern E, Guerrini R, Rubboli G, Cooper GM, Baulac S (2016) Germline and somatic mutations in the MTOR gene in focal cortical dysplasia and epilepsy. Neurol Genet 2:e118. https://doi.org/10.1212/NXG.0000000000000118
Murtaza M, Jolly LA, Gecz J, Wood SA (2015) La FAM fatale: USP9X in development and disease. Cell Mol Life Sci 72:2075–2089. https://doi.org/10.1007/s00018-015-1851-0
Nakashima M, Saitsu H, Takei N, Tohyama J, Kato M, Kitaura H, Shiina M, Shirozu H, Masuda H, Watanabe K, Ohba C, Tsurusaki Y, Miyake N, Zheng Y, Sato T, Takebayashi H, Ogata K, Kameyama S, Kakita A, Matsumoto N (2015) Somatic mutations in the MTOR gene cause focal cortical dysplasia type IIb. Ann Neurol 78:375–386. https://doi.org/10.1002/ana.24444
Nehal KS, PeBenito R, Orlow SJ (1996) Analysis of 54 cases of hypopigmentation and hyperpigmentation along the lines of Blaschko. Arch Dermatol 132:1167–1170
Reijnders MR, Zachariadis V, Latour B, Jolly L, Mancini GM, Pfundt R, Wu KM, van Ravenswaaij-Arts CM, Veenstra-Knol HE, Anderlid BM, Wood SA, Cheung SW, Barnicoat A, Probst F, Magoulas P, Brooks AS, Malmgren H, Harila-Saari A, Marcelis CM, Vreeburg M, Hobson E, Sutton VR, Stark Z, Vogt J, Cooper N, Lim JY, Price S, Lai AH, Domingo D, Reversade B, Study DDD, Gecz J, Gilissen C, Brunner HG, Kini U, Roepman R, Nordgren A, Kleefstra T (2016) De novo loss-of-function mutations in USP9X cause a female-specific recognizable syndrome with developmental delay and congenital malformations. Am J Hum Genet 98:373–381. https://doi.org/10.1016/j.ajhg.2015.12.015
Ruggieri M, Pavone L (2000) Hypomelanosis of Ito: clinical syndrome or just phenotype? J Child Neurol 15:635–644. https://doi.org/10.1177/088307380001501001
Sakamoto M, Iwama K, Sekiguchi F, Mashimo H, Kumada S, Ishigaki K, Okamoto N, Behnam M, Ghadami M, Koshimizu E, Miyatake S, Mitsuhashi S, Mizuguchi T, Takata A, Saitsu H, Miyake N, Matsumoto N (2021) Novel EXOSC9 variants cause pontocerebellar hypoplasia type 1D with spinal motor neuronopathy and cerebellar atrophy. J Hum Genet 66:401–407. https://doi.org/10.1038/s10038-020-00853-2
Salas-Labadia C, Gomez-Carmona S, Cruz-Alcivar R, Martinez-Anaya D, Del Castillo-Ruiz V, Duran-McKinster C, Ulloa-Aviles V, Yokoyama-Rebollar E, Ruiz-Herrera A, Navarrete-Meneses P, Lieberman-Hernandez E, Gonzalez-Del Angel A, Cervantes-Barragan D, Villarroel-Cortes C, Reyes-Leon A, Suarez-Perez D, Pedraza-Melendez A, Gonzalez-Orsuna A, Perez-Vera P (2019) Genetic and clinical characterization of 73 Pigmentary Mosaicism patients: revealing the genetic basis of clinical manifestations. Orphanet J Rare Dis 14:259. https://doi.org/10.1186/s13023-019-1208-0
Shaffer JV, Chernoff KA (2020) Pigmentary mosaicism (hypomelanosis of Ito). In: Levy ML, Hand JL (eds) UpToDate, Waltham
Sybert VP (1994) Hypomelanosis of Ito: a description, not a diagnosis. J Invest Dermatol 103:141S-143S. https://doi.org/10.1111/1523-1747.ep12399466
Thomas IT, Frias JL, Cantu ES, Lafer CZ, Flannery DB, Graham JG Jr (1989) Association of pigmentary anomalies with chromosomal and genetic mosaicism and chimerism. Am J Hum Genet 45:193–205
