Abstract
The use of messenger RNA (mRNA) profiling is considered a promising method in the identification of forensically relevant body fluids which can provide crucial information for reconstructing a potential crime. However, casework samples are usually of limited quantity or have been subjected to degradation, which requires improvement of body fluid identification. Circular RNAs (circRNAs), a class of products from the backsplicing of pre-mRNAs, are shown to have high abundance, remarkable stability, and cell type-specific expression in human cells. In this study, we investigated whether the inclusion of circRNAs in mRNA profiling improve the detection of biomarkers including δ-aminolevulinate synthase 2 (ALAS2) and matrix metallopeptidase 7 (MMP7) in body fluid identification. The major circRNAs of ALAS2 and MMP7 were first identified and primer sets for the simultaneous detection of linear and circular transcripts were developed. The inclusion of circRNAs in mRNA profiling showed improved detection sensitivity and stability of biomarkers revealed by using serial dilutions, mixed samples, and menstrual bloodstains as well as degraded and aged samples. Therefore, the inclusion of circRNAs in mRNA profiling should facilitate the detection of mRNA markers in forensic body fluid identification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The identification of body fluids in the forensic context can be important, which helps establish a link between crime scenes and criminal acts. Traditionally, enzymatic, immunological, or chemical detection tests are routinely performed to identify the biological origin of body fluids. However, most conventional tests have disadvantages, such as a lack of specificity and an inability to discriminate mixed body fluids [1]. Therefore, various kinds of biomarkers were introduced to enhance the identification of body fluids, such as tissue-specific messenger RNAs (mRNAs) [2, 3] and microRNAs [4, 5], methylation markers of genomic DNA [6, 7], and microbial markers [8, 9]. As a promising method, mRNA profiling benefits from the simultaneous DNA/RNA coextraction and the detection of a set of biomarkers in one polymerase chain reaction (PCR) [10, 11]. In recent years, tissue-specific mRNA markers have been intensively investigated and multiplex assays have been developed [12, 13], which advances the identification of body fluids.
Casework samples are usually of limited quantity or have been subjected to degradation, which represents a significant challenge in forensic body fluid identification. To this end, sensitive and stable biomarkers have been investigated, such as microRNAs. MicroRNAs are thought to be less prone to degradation because of their small size and the stabilizing effect of Argonaute proteins [5, 14, 15]. In mRNA profiling, technical improvements and the utilization of stable regions of transcripts as targets can facilitate the detection of biomarkers, which helps improve the body fluid identification of trace and degraded samples [16,17,18]. Technical improvements, such as the increased recovery of transcripts and the purification of PCR products, indeed do not increase intrinsic transcripts of biomarkers [16, 17]. Although the use of degraded RNA transcript stable regions (StaRs) as targets was recently shown to enhance the body fluid identification of degraded samples, a small number of biomarkers used in forensic body fluid identification were identified to harbor StaRs [18]. Therefore, the improvement in the sensitivity and stability of biomarkers still requires further investigation.
Recently, circular RNAs (circRNAs), a class of noncoding RNA molecules, were discovered to be abundantly expressed in human cells [19,20,21,22]. These circular transcripts share exons with their linear counterparts and are generally formed by alternative backsplicing of pre-mRNA, in which a downstream 3′ end of an exon is covalently linked with an upstream 5′ end of an exon [23,24,25]. The closed circular structure of circRNAs confers them remarkable stability and resistance to the treatment of RNase R, an exonuclease that degrades linear RNA molecules [19, 21]. The expression of circRNAs is shown to be tissue and cell-type specific although the regulation of circRNA expression remains to be further clarified [19, 26]. The compelling features of circRNAs including high abundance, remarkable stability, and tissue-specific expression enable them to be promising targets in the detection of biomarkers.
Since pre-mRNAs from a single gene can be alternatively spliced into linear and circular transcripts, we postulated that the inclusion of circRNAs in mRNA profiling could enhance the detection sensitivity and stability of biomarkers in forensic body fluid identification. In this study, we investigated the circular transcripts of peripheral and menstrual blood-specific biomarkers and evaluated the sensitivity and stability of biomarkers with the inclusion of circRNAs in mRNA profiling.
