Abstract
Vigna is a large, pan-tropic and highly variable group of the legumes family which is known for its > 10 cultivated species having significant commercial value for their nutritious grains and multifarious uses. The wild vignas are considered a reservoir of numerous useful traits which can be deployed for introgression of resistance to biotic and abiotic stresses, seed quality and enhanced survival capability in extreme environments. Nonetheless, for their effective utilization through introgression breeding information on their genetic diversity, population structure and crossability is imperative. Keeping this in view, the present experiment was undertaken with 119 accessions including 99 wild Vigna accessions belonging to 19 species and 18 cultivated genotypes of Vigna and 2 of Phaseolus. Total 102 polymorphic SSRs were deployed to characterize the material at molecular level which produced 1758 alleles. The genotypes were grouped into four major clusters which were further sub-divided in nine sub-clusters. Interestingly, all cultivated species shared a single cluster while no such similarities were observed for the wild accessions as these were distributed in different groups of sub-clusters. The co-dominant allelic data of 114 accessions were then utilized for obtaining status of the accessions and their hybrid forms. The model-based population structure analysis categorized 114 accessions of Vigna into 6 genetically distinct sub-populations (K = 6) following admixture-model based simulation with varying levels of admixture. 91 (79.82%) accessions resembled their hierarchy and 23 (20.18%) accessions were observed as the admixture forms. Maximum number of accessions (25) were grouped in sub-population (SP) 6 and the least accessions were grouped in SP3 and SP5 (11 each). The population genetic structure, therefore, supported genetic diversity analysis and provided an insight into the genetic lineage of these species which will help in effective use of germplasm for development of cultivars following selective prebreeding activities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vigna is an important genus of flowering plants in the legumes family which has a pan-tropic distribution. This genus comprises > 200 species (Pratap et al. 2014a) encompassed in five sub-genera (Ceratropis, Haydonia, Lasiospron, Plectrotropis and Vigna) (Takahashi et al. 2016) including ten domesticated species having significant agronomic potential. Among these, seven species belonging to the sub-genus Ceratotropis are also known as the Asiatic Vigna (Takahashi et al. 2016; Pratap et al. 2015) and include the versatile crops viz., mung bean (V. radiata), black gram (V. mungo (L.) Hepper), moth bean (V. aconiotifolia (Jacq.) Marechal), minni payaru (V. stipulacea Kuntze), creole bean (V. reflexo-pilosa Hayata), adzuki bean (V. angularis (Willd.) Ohwi & Ohashi) and rice bean (V. umbellata (Thunb)). These crops are mainly cultivated in South and South-east Asia and Africa (Pratap et al. 2014a, 2021a) and contribute significantly in the food and nutritional security and environmental sustainability. Rice bean (V. umbellata (Thumb.) Ohwi & Ohashi), tua pea (V. glabrescens) and creole bean (V. reflexo-pilosa) have been reported to be domesticated and consumed mostly in South Africa (Pratap et al. 2014a, 2015; Chankaew et al. 2014) and V. angularis (Willd) Ohwi & Ohashi (small red bean or adzuki bean) in East Asia, most probably in Japan (Tomooka 2009). An abundance of their wild and weedy types have been reported to flourish in the Savannah and the forested zones (Harlan 1971; Rawal 1975). At the other hand, V. trilobata and V. stipulacea have been reported as candidates for neo-domestication for drought tolerance and disease and pest resistance, respectively (Gore et al. 2019; Pratap et al. 2015).
Most of the Vigna crops are grown for their nutritious seeds which have a high amount of proteins and micro-nutrients. These crop species, although considered as minor, are important sources of dietary protein and also several micro-elements viz., iron, potassium, zinc, Vitamin A, Vitamin B, folate and thymine in the developing and underdeveloped countries, especially in the predominantly vegetarian diets (Vaz Patto et al. 2015). Many of these species are also valued as forage, cover, and green manure crops in many parts of the world. Nonetheless, while the domesticated and edible Vigna are only a few, the non-domesticated or semi-domesticated species are many. A few of these are also found to inhabit rough environments such as non-fertile rocky and mountainous tracts, sandy and salty beaches and uninhabited marshy and swampy lands and, therefore, these are expected to habour many survival-related traits (Pratap et al. 2014b). These species are, therefore, considered as a reservoir of valuable genetic resources for biotic and abiotic stress tolerance and seed quality traits (Douglas et al. 2020; Pratap et al. 2020, 2021b; Nair et al. 2019). Furthermore, many wild species are highly tolerant to extreme environmental conditions including drought and water logging (Bisht et al. 2005; Tomooka et al. 2014), high-salinity (Yoshida et al. 2016), heat and cold stress (HanumanthaRao et al. 2016), and acidic or alkaline soils (Soares et al. 2014) while others have resistance to bruchids (Kaewwongwal et al. 2017), cercospora leaf spot (Singh et al. 2017; Chankaew et al. 2013), and yellow mosaic disease (for a review, please see Singh et al. 2020). Many wild species also serve as a potential source for superior agronomic traits (Kajonphol et al. 2012; Aidbhavi et al. 2021), and photo-thermo insensitivity (Pratap et al. 2014b; Basu et al. 2019). Cross-compatibility studies and identification of tolerant and causative genes facilitating conventional genetics and breeding towards the new breeding concepts such as ‘neo-domestication’ and ‘reverse-breeding’ are imperative to harness the desirable traits of wild species for crop improvement (Palmgren et al. 2015). Further, to use wild species in crop improvement programmes through pre-breeding, precise information on their genetic architecture, population structure and relationship with other Vigna species are important. Morphological evaluation is highly environment-dependent, especially in Vigna crops and, therefore, may give variable results across different environments. Hence classifying the Vigna species using highly abundant molecular markers such as multi-allelic SSRs is indeed necessary and has been used in classifying wild species of many crops earlier (Wang et al. 2008; Gwag et al. 2010; Pratap et al. 2015; Sarr et al. 2020). Due to high polymorphism, multiple allelism, reproducibility, co-dominant nature and user-friendliness, the SSR markers are preferred genetic markers to recognize diversity in microsatellite variation (Weber and May 1989). Further, microsatellites or simple sequence repeats (SSRs) are widely distributed across plant genomes and have high sensitivity to detect polymorphisms (Parker et al. 1998). As a result, these have been abundantly deployed in discerning genetic diversity, phylogeny studies and population genetic structure analysis (Sarr et al. 2020; Kempf et al. 2016; Gwag et al. 2010; Pratap et al. 2015) in Vigna crops. Besides, SSR markers have been tremendously useful in successful marker-assisted breeding in food legumes (Varshney et al. 2014; Pratap et al. 2017). Several workers successfully utilized the SSR markers in developing mungbean maps (Chankaew et al. 2014; Isemura et al. 2012; Kitsanachandee et al. 2013), which indicated availability of reliable markers for marker-assisted selection and identification of QTLs for desired traits. The present investigation aimed to evaluate the genetic diversity among different accessions of a comprehensive set of Asiatic Vigna species, study their population genetic structure and interpret their inter-relationship for devising an effective pre-breeding programme.
