Abstract
Background and objective
Pathogenic variants in KCNT1 have been associated with severe forms of epilepsy, typically sleep-related hypermotor epilepsy or epilepsy of infancy with migrating focal seizures. To show that pathogenic variants in KCNT1 can be associated with mild extra-frontal epilepsy, we report a KCNT1 family with a wide spectrum of phenotypes ranging from developmental and epileptic encephalopathy to mild focal epilepsy without cognitive regression and not consistent with sleep-related hypermotor epilepsy.
Methods
A large Canadian family of Caucasian descent including 9 affected family members was recruited. Family members were phenotyped by direct interview and review of existing medical records. Clinical epilepsy gene panel analysis and exome sequencing were performed.
Results
Phenotypic information was available for five family members of which two had developmental and epileptic encephalopathy and three had normal development and focal epilepsy with presumed extra-frontal onset. All three had predominantly nocturnal seizures that did not show hyperkinetic features. All three reported clusters of seizures at night with a feeling of being unable to breathe associated with gasping for air, choking and/or repetitive swallowing possibly suggesting insular or opercular involvement. Genetic analysis identified a heterozygous KCNT1 c.2882G > A, p.Arg961His variant that was predicted to be deleterious.
Discussion
This family demonstrates that the phenotypic spectrum associated with KCNT1 pathogenic variants is broader than previously assumed. Our findings indicate that variants in KCNT1 can be associated with mild focal epilepsy and should not be excluded during variant interpretation in such patients based solely on gene–disease validity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathogenic variants in KCNT1, which encodes a sodium-activated potassium channel, have been described to be mainly associated with two distinct phenotypic presentations, a severe form of sleep-related hypermotor epilepsy (SHE, previously autosomal dominant nocturnal frontal lobe epilepsy) [1] and a developmental and epileptic encephalopathy (DEE) [2]. The latter often fulfills diagnostic criteria for epilepsy of infancy with migrating focal seizures (EIMFS) with seizure onset in the first 6 months of life and random and consecutive involvement of multiple cortical regions during a single seizure as well as significant developmental delay [2]. However, myoclonic atonic epilepsy and other developmental and epileptic encephalopathies such as Ohtahara syndrome have been observed as well [3,4,5,6,7]. This more severe phenotype is typically due to de novo variants [2, 3].
Sleep-related hypermotor epilepsy shows different phenotypic features. This is a focal epilepsy syndrome that is characterized by clusters of seizures occurring at night, typically with hyperkinetic seizures consistent with a frontal origin. SHE due to KCNT1 variants shows autosomal dominant inheritance and is often associated with intellectual disability and/or psychiatric features [1, 8] but the cognitive impact is less severe than in DEE/EIMFS.
Many patients with DEE/EIMFS due to KCNT1 variants have been reported in the literature, but the familial SHE phenotype is less clear. We identified seven families reported in the literature, including at least one individual with SHE [1, 8,9,10,11] (Table 1). In five families, the phenotype of all affected family members was consistent with SHE, typically associated with psychiatric features or intellectual disability. However, two families included family members consistent with SHE and others with EIMFS due to the same familial KCNT1 pathogenic variants [8, 10]. This suggests that additional factors modify the expression of the phenotype. Furthermore, multiple sporadic cases with a SHE phenotype have been reported [1, 7,8,9, 12].
Knowledge about the phenotypic spectrum that is associated with a particular gene is of utmost importance during variant interpretation. Only variants in genes that are associated with the particular disease phenotype of the patient, i.e., for which gene–disease validity exists, can be interpreted as causative. Knowledge of the level of variable expressivity possible for genes and associated variants is also critical, as inheritance of variants is often used to determine variant pathogenicity. Previous studies reported cognitive regression as a consistent phenotypic feature in KCNT1-related epilepsy [1, 7, 8]. Here, we report a KCNT1 family with a wide spectrum of phenotypes, including individuals with DEE but also individuals with milder focal epilepsy without psychiatric features, normal intelligence and not consistent with SHE.
