Abstract
Rodents have been widely studied as intermediate hosts of Sarcocystis; however, only a few reports on these parasites in the black rat (Rattus rattus) are known. Having examined 13 black rats captured in Latvia, sarcocysts were found in skeletal muscles of two mammals and were described as Sarcocystis ratti n. sp. Under a light microscope, sarcocysts were ribbon-shaped, 0.9–1.3 × 0.09–0.14 mm in size and had a thin (0.8–1.3 μm) and smooth cyst wall. The lancet-shaped bradyzoites were 8.3 × 4.3 (7.5–9.3 × 3.9–4.8) μm. Under a transmission electron microscope, the cyst wall was up to 1.3 μm thick, wavy, the ground substance appeared smooth, type 1a-like. Morphologically, sarcocysts of S. ratti were somewhat similar to those of S. cymruensis, S. rodentifelis, and S. dispersa-like previously identified in the brown rat (Rattus norvegicus). On the basis of 18S rDNA, 28S rDNA, and cox1, significant genetic differences (at least 2.3, 4.5, and 5.8%, respectively) were observed when comparing S. ratti with other Sarcocystis species using rodents as intermediate hosts. While ITS1 sequences of S. ratti were highly distinct from other Sarcocystis species available in GenBank. Phylogenetic and ecological data suggest that predatory mammals living near households are definitive hosts of S. ratti.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Members of the genus Sarcocystis are worldwide distributed apicomplexan parasites of reptiles, birds, and mammals. They are characterised by an obligatory two-host prey-predator life cycle. Sexual multiplication takes place in the small intestine of the definitive host, whereas asexual stages including sarcocysts develop in the extra-intestinal tissues of the intermediate host. Some Sarcocystis species are intermediate host specific, whereas others have a wide range of hosts (Dubey et al. 2016).
The brown rat was the most comprehensively examined for Sarcocystis infection among rat species and this rodent was shown to be an intermediate host for S. cymruensis Ashford, 1978, S. murinotechis Munday and Mason, 1980, S. rodentifelis Grikienienė et al., 1993, S. singaporensis Zaman and Colley (1975) 1976, S. villivillosi Beaver and Maleckar, 1981, S. zamani Beaver and Maleckar, 1981, S. zuoi Hu et al., 2005, and S. dispersa-like (Munday and Mason 1980; Beaver and Maleckar 1981; Munday 1983; Hu et al. 2005, 2011, 2012; Zaman and Colley 1975, 1976). The synonymy of S. cymruensis and S. rodentifelis has been discussed in the recent studies (Dubey et al. 2016; Antunes Murata et al. 2018). Snakes, birds of prey, and cats serve as definitive hosts of Sarcocystis species found in the muscles of rats (Munday 1977, 1983; Matuschka 1987; Jäkel et al. 1997; Koudela and Modrý 2000). Interestingly, rats can act as both the intermediate and definitive hosts of S. cymruensis and S. rodentifelis (Hu et al. 2011). Hence, Sarcocystis species from rats have a large variety of final hosts.
Two rat species, the black rat (Rattus rattus Linnaeus, 1758) and the brown rat, are known in Latvia. Both species dwell near human housing where their enemies are domestic animals, the European polecat (Mustela putorius Linnaeus, 1758), the beech marten (Martes foina Erxleben, 1777), and birds of prey such as owls. Despite an omnivore lifestyle, black rats prefer food of plant origin, while brown rats choose animal food (Burnie and Wilson 2006; Kampe-Pērsone 2017). Limited data are available on Sarcocystis in black rats. Sarcocysts similar to S. singaporensis were detected in the Malaysian black rat (Kan and Dissanaike 1977), meanwhile Thailand black rat harboured S. singaporensis and S. zamani (Jäkel et al. 1997). Also, sarcocysts of Sarcocystis sp. were detected in skeletal muscles of black rats from Lithuania; however, no morphology of cysts was described (Grikienienė et al. 2001). In this paper, a new Sarcocystis species found in skeletal muscles of the black rat in Latvia is described based on morphological and DNA investigations.