Materials and methods
Plant materials
The present study was conducted on a panel of 119 diverse Vigna accessions (acc.) including 99 wild Vigna accessions belonging to 19 different species, 9 released cultivars of mungbean, 4 of blackgram, 2 cultivated accessions each of Phaseolus vulgaris, V. unguiculata ssp. sequipedalis and V. umbellata and 1 accession of V. unguiculata. The wild accessions were collected from diversity rich hotspots of Western Ghats, Himalayan region, Central pleateau, and North-Eastern regions of India (Table 1). All the accessions were grown in cemented pots of 1 m diameter during Kharif (monsoon) and Spring/Summer season of 2017–2018 and 2018–2019 at the Main Research Farm, ICAR-Indian Institute of Pulses Research, Kanpur. The recommended package of practices for growing Vigna crops in the region was followed to raise healthy plants. To counter staggered/reduced germination in wild accessions of Vigna due to their hard and waxy seed coat, seed scarification was done following Pratap et al. (2015).
Microsatellite analysis
Total genomic DNA was extracted from fresh young leaves of one plant per accession at early vegetative stage (within 10–12 days of sowing) following the CTAB method (Doyle and Doyle 1990) with minor modifications (Pratap et al. 2015). The quality of extracted DNA was analysed on 0.8% agarose gel. The quantity of DNA was determined using a Nanodrop spectrophotometer ND 1000 (Nanodrop Technologies, DE, USA). Finally, the DNA of each sample was normalized to a concentration of 20–30 ng/μl for Polymerase Chain Reaction (PCR) analysis. Initially, 384 microsatellite markers from different Vigna backgrounds viz., cowpea (Li et al. 2001), adzuki bean (Wang et al. 2004), mungbean (Kumar et al. 2002a, b; Somta et al. 2009) and common bean (Gaitan-Solis et al. 2002; Blair et al. 2003) were used to identify polymorphic markers on a panel of 20 diverse Vigna genotypes. Out of these, 300 SSR primers showed amplification and 102 primer pairs revealed allele polymorphism (Supplementary Table 1). These 102 polymorphic SSRs were used for genotyping of 119 Vigna accessions.
The PCR amplification was carried out using a 96 well Tetrad thermocycler in a reaction volume of 20 μl containing 50–60 ng template DNA, 10 mM dNTPs, 0.6 U of Taq DNA polymerase (Fermentas, Mumbai), 10X Taq buffer A (Fermentas, Mumbai) with MgCl2, and 5 pmol each of forward and reverse primers (ILS, India). PCR amplifications were performed at an initial denaturation for 5 min at 95 °C, followed by 35 cycles of denaturation for 15 s at 95˚C, primer-specific annealing for 15 s at 45–55 °C, and extension at 68–72 °C for 1 min and the final extension at 72 °C for 10 min. The PCR products were separated by horizontal gel electrophoresis on 3% agarose gel in 1X TAE buffer for 3–4 h at 80–100 Volt and stained with ethidium bromide. The gels were documented using gel documentation system (Uvitech, Cambridge). Alleles were recorded on all genotypes according to their fragment sizes (in base pairs). Rare alleles were validated by repeated microsatellite analysis.
Genotypic diversity analysis
The allelic data of 102 polymorphic SSRs were subjected to statistical analysis using GenAlEx version 6.51b2 to calculate the total number of alleles (Na), effective alleles (Ne), private alleles, Shannon information Index (I), observed heterozygosity (Ho), expected heterozygosity/genetic diversity (He), genetic differentiation indices, Pairwise population Nei genetic identity and AMOVA (analysis of molecular variance). The polymorphic information content (PIC) of each marker was calculated using the formula PIC = 1 − ∑(Pij)2 where Pij denotes the frequency of ith allele of a jth locus summed across all alleles revealed by jth locus primer in a set of 119 genotypes (Botstein et al. 1980).
To establish the ancestry relationship of the ecological and reproductive characteristics of the species on their genetic diversity, genotypic data of 102 SSR markers on 119 genotypes of 19 Vigna species were used to generate genetic distance (GD) following distance based unweighted neighbour joining (UNJ) tree using Darwin V5.4. The co-dominant allelic data of each species were run at 30,000 bootstrap to draw the phylogenetic tree. Later the phylogeny was used as the robust signal for explaining the genetic diversity of the wild relatives and to predict evolutionary history.
Population structure analysis
To determine the genetic structure and define the number of clusters (gene pools), model-based cluster analysis was done using the software STRUCTURE, version 2.3.4 (Pritchard et al. 2000). The number of presumed population (K) was set from 2 to 10 and the program was run with ten independent runs for each cluster (K) following admixture model and correlated allele frequencies. The program was run with 30,000 burn-in-period and 100,000 Markov Chain Monte Carlo iterations. The optimum number of sub-populations (k) was determined using Structure Harvester web v0.6.94 (Earl and vonHoldt 2012) with structure output files based on the adhoc criterion (Delta K) proposed by Evanno et al. (2005).
Principal coordinates analysis (PCoA)
The principal coordinate analysis (PCoA) was performed with six sub-populations along with one admixture population identified from structure analysis to find and plot the major patterns within a multivariate dataset genotyped with many SSR loci. Distance matrix was calculated following ‘Distance’ option and the outcome matrix was used as an input for PCoA analysis following Distance-Standardised method available in GenAlEx 6.5 tool.
Results
Allelic diversity
A total of 102 polymorphic SSRs were used to characterize 119 wild and cultivated accessions belonging to 19 Vigna species (Table 1). Most of the primer pairs amplified with varying allele sizes between 130 and 285 bp in wild accessions, 150 and 250 bp in cultivated Vigna species, 170 and 240 bp in Phaseolus and 130–225 bp in the large beans (Supplementary Fig. 1). All the 102 SSR markers showed different degrees of polymorphism at each locus producing a total of 1758 alleles, as the number of different alleles at each locus (Na) varied from 9 (BMD-6 and BMD-50) to 31 (CP00226) with an average of 17 alleles per locus. Maximum of 13 loci produced 15 alleles per locus followed by 9 loci each producing 16 and 17 alleles per locus. The polymorphic information content (PIC) value of SSRs ranged between 0.78 and 0.93 with an average of 0.882 (Pl see supplementary information). The maximum PIC value of 0.93 was recorded for the SSRs CEDG096A, CP00226 and BM212 followed by 0.92 for the markers PvM03, J01263, BMD-13, SSR-IAC-188, VR022 and X34. The lowest PIC value of 0.77 was recorded for SSRs BMD-6 and VR024. Among the 102 markers used, 100 SSR markers (98.04%) showed high discriminating power in all Vigna species i.e. a high PIC value of > 0.80. The number of effective alleles varied from 4 to 16 (CEDG096A). The Shannon’s information index varied from 1.683 to 3.093 and the fixation index value ranged from 0.806 to 1.0. Total 47 SSR loci revealed the fixation index value of 1.0. Heterozogosity was observed in 54 SSR loci and the observed heterozygosity ranged from 0.08 to 0.168 (CEDG176). The expected heterozygosity varied between 0.775 and 0.94.
Cluster-based genetic diversity
The molecular data generated through SSR profiling of 119 accessions at 102 loci were used to study genetic inter-relationship between the different Vigna accessions. The Unrooted neighbour joining (UNJ) clearly separated these 119 genotypes into four major clusters (cluster A–D) (Fig. 1). Among these, Cluster B was the largest with 38 (31.93%) accessions followed by cluster A with 32 (26.89%) and cluster C with 25 (21%) accessions. Cluster D was the smallest one represented by 24 (20.16%) accessions. Cluster A could be further divided into three sub-clusters viz., AI, AII and AIII with 4, 13, and 15 accessions, respectively. Sub-cluster AIII comprised of V. trilobata (3 acc.), V. stipulaceae (2 acc.), V. unguiculata (3 acc.), V. umbellata (3 acc.) and V. radiata (1 acc.) which are locally cultivated and V. glabrescence, V. aconitifolia and V. khandalensis (1 acc. each) whose cultivation status is not known. Interestingly, sub-cluster AII accommodated all released cultivars of mungbean and urdbean, irrespective of the place where they were bred while all the 4 outliers (2 cultivars each of P. vulgaris and V. unguiculata ssp. sesquipedalis) were grouped in sub-cluster AI.
