Abstract
Purpose
Abnormal expression of miRNAs is closely related to a variety of human cancers. The purpose of this study is to identify new tumor suppressor miRNA and elucidate its physiological function and mechanism in renal cell carcinoma (RCC).
Methods
The expression of miR-145 in 45 RCC and adjacent normal tissues was performed by quantitative RT-PCR. Cell proliferation, migration, invasion, apoptosis and cycle assays were carried out for functional analysis after miR-145 transfection. Two target genes of miR-145 were identified by luciferase reporter assay. The altered expression of 84 epithelial to mesenchymal transition (EMT)-related genes after miR-145 transfection was detected by RT2 Profiler EMT PCR array.
Results
The expression of miR-145 was downregulated in RCC compared to their normal adjacent tissues. Restoring miR-145 expression in RCC cell lines dramatically suppressed cell proliferation, migration and invasion, and induced cell apoptosis and G2-phase arrest. We further validated those miR-145 targets two oncogenes, ANGPT2 and NEDD9 in RCC. In addition, miR-145 was found to regulate numerous genes involved in the EMT.
Conclusions
These findings demonstrate that miR-145 functions as tumor suppressor in RCC, suggesting that miR-145 may be a potential therapeutic target for RCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In addition to prostate and bladder cancer, renal cell carcinoma (RCC) is the third most common urological cancer with the highest mortality rate at over 40 % (Redova et al. 2012). Clear cell renal cell carcinoma is the most common subtype of RCC, responsible for approximately 80 % of cases (Rini et al. 2009). The 5-year survival rate of RCC is estimated to be approximately 55 %, and that of metastatic RCC is approximately 10 % (Pascual and Borque 2008; Reeves and Liu 2009). Despite increased early detection of RCC and more frequent surgery, the mortality rate has not changed significantly and 20–40 % patients will develop recurrence (Redova et al. 2012). Therefore, an urgent need of searching new sensitive, reliable biomarkers and developing new targeted therapies is emphasized for renal cancer.
In current, one of the most popular and progressive approaches for molecular characterization of tumors is based on microRNAs (miRNAs) expression profiles. miRNAs are small non-coding RNAs (~22 nucleotides in length), which can play important regulatory roles in a variety of biological processes, such as cell cycle, proliferation, differentiation, metabolism and apoptosis (Ha 2011). They are genome-encoded, endogenous negative regulators of translation and mRNA stability originating from long primary transcripts with local hairpin structures (Chen et al. 2012). The miRNAs initially bind to the 3′UTR of target gene mRNA and repress protein translation or induce mRNA cleavage (Inui et al. 2010), thereby inhibiting translation from mRNA to protein, which control both physiological and pathological processes. Approximately 1,000 miRNAs have been predicted, and bioinformatic studies have estimated that miRNAs may regulate up to a half of all human genes and that each miRNA may control hundreds of target genes (Berezikov et al. 2005).
Plenty of evidence suggests that miRNAs are aberrantly expressed in many types of human cancers (Zaman et al. 2010; Chiyomaru et al. 2010; Noh et al. 2013), and they play pivotal roles in tumor initiation, development and metastasis. Some highly expressed miRNAs function as oncogenes, whereas lowly level miRNAs could act as tumor suppressors by negatively regulating oncogenes. Among them, miR-145 is one of the most frequently studied miRNAs in cancer biology. Downregulation of miR-145 had been reported extensively in human cancers, suggesting that miR-145 may function as a suppressor in a variety of tumors. Our previous studies successfully identified numerous aberrant expressions of miRNAs in RCC by massively parallel sequencing technology and discovered that miR-145 was significantly downregulated in RCC, which was consistent with other human cancers (Zhou et al. 2010). However, the biological role of miR-145 in RCC is not fully elucidated.
The aim of our study was to analyze the expression pattern of miR-145 in clinical RCC samples and examine the effects of miR-145 on proliferation, migration, invasion, apoptosis and cell cycle in two RCC cell lines. To better understand its regulatory mechanism, we validated two target genes, angiopoietin-2 (ANGPT2) and neural precursor cell expressed developmentally downregulated 9 (NEDD9), by luciferase reporter assay. In addition, we evaluated the effect of miR-145 on the expression levels of 84 key genes that are documented to be altered during epithelial to mesenchymal transition (EMT) by human EMT PCR array.