Methods
A large Canadian family of Caucasian descent including 9 affected family members was recruited. Family members were phenotyped by direct interview and review of existing medical records. EEG reports were available in three patients. Two patients had additionally undergone video-EEG monitoring which were reviewed for this study. MRI was performed in three family members.
Clinical epilepsy gene panel analysis was performed in individuals III-14 and IV-5 via Prevention Genetics. Individual IV-5 also underwent exome sequencing using the MITO-FIND research study protocol [13]. The Nextera Flex for Enrichment Kit in combination with IDT xGen® Exome Research Panel (v1.0) probes was used for library preparation. Paired-end sequencing was performed using the NextSeq 500/550 High Output Kit v2.5 (300 Cycles). The Illumina Enrichment (v3.1.0) app workflow was used to generate variant call files using the GRCh37/hg19 assembly. The variant call file was imported into VarSeqTM 2.2.0 (Golden Helix®, Bozeman, United States of America) for tertiary analysis. The variant filtering process utilized databases and in silico predictors to annotate and evaluate the impact of the variants in the context of human disease, and variants were filtered based on quality (read depth and genotype quality), biological impact, clinical significance and minor allele frequency less than 5%. An internal assessment catalog and American College of Medical Genetics (ACMG) [14] classifier filter and Human Phenotype Ontology (HPO) terms [15] were used to assist in prioritizing variants (Phevor algorithm [16]). Variants matching HPO terms were then manually reviewed. Clinical segregation analysis was performed by Sanger sequencing in three other family members, two affected (III-11, II-5) and one unaffected (II-6).
Results
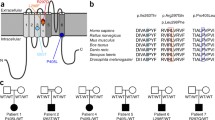
The family included 9 affected individuals over 3 generations (Fig. 1, Table 2). Phenotypic information was available for five family members of which two had DEE and three had focal epilepsy with normal development. Age of seizure onset in individuals with DEE was early (3–7 months), whereas family members with focal epilepsy had later onset (10–18 years). Individuals with DEE had significant developmental delay, whereas family members with focal epilepsy had normal development and showed no associated clinical features of psychiatric or learning disabilities. Neuropsychological assessment in individual III-14 with severe drug-resistant focal epilepsy described impaired to borderline scores for general intellect. However, the results were difficult to interpret due to considerable variability within and between cognitive domains and concomitant intake of topiramate at the time of testing. They were inconsistent with the patient’s presentation of normal intellect (she completed secondary school in a normal program with low average grades and maintains stable employment, Table 2).
Seizure semiology and EEG recordings in individual IV-5 with DEE were consistent with EIMFS with multifocal epileptiform discharges, focal and generalized onset seizures and propagation to different cortical areas in the same seizure (Online Resource 1). Her MRI did not show an epileptogenic lesion. No acquired epilepsy risk factors for IV-5 were recognized. Her mother was on carbamazepine and had two seizures but no status epilepticus during pregnancy. The pregnancy was otherwise unremarkable. Individual IV-5 was delivered at term without requiring resuscitation and discharged at two days of life. There was no history of brain injury or infection. The other individual with DEE (III-13) had passed away at age 28 years. She had been diagnosed with Lennox–Gastaut syndrome, but no further details on seizure semiology were available.
The three individuals with focal epilepsy (II-5, III-11, III-14) were most consistent with extra-frontal onset. All had predominantly nocturnal seizures but did not show hyperkinetic features associated with their seizures. All three reported episodes they had termed “breathing spells” which typically occurred in clusters at night. These were characterized by the feeling of being unable to breathe associated with gasping for air, choking and/or repetitive swallowing. Additional seizure types in these individuals were consistent with a temporal or insular/opercular onset (Table 1). Individual III-14 had undergone pre-surgical work-up including ictal subtraction SPECT and PET which was most consistent with a left temporal onset (Online Resource 2). A routine EEG in her sister III-11 had shown a single right temporal sharp transient that was not definitively epileptiform (Online Resource 3). MRI in both patients did not reveal an epileptogenic lesion.