Cluster-based diversity depicting the grouping of 119 accessions of 19 Vigna and 1 Phaseolus species. The numbers depict different Vigna accessions as decribed in Table 1
The clusters B, C and D were further divided into 2 sub-clusters each. Sub-cluster BI comprised of 11 accessions belonging to V. silvestris (4 acc.), V. trinervia var. bourneae (3 acc.), and V. radiata var. setulosa and V. pilosa (2 acc. each). Sub-cluster BII accommodated 27 accessions including 9 of V. mungo, 5 of V. radiata, 6 of V. radiata var. radiata, 6 of V. radiata var. sublobata and 1 accession of V. silvestris. Sub-cluster CI comprised of 14 accessions including 6 of V. trilobata, 3 of V. dalzelliana, 2 each of V. umbellata and V. vexillata and 1 accession of V. pilosa. Likewise, sub-cluster CII consisted of 11 accessions belonging to V. aconitifolia (5 acc.), V. unguiculata (1 acc.), V. umbellata (2 acc.) and V. trilobata (3 acc.). Sub-cluster DI consisted of 10 accessions belonging to V. umbellata (9 acc.) and V. trilobata (1 acc.). Sub-cluster DII consisted of 14 accessions including 7 of V. trilobata, 4 of V. hainiana, 2 of V. aconitifolia and 1 accession of V. trinervia. All accessions of V. hainiana grouped together in cluster DII.
Interestingly, all the wild accessions belonging to V. radiata, V. radiata var. radiata, V. sublobata, V. mungo, V. silvestris, V. setulosa, V. pilosa and V. trinervia var. bourneae were grouped in cluster B except that one accession each of V. pilosa (IC210580) and V. trinervia (JAP/10-51) shared cluster C and D, respectively. The accessions of V. umbellata, V. aconitifolia, and V. trilobata were highly variable and were found to be distributed in all the three major clusters namely, A, C and D.
Population genetic structure
Cluster diversity study singularized two accessions each of P. vulgaris and V. unguiculata ssp. sesquipedalis in a separate sub cluster (AIII). Furthermore, one accession of V. unguiculata (Goa Cowpea 3) in AIII sub-cluster was recorded to have comparatively longer pods, bold seeds and higher 100-seed weight as compared to all other accessions of different Vigna species under study. These five accessions stood apart at molecular as well as morphological level and were considered as outliers. Therefore, these were removed from the panel for the further study. To determine accurate and reliable population genetic structure, a model-based population structure analysis was used to detect the ancestral and hybrid forms within 114 accessions of Vigna. All 114 accessions were categorized into 6 genetically distinct sub-populations (K = 6) (Fig. 3) following admixture-model based simulation (Fig. 2) with varying levels of admixture. In total, 91 (79.82%) accessions resembled their hierarchy and 23 (20.18%) accessions were observed as the admixture forms. Maximum number of accessions (25) were grouped in sub-population (SP) 6 and the minimum number of accessions were grouped in SP3 and SP5 (11 each).
SP1 comprised of 17 (14.91%) accessions including 7 of V. umbellata, 5 of V. trilobata, 4 of V. hainiana, and 1 accession of V. aconitifolia. These 17 accessions were altogether clustered in the major cluster D and the remaining 7 accessions of the major cluster D were grouped in the admixture class. Sub population 3 (SP3) comprised of 11 (9.65%) accessions which belonged to V. umbellata (3 acc.), V. trilobata (3 acc.), V. stipulaceae (2 acc.), V. unguiculata (2 acc.) and V. radiata (1 acc.). Most of the semi-cultivated accessions belonging to V. radiata (Mung Seed 1), V. stipulaceae (Trichy Local-1 and Trichy Local-2), and V. trilobata (Trichy Local and Kumar Local) were grouped in SP3 in the population structure analysis and clustered in AIII as per UNJ tree. Further, the model based ancestry synteny corresponded well with the cluster diversity in case of cultivated Vigna accessions which comprised of released cultivars of mungbean and blackgram and these were assigned to SP 4 without any admixture (Table 2). Interestingly, all these genotypes clustered only in sub-cluster AII, falling in Cluster A specifically as per UNJ tree.
The SP2 comprised of 14 (12.29%) accessions belonging to V. trilobata (5 acc.), V. dalzelliana (3 acc.), and 2 accessions each of V. pilosa, V. umbellata and V. vexillata. Similarly, SP5 consisted of 11 (9.65%) accessions including 5 of V. aconitifolia, 3 of V. trilobata, 2 of V. umbellata, and 1 of V. unguiculata. 13 accessions of SP2 and 11 accessions of SP5 represented cluster CI and CII in UNJ tree, respectively. The exception was only one accession, IC210576, of V. pilosa which was grouped in SP2 and categorised in cluster C of UNJ tree. On contrary, LRM/13-32 of V. trilobata belonging to cluster C as identified from UNJ tree was grouped in admixture class in the model-based study. Interestingly, 10 out of 11 accessions grouped in cluster BI (except IC210576 of V. pilosa) were grouped in the admixture class. The 25 accessions including 9 accessions of V. mungo, 5 accessions each of V. radiata, V. radiata var. radiata, and V. sublobata, and 1 accession of V. silvestris shared single sub-population (SP6). Importantly, these genotypes represent the primary and secondary gene pool (GP 1 and II) of Vigna species and clustered majorly in BII as identified from UNJ tree. The accession IC253920 of V. radiata var. sublobata and IC251431 of V. radiata var. radiata clustered in BII were grouped as admixture class in model-based analysis.
The accessions which recorded likelihood thresholds lower than 0.70 were considered as admixture forms. A total of 23 (19.33%) accessions including 4 accessions each of V. trilobata and V. silvestris, 3 of V. trinervia var. baourneae, 2 each of V. umbellata, V. radiata var. setulosa, V. aconitifolia, and 1 each of V. pilosa, V. sublobata, V. trinervia, V. radiata var. radiata, V. glabrescence and V. khandalensis were representing two or more ancestories of different sub-populations and hence were considered as the admixture class. All these accessions with an admixture status were clustered in AIII (3 acc.), BI (10 acc.), BII (2 acc.), CI (1 acc.), DI (3 acc.) and DII (4 acc.).
Additionally, the Vigna species viz., V. umbellata (16 acc.), and V. trilobata (20 acc.) were observed as highly variable and these were distributed in 4 different sup-populations namely SP1, 2, 3 and 5 as well as in the admixture class (Table 2). The same holds true in cluster analysis where these clustered in A, C and D of UNJ tree. All accessions of V. hainiana, 43.75% of V. umbellata, 25% of V. trilobata and 12.5% accessions of V. aconitifolia grouped in SP1.