Materials and methods
Preparation of samples
All fresh body fluid samples were collected from healthy volunteers. Peripheral blood samples were collected by venipuncture into tubes containing an anticoagulant. Each bloodstain was prepared by pipetting 100 μl of peripheral blood on a piece of sterile gauze. For environmental challenge, bloodstains were placed under a clump of bushes which provided a humid condition and the protection from loss, for 1, 4, 8, 13, and 18 days, respectively, and three samples were collected every period. The average ambient temperature was approximately 10 with 17 °C for the highest and 2 °C for the lowest. Freshly ejaculated semen in sterile tubes was used to prepare dried stains on sterile filter papers at room temperature. Aged peripheral bloodstains from different donors on sterile filter papers were stored at room temperature in a dark, dust-free, and non-humid place for approximately 6 to 7 years. Menstrual blood samples on sanitary napkins were collected from the first to the third day of the menstrual cycle and dried at room temperature followed by preparation as 1 × 1 cm2 pieces. For artificial thermal treatment to eliminate RNA, menstrual bloodstains in sterile tubes were placed at 37 °C for 1, 4, 8, 13, and 18 days, respectively, and three samples were collected every period. After collection, entire stains were cut into pieces followed by RNA extraction or storage at − 80 °C before use. In this study, none of the samples was treated with RNA stabilization reagents. All procedures were approved by the ethics committee of Shanghai Medical College, Fudan University, and all donors volunteered for this study based on informed consent.
RNA preparation
Total RNAs were isolated in a final volume of 40 μl of elution buffer using an RNeasy® Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions and were quantified using a NanoDrop ND-2000 spectrophotometer (Thermo Scientific, USA). The removal of potential genomic DNA was performed with RNase-Free DNase Set (Qiagen, Germany) according to the manufacturer’s instructions. The removal of linear transcripts was carried out with the treatment of RNase R (Epicentre, USA) at 37 °C for 4 h with a dose of 3 U per 1 μg of RNA [21]. In the sensitivity test, the indicated dilution series of input total RNA were prepared followed by reverse transcription. The mixture of total RNA from peripheral or menstrual blood with that from semen stains was prepared at a concentration ratio of 1:100 followed by reverse transcription.
Reverse transcription
Random hexamers were used to synthesize cDNA in a final volume of 20 μL using a Transcriptor First Stand cDNA Synthesis Kit (Roche, USA) according to the manufacturer’s instructions and each reaction contained 1 ng, the indicated amounts, or the indicated volumes of total RNA. Reverse transcription (RT) minus controls without reverse transcriptase were used to rule out potential contamination of genomic DNA.
Primer design
The mRNA markers for peripheral and menstrual blood were acquired from previous reports [27, 28]. The circRNAs of δ-aminolevulinate synthase 2 (ALAS2) and hemoglobin alpha (HBA) were obtained by the analysis of previous sequencing data with TopHat-Fusion and CIRCexplorer [23, 26]. Primer sets were designed using Primer Premier v5.0 (Premier Biosoft, USA) [29]. Outward-facing primer sets, of which the binding sites of forward primers in the coding sequence are located downstream from the binding sites of reverse primers (Fig. S1A), were developed for detecting circular transcripts of ALAS2, HBA, matrix metalloproteinase 7 (MMP7), and matrix metalloproteinase 11 (MMP11). Conventional primer sets, of which the binding sites of forward primers in the coding sequence are located upstream from the binding sites of reverse primers, were developed based on exons shared by linear and circular transcripts (LC-primers, Fig. S1B) for simultaneously detecting linear and circular transcripts of ALAS2 or MMP7. Reported primer sets for the detection of ALAS2 and MMP7 were acquired from previous reports [28, 30]. Reported primer sets appeared to mainly amplify linear transcripts after the investigation of circRNAs of ALAS2 and MMP7 and thus were designed as L-primers in this study. For obtaining products of full-length coding sequences of ALAS2 and MMP7, primer sets, designed as FL-primers, were developed. FAM-labeled or unlabeled primer sets were synthesized in Sangon Biotech., Shanghai. The primer sequences and concentrations used in this study as well as product sizes are listed in Table S1.
PCR amplification
In singleplex PCR, 25 μl of reaction mix contained 1 μl of cDNA or equal amount of sterile water as the non-template control, 2 μl of dNTPs (Takara, Japan), 2.5 μl of 10× PCR buffer, and 1 U AmpliTaq Gold DNA polymerase (Thermofisher, USA). After an initial denaturation at 94 °C for 2 min in a Mastercycler® nexus GSX1 (Eppendorf, USA), amplification was carried out at the following conditions: denaturation at 94 °C for 30 s, annealing at the temperature indicated in Table S1, extension at 72 °C for 1 min, and a final extension at 72 °C for 10 min. The outward-facing primer sets and 34 cycles of PCR amplification were used for the detection of circular transcripts. FL-primers and 31 cycles of PCR amplification were used for obtaining products of full-length coding sequences. LC-primers or L-primers as well as 28 cycles of PCR amplification were used for characterizing the mRNA profiling or the evaluation of primer sets by capillary electrophoresis (CE). PCR products of circRNAs were separated by agarose gel electrophoresis (AGE) followed by purification and the junctions of head-to-tail products were determined by Sanger sequencing. PCR products of full-length coding sequences were separated by AGE followed by purification and quantification.