Materials and methods
Sample collection and morphological examination
Thirteen black rats (Rattus rattus) captured in a trap near the farm from Latgale Region in November 2015 were necropsied. Skeletal muscles and such internal organs as the kidneys, the heart, the liver, and the lungs were examined for Sarcocystis infection.
To detect sarcocysts, fragments of muscle tissue were stained with 0.2% methylene blue solution, lightened with 1.5% acetic acid solution, placed in a glass compressor, and studied under a stereomicroscope at × 20 magnification. Sarcocystis infection intensity was evaluated by counting sarcocysts found in methylene blue–stained 28 oat-size pieces of muscle (~ 1 g).
Morphological analysis of sarcocysts observed was performed in fresh-squashed samples of the muscle. Sarcocysts with a small amount of host tissue were excised with the help of two preparation needles, and afterwards were characterised morphologically. Sarcocysts were described according to the size and shape of the cyst, the structure of the cyst wall, and morphometric parameters of bradyzoites. Cysts were measured under a stereomicroscope at × 20 magnification, and a detailed morphological characterisation was carried out under a light microscope (LM) at × 40–1000 magnification.
A single isolated sarcocyst was fixed in 2% glutaraldehyde and subjected to transmission electron microscopy (TEM) analysis. Sarcocyst was postfixed in 1% osmium tetroxide, dehydrated, and infiltrated in epoxy resin. Sections were cut on a Leica UC6 ultramicrotome and stained with 4% uranyl acetate and 3% lead citrate. Grids were imaged at 100 kV with the Morgagni 268 TEM (FEI, Hillsboro, OR, USA).
Molecular analysis
Two sarcocysts from two individuals of black rats were excised from fresh muscle preparations, preserved in individual microcentrifuge tubes containing 96% ethanol and kept at − 20 °C until molecular examination. The isolated sarcocysts were molecularly characterised at four genetic loci (18S rDNA, 28S rDNA, ITS1, and cox1).
Genomic DNA was extracted from sarcocysts using the QIAamp® DNA Micro Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Nearly complete 18S rDNA was amplified using SarAF/SarBR and SarCF/SarDR primer pairs; partial 28S rDNA sequences were amplified using KL-P1F/KL-P2R primer pair (Kutkienė et al. 2010) and partial cox1 sequences were amplified with the help of SF1/SR5 primers (Gjerde 2013) and the complete ITS1 region was amplified using SU1F/5.8SR2 primer pair (Gjerde 2014). PCRs for 18S rDNA, 28S rDNA, and cox1 were performed in a final 25-μL volume consisting of 0.5 μM of each primer, 12.5-μL DreamTaq PCR Master Mix (Thermo Fisher Scientific Baltics, Vilnius, Lithuania), 0.04 μg template DNA, and nuclease-free water. Amplification reactions were carried out, starting with the initial hot start at 95 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 45 s, annealing at 54–60 °C depending on the primer pair for 45 s, elongation at 72 °C for 80 s, and ended with the final extension at 72 °C for 10 min. PCR amplification of ITS1 region was unsuccessful with DreamTaq polymerase; therefore, Platinum™ II Hot-Start Green PCR Master Mix (Thermo Fisher Scientific Baltics, Vilnius, Lithuania) was used. PCR for ITS1 was performed in a final 20-μL volume consisting of 0.5 μM of each primer, 10-μL Platinum II Hot-Start Green PCR Master Mix, 0.04 μg template DNA, and nuclease-free water. The cycling conditions began with 1 cycle at 94 °C for 2 min, followed by 30 cycles at 94 °C for 15 s, at 60 °C for 15 s, and at 68 °C for 15 s. The amplified products were visualised using 1.5% agarose gel electrophoresis and purified with the help of ExoI and FastAP (Thermo Fisher Scientific Baltics, Vilnius, Lithuania). The PCR product visualisation, purification, and sequencing were carried out in the previously described way (Prakas et al. 2016).