Genetic diversity within structured Vigna populations
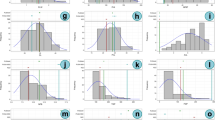
The number of alleles in six populations along with the admixture class varied from 4.9 (SP4) to 9.8 (admixture class). The mean expected heterozygosity (mHe) was quite high in each population which varied from 0.67 (SP4) to 0.825 (admixture group). The mean Shannon’s information index value varied from 1.34 (SP) to 2 (admixture group). All loci revealed polymorphism in accessions belonging to all populations except in SP4 where one locus (GMES0211) did not reveal polymorphism across the accessions (Table 3). All 114 accessions recorded private alleles which varied from 1 (IC331456 at locus CEDG036; IC251431 at locus CP00226; IC277021 at locus VR011) to 16 (Pant U39 at loci PV-ag005, BMD-55, X49, VR016, VR048, CEDG291, CEDG220, CEDC139, CEDG185, CEDC033, DMBSSR001, CEDG225, CEDG073, VM27, BM212 and BM149) (Supplementary Table 2,). Population-wise, the maximum number of private alleles were observed for SP6 (76), followed by SP4 (75), SP3 (71), SP1 (67), SP5 (48) and SP2 (44) while the admixture population group recorded 80 private alleles (Supplementary Table 3). The Pair-wise population Nei genetic identity value ranged from 0.26 (SP4 vs SP5) to 0.624 (SP1 vs SP7). All the 6 populations alongwith the admixture class recorded mostly low (0.36; SP4) to moderate (0.62; SP1) similarity values (Table 4).
The genetic differentiation indices between populations (Fst) were observed as moderate to high which varied from 0.048 (between SP6 and SP7) to 0.132 (between SP4 vs SP5). Accessions of SP4 had high differentiation indices with rest of the populations including the admixture class. Similarly, admixture class had moderate Fst values (0.046 to 0.071) between rest of the populations (Table 5). An analysis of the molecular variance (AMOVA) was performed using raw data for genetic differentiations (Table 6). The overall genetic variation was divided among population (9%), among individuals within populations (88%), and within individuals (2%). The diversity within individuals of a population was greater than the diversity between the populations. The observed Fst value was 0.094, suggesting moderate differentiation of Vigna sub-populations.
On the basis of principal coordinates analysis (PCoA), it was observed that Population 1 comprising of the accessions of V. umbellata, V. trilobata V. hainiana, and V. aconitifolia clustered in two groups. The population 2 comprising of the accessions belonging to V. trilobata, V. dalzelliana, V. pilosa, V. umbellata and V. vexillata were scattered as population 1. Similarly, the accessions V. aconitifolia, V. trilobata, V. umbellata, and V. unguiculata belonging to population five were also scattered. On contrary, the population three comprising of accessions belonging to V. umbellata, V. trilobata, V. stipulaceae, V. unguiculata and V. radiata were clustered together except Mung seed-1 and TCR279. Similarly the released varieties of Vigna representing population four were uniquely clustered without any admixture as also observed in structure analysis. Population six having the accessions from primary and secondary gene-pool of Vigna viz., V. mungo, V. radiata, V. radiata var. radiata, V. sublobata and V. silvestris were clustered together except IC251426A, IC251425, IC251387 and IC251390. As expected, accessions belonging to admixture groups were scattered all around (Supplementary Fig. 2).
Discussion
The genus Vigna is large, genetically variable and pan-tropic in distribution having a considerable agronomic and environmental significance. Many of the Vigna species viz., mungbean (V. radiata), urdbean (V. mungo), moth bean (V. aconitifolia) and ricebean (V. umbellata) have their centre of origin as well as diversity in India (de Candolle 1884; Vavilov 1926; Zukovskij 1962; Smartt 1985; Pratap and Kumar 2011) and their wild and cultivated forms are found variably distributed across the Himalayan region, central plateau, western ghats and the north-eastern hill regions. Nonetheless, the Asiatic Vignas are considered to be recently evolved and hence differentiation of taxa and sub-specific classification using morphological marker is limited and more complex (Baudoin and Marechal 1988). Therefore, classifying this species using highly abundant molecular markers such as multi-allelic SSRs is indeed necessary and has been used to some extent in classifying wild species earlier (Sarr et al. 2020; Kempf et al. 2016; Wang et al. 2008; Gwag et al. 2010; Pratap et al. 2015).
Assessment of genetic diversity helps in identifying the appropriate donors in a pre-breeding activities and breeding programme. The 102 SSRs used in this study revealed high degree of polymorphism by amplifying 1758 alleles from 119 accessions belonging to 19 Vigna species with the number of alleles varying between 9 and 31 and the effective number of alleles varying from 4.50 to 15.66. Number of alleles per locus detected in this study is comparatively higher than the earlier reports. Pratap et al. (2015) reported 4–16 alleles per locus in a set of 53 Asiatic Vigna accessions including 41 wild accessions belonging to 13 species and 12 commercial cultivars of mungbean, blackgram and rice bean using 53 SSRs. In other similar studies, 6–20 alleles with an average of 13.65 alleles per locus in 422 wild and cultivated genotypes of zombi pea (V. vexillata (L.) A. Rich) (Dachapak et al. 2017), 2 to 15 alleles with an average of 6.2 alleles per marker locus in 737 samples of cowpea (V. unguiculata (L.) Walp.) (Sarr et al. 2020), and 8–26 alleles per locus in 127 genotypes of mungbean (Singh et al. 2020) have been reported. A higher number of alleles per locus detected in the present study could be primarily attributed to use of a large number of highly diverse accessions of 19 Vigna species. These accessions have been collected from diversity rich endemic hot spots of India which have been reported to be centre of origin/diversity for many of the Vigna species (Sangiri et al. 2007; de Candolle 1884; Vavilov 1926; Zukovskij 1962). Additionally this could also be attributed to the deployment of a high number of polymorphic SSRs than the earlier studies (102 SSRs in the present study against 20–53 SSRs in earlier studies). Higher estimates of polymorphism (PIC: 0.78–0.93), Shannon information index (1.683–3.93), number of effective alleles (4–16) and value of expected heterozygosity (0.775–0.94) as compared to the previous studies (Wang et al. 2008; Dachapak et al. 2017; Singh et al. 2020; Pratap et al. 2015; Sarr et al. 2020) also indicates high genetic diversity among the accessions studied. This finding also suggests high usefulness of the material under study for generating additional genetic variability in different Vigna crops. A few marker loci (CEDG176, BMD-26, X87 and VR011) revealed higher heterozygosity and, therefore, could be deployed in studying the hybridity of wild relatives. Therefore, it is evident that the SSRs used in this experiment have a great potential for germplasm characterisation and identifying trait-linked markers that could be eventually utilised in marker-assisted breeding programme.
Cluster diversity
All the cultivated Vigna accessions clustered in a single sub cluster-AII, whereas the outliers such as Amber and Utkarsh belonging to the genus Phaseolus, and 2 accessions of the large beans (V. unguiculata ssp. sesquipedalis) separated in sub cluster-AI. At morphological level also these accessions were observed to have comparatively longer pods, bold seeds and high 100-seed weight as compared to all other accessions under study. Phaseolus is highly diverse from Vigna species although it shares high genetic similarity with V. unguiculata, justifying their clubbing together in sub cluster AI. Nonetheless, P. vulgaris is related with the Vigna species in the context of having the same chromosome number with almost similar genome sizes. Vasconcelos et al. (2015) reported numerous breaks of macrosynteny between P. vulgaris and V. unguiculata which could be due to larger sequence divergence between them. V. glabrescence and V. khandalensis are not in cultivation although they clustered with the semi-cultivated group in AIII sub-cluster. Earlier, V. glabrescence had been reported to be highly cross compatible with mungbean and urd bean (Dana 1968; Krishnan and De 1968; Chen et al. 1989) which indicates that it is genetically very similar to the cultivated species. V. khandalensis and V. stipulacea, belonging to the section Aconitifoliae are closely related to each other and cross compatible with mungbean (Takahashi et al. 2016; Chavan et al. 1966).