Capillary electrophoresis and profile analysis
PCR products obtained using LC-primers and L-primers were detected by CE using POP-7 on an ABI PRISM 3130xL Genetic Analyzer (Thermofisher, USA). Briefly, 1 μl of PCR products, 9 μL of HiDi formamide (Thermofisher, USA), and 1 μL of CC5 Internal Lane Standard 500 (Promega, USA) were mixed. Then, samples were denatured at 95 °C for 5 min followed by snap cooling on ice for 5 min. The electrophoresis conditions included a 10-s injection time, 2-kV injection voltage, 15-kV run voltage, and 30-min run time at 60 °C. Profiles were analyzed using GeneMapper ID software v3.2 and signals equal to or above 100 RFU were interpreted as positive results. Negative controls did not show amplification signals. For samples with negative results in the sensitivity and stability analysis, PCR amplification using LC-primers and L-primers were replicated followed by CE. Biomarkers were considered to have positive expression if signals were equal to or above 100 RFU.
Evaluation of primer sets
The copy numbers of full-length ALAS2 and MMP7 per microliter of elution buffer were calculated based on the concentration quantified using a NanoDrop ND-2000 spectrophotometer and the molecular weights (325 Da as the average molecular weight of dNMP), followed by the preparation of the dilution series of full-length ALAS2 and MMP7 DNA. The efficiency of LC-primers and L-primers in PCR amplification was compared by PCR-CE assay and quantitative PCR (qPCR). qPCR was performed using QuantiNova™ SYBR® Green PCR Kit (Qiagen, Germany) in a final reaction volume of 10 μl. The reaction mix with LC-primers or L-primers was run on an ABI Prism 7500 fluorescence quantitative PCR instrument (Thermofisher, USA) according to the conditions consisting of an initial denaturation at 95 °C for 2 min, followed by 40 cycles of 5 s at 95 °C, and 34 s at 60 °C. The Ct values were calculated using SDS software with an automatic baseline and a threshold of 0.2. Then, standard curves were constructed.
Results
Identification of circRNAs of peripheral and menstrual blood markers
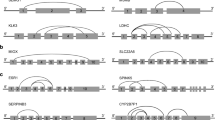
Analysis of previous sequencing data showed that ALAS2 and HBA, two peripheral blood-specific biomarkers, had circular transcripts (Fig. 1a and Fig. S2A). Electrophoresis of PCR products from amplification using outward-facing primer sets of ALAS2 and HBA showed clear bands in all four samples (Fig. S2B and C), which validates the expression of circular transcripts of ALAS2 and HBA. The covalent junction in circular transcripts of ALAS2 and HBA was further revealed by sequencing (Fig. 1b, Fig. S2D and E).
Identification of ALAS2 and MMP7 circRNAs. a Three putative circRNAs of ALAS2 revealed by bioinformatics analysis are indicated by the triangles. The arrows indicate the binding sites of primers and the asterisk indicates the transcript identified in (b). b The junction sequence of the head-to-tail ALAS2 circRNA detected in Fig. S2B. The arrow indicates the junction site. c Two identified circRNAs of MMP7 by sequencing PCR products in this study. The arrows indicate the binding sites of primers. The asterisk indicates the transcript identified in (d) and the cross indicates the circRNA with a low expression level. d The junction sequence of the head-to-tail MMP7 circRNA detected in Fig. S2F. The arrow indicates the junction site
The expression of circRNAs from two menstrual blood-specific biomarkers, including MMP7 and MMP11, was subsequently investigated using outward-facing primer sets. As shown in Fig. S2F, PCR products from circular transcripts of MMP7 could be clearly observed in all four samples. The covalent junction in the circular transcript of MMP7 was further revealed by sequencing (Fig. 1c, d). Additionally, other outward-facing primer sets based on the combination of exons were used to further determine potential circular transcripts of MMP7. Results from endpoint PCR and sequencing revealed another circular transcript of MMP7 that might have a low expression level (Fig. 1c and data not shown). In contrast, we detected no products from the backsplicing of MMP11 pre-mRNA using a set of outward-facing primer sets based on the combination of exons (data not shown).