The resulting sequences were compared with those of various Sarcocystis spp. using Nucleotide BLAST program megablast and blastn options (http://blast.ncbi.nlm.nih.gov/). Multiple sequence alignments were obtained with the help of MUSCLE algorithm implemented in the MEGA7 (Kumar et al. 2016). Selection of nucleotide substitution model and phylogenetic analyses under Bayesian inference were conducted using TOPALi v2.5 (Milne et al. 2004).
Results
Infection rates and morphological characteristics of S. ratti
Sarcocystis infection was detected in 15.4% (2/13) of rats examined. Sarcocysts were found in skeletal muscles, while no cysts were discovered in internal organs. The intensity of Sarcocystis infection varied in two animals and numbered from 2 to 67 sarcocysts in 1 g of muscle samples.
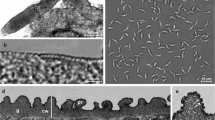
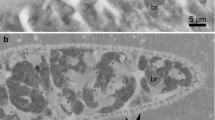
Under a light microscope, one morphological type of sarcocysts was observed. Sarcocysts were microscopic, ribbon-shaped with round tips, and measured 0.9–1.3 × 0.09–0.14 (n = 5) mm. The cyst wall was thin (0.8–1.3 μm; n = 3) and seemed smooth without visible protrusions (Fig. 1a). Lancet-shaped bradyzoites were 8.3 × 4.3 (7.5–9.3 × 3.9–4.8; n = 35) μm in size (Fig. 1b). Under the TEM, the cyst wall reached up to 1.3 μm in thickness, lacked protrusions, and appeared slightly wavy (Fig. 1c). The parasitophorous vacuolar membrane was about 100 nm thick and had small knob-like blebs (Fig. 1d). The ground substance appeared smooth and continued into the interior of the cyst as septae that subdivided the cyst in chambers filled with bradyzoites. The cyst wall was type 1a-like (Dubey et al. 2016). Based on DNA sequence comparison, sarcocysts found in black rats in Latvia were proposed as S. ratti n. sp.
Morphology of Sarcocystis ratti n. sp. from muscles of the black rat (Rattus rattus). a, b LM micrographs. Fresh preparations. a Portion of sarcocyst showing a thin and apparently smooth cyst wall (arrow). b Lancet-shaped bradyzoites. c, d TEM micrographs. c Fragment of a slightly wavy cyst wall (arrows); note muscular host cell (hc), bradyzoite (br), septae (se), and ground substance (g). d High magnification of the cyst wall; a parasitophorous vacuolar membrane has knob-like blebs (arrowheads), the ground substance seems smooth
Molecular characteristics of S. ratti and phylogeny
Two S. ratti isolates obtained from two black rats were identical at 1757-bp-long 18S rDNA (MK425189-MK425190), 1475-bp-long 28S rDNA (MK425192-MK425193), 1053-bp-long cox1 (MK430072-MK430073), and 902-bp-long ITS1 (MK910965-MK910966). Considerable genetic differences, from 2.3, 4.5, and 5.8% within 18S rDNA, cox1, and 28S rDNA, respectively, were observed when comparing S. ratti with other Sarcocystis species using the rodent as intermediate hosts. While significant match was not obtained comparing ITS1 sequences of S. ratti with those of other Sarcocystis spp. available in GenBank. Thus, S. ratti is clearly molecularly separable at all four loci examined from other Sarcocystis species using rodents as intermediate hosts. The lowest interspecific variability was detected comparing 18S rDNA sequences. Based on this locus, S. ratti showed the greatest similarity (98.3%) to S. rileyi Stiles, 1893 (Dubey et al. 2003) (GU120092) and displayed > 97% similarity to more than 20 Sarcocystis species. At cox1, sequences of S. ratti demonstrated 95.5% identity with those of S. strixi Verma et al., 2017 (MF162317) and S. cymruensis (MG571085), 95.2% identity with S. lutrae Gjerde and Josefsen, 2015 (MF596285), S. lari Prakas et al., 2014 (MF596283), and S. speeri Dubey and Lindsay, 1999 (KT207461). Based on the 28S rDNA sequences, S. ratti had the strongest identity (94.2%) with S. cymruensis (MH564724) and disclosed 93.1–93.4% identity with S. glareoli (Erhardrová, 1955) Odening, 1997 (AF044251), S. jamaicensis Verma et al., 2017 (KY994650), S. lari (MF946611), S. muris (Railliet, 1886) Labbe, 1899 (AF012883), S. fulicae Prakas et al., 2018 (MG273672), and S. turdusi Kutkienė et al., 2012 (JF975682). In summary, S. ratti showed the greatest genetic similarity to Sarcocystis species distinguished by the rodent-cat (S. cymruensis, S. muris, and S. rodentifelis), the rodent-bird (S. glareoli, S. jamaicensis, and S. strixi), the rodent-opossum (S. speeri), the bird-bird (for instance, S. halieti Gjerde et al., 2018), the bird-carnivorans (for instance, S. rileyi), and the carnivorans-unknown (for instance, S. lutrae Gjerde and Josefsen, 2015) life cycle.