The progenitors of mungbean and urdbean namely V. radiata var. sublobata and V. mungo var. silvestris, characterized in Gene pool I, also clustered together in cluster BII as expected along with V. mungo and V. radiata. The accessions of V. silvestris and V. radiata var. setulosa belonging to secondary gene pool of mungbean and urdbean, respectively clustered with V. pilosa (2 acc.), and V. trinervia var. bourneae (3 acc.) in Sub-cluster BI. In this subcluster, all accessions belonged to diploid Vigna species except V. pilosa which is a tetraploid (2n = 4x = 44) (Chankaew et al. 2014).
Sub-cluster CI represented the accessions belonging to secondary (V. trilobata) and tertiary gene-pools (V. dalzelliana, V. umbellata, and V. vexillata) along with one accession of V. pilosa. The geographical distribution of V. dalzelliana (O. Kuntze) Verdcourt was limited to India, especially the Andaman Islands, and Sri Lanka (John et al. 2009; Tomooka et al. 2002) and it could be one of the ancestral species of the section Angulares. V. vexillata (L.) A. Rich, is reported to be widely distributed in pantropical regions, including Africa, Asia, Oceania, and America while V. vexillata and Cowpea (V. unguiculata) were observed to be relatively closer at the molecular level (Sonnante et al. 1996) and also cross-compatible to produce an interspecific hybrid (Gomathinayagam et al. 1998). Sub-cluster CII had an array of accessions belonging to the secondary (V. aconitifolia and V. trilobata) and tertiary (V. umbellata) gene-pools along with accessions of unknown gene pool (V. unguiculata). Most of the accessions representing major cluster-II were collected from peninsular region (Western Ghats) of India.
The sub-cluster DI consisted 90% of the accessions belonging to the tertiary gene-pool. Sub-cluster DII consisted of 9 accessions representing secondary gene pool (7 of V. trilobata, and 2 of V. aconitifolia) along with 4 accessions of V. hainiana and 1 of V. trinervia. Pratap et al. (2015) reported that two accessions of V. hainiana (IC251381, IC251376) clustered alongwith V. umbellata and V. glabrescens while the remaining two accessions (IC331448; IC331450) clustered with V. trilobata. This could be due to less number of polymorphic markers employed by them in a smaller panel of genotypes. The accessions of V. umbellata, V. aconitifolia, and V. trilobata were highly variable and distributed in all the three major clusters namely A, C and D. Among these, mothbean (V. aconitifolia) is considered as one of the most primitive Vigna crop in respect of its evolution (Smartt 1985) and its wild ancestor primarily occur in south-eastern India (Arora et al. 1984). It is reported to be cross compatible with mungbean (Pandiyan et al. 2010) and serves as a source for drought and heat tolerance in the subgenus Ceratotropis.
The observed clustering pattern of the accessions belonging to different Vigna species clearly showed their closeness with the corresponding gene-pools. Nonetheless a few accessions from diverse Vigna species also clustered together indicating that there could be a possibility of taxonomic misclassification of a few accessions based only on phenotypic observations which might have led to their assignment to an incorrect species. This could also have occurred during extensive germplasm exchange. It may also be possible that some of these Vigna species might have been genetically close to other species leading to spontaneous hybridization and subsequent exchange of genetic materials to produce new species through the process of natural selection without compromising much with their original identity. Moreover, since most of the Vigna species are diploid in nature that could be the cause for less genetic divergence among species of interspecific hybrids than the allopolyploid hybrid species (Chapman and Burke 2007). Species that co-occur in the same geographical location have a better chance for interspecific gene flow (Abbott et al. 2008). Moreover ecotypic variation is also one of the important steps in speciation where some degree of reproductive isolation is maintained for the transition of ecotypes to species. Therefore, cross-compatibility studies and a combination of phenotype- and genotype-based classification of the wild accessions must be resorted to establish the closeness of these species with their corresponding clusters.
Population genetic structure
Model-based clustering is tremendously useful to visualise the genetic ancestry and assigning individuals to a defined population in plants, animals as well as humans (Pritchard et al. 2000). In the present study, Bayesian algorithm with ADMIXTURE model divided the 114 Vigna accessions including 13 released cultivars into 6 genetically distinct sub-populations with a few admixtures. The results obtained from both DARWIN and STRUCTURE indicated that the grouping of all accessions did not strictly follow their geographical distribution as well as their species lineage. This could be attributed to several reasons including species misclassification, spontaneous hybridization and mutation among the accessions. Spontaneous hybridization may lead to development of new forms hitherto not found in nature resulting in the process of speciation. Likewise, inter-specific outcrossing, although difficult to notice and characterize, is also expected to occur in plants naturally and even a single backcross may develop plants which are morphologically similar to the species with which they were backcrossed (Anderson 1948).
All accessions of V. radiata and V. mungo and their progenitors viz., V. sublobata and V. silvestris were categorized together in one group (SP 6 and sub-cluster BII). Similar findings have been reported earlier by Chandel and Laster (1991), Dana and Karmakar (1990), Kumar et al. (2004) and Pandiyan et al. (2010) based upon morphological observations. More interestingly, all the released cultivars of mungbean and blackgram developed at quite distinct geographical locations of Kanpur, Pantnagar, Hisar (north India) and Vamban (south India) were assigned to the same sub-cluster AII in UNJ tree and likewise in SP 4, as also reported in earlier studies (Singh et al. 2020; Pratap et al. 2015). This indicates the inheritance of similar genetic sequences from common lineage/parenatge across these cultivars. Their grouping in the same population group may also be attributed to involvement of common ancestors in their past history (Pratap et al. 2015).
V. hainiana was grouped distinctively with V. umbellata and V. trilobata despite showing phenotypic similarity with wild types of V. mungo and V. radiata and also being close to V. radiata var. sublobata and V. mungo var. silvestris with respect to pod characteristics (Arora et al. 1973; Chandel et al. 1984; Bisht et al. 2005). However, V. hainiana is morphologically more primitive than V. mungo var. silvestris and V. radiata var. sublobata with comparatively small flowers and very small seeds and, therefore, could play a significant role in phylogeny and evolutionary studies particularly in the mungo-radiata complex (Bisht et al. 2005). Other Vigna species such as V. umbellata, V. dalzelliana, V. mungo var. silvestris, V. radiata var. sublobata and V. setulosa were also observed to be closely related to each other and exhibited sympatric distribution (Bisht et al. 2005).