Evaluation of primer sets in the detection of biomarkers
ALAS2 and MMP7 were used for the next investigation in this study and HBA was excluded due to its high level of expression. LC-primers were used for simultaneous detection of linear and circular transcripts of ALAS2 and MMP7 (Fig. 2a, b). The investigation of circRNAs of ALAS2 and MMP7 revealed that L-primers from previous reports mainly amplify linear transcripts (Fig. 2a, b). The PCR amplification using LC-primers and L-primers was carried out on the dilution series of full-length double-stranded DNA of ALAS2 and MMP7. Although the amplification using L-primers of ALAS2 seemed to be lower than that with LC-primers of ALAS2, the amplification using L-primers of ALAS2 on one copy could obtain significantly detectable signals in CE (Fig. 2c). In fact, results from the dissociation curve of L-primers of ALAS2 might imply the generation of primer dimers in qPCR (Fig. S3A and B). Compared with amplification using L-primers of MMP7, the lower amplification signals using LC-primers of MMP7 might be due to longer PCR products in endpoint PCR (Fig. 2d). In qPCR, the amplification efficiencies of L-primers and LC-primers of MMP7 were comparable (Fig. S3B).
Evaluation of primer sets in the detection of biomarkers. a The horizontal lines cover the regions L-primers (lower) and LC-primers (upper) of ALAS2 amplify. b The horizontal lines cover the regions L-primers (lower) and LC-primers (upper) of MMP7 amplify. c Detection of ALAS2 using L-primers and LC-primers of ALAS2 by PCR-CE on the indicated copy number of double-stranded DNA. Data represent mean ± SD from four replicate samples. d Detection of MMP7 using L-primers and LC-primers of MMP7 by PCR-CE on the indicated copy number of double-stranded DNA. Data represent mean ± SD from four replicate samples
The detection sensitivity of biomarkers with the inclusion of circRNAs
Sensitivity testing of ALAS2 and MMP7 was assessed in a quantitative approach (input total RNA). A dilution series of manually extracted total RNA from different individuals (0.2–0.003 ng) was reversely transcribed. PCR amplification was performed using L-primers and LC-primers, respectively, and results are summarized in Table 1. The detection limits of ALAS2 and MMP7 determined using L-primers were up to 0.012 and 0.025 ng of input total RNA, respectively (Table 1). In contrast, ALAS2 and MMP7 could be detectable with as little as 0.003 ng of input total RNA using LC-primers in all tested samples (Table 1). Additionally, in contrast to L-primers, the amplification using LC-primers on mixtures of total RNA from peripheral or menstrual blood with that from semen at a concentration ratio of 1:100 produced considerably higher peaks (Fig. 3a, b; Fig. S4A and B). This is probably correlated with a higher number of target molecules by the inclusion of circRNAs.
The detection sensitivity of ALAS2 and MMP7 with the inclusion of circRNAs. (A, B) L-primers and LC-primers of ALAS2 (a) and MMP7 (b) were used, respectively, for PCR amplification on the mixtures of total RNA from peripheral (a) or menstrual (b) blood with that from semen at a ratio of 1:100. The electrophoretogram represents one out of triplicate samples. c One representative electrophoretogram from Table 2 and the arrow indicates the low amplification signal of MMP7 using L-primers
Previous studies have shown that the mRNA expression level of MMP7 fluctuates over the days of menstruation in individuals [31]. Results from amplification using L-primers showed that the mRNA expression of MMP7 was negative in some samples (Table 2 and Fig. 3c). Although the simultaneous detection of linear and circular transcripts showed a low expression of MMP7 in linear mRNA-negative samples, the detection of MMP7 could be considered as positive results in mRNA profiling (Table 2 and Fig. 3c), which might result from the increase of targets by the inclusion of circular transcripts.
The stability of biomarkers with the inclusion of circRNAs
To determine the stability of biomarkers with the inclusion of circRNAs in the detection of artificially degraded samples, total RNA from fresh stains were treated with RNase R followed by reverse transcription. PCR amplification using RT products of intact total RNA as templates showed significantly detectable amplification signals of ALAS2 and MMP7 (Fig. 4a, b). However, treatment with RNase R resulted in undetectable amplicons of ALAS2 and MMP7 when amplification was performed using L-primers (Fig. 4c, d). In contrast, amplification using LC-primers demonstrated detectable amplicons of ALAS2 and MMP7 in RNase R-treated samples (Fig. 4c, d).