The fragment of 18S rDNA analysed was not variable enough to robustly resolve phylogenetic relationships of selected Sarcocystis species closely related to S. ratti (Fig. 2). The newly described species from the black rat formed a separate branch in the phylogenetic tree obtained using cox1 sequences. It should be noted that 18S rDNA and 28S rDNA phylogenetic trees did not demonstrate any close relationship between S. ratti and Sarcocystis species employing rodents as intermediate and snakes as definitive hosts (S. atheridis Slapeta et al., 1999, S. singaporensis, S. zamani, S. zuoi, and Sarcocystis sp. AF513490). Based on 28S rDNA, S. ratti was a sister species to S. muris and S. cymruensis.
Phylogenetic placement of S. ratti based on 18S rDNA (a), cox1 (b), and 28S rDNA (c) sequences. The trees have been constructed using the Bayesian methods, scaled according to the branch length, and rooted on Toxoplasma gondii (a and b) or S. cruzi (c). Sarcocystis species using rodents as natural and experimental intermediate hosts are in boldface and underlined, respectively
Taxonomic summary of S. ratti n. sp.
Type intermediate host: The black rat (Rattus rattus).
Definitive host: Unknown.
Locality: Eastern Latvia, Latgale region.
Specimens deposited: TEM material deposited at the National Centre of Pathology, Vilnius, Lithuania. Sequences deposited in NCBI GenBank with accession numbers MK425189-MK425190 (18S rDNA), MK425192-MK425193 (28S rDNA), MK430072-MK430073 (cox1), and MK910965-MK910966 (ITS1).
Etymology: the Latin name of genus Rattus is used for the species name.
Recorded in URN as urn:lsid:zoobank.org:act:28CFF2F6-D557-46FC-8F01-BEF9A2AAE88A.