Epilepsy severity was variable in the family. The two individuals with DEE and individual III-14 with focal epilepsy had severe drug-resistant epilepsy. The latter had frequent focal aware and impaired awareness seizures and rare focal to bilateral tonic–clonic seizures despite a combination of three anti-seizure medications. Individual III-11 had less severe epilepsy with focal aware seizures and focal to bilateral tonic–clonic seizures only from sleep on a monotherapy with carbamazepine. Individual II-5 had not been diagnosed with epilepsy previously. She experienced an episode with loss of awareness at age 14 years of uncertain etiology but reported similar “breathing spells” as her daughters which were considered focal aware seizures. She had been treated with phenobarbital for two years after the event at age 14 years but had not taken anti-seizure medication since. The family members with focal epilepsy did not show significant changes in seizure semiology, severity and frequency over time.
Four additional family members had been reported to have epileptic seizures but were unavailable for direct study (Fig. 1). No cardiac or respiratory comorbidities had been identified in the family.
Genetic analysis identified a heterozygous conserved (PhyloP and GERP + +) missense variant with a z-score > 1 (3.52) exhibiting an autosomal dominant inheritance pattern in KCNT1 (g.138675910; NM_020822.2:c.2882G > A; pArg961His). The variant was predicted to be damaging (MSA-SIFT and MSA-PolyPhen2) but has not, to our knowledge, been functionally evaluated yet. The variant was not present in control cohorts including the Genome Aggregation Database (gnomAD) [17] and had been reported previously in seven patients with DEE and SHE [3, 7, 18, 19]. Based on clinical and family history, previous reports, allele frequency and in silico predictors, we classified the variant as likely pathogenic (PS1, PM2, PP1, PP3) using the ACMG recommendations [14]. This is consistent with previous ClinVar reports.
A review of variants of uncertain significance identified no other candidate gene that was related to the phenotype using ACMG criteria [14]. Particularly, there was no evidence for a second contributing variant in individual IV-5 as an explanation for the more severe phenotype.
Discussion
This multigenerational family segregating a KCNT1 pathogenic variant shows significant variability in epilepsy severity and epilepsy syndrome. Intra-familial variability has been well described for other genes such as SCN1A ranging from simple febrile seizures to Dravet syndrome [20]. However, only two KCNT1 families have been described previously with differing epilepsy severity between family members, ranging from the more severe EIMFS to the generally less severe SHE [8, 10]. Specific single variants may also be associated with either EIMFS or SHE in unrelated individuals [6, 7, 12].
Most affected individuals with KCNT1 pathogenic variants reported so far have been either consistent with DEE/EIMFS or SHE. One patient was reported with overlapping features of both conditions [11]. A recent study described a patient with drug-resistant bilateral temporal lobe epilepsy confirmed by invasive pre-surgical evaluation [7]. In others, the available phenotypic information was insufficient to assign a clear phenotype. The latter includes a patient with nocturnal focal epilepsy without further published details on seizure semiology and EEG findings [8], a patient with multifocal epilepsy who also had anti-GAD65 antibodies [8] and a patient with diurnal bilateral tonic–clonic seizures with normal EEG and MRI [7]. Additional described phenotypes are myoclonic atonic epilepsy [6] and temporal lobe epilepsy with intellectual disability and ataxia [21]. However, in the last case, pathogenicity of the underlying variant is questionable as 23 heterozygotes are reported in gnomAD. A consistent feature was cognitive regression after seizure onset [7].
Our family is unique as it includes one individual (II-5) who had mild epilepsy with only focal aware seizures and one single event with loss of awareness. All three individuals with focal epilepsy had seizures that were not consistent with SHE or nocturnal frontal lobe epilepsy based on seizure semiology, routine EEG, video-EEG monitoring and functional imaging. In addition, none of the family members with focal epilepsy had intellectual disability, although we cannot exclude subtle cognitive dysfunction based on the neuropsychological testing in individual III-14 with severe drug-resistant epilepsy.