Tsuchida N, Nakashima M, Kato M, Heyman E, Inui T, Haginoya K, Watanabe S, Chiyonobu T, Morimoto M, Ohta M, Kumakura A, Kubota M, Kumagai Y, Hamano SI, Lourenco CM, Yahaya NA, Ch’ng GS, Ngu LH, Fattal-Valevski A, Weisz Hubshman M, Orenstein N, Marom D, Cohen L, Goldberg-Stern H, Uchiyama Y, Imagawa E, Mizuguchi T, Takata A, Miyake N, Nakajima H, Saitsu H, Miyatake S, Matsumoto N (2018) Detection of copy number variations in epilepsy using exome data. Clin Genet 93:577–587. https://doi.org/10.1111/cge.13144
Uchiyama Y, Yamaguchi D, Iwama K, Miyatake S, Hamanaka K, Tsuchida N, Aoi H, Azuma Y, Itai T, Saida K, Fukuda H, Sekiguchi F, Sakaguchi T, Lei M, Ohori S, Sakamoto M, Kato M, Koike T, Takahashi Y, Tanda K, Hyodo Y, Honjo RS, Bertola DR, Kim CA, Goto M, Okazaki T, Yamada H, Maegaki Y, Osaka H, Ngu LH, Siew CG, Teik KW, Akasaka M, Doi H, Tanaka F, Goto T, Guo L, Ikegawa S, Haginoya K, Haniffa M, Hiraishi N, Hiraki Y, Ikemoto S, Daida A, Hamano SI, Miura M, Ishiyama A, Kawano O, Kondo A, Matsumoto H, Okamoto N, Okanishi T, Oyoshi Y, Takeshita E, Suzuki T, Ogawa Y, Handa H, Miyazono Y, Koshimizu E, Fujita A, Takata A, Miyake N, Mizuguchi T, Matsumoto N (2021) Efficient detection of copy-number variations using exome data: batch- and sex-based analyses. Hum Mutat 42:50–65. https://doi.org/10.1002/humu.24129
Vabres P, Sorlin A, Kholmanskikh SS, Demeer B, St-Onge J, Duffourd Y, Kuentz P, Courcet JB, Carmignac V, Garret P, Bessis D, Boute O, Bron A, Captier G, Carmi E, Devauchelle B, Genevieve D, Gondry-Jouet C, Guibaud L, Lafon A, Mathieu-Dramard M, Thevenon J, Dobyns WB, Bernard G, Polubothu S, Faravelli F, Kinsler VA, Thauvin C, Faivre L, Ross ME, Riviere JB (2019) Postzygotic inactivating mutations of RHOA cause a mosaic neuroectodermal syndrome. Nat Genet 51:1438–1441. https://doi.org/10.1038/s41588-019-0498-4
Yigit G, Saida K, DeMarzo D, Miyake N, Fujita A, Yang Tan T, White SM, Wadley A, Toliat MR, Motameny S, Franitza M, Stutterd CA, Chong PF, Kira R, Sengoku T, Ogata K, Guillen Sacoto MJ, Fresen C, Beck BB, Nurnberg P, Dieterich C, Wollnik B, Matsumoto N, Altmuller J (2020) The recurrent postzygotic pathogenic variant p.Glu47Lys in RHOA causes a novel recognizable neuroectodermal phenotype. Hum Mutat 41:591–599. https://doi.org/10.1002/humu.23964
Zhang X, Fan Y, Liu X, Zhu MA, Sun Y, Yan H, He Y, Ye X, Gu X, Yu Y (2019) A novel nonsense mutation of PHF6 in a female with extended phenotypes of Borjeson-Forssman-Lehmann syndrome. J Clin Res Pediatr Endocrinol 11:419–425. https://doi.org/10.4274/jcrpe.galenos.2019.2018.0220
Zou Z, Tao T, Li H, Zhu X (2020) mTOR signaling pathway and mTOR inhibitors in cancer: progress and challenges. Cell Biosci 10:31. https://doi.org/10.1186/s13578-020-00396-1
Zweier C, Kraus C, Brueton L, Cole T, Degenhardt F, Engels H, Gillessen-Kaesbach G, Graul-Neumann L, Horn D, Hoyer J, Just W, Rauch A, Reis A, Wollnik B, Zeschnigk M, Ludecke HJ, Wieczorek D (2013) A new face of Borjeson-Forssman-Lehmann syndrome? De novo mutations in PHF6 in seven females with a distinct phenotype. J Med Genet 50:838–847. https://doi.org/10.1136/jmedgenet-2013-101918
Acknowledgements
We thank the individuals and their families for their participation in this study. The authors thank Ms. Kaori Takabe, Mr. Takabumi Miyama, Ms. Nobuko Watanabe, Ms. Mai Sato, and Ms. Sayaka Sugimoto at Department of Human Genetics, Yokohama City University Graduate School of Medicine for their technical assistance.