The stability of ALAS2 and MMP7 with the inclusion of circRNAs. a, b The detection of ALAS2 in peripheral bloodstains (a) and MMP7 in menstrual bloodstains (b) using L-primers or LC-primers. c, d The detection of ALAS2 in RNase R-treated total RNA from peripheral bloodstains (c) and MMP7 in RNase R-treated total RNA from menstrual bloodstains using L-primers or LC-primers (d). Reverse transcription was performed using 2 ng of total RNA treated with RNase R and 1 μl of RT products was used for 28 cycles of PCR amplification. e, g The detection of ALAS2 in 13-day-old (e) and 18-day-old (f) peripheral bloodstains using L-primers or LC-primers. Reverse transcription was performed using 4 μl of total RNA and 1 μl of RT products was used for 28 cycles of PCR amplification. f, h The detection of MMP7 in 13-day-old (f) and 18-day-old (h) menstrual bloodstains using L-primers or LC-primers. Reverse transcription was performed using 4 μl of total RNA and 1 μl of RT products was used for 28 cycles of PCR amplification. Each electrophoretogram represents one out of three samples. The font of scales was enlarged due to the low resolution of original figures
Peripheral bloodstains were exposed to environmental conditions and menstrual bloodstains were subjected to artificial thermal treatment for different periods. In 1- to 8-day-old samples, amplification using L-primers and LC-primers could obtain detectable amplicons of ALAS2 and MMP7 (data not shown). In 13- and 18-day-old samples, weak or no amplification signals of ALAS2 (Fig. 4e, g; Fig. S5A and B) and MMP7 (Fig. 4f, h; Fig. S5C and D) were observed when performing amplification using L-primers. In contrast, there were detectable amplicons of ALAS2 (Fig. 4e, g; Fig. S5A and B) and MMP7 (Fig. 4f, h; Fig. S5C and D) in 13- and 18-day-old samples when performing amplification using LC-primers.
Aged dried bloodstains stored at room temperature were used to compare the detection of ALAS2 using L-primers and LC-primers and results are summarized in Table 3. There were no detectable amplification signals in mRNA profiling when performing amplification using L-primers. However, amplicons could be observed by CE when performing amplification using LC-primers. These results suggest that the inclusion of circRNAs could facilitate the detection of ALAS2 in aged samples.
Discussion
The limited quantity and degradation of casework samples are challenging the current methods for forensically relevant body fluid identification. The development of sensitive and stable biomarkers as well as technical improvements in the detection of biomarkers can facilitate the identification of body fluids. As circRNAs and mRNAs are products of pre-mRNA splicing, the backsplicing of pre-mRNAs indeed decreases the expression level of mRNAs in cells. Although not all biomarkers include the expression of circRNAs, the discard of stable transcripts in the detection of biomarkers with the expression of circRNAs might underestimate the sensitivity and stability of biomarkers. Published primer sets do not appear to amplify the major circRNAs of ALAS2 and MMP7 identified in this study [27, 28, 30, 32,33,34,35]. Therefore, the selection of regions as targets requires more attention for improving the detection of biomarkers. In this study, we performed the mRNA profiling with the inclusion of circRNAs in the detection of biomarkers. Results from our investigation on the detection sensitivity and stability of biomarkers using L-primers and LC-primers show that the utility of regions shared by linear and circular transcripts as targets could improve the detection of biomarkers in mRNA profiling.
Pre-mRNA splicing is essential for eukaryotic gene expression, which can be regulated to produce multiple mRNA molecules from a single gene. A number of alternative splicing events are shown to be tissue-specific, which plays an important role in cellular differentiation and organismal development. Similarly, alternative backsplicing of pre-mRNA was recently revealed to generate multiple circRNAs from a single gene within cells [19, 20]. Since the mechanisms of backsplicing remain elusive, it is difficult to predict the combination of exons circRNAs employ, which is crucial to the purposeful design of primer sets in the detection of circRNAs. Currently, the expression profiles of circRNAs in forensically relevant body fluids are not available, but biochemical approaches can be applied to determine circRNAs from genes of interest [29]. In this study, circRNAs of ALAS2 and MMP7 were revealed using outward-facing primer sets for amplification and the head-to-tail junctions of circRNAs were further determined by sequencing. Interestingly, some circRNAs consisted of incomplete exons (Fig. 1d and Fig. S2D). It is possible that the rule of backsplicing is different from that of alterative splicing.