Our findings therefore expand the phenotypic spectrum associated with KCNT1 pathogenic variants and show that KCNT1 can also cause mild focal epilepsy without cognitive regression. The strength of evidence that variation in a particular gene is causative for a specific disease or seizure type (i.e., the concept of gene–disease validity) is critical for variant interpretation [22]. Therefore, our finding that KCNT1 variants are causative for mild focal epilepsy means that KCNT1 variants that may have previously been disregarded in this context should now be considered. Although we cannot exclude that the association with mild extra-frontal epilepsy may be a feature specific for the p.Arg961His variant, we suspect that other KCNT1 variants may show similar features. It is possible that preferential genetic workup in more severe epilepsies has precluded the identification of milder phenotypes in the past. It is also possible that inherited variants that may have been disregarded previously should be reconsidered given further evidence that intra-familial variability can be quite striking for KCNT1-related epilepsy.
The identified KCNT1 c.2882G > A, p.Arg961His variant has been reported previously in seven patients [3, 7, 18, 19]. Three of these individuals had DEE (1 de novo, 1 inherited and 1 unknown inheritance) and four had SHE (3 de novo, 1 inheritance unknown). For two individuals with DEE, detailed phenotypic information was reported, with onset within the first 3 months of life and multiple seizures daily [3, 7]. The phenotype of these reported individuals was comparable to the two family members with DEE in our family (III-13, IV-5). However, individual IV-5 had typical features for EIMFS which had not been described in the previously reported patients. Detailed information was also reported on three patients with SHE due to the Arg961His variant [7]. These individuals had onset between 6 and 11 years and mild to moderate intellectual disability. They were different to the family members with focal epilepsy in our family (II-5, III-11, III-14), who had later seizure onset (10–18 years) with no typical features of SHE and no intellectual disability.
An interesting observation was that the three family members with focal epilepsy all reported focal sensory aware seizures with unique semiology which they had termed “breathing spells”. These typically occurred in clusters at night and were associated with a feeling of being unable to breathe associated with gasping for air, choking and/or repetitive swallowing. The described semiology may suggest an involvement of the insular or opercular cortex in the seizures although this could not be substantiated with EEG data. However, it is consistent with a previous report of insular onset of SHE [23] of which one patient was later found to have a KCNT1 pathogenic variant [11].
The significant variability in severity and epilepsy syndrome within our and other families suggests that additional factors influence the phenotype. This may include modifier genes but also environmental factors. We did not identify a second contributing rare variant in the most severely affected individual, but it may well be possible that common variants, or rare variants in other so far unknown genes, may modify epilepsy severity. Identification of these factors would significantly advance our understanding of the pathophysiology in genetic epilepsies. However, this remains challenging as the few individuals known with similar variants preclude statistical comparisons.
A limitation of our study is that not all affected family members participated in the research study and detailed EEG and MRI data were unavailable in two family members including one deceased individual. Although extra-frontal onset was highly suspected based on semiology and non-invasive data, it could not definitely be confirmed without intracranial EEG evaluation.
In conclusion, we report a new KCNT1 family with variable phenotype including individuals with severe DEE and mild focal epilepsy suggesting an extra-frontal onset. This family shows that the phenotype associated with KCNT1 pathogenic variants is broader than previously assumed. This is an important observation for variant interpretation as it indicates that KCNT1 variants may be responsible for mild focal epilepsy. Identified KCNT1 variants in patients with mild focal epilepsy should, therefore, assessed according to ACMG criteria [14] and not excluded based on gene–disease validity. This is particularly important as quinidine may be a precision medicine treatment for KCNT1-related epilepsy [24] although the outcome has been variable [25, 26].
Availability of data and material
All de-identified data is included in the article.
Code availability
Not applicable.