Materials and methods
RCC clinical specimens
Adjacent normal and cancerous renal tissues (36 clear cell RCC, 5 papillary RCC and 4 chromophobe RCC) were obtained from a total of 45 patients undergoing nephrectomy for RCC from the Sun Yat-Sen university cancer center and the first affiliated hospital of Anhui Medical University. Histopathological diagnoses were made according to the WHO classification system, and nuclear grade was scored according to the Fuhrman classification system (Fuhrman et al. 1982). The clinicopathological information of the patients is shown in Table 1. All specimens were obtained on the basis of their availability for research purpose and under a protocol approved by the local medical ethics committee. Written consent was obtained from the patients in the study.
Quantitative RT-PCR
Total RNA was extracted from tissue samples using the miRNeasy Mini Kit (Qiagen, Valencia, CA, USA). All isolated RNA was quantified by Nano-Drop® spectrophotometer. Only the RNA samples with 260/280 ratios of 1.8–2.0 were used for further investigation. cDNA was generated from 1 μg total RNA using the miScript II RT Kit (Qiagen), and quantitative RT-PCR was performed using the miScript SYBR® Green PCR Kit (Qiagen) according to the manufacturer’s instructions on the Roche Lightcycler 480 Real-Time PCR System. To normalize the data for quantification of miRNAs, U6 small nuclear RNA was selected as the endogenous control. The miRNAs expression level was determined using the delta–delta Ct method.
Cell culture and miRNAs transfection
Human renal carcinoma cell lines ACHN and 786-O cells were purchased from the American type culture collection (ATCC, Manassas, VA, USA). Both of them were maintained in DMEM media supplemented with 10 % fetal bovine serum and 1 % antibiotics (100 μ/ml penicillin and 100 mg/ml streptomycin sulfates) and were incubated in humidified atmosphere (5 % CO2) at 37 °C.
ACHN and 786-O cells were transfected with the indicated amounts of miR-145 mimics or negative control (RIBOBIO, Guangzhou, China) using Lipofectamine 2,000 (Invitrogen, Carlsbad, CA, USA).
Cell proliferation assay
Cell proliferation was measured by the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay (MTT, Sigma-Aldrich). Cells (approximately 6 × 103 cells) were seeded into 96-well culture plate 24 h prior to transfection with miR-145 mimics or mimics negative control. The blank control was set up with medium only; 0, 24, 48 or 72 h post-transfection, 20 μl of MTT (5 mg/ml) was added to each well, and plates were incubated for 4 h at 37 °C. Then, the MTT medium mixtures were discarded, and 150 μl of dimethyl sulfoxide (DMSO) was added to each well, shaking for 10 min at room temperature to solubilize the crystals. Absorbance was measured at a wavelength of 490 nm (with 630 nm as the reference wavelength) using ELISA microplate reader (Bio-Rad, Hercules, CA, USA). Assays were repeated at least three times.
Cell migration assay
Cell migration was examined by scratch assay according to the methods previously described (Liang et al. 2007). One day before scratch, stable cell lines of ACHN and 786-O cells were trypsinized and seeded equally into 6-well culture plates and grew to reach almost total confluence in 24 h. Using Lipofectamine 2,000, cells were transfected with 50 nM miR-145 mimics or the mimics negative control. At 6 h after transfection, an artificial homogenous wound was created onto the monolayer with a sterile 200-μl tip. After scratching, the cells were washed with PBS medium three times and incubated at 37 °C. Images of cells migrating into the wound were captured at time points of 0, 6, 12 18 and 24 h by digital camera system. The software program MIAS-2000 was used to determine the migration distance. Experiments were run in three independent repeats in triplicate and analyzed in a double-blind fashion by at least two observers.