Discussion
Here we describe S. ratti from the black rat in Latvia characterised by sarcocyst having a thin (up to 1.3 μm) cyst wall without clearly visible protrusions (Fig. 1). Ribbon-shaped sarcocysts of S. ratti were up to 1.3 × 0.14 mm in size, and lancet-shaped bradyzoites measured 8.3 × 4.3 (7.5–9.3 × 3.9–4.8) μm. Despite numerous reports on Sarcocystis in rodents, limited data have been acquired of Sarcocystis in the black rat. The reticulated python (Python reticulatus Schneider, 1801), definitive hosts of two species identified in the black rat, S. singaporensis and S. zamani (Jäkel et al. 1997), does not dwell in the territory of Latvia. Furthermore, the cyst wall structure of both species apparently differs from the detected one in the present investigation. Whereas, three Sarcocystis species (S. cymruensis, S. rodentifelis, and S. dispersa-like), which also have a thin and smooth sarcocyst wall, were found in the brown rat. Under TEM, S. ratti and S. cymruensis had a similar sarcocyst wall structure, and their parasitophorous vacuolar membranes contained numerous small blebs (Hu et al. 2011; Antunes Murata et al. 2018). However, there is a notable difference in the sizes of bradyzoites, as those of S. cymruensis were more elongated, measured 11.0–13.5 × 3.0–5.0 μm, in size (Hu et al. 2011). Under LM, a smooth cyst wall of S. rodentifelis was up to 1.0 μm in thickness, whereas, banana-like bradyzoites of this species were slightly longer and thinner (11.6–14.7 × 2.2–4.2 μm) than those belonging to S. ratti (Grikienienė et al. 1993). By TEM, sarcocysts of S. dispersa-like had simple wall structure, without protrusions on the surface, but with invaginations (Munday 1983). The DNA sequence comparison performed in this study indicated significant genetic differences between S. cymruensis, S. rodentifelis, S. dispersa-like, and S. ratti at 18S rDNA, 28S rDNA, and cox1. Based on the 18S rDNA sequences, S. ratti showed 97.5% identity to S. dispersa. Having compared sequences of S. ratti and S. rodentifelis, 97.1% (18S rDNA) and 89.9% (28S rDNA) identity was observed. Meanwhile sequences of S. ratti and S. cymruensis differed by 3.5%, 5.8%, and 4.5% within 18S rDNA, 28S rDNA, and cox1, respectively.
Despite a wide variety of Sarcocystis species using rodents as an intermediate host (Prakas and Butkauskas 2012), only some of them (S. atheridis, S. cymruensis, S. dispersa, S. glareoli, S. jamaicensis, S. microti (Findlay and Middleton, 1934) Modrý et al. 2004, S. muris, S. rodentifelis, S. singaporensis, S. speeri, S. strixi, S. zamani, S. zuoi, and several Sarcocystis sp.) have been characterised genetically. Predominantly, rDNA sequences were used for a genetic description and identification of these Sarcocystis spp. (Votýpka et al. 1998; Dolezel et al. 1999; Mugridge et al. 1999, 2000; Slapeta et al. 2001, 2003; Hu et al. 2012). Whereas other genes, like cox1 and ITS1, were only occasionally applied to Sarcocystis using rodents as intermediate hosts (Dubey et al. 2015; Verma et al. 2017; Watthanakaiwan et al. 2017; Antunes Murata et al. 2018). Interestingly, in the present study, obtained ITS1 sequences of S. ratti had no considerable similarity to other Sarcocystis species; therefore, further molecular studies on ITS1 of Sarcocystis species from rodents are highly preferable. The current study indicated higher variability of Sarcocystis from rodents within cox1 and 28S rDNA as compared with 18S rDNA. Furthermore, 18S rDNA sequences, in contrast to 28S rDNA and cox1, did not establish robust phylogenetic relationships among Sarcocystis spp. closely related to S. ratti (Fig. 2).
A phylogenetic analysis demonstrates no close relationship between S. ratti and Sarcocystis species characterised by a rodent-bird or rodent-snake life cycle (Fig. 2). Based on 28S rDNA, S. ratti was most closely related to S. muris and S. cymruensis using the cat as the natural definitive host. Whereas, S. ratti was placed separately from S. cymruensis in the cox1 phylogenetic tree. Thus, on the basis of phylogenetic results, predatory mammals are presumed to be definitive hosts of S. ratti. Since the black rat is synanthropic species, possible definitive hosts of S. ratti could be predators living near households, such as the domestic cat, the domestic dog, the European polecat, or the beech marten.