The expression of genes is tightly regulated in cells. As a consequence, most of mRNA molecules have a tissue-specific expression pattern. Previous studies show that the expression of circRNAs is abundant and cell-type specific in thousands of human genes. In some cases, the expression of circRNAs even exceeds the abundance of traditional linear transcripts [20]. However, the expression levels of different circRNAs from a single gene might be regulated in cells since results from endpoint PCR demonstrated one major circular transcript of ALAS2 and MMP7, respectively (Fig. S2B and C). In fact, different expression levels of circRNAs from a single gene were observed in previous reports [19, 23]. Furthermore, the backsplicing of circRNAs is shown to compete with pre-mRNA splicing [36]. Therefore, simultaneous detection of linear and circular transcripts indeed fully utilizes transcripts of biomarkers in mRNA profiling. Apart from an increased level of targets by the inclusion of circRNAs in mRNA profiling, the detection sensitivity of biomarkers is susceptible to the amplification efficiency of primer sets. As the amplification efficiency of primer sets has a substantial impact on the evaluation of the Ct value in qPCR, it is difficult to conduct a comparison in the detection of biomarkers by qPCR using L-primers and LC-primers. Furthermore, the △Ct value is theoretically small even though circRNAs account for approximately 40% of all transcripts from a single gene, which might result in an inaccurate evaluation. In this study, since the PCR amplification using L-primers and LC-primers on one copy of double-stranded DNA obtained significantly detectable signals in CE, the improvement in the detection sensitivity of biomarkers might be mainly attributed to the increased level of targets. Therefore, the inclusion of circRNAs in mRNA profiling could help in the detection of biomarkers. However, it should be noted that some RNA markers in potential non-target tissues might have quite a weak expression and the inclusion of circRNAs in the detection of these markers might result in a positive interpretation in non-target tissues, which will lower the specificity of biomarkers. Therefore, if circRNAs are included in forensic body fluid identification, the specificity of biomarkers with the expression of circRNAs, such as ALAS2 and MMP7, still requires to be further verified.
Forensic casework samples are often subjected to environmental insults, such as UV, humidity, heat, and enzymes. Apart from the expression level of biomarkers, the stability of transcripts is crucial to the identification of degraded or aged biological stains in mRNA profiling. The structure of RNA molecules and RNA-binding partners might have important roles in the maintenance of their stability in cells [37]. CircRNAs have been shown to have a longer decay half-life than mRNAs in cells, which might be due to the lack of free ends. In this study, our results showed that the inclusion of circRNAs in mRNA profiling could improve the detection of ALAS2 and MMP7 in samples with artificial treatment and environmental insults. Although we did not quantify the remnants of mRNAs and circRNAs in the degraded and aged samples, PCR-CE technology can reveal a difference in the copy numbers of targets since the singleplex PCR amplification using L-primers and LC-primers is similarly efficient. The unsuccessful detection of ALAS2 and MMP7 using L-primers might result from targets with a copy number that is less than the detection limit of mRNA profiling. The improvement in the stability of biomarkers might result from the increased level of targets and the inclusion of stable transcripts.
In conclusion, the inclusion of circular transcripts in mRNA profiling might be an efficient strategy for improving the detection sensitivity and stability of biomarkers in body fluid identification. Subsequent studies should determine the expression profile of circRNAs in all of forensically relevant body fluids and characterize a set of biomarkers with the expression of circRNAs, which might help with the development of more robust multiplex assays for body fluid identification.
References
Virkler K, Lednev IK (2009) Analysis of body fluids for forensic purposes: from laboratory testing to non-destructive rapid confirmatory identification at a crime scene. Forensic Sci Int 188:1–17. https://doi.org/10.1016/j.forsciint.2009.02.013
Fleming RI, Harbison S (2010) The development of a mRNA multiplex RT-PCR assay for the definitive identification of body fluids. Forensic Sci Int Genet 4:244–256. https://doi.org/10.1016/j.fsigen.2009.10.006
Richard ML, Harper KA, Craig RL, Onorato AJ, Robertson JM, Donfack J (2012) Evaluation of mRNA marker specificity for the identification of five human body fluids by capillary electrophoresis. Forensic Sci Int Genet 6:452–460. https://doi.org/10.1016/j.fsigen.2011.09.007