References
Heron SE, Smith KR, Bahlo M et al (2012) Missense mutations in the sodium-gated potassium channel gene KCNT1 cause severe autosomal dominant nocturnal frontal lobe epilepsy. Nat Genet 44:1188–1190. https://doi.org/10.1038/ng.2440
Barcia G, Fleming MR, Deligniere A et al (2012) De novo gain-of-function KCNT1 channel mutations cause malignant migrating partial seizures of infancy. Nat Genet 44:1255–1259. https://doi.org/10.1038/ng.2441
Borlot F, Abushama A, Morrison-Levy N et al (2020) KCNT1-related epilepsy: an international multicenter cohort of 27 pediatric cases. Epilepsia 61:679–692. https://doi.org/10.1111/epi.16480
Ohba C, Kato M, Takahashi N et al (2015) De novo KCNT1 mutations in early-onset epileptic encephalopathy. Epilepsia 56:e121–e128. https://doi.org/10.1111/epi.13072
McTague A, Appleton R, Avula S et al (2013) Migrating partial seizures of infancy: expansion of the electroclinical, radiological and pathological disease spectrum. Brain : a journal of neurology 136:1578–1591. https://doi.org/10.1093/brain/awt073
Routier L, Verny F, Barcia G et al (2019) Exome sequencing findings in 27 patients with myoclonic-atonic epilepsy: is there a major genetic factor? Clin Genet 96:254–260. https://doi.org/10.1111/cge.13581
Bonardi CM, Heyne HO, Fiannacca M et al (2021) KCNT1-related epilepsies and epileptic encephalopathies: phenotypic and mutational spectrum. Brain. https://doi.org/10.1093/brain/awab219
Møller RS, Heron SE, Larsen LHG et al (2015) Mutations in KCNT1 cause a spectrum of focal epilepsies. Epilepsia 56:e114–e120. https://doi.org/10.1111/epi.13071
Rubboli G, Plazzi G, Picard F et al (2019) Mild malformations of cortical development in sleep-related hypermotor epilepsy due to KCNT1 mutations. Ann Clin Transl Neurol 6:386–391. https://doi.org/10.1002/acn3.708
Barcia G, Chemaly N, Kuchenbuch M et al (2019) Epilepsy with migrating focal seizures: KCNT1 mutation hotspots and phenotype variability. Neurol Genet 5:e363. https://doi.org/10.1212/NXG.0000000000000363
Cataldi M, Nobili L, Zara F et al (2019) Migrating focal seizures in autosomal dominant sleep-related hypermotor epilepsy with KCNT1 mutation. Seizure 67:57–60. https://doi.org/10.1016/j.seizure.2019.02.019
Licchetta L, Pippucci T, Baldassari S et al (2020) Sleep-related hypermotor epilepsy (SHE): contribution of known genes in 103 patients. Seizure 74:60–64. https://doi.org/10.1016/j.seizure.2019.11.009
Kerr M, Hume S, Omar F et al (2020) MITO-FIND: a study in 390 patients to determine a diagnostic strategy for mitochondrial disease. Mol Genet Metab 131:66–82. https://doi.org/10.1016/j.ymgme.2020.08.009
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17:405–424. https://doi.org/10.1038/gim.2015.30
Köhler S, Carmody L, Vasilevsky N et al (2019) Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources. Nucleic Acids Res 47:D1018–D1027. https://doi.org/10.1093/nar/gky1105
Singleton MV, Guthery SL, Voelkerding KV et al (2014) Phevor combines multiple biomedical ontologies for accurate identification of disease-causing alleles in single individuals and small nuclear families. Am J Human Genet 94:599–610. https://doi.org/10.1016/j.ajhg.2014.03.010
Genome Aggregation Database Consortium, Karczewski KJ, Francioli LC et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581:434–443. https://doi.org/10.1038/s41586-020-2308-7
Zhu X, Padmanabhan R, Copeland B et al (2017) A case-control collapsing analysis identifies epilepsy genes implicated in trio sequencing studies focused on de novo mutations. PLoS Genet 13:e1007104. https://doi.org/10.1371/journal.pgen.1007104
Parrini E, Marini C, Mei D et al (2017) Diagnostic targeted resequencing in 349 patients with drug-resistant pediatric epilepsies identifies causative mutations in 30 different genes. Hum Mutat 38:216–225. https://doi.org/10.1002/humu.23149
Scheffer IE, Zhang Y-H, Jansen FE, Dibbens L (2009) Dravet syndrome or genetic (generalized) epilepsy with febrile seizures plus? Brain Develop 31:394–400. https://doi.org/10.1016/j.braindev.2009.01.001