References
Antunes Murata FH, Cerqueira-Cézar CK, Thompson PC, Tiwari K, Mowery JD, Verma SK, Rosenthal BM, Sharma RN, Dubey JP (2018) Sarcocystis cymruensis: discovery in Western hemisphere in the Brown rat (Rattus norvegicus) from Grenada, West Indies: redescription, molecular characterization, and transmission to IFN-γ gene knockout mice via sporocysts from experimentally infected domestic cat (Felis catus). Parasitol Res 117:1195–1204. https://doi.org/10.1007/s00436-018-5799-5
Beaver PC, Maleckar JR (1981) Sarcocystis singaporensis Zaman and Colley, (1975) 1976, Sarcocystis villivilliso sp. n., and Sarcocystis zamani sp. n.: development, morphology, and persistence in the laboratory rat, Rattus norvegicus. J Parasitol 67:241–256
Burnie D, Wilson DE (2006) Animal: the definitive visual guide. Mammals Part I. Dorling Kindersley Limited, London
Dolezel D, Koudela B, Jurků M, Hypsa V, Obornik M, Votýpka J, Modrý D, Slapeta JR, Lukes J (1999) Phylogenetic analysis of Sarcocystis spp. of mammals and reptiles supports the coevolution of Sarcocystis spp. with their final hosts. Int J Parasitol 29:795–798. https://doi.org/10.1016/S0020-7519(99)00018-1
Dubey JP, Cawthorn RJ, Speer CA, Wobeser GA (2003) Redescription of the sarcocysts of Sarcocystis rileyi (Apicomplexa: Sarcocystidae). J Eukaryot Microbiol 50:476–482. https://doi.org/10.1111/j.1550-7408.2003.tb00274.x
Dubey JP, Verma SK, Dunams D, Calero-Bernal R, Rosenthal BM (2015) Molecular characterization and development of Sarcocystis speeri sarcocysts in gamma interferon gene knockout mice. Parasitology 142:1555–1562. https://doi.org/10.1017/S0031182015001109
Dubey JP, Calero-Bernal R, Rosenthal BM, Speer CA, Fayer R (2016) Sarcocystosis of animals and humans, 2nd edn. CRC Press, Boca Raton
Gjerde B (2013) Phylogenetic relationships among Sarcocystis species in cervids, cattle and sheep inferred from the mitochondrial cytochrome c oxidase subunit I gene. Int J Parasitol 43:579–591. https://doi.org/10.1016/j.ijpara.2013.02.004
Gjerde B (2014) Molecular characterisation of Sarcocystis rileyi from a common eider (Somateria mollissima) in Norway. Parasitol Res 113:3501–3509. https://doi.org/10.1007/s00436-014-4062-y
Gjerde B, Vikøren T, Hamnes IS (2018) Molecular identification of Sarcocystis halieti n. sp., Sarcocystis lari and Sarcocystis truncata in the intestine of a white-tailed sea eagle (Haliaeetus albicilla) in Norway. Int J Parasitol Parasites Wildl 7:1–11. https://doi.org/10.1016/j.ijppaw.2017.12.001
Grikienienė J, Arnastauskienė T, Kutkienė L (1993) On some disregarded ways of Sarcosporidians’ circulation and remarks about systematics of the genus Sarcocystis Lankester, 1882 with the description of the new species from rodents. Ekologija 1:16–24
Grikienienė J, Malakauskas M, Mažeikytė R, Balčiauskas L, Senutaitė J (2001) Muscle parasites (Sarcocystis, Trichinella, Alaria) of wild mammals in Lithuania. Theriol Lituanica 1:29–46
Hu JJ, Ma TC, Li XR (2005) A new species of sarcocysts (Sporozoea, Eucoccidiida) from Rattus norvegicus. Acta Zootaxon Sin 30:287–290. https://doi.org/10.1645/GE-2831.1
Hu JJ, Liao JY, Meng Y, Guo YM, Chen XW, Zuo YX (2011) Identification of Sarcocystis cymruensis in wild Rattus flavipectus and Rattus norvegicus from Peoples Republic of China and its transmission to rats and cats. J Parasitol 97:421–424. https://doi.org/10.1645/GE-2633.1