Park JL, Park SM, Kwon OH et al (2014) Microarray screening and qRT-PCR evaluation of microRNA markers for forensic body fluid identification. Electrophoresis 35:3062–3068. https://doi.org/10.1002/elps.201400075
Sauer E, Reinke AK, Courts C (2016) Differentiation of five body fluids from forensic samples by expression analysis of four microRNAs using quantitative PCR. Forensic Sci Int Genet 22:89–99. https://doi.org/10.1016/j.fsigen.2016.01.018
An JH, Choi A, Shin KJ, Yang WI, Lee HY (2013) DNA methylation-specific multiplex assays for body fluid identification. Int J Legal Med 127:35–43. https://doi.org/10.1007/s00414-012-0719-1
Park JL, Kwon OH, Kim JH et al (2014) Identification of body fluid-specific DNA methylation markers for use in forensic science. Forensic Sci Int Genet 13:147–153. https://doi.org/10.1016/j.fsigen.2014.07.011
Giampaoli S, Berti A, Valeriani F et al (2012) Molecular identification of vaginal fluid by microbial signature. Forensic Sci Int Genet 6:559–564. https://doi.org/10.1016/j.fsigen.2012.01.005
Choi A, Shin KJ, Yang WI, Lee HY (2014) Body fluid identification by integrated analysis of DNA methylation and body fluid-specific microbial DNA. Int J Legal Med 128:33–41. https://doi.org/10.1007/s00414-013-0918-4
Haas C, Hanson E, Anjos MJ et al (2012) RNA/DNA co-analysis from blood stains—results of a second collaborative EDNAP exercise. Forensic Sci Int Genet 6:70–80. https://doi.org/10.1016/j.fsigen.2011.02.004
Haas C, Hanson E, Anjos MJ et al (2013) RNA/DNA co-analysis from human saliva and semen stains—results of a third collaborative EDNAP exercise. Forensic Sci Int Genet 7:230–239. https://doi.org/10.1016/j.fsigen.2012.10.011
Lindenbergh A, van den Berge M, Oostra RJ et al (2013) Development of a mRNA profiling multiplex for the inference of organ tissues. Int J Legal Med 127:891–900. https://doi.org/10.1007/s00414-013-0895-7
Song F, Luo H, Hou Y (2015) Developed and evaluated a multiplex mRNA profiling system for body fluid identification in Chinese Han population. J Forensic Legal Med 35:73–80. https://doi.org/10.1016/j.jflm.2015.08.006
Lux C, Schyma C, Madea B, Courts C (2014) Identification of gunshots to the head by detection of RNA in backspatter primarily expressed in brain tissue. Forensic Sci Int 237:62–69. https://doi.org/10.1016/j.forsciint.2014.01.016
Winter J, Diederichs S (2011) Argonaute proteins regulate microRNA stability: increased microRNA abundance by Argonaute proteins is due to microRNA stabilization. RNA Biol 8:1149–1157. https://doi.org/10.4161/rna.8.6.17665
Setzer M, Juusola J, Ballantyne J (2008) Recovery and stability of RNA in vaginal swabs and blood, semen, and saliva stains. J Forensic Sci 53:296–305. https://doi.org/10.1111/j.1556-4029.2007.00652.x
Sirker M, Schneider PM, Gomes I (2016) A 17-month time course study of human RNA and DNA degradation in body fluids under dry and humid environmental conditions. Int J Legal Med 130:1431–1438. https://doi.org/10.1007/s00414-016-1373-9
Lin MH, Albani PP, Fleming R (2015) Degraded RNA transcript stable regions (StaRs) as targets for enhanced forensic RNA body fluid identification. Forensic Sci Int Genet 20:61–70. https://doi.org/10.1016/j.fsigen.2015.09.012
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 7:e30733. https://doi.org/10.1371/journal.pone.0030733
Jeck WR, Sorrentino JA, Wang K et al (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19:141–157. https://doi.org/10.1261/rna.035667.112
Memczak S, Jens M, Elefsinioti A et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338. https://doi.org/10.1038/nature11928
Guo JU, Agarwal V, Guo H, Bartel DP (2014) Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15:409. https://doi.org/10.1186/s13059-014-0409-z
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L (2014) Complementary sequence-mediated exon circularization. Cell 159:134–147. https://doi.org/10.1016/j.cell.2014.09.001
Starke S, Jost I, Rossbach O et al (2015) Exon circularization requires canonical splice signals. Cell Rep 10:103–111. https://doi.org/10.1016/j.celrep.2014.12.002
Chen LL, Yang L (2015) Regulation of circRNA biogenesis. RNA Biol 12:381–388. https://doi.org/10.1080/15476286.2015.1020271
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO (2013) Cell-type specific features of circular RNA expression. PLoS Genet 9:e1003777. https://doi.org/10.1371/journal.pgen.1003777
Haas C, Hanson E, Bar W et al (2011) mRNA profiling for the identification of blood—results of a collaborative EDNAP exercise. Forensic Sci Int Genet 5:21–26. https://doi.org/10.1016/j.fsigen.2010.01.003
Haas C, Hanson E, Kratzer A, Bar W, Ballantyne J (2011) Selection of highly specific and sensitive mRNA biomarkers for the identification of blood. Forensic Sci Int Genet 5:449–458. https://doi.org/10.1016/j.fsigen.2010.09.006