Accessions of V. umbellata and V. trilobata were highly scattered and grouped in sub populations 1, 2 and 3 alongwith the accessions of V. hainiana, V. aconitifolia (LRM/13-36), V. dalzelliana, V. pilosa, V. vexillata, V. stipulaceae and V. unguiculata. The wild types of V. umbellata have been observed to be widely distributed than V. dalzelliana, a closely related species. Bisht et al. (2005) reported two distinct overlapping groups of V. umbellata and V. dalzelliana accessions which is also evident from molecular grouping of V. umbellata accessions. Likewise, they also documented more widespread distribution and highly diverse nature of V. trilobata accessions collected from diverse agro-ecologies of India at phenotypic level. This is more evident from clustering of accessions at molecular level in the present study. A large admixture group (23 accessions) which shared two or more ancestries indicates the possibility of historical inter- and intra-specific gene transfer and is in agreement with previous studies (Pratap et al. 2015; Sarr et al. 2020). The only accession of V. glabrescense was grouped in the admixture group as also in the earlier study (Pratap et al. 2015). It could be corroborated with the earlier observations of Dana (1964) and Bisht et al. (2005) that V. glabrescens is probably an amphidiploid combining the genomes of V. radiata and V. umbellata. However, Goel et al. (2001) reported that V. glabrescens is a derivative from V. umbellata and V. angularis based on the variation observed in rDNA sequences. The accessions of V. umbellata and V. trilobata used in this study were collected from several diverse locations including the Himalayan region, Western Ghats, Southern Eastern India, North-East region and Central pleateu. Going into the history, it could be suggested that probably their seeds were dispersed historically by monks, merchants and villagers because of their movement from one place to another. In due course of time, adaptation and acclimatization in new environment might have happened accompanied by inter- and intra-specific recombination and natural selection. This might also have happened due to genetic drift, domestication, mutation and background selection (Sangiri et al. 2007).
Genetic diversity in structured Vigna populations
The mean expected heterozygosity (mHe) was observed to be quite high in each sub-population which varied from 0.67 to 0.825 and was higher than the observed heterozygosity. These estimates were higher than the previous reports in V. unguiculata, V. radiata and other Asiatic Vigna species (Sarr et al. 2020; Badiane et al. 2012; Pratap et al. 2015). This could be attributed to the fact that India is the centre of origin as well as diversity for many of the Vigna species.
AMOVA showed higher genetic diversity (88.33%) among individuals within the populations and it could be primarily due to representation of diverse Vigna species in each population. On the other hand, the low genetic diversity between the populations could be due to the distribution of the similar Vigna species in each population.
The genetic differentiation indices observed between populations were moderate to high and the SP4 was observed to have high differentiation indices with rest of the populations. Noticeably, this sub-population consists mainly of released cultivars of mungbean and urdbean and these cultivars were also distinctively clustered in sub-cluster AII. The highest genetic differentiation indices of 0.132 were observed between SP4 and SP5 followed by 0.12 between SP 4 and SP 6. The SP5 consisted of accessions belonging to V. aconitifolia, V. trilobata, V. umbellata and V. ungiculata and these accessions/species are quite diverse from released cultivars of mungbean and urdbean representing SP4. Similarly SP6 comprised of V. mungo var. mungo, V. radiata var. radiata and their respective wild ancestors. The low genetic differentiation indices (0.077) between SP1 and SP2 could be due to the fact that both the populations shared majority of accessions belonging to V. trilobata and V. umbellata. Similarly the low to moderate genetic identity observed between different populations was attributed to the representation of accessions belonging to either released cultivars (SP4) or close wild relatives of each species (SP6).
Conclusion
In summary, the 102 SSR markers used in this study successfully detected a high amount of genetic variability in 119 wild and cultivated accessions belonging to 19 different species of Vigna. Keeping in view that a large set of wild Vigna accessions was deployed in this study, the genotypic phylogenetic data provided an insight into the diversity status and interrelationship of different species which will be of immense use in their utilization in introgression breeding and the subsequent improvement of the cultivated types. This study also suggested that India is most likely the primary centre of origin and centre of diversity of many of the Vigna species. Simultaneously, this study also underlined the importance to generate more genotypic as well as phenotypic data to clearly distinguish different Vigna species and unleash their genetic potential for improvement of cultivated types.
References
Abbott RJ, Ritchie MG, Hollingsworth PM (2008) Introduction. Speciation in plants and animals: pattern and process. Phil Trans R Soc B 363:2965–2969
Aidbhavi R, Pratap A, Verma P, Lamichaney A, Bandi SM, Nitesh SD, Akram M, Rathore M, Singh B, Singh NP (2021) Screening of endemic wild Vigna accessions for resistance to three bruchid species. J Stored Prod Res 93:101864
Anderson E (1948) Hybridization of the habitat. Evolution 2:1–9
Arora RK, Nayar ER (1984) Wild relatives of crop plants in India NBPGR Sci. Monograph No. 7. National Bureau of Plant Genetic Resources, New Delhi, India
Arora RK, Chandel KPS, Joshi BS (1973) Morphological diversity in Phaseolus sublobatus Roxb. Curr Sci 42:359–361
Badiane FA, Gowda BS, Cissé N, Diouf D, Sadio O, Timko MP (2012) Genetic relationship of cowpea (Vigna unguiculata) varieties from Senegal based on SSR markers. Genet Mol Res 11:292–304. https://doi.org/10.4238/2012
Basu PS, Pratap A, Gupta S, Sharma K, Tomar R, Singh NP (2019) Physiological traits for shortening crop duration and improving productivity of greengram (V. radiata (L.) Wilczek) under high temperature. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01508
Baudoin JP, Marechal R (1988) Taxonomy and evolution of the genus Vigna. In: Shanmugasundaram S, McLean BT (eds) Mungbean: Proceedings of the Second International Symposium. Taiwan: Asian Vegetable Research and Development Center, pp 2–12
Bisht IS, Bhat KV, Lakhanpaul S, Latha M, Jayan PK, Biswas BK, Singh AK (2005) Diversity and genetic resources of wild Vigna species in India. Genet Res Crop Evol 52:53–68
Blair MW, Pedraza F, Buendia HF, Gaitan-Solis E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L). Theor Appl Genet 107:1362–1374
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Chandel KPS, Laster RN (1991) Origin and evolution of Asiatic Vigna species. In: Sharma B, Mehra RB (eds) Golden jubilee celebration symposium on grain legumes. IARI, New Delhi, India, pp 25–45