Hu JJ, Meng Y, Guo YM, Liao JY, Song JL (2012) Completion of the life cycle of Sarcocystis zuoi, a parasite from the Norway rat, Rattus norvegicus. J Parasitol 98:550–553. https://doi.org/10.1645/GE-2831.1
Jäkel T, Khoprasert Y, Sorger I, Kliemt D, Seehabutr V, Suasa-ard K, Hongnark S (1997) Sarcosporidiasis in rodents from Thailand. J Wildl Dis 33:860–867. https://doi.org/10.7589/0090-3558-33.4.860
Kampe-Pērsone G (2017) Latvijas zīdītāji. Pilnīgs sugu apraksts. Zvaigzne ABC, Riga
Kan SP, Dissanaike AS (1977) Ultrastructure of Sarcocystis sp. from the Malaysian house rat, Rattus rattus diardii. Z Parasitenkd 52:219–227
Koudela B, Modrý D (2000) Sarcocystis muris possesses both diheteroxenous and dihomoxenous characters of life cycle. J Parasitol 86:877–879. https://doi.org/10.1645/0022-3395(2000)086[0877:SMPBDA]2.0.CO;2
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kutkienė L, Prakas P, Sruoga A, Butkauskas D (2010) The mallard duck (Anas platyrhynchos) as intermediate host for Sarcocystis wobeseri sp. nov. from the barnacle goose (Branta leucopsis). Parasitol Res 107:879–888. https://doi.org/10.1007/s00436-010-1945-4
Kutkienė L, Prakas P, Butkauskas D, Sruoga A (2012) Description of Sarcocystis turdusi sp. nov. from the common blackbird (Turdus merula). Parasitology 139:1438–1443. https://doi.org/10.1017/S0031182012000819
Matuschka FR (1987) Reptiles as intermediate and/or final hosts of Sarcosporidia. Parasitol Res 73:22–32
Milne I, Wright F, Rowe G, Marshall D, Husmeier D, McGuire G (2004) TOPALi: software for automatic identification of recombinant sequences within DNA multiple alignments. Bioinformatics 20:1806–1807. https://doi.org/10.1093/bioinformatics/bth155
Modrý D, Votýpka J, Svobodová M (2004) Note on the taxonomy of Frenkelia microti (Findlay & Middleton, 1934) (Apicomplexa: Sarcocystidae). Syst Parasitol 58:185–187. https://doi.org/10.1023/b:sypa.0000032924.63708.57
Mugridge NB, Morrison DA, Johnson AM, Luton K, Dubey JP, Votýpka J, Tenter AM (1999) Phylogenetic relationships of the genus Frenkelia: a review of its history and new knowledge gained from comparison of large subunit ribosomal ribonucleic acid gene sequences. Int J Parasitol 29:957–972. https://doi.org/10.1016/S0020-7519(99)00062-4
Mugridge NB, Morrison DA, Jäkel T, Heckeroth AR, Tenter AM, Johnson AM (2000) Effects of sequence alignment and structural domains of ribosomal DNA on phylogeny reconstruction for the protozoan family sarcocystidae. Mol Biol Evol 17:1842–1853. https://doi.org/10.1093/oxfordjournals.molbev.a026285
Munday BL (1977) A species of Sarcocystis using owls as definitive hosts. J Wildl Dis 13:205–207
Munday BL (1983) An isosporan parasite of masked owls producing sarcocysts in rats. J Wildl Dis 19:146–147
Munday BL, Mason RW (1980) Sarcocystis and related organisms in Australian wildlife: III. Sarcocystis murinotechis sp. n. life cycle in rats (Rattus, Pseudomys and Mastocomys spp.) and tiger snakes (Notechis ater). J Wildl Dis 16:83–88
Prakas P, Butkauskas D (2012) Protozoan parasites from genus Sarcocystis and their investigations in Lithuania. Ekologija 58:45–58
Prakas P, Kutkienė L, Butkauskas D, Sruoga A, Žalakevičius M (2014) Description of Sarcocystis lari sp. n. (Apicomplexa: Sarcocystidae) from the great black-backed gull, Larus marinus (Charadriiformes: Laridae), on the basis of cyst morphology and molecular data. Folia Parasitol 61:11–17. https://doi.org/10.14411/fp.2014.002