Cell invasion assay
Matrigel (1:4, 60 μl/well, BD Bioscience, San Jose, CA, USA) was added to Transwell chambers (8.0 μm pore size, Corning, NY) in a 24-well plate. ACHN and 786-O cells transfected with miR-145 or negative control were harvested 24 h post-transfection and resuspended in serum-free DMEM medium. Aliquots (2 × 104 cells) of the prepared cell suspension were added into the upper chamber, and the lower chamber was filled with 1 ml DMEM supplemented with 10 % FBS. Cells were allowed to invade for 48 h at 37 °C with 5 % CO2. After incubation, non-invading cells in the upper chamber were removed with a cotton swab and washed three times with PBS. Then, the invasive cells were fixed in 100 % ethanol and stained with 0.1 % crystal violet. Invading cells were counted by taking photomicrographs in three fields. Experiments were performed in triplicate.
Cell apoptosis assay
The extent of apoptosis was evaluated by annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) detection kit (Invitrogen). ACHN and 786-O cells were transfected with miR-145 mimics or negative control. Cells were collected, washed twice with pre-chilled PBS and re-suspended in 1 × binding buffer, 48 h after treatment. Aliquots were mixed with 5 μl annexin V-FITC and 5 μl PI at room temperature for 15 min. The apoptosis assay was performed using a flow cytometry (EPICS, XL-4, Beckman, CA, USA). Each experiment was done at least three times.
Cell cycle analysis
786-O cells were transfected with miR-145 mimics or a negative control. Cells were harvested, washed twice with PBS and fixed with pre-chilled 70 % ethanol, 48 h after treatment. After centrifugation, cells were resuspended and incubated with 100 μl DNA PREP LPR and 500 μl DNA PREP STAIN for 30 min. Then, cell cycle analysis was performed using flow cytometry. Experiments were performed in three independent repeats.
Luciferase reporter assay
For production of a reporter construct, the miRNAs target sequences were inserted between the XhoI–NotI restriction sites in the 3′UTR of the hRluc gene in the psiCHECKTM-2 luciferase vector (Promega, Madison, WI, USA). Primer sequences for the 3′UTR of ANGPT2 mRNA (forward primer 5′-CCGCTCGAGAACTTGCATCACTTAACGGACCA-3′ and reverse primer 5′-ATTTGCGGCCGCTAGTCCCGAGTATAAAGCTGT-3′) and NEDD9 mRNA (forward primer 5′-CCGCTCGAGAAAGAGGAAGGGGACTGCGTTA-3′ and reverse primer 5′-ATTTGCGGCCGCTGAAATGTCTCCACTTAGCGTA-3′) were designed. In order to verify the binding sites of the miR-145, we synthesized two short fragments of ANGPT2 and NEDD9 3′UTR, which contains the potential binding sites. Then, we manually mutated the potential binding sites by exchanging the G and T, A and C. The sequences of the short fragments are shown in Fig. 4c, d. These four short fragments were all cloned into psiCHECK™-2 luciferase vector, respectively, and all the constructs were verified by sequencing. Then, the luciferase reporter constructs, together with miR-145 mimics or a negative control, were transfected into ACHN cells. Luciferase activity was detected using the dual luciferase assay system (Promega, Madison, WI, USA) according to the manufacturer’s instructions at 48 h after transfection. Normalized data were calculated as the quotient of Renilla/firefly luciferase activities. The experiments were performed in duplicate and repeated at least three times.
Epithelial to mesenchymal transition (EMT) PCR array
For the identification of the EMT-related genes, a RT2 Profiler PCR array for human EMT (Qiagen/SABiosciences) was used, which consists of 84 EMT-related genes. Total RNA isolated from the ACHN cells that transfected with miR-145 mimics or negative control was used for screening by real-time PCR according to the manufacturer’s instructions.
Statistical analysis
Statistical significance was determined using Student’s t test except that MTT was analyzed using ANOVA. p < 0.05 was considered statistically significant.