Hansen N, Widman G, Hattingen E et al (2017) Mesial temporal lobe epilepsy associated with KCNT1 mutation. Seizure 45:181–183. https://doi.org/10.1016/j.seizure.2016.12.018
Rehm HL, Berg JS, Brooks LD et al (2015) ClinGen–the clinical genome resource. N Engl J Med 372:2235–2242. https://doi.org/10.1056/NEJMsr1406261
Proserpio P, Cossu M, Francione S et al (2011) Insular-opercular seizures manifesting with sleep-related paroxysmal motor behaviors: a stereo-EEG study. Epilepsia 52:1781–1791. https://doi.org/10.1111/j.1528-1167.2011.03254.x
Milligan CJ, Li M, Gazina EV et al (2014) KCNT1 gain of function in 2 epilepsy phenotypes is reversed by quinidine. Ann Neurol 75:581–590. https://doi.org/10.1002/ana.24128
Mullen SA, Carney PW, Roten A et al (2018) Precision therapy for epilepsy due to KCNT1 mutations: a randomized trial of oral quinidine. Neurology 90:e67–e72. https://doi.org/10.1212/WNL.0000000000004769
Mikati MA, Jiang YH, Carboni M et al (2015) Quinidine in the treatment of KCNT1-positive epilepsies. Ann Neurol 78:995–999. https://doi.org/10.1002/ana.24520
Acknowledgements
We thank the family for participation in our study. The study was supported by the Department of Clinical Neurosciences, Hotchkiss Brain Institute and Alberta Children’s Hospital Research Institute, University of Calgary, to KMK.
Funding
The study was supported by the Department of Clinical Neurosciences, Hotchkiss Brain Institute and Alberta Children’s Hospital Research Institute, University of Calgary, to KMK.
Author information
Authors and Affiliations
Contributions
CC: analyzed the data; drafted the manuscript for intellectual content. JPA: Designed and conceptualized study; analyzed the data; revised the manuscript for intellectual content. Setareh Ashtiani: Major role in the acquisition of data; revised the manuscript for intellectual content. PF: major role in the acquisition of data; analyzed the data; revised the manuscript for intellectual content. CPM: major role in the acquisition of data; analyzed the data; revised the manuscript for intellectual content. MK: major role in the acquisition of data; analyzed the data; revised the manuscript for intellectual content. AK: major role in the acquisition of data; analyzed the data; revised the manuscript for intellectual content. P-YBA: designed and conceptualized study; analyzed the data; revised the manuscript for intellectual content. KMK: designed and conceptualized study; analyzed the data; drafted the manuscript for intellectual content.
Corresponding author
Ethics declarations
Conflicts of interest
C. Cherian reports no disclosures relevant to the manuscript. J.P. Appendino reports personal and educational fees from UCB Pharma, PendoPharm Inc., and Sunovion Pharmaceuticals Inc. S. Ashtiani reports no disclosures relevant to the manuscript. P. Federico reports no disclosures relevant to the manuscript. C.P. Molnar reports no disclosures relevant to the manuscript. A. Khan and M. Kerr received part of the funding for the MITO-FIND project from MITOCANADA through the Alberta Children’s Hospital Research Foundation. P. Y. B. Au reports no disclosures relevant to the manuscript. K.M. Klein reports personal fees from UCB Pharma, Novartis Pharma AG, Eisai, and GW Pharmaceuticals, grants from the federal state Hessen, Germany, through the LOEWE program and from the Canadian Institutes of Health Research.
Ethical approval
The study was approved by the Conjoint Health Research Ethics Board at University of Calgary and have therefore been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments.
Consent to participate
All participants or their legal guardians provided written informed consent prior to inclusion in the study.
Consent for publication
The consent includes publication of de-identified health information.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cherian, C., Appendino, J.P., Ashtiani, S. et al. The phenotypic spectrum of KCNT1: a new family with variable epilepsy syndromes including mild focal epilepsy. J Neurol 269, 2162–2171 (2022). https://doi.org/10.1007/s00415-021-10808-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-021-10808-y