Chandel KPS, Lester RN, Starling RJ (1984) The wild ancestors of urd and mungbeans [Vigna mungo (L.) Hepper and V. radiata (L.) Wilczek]. Bot J Linn Soc 89:85–96
Chankaew S, Somta P, Isemura T, Tomooka N, Kaga A, Vaughan DA, Srinives P (2013) Quantitative trait locus mapping reveals conservation of major and minor loci for powdery mildew resistance in four sources of resistance in mungbean [Vigna radiata (L.) Wilczek]. Mol Breed 32(1):121–130
Chankaew S, Isemura T, Isobe S, Kaga A, Tomooka N, Somta P, Hirakawa H, Shirasawa K, Vaughan DA, Srinives P (2014) Detection of genome donor species of neglected tetraploid crop Vigna reflexo-pilosa (creole bean) and genetic structure of diploid species based on newly developed EST-SSR markers from Azuki bean (Vigna angularis). PLoS ONE 9(8):e104990
Chapman MA, Burke JM (2007) Genetic divergence and hybrid speciation. Evolution 61:1773–1780. https://doi.org/10.1111/j.1558-5646.2007.00134.x
Chavan VM, Patil GD, Bhapkar DG (1966) Improvement of cultivated Phaseolus species-need for interspecific hybridization. Indian J Genet Plant Breed 26:152–154
Chen HK, Mok MC, Shanmugasundaram S, Mok DWS (1989) Interspecific hybridization between Vigna radiata (L.) Wilczek and V. glabrescens. Theor Appl Genet 78:641–647
Dachapak S, Somta P, Poonchaivilaisak S, Yimram T, Srinives P (2017) Genetic diversity and structure of the zombi pea (Vigna vexillata (L.) A. Rich) gene pool based on SSR marker analysis. Genetica 145:189–200. https://doi.org/10.1007/s10709-017-9957-y
Dana S (1964) Interspecific cross between tetraploid Phaseolus species and P. ricciardianus. Nucleus 7:1–10
Dana S (1968) Hybrid between Phaseolus mungo and tetraploid Phaseolus species. Japan J Genet 43:153–155
Dana S, Karmakar PG (1990) Species relation in Vigna subgenus Ceratotropis and its implications in breeding. Plant Breed Rev 8:19–42
de Candolle A (1884) Origin of cultivated plants. Hafner, New York
Douglas C, Pratap A, HanumanthaRao B, Manu B, Dubey S, Singh P, Tomar R (2020) Breeding progress and future challenges: abiotic stresses. In. Nair RM et al (eds) The mungbean genome, compendium of plant genomes. https://doi.org/10.1007/978-3-030-20008-4_6
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Gaitan-Solis E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in ommon bean (Phaseolus vulgaris): isolation, characterization and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136
Goel S, Raina SN, Ogihara Y (2001) Molecular evolution and phylogenetic implications of internal transcribed spacer sequences of nuclear ribosomal DNA in the Phaseolus-Vigna complex. Mol Phylogenet Evol 1037:1–19
Gomathinayagam P, Ram SG, Rathnaswamy R, Ramaswamy NM (1998) Interspecific hybridization between Vigna unguiculata and V. vexillata through in vitro embryo culture. Euphytica 102:203–209
Gore PG, Tripathi K, Pratap A, Bhat KV, Umdale SD, Gupta V, Pandey A (2019) Delineating taxonomic identity of two closely related Vigna species of section Aconitifoliae: V. trilobata (L.) Verdc. and V. stipulacea (Lam.) Kuntz in India. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-019-00767-9
Gwag JG, Dixit A, Park YJ, Ma KH, Kwon SJ, Cho GT, Lee GA, Lee SY, Kang HK, Lee SH (2010) Assessment of genetic diversity and population structure in mungbean. Genes Genomics 32:299–308
HanumanthaRao B, Nair RM, Nayyar H (2016) Salinity and high temperature tolerance in Mungbean [Vigna radiata (L.) Wilczek] from a physiological perspective. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00957
Harlan JR (1971) Agricultural origins: centers and noncenters. Science 174:468–474
Isemura T, Kaga A, Tabata S, Somta P, Srinives P, Shimizu T, Jo U, Vaughan DA, Tomooka N (2012) Construction of a genetic linkage map and genetic analysis of domestication related traits in mungbean. PLoS ONE 7:41304. https://doi.org/10.1371/journal.pone.0041304
John KJ, Latha M, Senthil KR, Asokan NR, Abraham Z, Mishra SK (2009) Vigna dalzelliana (O. Kuntz) Verdc: a new distributional record from Andaman Islands, India. Indian J Plant Genet Resour 22:138–140
Kaewwongwal A, Chen J, Somta P, Kongjaimun A, Yimram T, Chen X, Srinives P (2017) Novel alleles of two tightly linked genes encoding polygalacturonase-inhibiting proteins (VrPGIP1 and VrPGIP2) associated with the Br locus that Confer Bruchid (Callosobruchus spp) resistance to Mungbean (Vigna radiata) accession V2709. Front Plant Sci 8:1692. https://doi.org/10.3389/fpls.2017.01692
Kajonphol T, Sangsiri C, Somta P, Toojinda T, Srinives P (2012) SSR map construction and quantitative trait loci (QTL) identification of major agronomic traits in mung bean (Vigna radiata (L.) Wilczek). SABRAO J Breed Genet 44:71–86. https://doi.org/10.1556/019.70.2019.09
Kempf K, Mora-Ortiz M, Smith LM, Kölliker R, Skøt L (2016) Characterization of novel SSR markers in diverse sainfoin (Onobrychis viciifolia) germplasm. BMC Genet 17(1):1–4
Kitsanachandee R, Somta P, Chatchawankanphanich O, Akhtar KP, Shah TM, Nair RM, Bains TS, Sirari A, Kaur L, Srinives P (2013) Detection of quantitative trait loci for mungbean yellow mosaic India virus resistance in mungbean in India and Pakistan. Breed Sci 63:367–373
Krishnan R, De DN (1968) Cytogenetical studies in Phaseolus II. Phaseolus mungo × tetraploid phaseolus species and the amphidiploid. Indian J Genet Plant Breed 28:23–30
Kumar SV, Tan SG, Quah SC, Yusoff K (2002a) Isolation of microsatellite markers in mungbean, Vigna radiata. Mol Ecol Notes 2:96–98
Kumar SV, Tan SG, Quah SC, Yusoff K (2002b) Isolation and characterization of seven tetranucleotide microsatellite loci in mungbean, Vigna radiata. Mol Ecol Notes 2:293–295
Kumar S, Gupta S, Chandra S, Singh BB (2004) How wide is the genetic base of pulse crops? In: Ali M, Singh BB, Kumar S, Dhar V (eds) Pulses in new perspective. Indian Society of Pulses Research and Development, Kanpur, pp 211–221
Li CD, Fatokun CA, Ubi B, Singh BB, Scoles GJ (2001) Determining genetic similarities and relationships among cowpea breeding lines and cultivars by microsatellite markers. Crop Sci 41:189–197
Nair RM, Pandey AK, War AR, Bindumadhava H, Shwe T, Alam AKMM, Pratap A, Malik SR, Karimi R, Mbeyagala EK, Douglas CA, Rane J, Schafleitener R (2019) Biotic and abiotic constraints in mungbean production—progress in genetic improvement. Front Plant Sci 10:1340. https://doi.org/10.3389/fpls.2019.01340
Palmgren MG, Edenbrandt AK, Vedel SE, Andersen MM, Landes X, Østerberg JT, Falhof J, Olsen LI, Christensen SB, Sandøe P, Gamborg C (2015) Are we ready for back-to-nature crop breeding? Trends Plant Sci 20(3):155–164
Pandiyan M, Senthil N, Ramamoorthi N, Muthiah AR, Tomooka N, Duncan V et al (2010) (2010) Interspecific hybridization of Vigna radiata × 13 wild Vigna species for developing MYMV donor. Electron J Plant Breed 1:600–610
Parker PG, Snow AA, Schug MD, Booton GC, Fuerst PA (1998) What molecules can tell us about populations: choosing and using a molecular marker. Ecology 79:361–382
Pratap A, Kumar J (2011) History origin and evolution. In: Pratap A, Kumar J (eds) Biology and breeding of food legumes. CABI, Oxfordshire, p 432