Results
miR-145 is downregulated in RCC
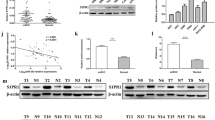
Expression of miR-145 has been reported to be suppressed in multiple cancers (Zaman et al. 2010; Chiyomaru et al. 2010; Noh et al. 2013). To analyze the expression of miR-145 renal cancer patients, total RNA was extracted from 45 pairs of RCC and their adjacent normal kidney tissues, and quantitative RT-PCR was performed. As shown in Fig. 1, the expression of miR-145 was decreased in 31 samples (68.9 %), with an overall average of 43.7 % downregulation (p = 0.008), suggesting that miR-145 may act as tumor suppressor gene in RCC.
Effects of miR-145 on cell proliferation, apoptosis and cell cycle in RCC cell lines
To investigate the biological role of miR-145, miR-145 mimics or negative control was transfected into ACHN and 786-O cells. The MTT assay demonstrated that the relative cell proliferation in miR-145 transfectants was significantly decreased by 32.8% (24 h), 27.2 % (48 h) and 24.4 % (72 h) in ACHN cells (p < 0.001). As for 786-O cells, the inhibition rates of cellular proliferation were 14.9 % (24 h), 16.9 % (48 h) and 11.6 % (72 h), respectively (p < 0.001) (Fig. 2a).
Effects of the miR-145 on cell proliferation, apoptosis and cell cycle in RCC cell lines. a miR-145 inhibited cell proliferation in ACHN cells (left panel) and 786-O cells (right panel). Cell growth was measured by MTT assay at different time intervals. ANOVA was used for the comparison of curves of cell growth. b miR-145 induced cell apoptosis in ACHN cells (upper panel) and 786-O cells (lower panel). Cell apoptosis is measured by flow cytometry analysis of Annexin V-FITC double-labeled. Representative images of flow cytometry analysis are shown. c miR-145 induced G2-phase arrest in 786-O cells. Cell cycle assay was performed by flow cytometry, and representative flow cytometric histograms are shown. The experiment was repeated three times. Data were the average of three independent experiments; bars, SD. *p < 0.05
To evaluate the effects of miR-145 on cell apoptosis, we therefore performed apoptosis experiments on ACHN and 786-O by flow cytometry. As shown in Fig. 2b, miR-145 significantly promoted cell apoptosis in ACHN cells (29.3 vs 12 %, p = 0.031) and in 786-O cells (30.4 vs 13 %, p = 0.018) compared with negative control.
Inhibition of cell growth in cancer cells is usually related to the cell cycle arrest. Therefore, we analyzed the function of miR-145 on cell cycle. The result showed an increase in the percentage of cells in G2 phase from 20.6 to 30.5 % in 786-O cells (Fig. 2c), indicating that miR-145 may lead to G2 cell cycle arrest in RCC cells.
Effects of miR-145 on cell migration and invasion in RCC cell lines
We analyzed the effect of miR-145 on cellular migration by wound healing assay. As shown in Fig. 3a, b, cell migration was significantly inhibited in the groups transfected with miR-145 compared with those in the negative control. The inhibition rates of migration were 39.3 % for ACHN cells (p < 0.001) and 30.4 % for 786-O cells (p < 0.001), indicating that miR-145 has a negative effect on cellular migration.
Effects of the miR-145 on cell migration and invasion in RCC cell lines. a miR-145 inhibited cell migration in ACHN cells. b miR-145 inhibited cell migration in 786-O cells. c miR-145 inhibited cell invasion in ACHN cells. d miR-145 inhibited cell invasion in 786-O cells. All experiments were performed three times, and a representative picture is shown. *p < 0.05. **p < 0.01
Since invasion is a key characteristic of malignant tumor, we next investigated the effects of miR-145 on tumor invasion in ACHN and 7i86-O cell lines. Invasion assay showed that cell invasion was obviously inhibited in miR-145-transfected cells compared with the negative control. The number of invading cells was decreased by 62.8 % (p < 0.001) in ACHN cells and was decreased by 44.3 % (p < 0.001) in 786-O cells, suggesting that miR-145 inhibited the invasive potential of renal cancer cells (Fig. 3c, d).