Funding
This work was supported by the Japan Agency for Medical Research and Development (AMED) under Grant Numbers JP21ek0109486, JP21ek0109549, and JP21ek0109493 (N. Matsumoto); JSPS KAKENHI under Grant Numbers JP21K07869 (E. Koshimizu), JP20K17428 (N. Tsuchida), JP21k15907 (Y. Uchiyama), JP20K16932 (K. Hamanaka), JP20K17936 (A. Fujita), JP20K08164 (T. Mizuguchi), JP20K07907 (S. Miyatake), JP19H03621 (N. Miyake); and the Takeda Science Foundation (N. Miyake, T. Mizuguchi, and N. Matsumoto).
Author information
Authors and Affiliations
Contributions
KS performed the genetic analysis, interpreted the data and wrote the first draft. All authors read and commented on previous versions of the manuscript. All authors approved the final manuscript. Patient recruitment, material preparation and data collection were performed by PFC, AS, RM, KN, HO, TS, KF, EO MK and YA. Skin biopsy was conducted by AI, HA, KO and KK. EK, NT, YU, KM, AF, TM, SM and N. Miyake contributed to genetic data analysis. N. Matsumoto supervised and organized the study and secured funding.
Corresponding author
Ethics declarations
Conflict of interests
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Institutional Review Board of the Yokohama City University Faculty of Medicine.
Consent to participate
Written informed consent was obtained from the patients’ parents.
Web resources
The Human Gene Mutation Database (HGMD): http://www.hgmd.cf.ac.uk/ac/index
Catalogue of Somatic Mutations in Cancer database (COSMIC): https://cancer.sanger.ac.uk/cosmic
gnomAD: http://gnomad.broadinstitute.org/
OMIM: https://www.omim.org/
SIFT: http://sift.jcvi.org/
PolyPhen-2: http://genetics.bwh.harvard.edu/pph2/
Mutation Taster: http://www.mutationtaster.org/
CADD: https://cadd.gs.washington.edu/snv
Primer3plus: https://dev.primer3plus.com/index.html
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Sanger sequencing of Patients 1–5. Somatic variants are detected in the skin DNA and sometimes blood DNA as a small peak of signal intensity (arrows). Supplementary Fig. 2 Family pedigrees of Patients 9 and 11. a Exome sequencing was performed in the proband. *Sanger sequencing is performed. The pathogenic variant, c.821G>A, p.Arg274Gln in PHF6, is detected in the proband (Patient 9) and the mother, but not in the grandmother. b The compound heterozygous GTF3C5 variants in Patient 11. His father and mother are both carriers of the respective variants. (PPTX 346 KB)
Rights and permissions
About this article
Cite this article
Saida, K., Chong, P.F., Yamaguchi, A. et al. Monogenic causes of pigmentary mosaicism. Hum Genet 141, 1771–1784 (2022). https://doi.org/10.1007/s00439-022-02437-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-022-02437-w