Jeck WR, Sharpless NE (2014) Detecting and characterizing circular RNAs. Nat Biotechnol 32:453–461. https://doi.org/10.1038/nbt.2890
Lindenbergh A, de Pagter M, Ramdayal G et al (2012) A multiplex (m)RNA-profiling system for the forensic identification of body fluids and contact traces. Forensic Sci Int Genet 6:565–577. https://doi.org/10.1016/j.fsigen.2012.01.009
Haas C, Klesser B, Maake C, Bar W, Kratzer A (2009) mRNA profiling for body fluid identification by reverse transcription endpoint PCR and realtime PCR. Forensic Sci Int Genet 3:80–88. https://doi.org/10.1016/j.fsigen.2008.11.003
Juusola J, Ballantyne J (2007) mRNA profiling for body fluid identification by multiplex quantitative RT-PCR. J Forensic Sci 52:1252–1262. https://doi.org/10.1111/j.1556-4029.2007.00550.x
Juusola J, Ballantyne J (2005) Multiplex mRNA profiling for the identification of body fluids. Forensic Sci Int 152:1–12. https://doi.org/10.1016/j.forsciint.2005.02.020
Roeder AD, Haas C (2013) mRNA profiling using a minimum of five mRNA markers per body fluid and a novel scoring method for body fluid identification. Int J Legal Med 127:707–721. https://doi.org/10.1007/s00414-012-0794-3
van den Berge M, Bhoelai B, Harteveld J, Matai A, Sijen T (2015) Advancing forensic RNA typing: on non-target secretions, a nasal mucosa marker, a differential co-extraction protocol and the sensitivity of DNA and RNA profiling. Forensic Sci Int Genet 20:119–129. https://doi.org/10.1016/j.fsigen.2015.10.011
Ashwal-Fluss R, Meyer M, Pamudurti NR et al (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56:55–66. https://doi.org/10.1016/j.molcel.2014.08.019
Houseley J, Tollervey D (2009) The many pathways of RNA degradation. Cell 136:763–776. https://doi.org/10.1016/j.cell.2009.01.019
Acknowledgements
The authors thank Dr. Xiao-Ou Zhang from the Chinese Academy of Sciences in Shanghai for his help with the analysis of circRNAs. This work was supported by the National Natural Science Foundation of China (81571853 and 31270862).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Figure S1
The strategy for the detection of linear and/ or circular transcripts. (A) Outward-facing primer sets (orange arrows) were designed based on exons shared by linear and circular transcripts. As for outward-facing primer sets, the binding sites of forward primers (F) in the coding sequence are located downstream from the binding sites of reverse primers (R), which result in the failure of PCR amplification on linear transcripts. Therefore, outward-facing primer sets were used for the detection of circRNAs in this study. (B) Conventional primer sets were designed based on at least one exon employed only by linear transcripts (L-primers, red arrows) for the detection of linear transcripts. Conventional primer sets were designed based on exons shared by linear and circular transcripts (LC-primers, green arrows) for the detection of linear and circular transcripts. F: forward primers; R: reverse primers. (GIF 13 kb)
Figure S2
(A) One putative circRNAs of HBA identified by bioinformatics analysis is indicated by the triangle and one circular transcript identified in (C) is indicated by the asterisk. (B and C) Detection of circRNAs of ALAS2 (B) and HBA (C) in the peripheral blood using outward-facing primer sets for PCR amplification followed by AGE. (D) The conjunct sequence of the circular HBA transcript identified in (C). The arrow indicates the junction site. (E) A longer conjunct sequence of the ALAS2 circRNA identified in (B) and the arrow indicates the junction site. (F) Detection of circRNAs of MMP7 in the menstrual blood using outward-facing primer sets for PCR amplification followed by AGE. (GIF 83 kb)
Figure S3
(A) Dissociation curve from the real-time PCR assay using L-primers (left) and LC-primers (right) of ALAS2. (B) Determination of the amplification efficiencies using LC-primers of ALAS2 (left) as well as L-primers (middle) and LC-primers (right) of MMP-7 by qPCR. The panels indicate the mathematical relationships of the Ct value and the copy number of full-length DNA. Each point represents the mean of three replicates. (GIF 43 kb)
Figure S5
(A, B) The RFU values from the detection of ALAS2 using L-primers and LC-primers in three 13 day-old (A) and 18 day-old (B) peripheral bloodstains were calculated. (C, D) The RFU values from the detection of MMP7 using L-primers and LC-primers in three 13 day-old (C) and 18 day-old (D) menstrual bloodstains were calculated. (GIF 41 kb)
Table S1
(DOCX 20 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Liu, B., Shao, C. et al. Evaluation of the inclusion of circular RNAs in mRNA profiling in forensic body fluid identification. Int J Legal Med 132, 43–52 (2018). https://doi.org/10.1007/s00414-017-1690-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-017-1690-7