ANGPT2 and NEDD9 are two target genes of miR-145
We used three computational algorithms—picTar, TargetScan and miRanda—to predict the targets of miR-145. Among the potential targets, we focused on two oncogenes, ANGPT2 and NEDD9. As illustrated in the Fig. 4a, ANGPT2 and NEDD9 revealed putative target sites at the position 316–338 and 32–54 of 3′UTR, respectively, and with an exact match in the seed region at position 331–337 and 47–53, respectively. These two sites are located in conserved regions of the ANGPT2 and NEDD9 3′UTR among several species (Fig. 4a). The luciferase reporter assay indicated that the activity of the reporter containing the 3′UTR of ANGPT2 and NEDD9 genes was decreased following treatment with miR-145 mimics (Fig. 4b, p = 0.014, p = 0.006, respectively). To further verify the binding sites, we generated four luciferase reporter constructs containing the wild-type or mutated-type target sites of ANGPT2 or NEDD9 3′UTR. As shown in Fig. 4c, d the luciferase activity was significantly inhibited by miR-145 in wild-type constructs (p = 0.036, p < 0.001, respectively), whereas it had no effects on the mutated ones. Taken together, our results demonstrated that ANGPT2 and NEDD9 are two target genes of miR-145 and identified the sites of interaction in the 3′UTR of ANGPT2 and NEDD9.
ANGPT2 and NEDD9 are two target genes of miR-145. a Putative duplex formation between miR-145 and the ANGPT2 3′UTR (left panel), and between miR-145 and the NEDD9 3′UTR (right panel). These two binding sites are highly conserved among several mammalian species. The seed-recognizing sites are marked in bold type and underlined. b The luciferase reporter constructs that contained 3′UTR of ANGPT2 or NEDD9, together with miR-145 mimics or a negative control, were transfected into ACHN cells. At 48 h after transfection, luciferase activity was detected. Normalized data were calculated as the quotient of Renilla/firefly luciferase activities. c, d We synthesized two 23-nucleotide-long fragments of ANGPT2 and NEDD9 3′UTR, which contained the wild-type (WT) potential binding sites. And then, the mutated-type (MU) fragments were generated by exchanging the G and T, A and C on the putative binding sites. The sequences of these four fragments are shown in the upper panels. Samples were assayed as in b. The experiments were performed in duplicate and repeated three times. *p < 0.05 **p < 0.01
miR-145 promotes alterations in the expression of the EMT markers
EMT is the key process that is involved in tumor metastasis. Since miR-145 significantly inhibited the migration and invasion of RCC, we examined whether the restoration of miR-145 affects the expression of genes involved in EMT on ACHN cells transfected with miR-145 or negative control by performing an EMT PCR array. Of the 84 genes tested, 29 genes were altered after miR-145 treatment (>twofold), including 25 downregulated genes and 4 up-regulated genes (Fig. 5).
miR-145 transfection in ACHN cells leads to differential expression of EMT-related genes. a Overexpression of the miR-145 in ACHN cells leads to differential expression of genes represented on an EMT PCR array. Gene expression is represented in a scatter plot with a twofold cut-off displaying fold-changes between miR-145 and negative control transfected cells. The X-axis and the Y-axis represent the relative gene expression of the control group and miR-145 transfected group, respectively. Data is shown as the Log10(2−ΔCT). Each dot represents the mean of 3 replicates. Genes represented as down- or upregulated in the plot are listed with fold-change and p values. b Heat map of the EMT PCR array. Green square (downregulated), Red square (upregulated), Black square (not changed), Gray square (undetermined)
Discussion
Despite the great improvement of cancer therapy, major limitations were still existed in managing RCC. RCC is characterized by its resistance to current standard therapies (Cohen and McGovern 2005). Searching for alternative treatment strategies is still the top priority.