Pratap A, Malviya N, Tomar R, Gupta DS, Kumar J (2014a) Vigna. In: Pratap A, Kumar J (eds) Alien gene transfer in crop plants, vol 2. Achievements and impacts. Springer Business+ Science Media, New York, pp 163–190
Pratap A, Basu PS, Gupta S, Malviya N, Rajan N, Tomar R, Latha M, Nadarajan N, Singh NP (2014b) Identification and characterization of sources for photo- and thermo-insensitivity in Vigna species. Plant Breeding 133:756–764
Pratap A, Gupta S, Malviya N, Rajan N, Tomar R, Latha M, John JK, Singh NP (2015) Genome scanning of asiatic Vigna species for discerning population genetic structure based on microsatellite variation. Mol Breed 35:178
Pratap A, Chaturvedi SK, Tomar R, Rajan N, Malviya N, Thudi M, Saabale PR, Prajapati U, Varshney RK, Singh NP (2017) Marker-assisted introgression of resistance to fusarium wilt race 2 in Pusa 256, an elite cultivar of desi chickpea. Mol Genet Genomics 292:137–1245. https://doi.org/10.1007/s00438-017-1343-z
Pratap A, Douglas C, Prajapati U, Kumari G, Was AR, Tomar R, Pandey AK, Dubey S (2020) Breeding progress and future challenges: Biotic stresses. In. Nair RM et al (eds) The Mungbean genome, compendium of plant genomes. https://doi.org/10.1007/978-3-030-20008-4_5
Pratap A, Gupta S, Rathore M, Basavaraja T, Singh CM, Prajapati U, Singh P, Singh Y, Kumari G (2021a) Mungbean. In: Pratap A, Gupta S (eds) The beans and the peas: from orphan to mainstream crops. Elsvier, Duxdorf, pp 1–32
Pratap A, Das A, Kumar S, Gupta S (2021b) Current perspectives on introgression breeding in food legumes. Front Plant Sci. https://doi.org/10.3389/fpls.2020.589189
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rawal KM (1975) Natural hybridization among wild, weedy and cultivated Vigna unguiculata L. Walp. Euphytica 24:699–707
Sangiri C, Kaga A, Tomooka N (2007) Genetic diversity of the mungbean (Vigna radiata) gene pool on the basis of microsatellite analysis. Aust J Bot 55:837–847
Sarr A, Bodian A, Gbedevi KM et al (2020) Genetic Diversity and population structure analyses of wild relatives and cultivated cowpea (Vigna unguiculata (L.) Walp.) from senegal using simple sequence repeat markers. Plant Mol Biol Rep. https://doi.org/10.1007/s11105-020-01232-z
Singh DP, Singh BB, Pratap A (2017) Genetic improvement of mungbean and urdbean and their role in enhancing pulse production in India. Indian J Genet Plant Breed 76:550–567
Singh B, Das A, Parihar AK, Bhagawati B, Singh D, Pathak KN, Dwivedi K, Das N, Keshari N, Midha RL, Kumar R, Pratap A, Kumar V, Gupta S (2020) Delineation of Genotype-by-Environment interactions for identification and validation of resistant genotypes in root-knot nematode (Meloidogyne incognita) using GGE biplot. Sci Rep 10(1):4108. https://doi.org/10.1038/s41598-020-60820-x
Smartt J (1985) Evolution of grain legumes. III. Pulses in the genus Vigna. Exp Agric 21:87–100
Soares BM, Ferreira PAA, de Oliveira-Longatti SM, Marra LM, Rufini M, de Andrade MJB, de Souza-Moreira FM (2014) Cowpea symbiotic efficiency, pH and aluminium tolerance in nitrogen-fixing bacteria. Sci Agric 71:171–180
Somta P, Seehalak W, Srinives P (2009) Development, characterization and cross-species amplification of mungbean (Vigna radiata) genic microsatellite markers. Conserv Genet 10:1939–1943
Sonnante G, Piergiovanni AR, Ng QN, Perrino P (1996) Relationships of Vigna unguiculata (L.) Walp., V. vexillata (L.) A. Rich. and species of section Vigna based on isozyme variation. Genet Resour Crop Evol 43:157–165
Takahashi Y, Somta P, Muto C, Iseki K, Naito K, Pandiyan M et al (2016) Novel genetic resources in the genus Vigna unveiled from gene bank accessions. PLoS ONE 11(1):e0147568. https://doi.org/10.1371/journal.pone.0147568
Tomooka N (2009) The origin of rice bean (Vigna umbellata) and azuki bean (V. angularis): the evolution of two lesser-known Asian beans. In: Akimiti T (ed) An illustrated eco-history of the Mekong River Basin. White Lotus Co, Bangkok
Tomooka N, Vaughan DA, Moss H, Maxted N (2002) The Asian Vigna: Genus Vigna subgenus Ceratotropis genetic resources. Kluwer Academic Publishers
Tomooka N, Naito K, Kaga A, Sakai H, Isemura T, Ogiso-Tanaka E, Iseki K, Takahashi Y (2014) Evolution, domestication and neo-domestication of the genus Vigna. Plant Genet Resour 2(S1):S168–S171
Varshney RK, Mohan SM, Gaur PM, Chamarthi SK, Singh VK, Srinivasan S, Swapna N, Sharma M, Pande S, Singh S, Kaur L (2014) Marker-assisted backcrossing to introgress resistance to fusarium wilt race 1 and Ascochyta blight in C 214, an elite cultivar of chickpea. Plant Genome 7:1–11
Vasconcelos EV, de Andrade Fonsêca AF, Pedrosa-Harand A, de Andrade Bortoleti KC, Benko-Iseppon AM, da Costa AF, Brasileiro-Vidal AC (2015) Intra- and interchromosomal rearrangements between cowpea [Vigna unguiculata (L.) Walp.] and common bean (Phaseolus vulgaris L.) revealed by BAC-FISH. Chromosome Res 23(2):253–266. https://doi.org/10.1007/s10577-014-9464-2
Vavilov NI (1926) Studies on the origin of cultivated plants. Leningrad. 1951
Vaz Patto MC, Amarowicz R, Aryee AN, Boye JI, Chung HJ, Martín-Cabrejas MA, Domoney C (2015) Achievements and challenges in improving the nutritional quality of food legumes. Crit Rev Plant Sci 134:105–143
Wang XW, Kaga A, Tomooka N, Vaughan DA (2004) The development of SSR markers by a new method in plants and their application to gene flow studies in azuki bean [Vigna angularis (Willd.) Ohwi & Ohashi]. Theor Appl Genet 109:352–360
Wang ML, Barkley NA, Gillaspie GA, Pederson GA (2008) Phylogenetic relationships and genetic diversity of the USDA Vigna germplasm collection revealed by gene-derived markers and sequencing. Genet Res 90:467–480. https://doi.org/10.1017/S0016672308009889
Weber JL, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am J Hum Genet 44:388
Yoshida Y, Marubodee R, Ogiso-Tanaka E et al (2016) Salt tolerance in wild relatives of adzuki bean, Vigna angularis (Willd.) Ohwi et Ohashi. Genet Resour Crop Evol 63:627–637. https://doi.org/10.1007/s10722-015-0272-0
Zukovskij PM (1962) Cultivated plants and their wild relatives. Commonwealth Agriculture Bureau, London
Funding
The authors acknowledge the funding support received from Department of Biotechnology, Government of India (Grant no. BT/Ag/Network/Pulses-I/2017-18) and the Uttar Pradesh Council of Agricultural Research, Lucknow (745/AP&AS/CROPS/RF/2014) in the form of funded research projects.
Author information
Authors and Affiliations
Contributions
GK and AP conceptualized, planned and executed the experiment. GK, SPS and AP analyzed the data and interpreted the results. GK, YS, BP and PS undertook field and laboratory experiments, LM provided seeds of some of the accessions. SG, GRP, NPS provided suggestions for improvement of the experiments and the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumari, G., Roopa Lavanya, G., Shanmugavadivel, P.S. et al. Genetic diversity and population genetic structure analysis of an extensive collection of wild and cultivated Vigna accessions. Mol Genet Genomics 296, 1337–1353 (2021). https://doi.org/10.1007/s00438-021-01825-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01825-7