Prakas P, Butkauskas D, Rudaitytė E, Kutkienė L, Sruoga A, Pūraitė I (2016) Morphological and molecular characterization of Sarcocystis taeniata and Sarcocystis pilosa n. sp. from the sika deer (Cervus nippon) in Lithuania. Parasitol Res 115:3021–3032. https://doi.org/10.1007/s00436-016-5057-7
Prakas P, Butkauskas D, Švažas S, Juozaitytė-Ngugu E, Stanevičius V (2018) Morphologic and genetic identification of Sarcocystis fulicae n. sp. (Apicomplexa: Sarcocystidae) from the Eurasian coot (Fulica atra). J Wildl Dis 54:765–771. https://doi.org/10.7589/2017-11-279
Slapeta JR, Modrý D, Koudela B (1999) Sarcocystis atheridis sp. nov., a new sarcosporidian coccidium from Nitsche’s bush viper, Atheris nitschei Tornier, 1902, from Uganda. Parasitol Res 85:758–764
Slapeta JR, Modrý D, Votýpka J, Jurků M, Koudela B, Lukes J (2001) Multiple origin of the dihomoxenous life cycle in sarcosporidia. Int J Parasitol 31:413–417. https://doi.org/10.1016/S0020-7519(01)00127-8
Slapeta JR, Modrý D, Votýpka J, Jurků M, Lukes J, Koudela B (2003) Evolutionary relationships among cyst-forming coccidian Sarcocystis spp. (Alveolata: Apicomplexa: Coccidea) in endemic African tree vipers and perspective for evolution of heteroxenous life cycle. Mol Phylogenet Evol 27:464–475. https://doi.org/10.1016/S1055-7903(03)00018-6
Verma SK, von Dohlen AR, Mowery JD, Cerqueira-Cézar CK, Rosenthal BM, Dubey JP, Lindsay DS (2017) Sarcocystis strixi n. sp. from barred owl (Strix varia) definitive host and interferon gamma gene knockout mice as experimental intermediate host. J Parasitol 103:768–777. https://doi.org/10.1645/16-173
Votýpka J, Hypsa V, Jirků M, Flegr J, Vávra J, Lukes J (1998) Molecular phylogenetic relatedness of Frenkelia spp. (Protozoa, Apicomplexa) to Sarcocystis falcatula Stiles 1893: is the genus Sarcocystis paraphyletic? J Eukaryot Microbiol 45:137–141. https://doi.org/10.1111/j.1550-7408.1998.tb05081.x
Watthanakaiwan V, Sukmak M, Hamarit K, Kaolim N, Wajjwalku W, Muangkram Y (2017) Molecular characterization of the ribosomal DNA unit of Sarcocystis singaporensis, Sarcocystis zamani and Sarcocystis zuoi from rodents in Thailand. J Vet Med Sci 79:1412–1418. https://doi.org/10.1292/jvms.16-0086
Zaman V, Colley FC (1975) Light and electron microscopic observations of the life cycle of Sarcocystis orientalis sp. n. in the rat (Rattus norvegicus) and the Malaysian reticulated python (Python reticulatus). Z Parasitenkd 47:169–185
Zaman V, Colley FC (1976) Replacement of Sarcocystis orientalis Zaman and Colley, 1975, by Sarcocystis singaporensis sp. n. Z Parasitenkd 51:137
Acknowledgements
The authors are grateful to Ms. S. Amšiejienė from the National Centre of Pathology (Vilnius, Lithuania) for her help in carrying out electron microscopy investigations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and institutional guidelines for the care and use of animals were followed.
Additional information
Section Editor: Daniel K. Howe
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Prakas, P., Kirillova, V., Gavarāne, I. et al. Morphological and molecular description of Sarcocystis ratti n. sp. from the black rat (Rattus rattus) in Latvia. Parasitol Res 118, 2689–2694 (2019). https://doi.org/10.1007/s00436-019-06393-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-019-06393-9