It is well known that miRNAs have added a layer of control to complex gene regulatory network in human (Lagos-Quintana et al. 2001). More and more evidence proved that the miRNAs have a versatile role in the regulation of genes during cancer development (Zhao et al. 2011). Among them, miR-145 is one of the most frequently studied miRNAs in human cancers, including liver, prostate and bladder cancer (Zaman et al. 2010; Chiyomaru et al. 2010; Noh et al. 2013). Downregulation of miR-145 and its functional analysis in various cancers suggest a tumor-suppressing role for this miRNA. However, there were few reports regarding the involvement of miR-145 in RCC. In our previous study, we noticed that miR-145 was downregulated in RCC by second-generation sequencing technology (Zhou et al. 2010). In the present study, we performed real-time PCR on 45 pairs of RCC samples to validate the sequencing data and found that miR-145 expression level was significantly lower in RCC compared with matched normal tissue. On the basis of these results, we assumed that miR-145 may have an important role as a tumor suppressor in kidney cancer. We therefore investigated its functional significance in RCC and found that miR-145 inhibited cell proliferation, migration and invasion, promoted cell apoptosis in RCC cells. Collectively, our results clearly illustrate the anti-cancer effect of miR-145, providing sufficient evidence to support the tumor suppressor role of miR-145 in RCC.
Although the evidence has highlighted the importance of miR-145 as a tumor suppressor in RCC, the precise molecular mechanisms remain largely unknown. To better understand the tumor suppressive effect of miR-145 in renal tumorigenesis, we used three computational algorithms to predict the targets of miR-145 and found two new target genes, ANGPT2 and NEDD9. We subsequently validated these two target genes as well as the binding sites of miR-145 on the ANGPT2 and NEDD9 3′UTR by the luciferase reporter assay.
ANGPT2 is a member of the angiopoietin family, which plays a pivotal role in angiogenesis during the development and expansion of human cancers. It is generally accepted that ANGPT2 is a natural antagonist of ANGPT1 that competes for binding to the Tie2 receptor and blocks ANGPT1-induced Tie2 autophosphorylation during angiogenesis (Hu and Cheng 2009). ANGPT1 is regarded to promote blood vessel maturation and stabilization, while ANGPT2 is thought to suppress Tie2 activity and destabilize blood vessels (Thurston and Daly 2012). ANGPT2 has been the focus of studies because of the close relation between ANGPT2 expression and tumor progression, growth and metastasis. Upregulation of ANGPT2 was reported in various human cancers, such as lung cancer (Naumnik et al. 2013), gastric cancer (Etoh et al. 2001) and RCC (Baldewijns et al. 2007; Bullock et al. 2010). More importantly, inhibiting the function of ANGPT2 resulted in suppression of cancer growth and invasion (Oliner et al. 2004), suggesting a potential inhibitor for antiangiogenesis treatment for cancer patients.
NEDD9, also known as HEF1 or Cas-L, is a non-catalytic scaffolding protein, which function as adaptor molecules that regulate and control cytoskeletal dynamics to promote multiple cellular processes, including proliferation, migration and invasion (Honda et al. 1998; Cabodi et al. 2010). In the previous studies, upregulation of NEDD9 has been strongly linked to enhanced invasion and metastasis in melanoma and glioblastoma (Natarajan et al. 2005; Kim et al. 2006), suggesting a pro-oncogenic role of NEDD9 in cancer development. Moreover, NEDD9 has been proposed as a candidate biomarker of invasiveness in lung cancer, breast cancer and melanoma (Chang et al. 2012; Izumchenko et al. 2009; Kim et al. 2006). Notably, negatively correlation between the expression of NEDD9 and miR-145 was reported in glioblastoma (Speranza et al. 2012).
In the present study, our data show that miR-145 regulates ANGPT2 and NEDD9 expression through directly binding to their 3′UTR, suggesting that miR-145 may has tumor suppressive functions through regulating these two oncogenic genes in RCC. Loss of miR-145 in RCC may lead to upregulation of ANGPT2 and NEDD9, which subsequently provide a selective growth and expansion advantage during renal carcinogenesis.
Gene regulation by miRNAs is a complicated process as they can have both direct and indirect targets. Indirect impacts of miRNAs can be downstream effectors of miRNAs targets. Since miR-145 has a significantly inhibitory effect on RCC cell migration and invasion, we set out to search the potential EMT-related pathway that is affected by miR-145 restoration by performing an EMT PCR array. The EMT array contains 84 genes including morphogenesis, extracellular matrix and cell adhesion, cytoskeletal genes and transcription factor genes that cause EMT. Our data indicated that miR-145 plays a critical role in the expression of a variety of genes involved in EMT, suggesting that the downregulation of miR-145 in RCC may provide advantages to tumor metastasis by modulating a set of genes required for EMT.
In summary, we found that miR-145 was frequently reduced in clinical specimens of RCC. By regulating direct or indirect target genes, miR-145 has a significant suppressive effect on tumor proliferation, migration and invasion, suggesting that miR-145 functions as a tumor suppressor in RCC. This novel molecular network may play a pivotal role in RCC oncogenesis and contribute to molecular-targeted therapy based on miRNAs.
References
Baldewijns M, Thijssen V, Van den Eynden G, Van Laere S, Bluekens A, Roskams T, Van Poppel H, De Bruine A, Griffioen A, Vermeulen P (2007) High-grade clear cell renal cell carcinoma has a higher angiogenic activity than low-grade renal cell carcinoma based on histomorphological quantification and qRT–PCR mRNA expression profile. Br J Cancer 96(12):1888–1895
Berezikov E, Guryev V, van de Belt J, Wienholds E, Plasterk RH, Cuppen E (2005) Phylogenetic shadowing and computational identification of human micro RNA genes. Cell 120(1):21–24
Bullock A, Zhang L, O’Neill A, Percy A, Sukhatme V, Mier J, Atkins M, Bhatt R (2010) Plasma angiopoietin-2 (ANG2) as an angiogenic biomarker in renal cell carcinoma (RCC). J Clin Oncol 2010(28):4630
Cabodi S, del Pilar Camacho-Leal M, Di Stefano P, Defilippi P (2010) Integrin signalling adaptors: not only figurants in the cancer story. Nat Rev Cancer 10(12):858–870
Chang J-X, Gao F, Zhao G-Q, Zhang G-J (2012) Expression and clinical significance of NEDD9 in lung tissues. Med Oncol 29(4):2654–2660
Chen B, Li H, Zeng X, Yang P, Liu X, Zhao X, Liang S (2012) Roles of microRNA on cancer cell metabolism. J Transl Med 10:228. doi:10.1186/1479-5876-10-228
Chiyomaru T, Enokida H, Tatarano S, Kawahara K, Uchida Y, Nishiyama K, Fujimura L, Kikkawa N, Seki N, Nakagawa M (2010) miR-145 and miR-133a function as tumour suppressors and directly regulate FSCN1 expression in bladder cancer. Br J Cancer 102(5):883–891
Cohen HT, McGovern FJ (2005) Renal-cell carcinoma. New Engl J Med 353(23):2477–2490
Etoh T, Inoue H, Tanaka S, Barnard GF, Kitano S, Mori M (2001) Angiopoietin-2 is related to tumor angiogenesis in gastric carcinoma possible in vivo regulation via induction of proteases. Cancer Res 61(5):2145–2153
Fuhrman SA, Lasky LC, Limas C (1982) Prognostic significance of morphologic parameters in renal cell carcinoma. Am J Surg Pathol 6(7):655–664
Ha T-Y (2011) MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw 11(3):135–154
Honda H, Oda H, Nakamoto T, Honda Z-I, Sakai R, Suzuki T, Saito T, Nakamura K, Nakao K, Ishikawa T (1998) Cardiovascular anomaly, impaired actin bundling and resistance to Src-induced transformation in mice lacking p130Cas. Nat Genet 19(4):361–365
Hu B, Cheng S-Y (2009) Angiopoietin-2: development of inhibitors for cancer therapy. Curr Oncol Rep 11(2):111–116
Inui M, Martello G, Piccolo S (2010) MicroRNA control of signal transduction. Nat Rev Mol Cell Bio 11(4):252–263
Izumchenko E, Singh MK, Plotnikova OV, Tikhmyanova N, Little JL, Serebriiskii IG, Seo S, Kurokawa M, Egleston BL, Klein-Szanto A (2009) NEDD9 promotes oncogenic signaling in mammary tumor development. Cancer Res 69(18):7198–7206
Kim M, Gans JD, Nogueira C, Wang A, Paik J-H, Feng B, Brennan C, Hahn WC, Cordon-Cardo C, Wagner SN (2006) Comparative oncogenomics identifies NEDD9 as a melanoma metastasis gene. Cell 125(7):1269–1281
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294(5543):853–858
Liang C–C, Park AY, Guan J-L (2007) In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc 2(2):329–333
Natarajan M, Stewart J, Golemis EA, Pugacheva E, Alexandropoulos K, Cox BD, Wang W, Grammer J, Gladson C (2005) HEF1 is a necessary and specific downstream effector of FAK that promotes the migration of glioblastoma cells. Oncogene 25(12):1721–1732
Naumnik W, Naumnik B, Niewiarowska K, Ossolinska M, Chyczewska E (2013) Angiogenic axis angiopoietin-1 and angiopoietin-2/tie-2 in non-small cell lung cancer: a bronchoalveolar lavage and serum study. Neurobiol Respir Spring 341–348
Noh JH, Chang YG, Kim MG, Jung KH, Kim JK, Bae HJ, Eun JW, Shen Q, Kim S-J, Kwon SH (2013) miR-145 functions as a tumor suppressor by directly targeting histone deacetylase 2 in liver cancer. Cancer Lett 335(2):455–462
Oliner J, Min H, Leal J, Yu D, Rao S, You E, Tang X, Kim H, Meyer S, Han SJ (2004) Suppression of angiogenesis and tumor growth by selective inhibition of angiopoietin-2. Cancer Cell 6(5):507–516
Pascual D, Borque A (2008) Epidemiology of kidney cancer. Adv Urol 2008:782381
Redova M, Poprach A, Besse A, Iliev R, Nekvindova J, Lakomy R, Radova L, Svoboda M, Dolezel J, Vyzula R, Slaby O (2012) MiR-210 expression in tumor tissue and in vitro effects of its silencing in renal cell carcinoma. Tumor Biol 34(1):481–491. doi:10.1007/s13277-012-0573-2
Reeves DJ, Liu CY (2009) Treatment of metastatic renal cell carcinoma. Cancer Chemoth Pharm 64(1):11–25
Rini BI, Campbell SC, Escudier B (2009) Renal cell carcinoma. Lancet 373(9669):1119–1132
Speranza MC, Frattini V, Pisati F, Kapetis D, Porrati P, Eoli M, Pellegatta S, Finocchiaro G (2012) NEDD9, a novel target of miR-145, increases the invasiveness of glioblastoma. Oncotarget 3(7):723
Thurston G, Daly C (2012) The complex role of angiopoietin-2 in the angiopoietin–tie signaling pathway. Cold Spring Harb Perspect Med 2(9):a006550
Zaman M, Chen Y, Deng G, Shahryari V, Suh S, Saini S, Majid S, Liu J, Khatri G, Tanaka Y (2010) The functional significance of microRNA-145 in prostate cancer. Br J Cancer 103(2):256–264
Zhao L, Chen X, Cao Y (2011) New role of microRNA: carcinogenesis and clinical application in cancer. Acta Biochim Biophys Sin 43(11):831–839
Zhou L, Chen J, Li Z, Li X, Hu X, Huang Y, Zhao X, Liang C, Wang Y, Sun L (2010) Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27. 3 associate with clear cell renal cell carcinoma. PLoS ONE 5(12):e15224
Acknowledgments
This project was supported by Science and Technology Planning Project of Shenzhen in China (No. 201102006) and Science and Technology Planning Project of Guangdong Province, China (No. 2011B031800381).
Conflict of interest
We declare that we have no conflict of interest in relation to this article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Ruijing Lu and Ziliang Ji have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Lu, R., Ji, Z., Li, X. et al. miR-145 functions as tumor suppressor and targets two oncogenes, ANGPT2 and NEDD9, in renal cell carcinoma. J Cancer Res Clin Oncol 140, 387–397 (2014). https://doi.org/10.1007/s00432-013-1577-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-013-1577-z