Abstract
Although it is known that deletions or mutations of the SMN1 gene on chromosome 5 cause decreased levels of the SMN protein in subjects with proximal autosomal recessive spinal muscular atrophy (SMA), the exact sequence of pathological events leading to selective motoneuron cell death is not fully understood yet. In this review, new findings regarding the dual cellular role of the SMN protein (translocation of β-actin to axonal growth cones and snRNP biogenesis/pre-mRNA splicing) were integrated with recent data obtained by detailed neuropathological examination of SMA and control subjects. A presumptive series of 10 pathogenetic events for SMA is proposed as follows: (1) deletions or mutations of the SMN1 gene, (2) increased SMN mRNA decay and reduction in full-length functional SMN protein, (3) impaired motoneuron axono- and dendrogenesis, (4) failure of motoneurons to form synapses with corticospinal fibers from upper motoneurons, (5) abnormal motoneuron migration towards ventral spinal roots, (6) inappropriate persistence of motoneuron apoptosis due to impaired differentiation and motoneuron displacement, (7) substantial numbers of motoneurons continuing to migrate abnormally (“heterotopic motoneurons”) and entering into the ventral roots, (8) attracted glial cells following these heterotopic motoneurons, which form the glial bundles of ventral roots, (9) impaired axonal transport of actin, causing remaining motoneurons to become chromatolytic, and (10) eventual death of all apoptotic, heterotopic and chromatolytic neurons, with apoptosis being more rapid and predominating in the earlier stages, with death of heterotopic and chromatolytic neurons occurring more slowly by necrosis during the later stages of SMA. According to this model, the motoneuron axonopathy is more important for pathogenesis than the ubiquitous nuclear splicing deficit. It is also supposed that individually variable levels of SMN protein, together with influences of other phenotype modifier genes and their products, cause the clinical SMA spectrum through differential degree of motoneuron functional loss.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The spinal muscular atrophies (SMAs) are a genetically heterogeneous group of inherited diseases that cause progressive muscle degeneration (Table 1) [20, 44, 55, 88]. In this review, we focus on the most common form, proximal autosomal recessive SMA, which is the second leading cause of neuromuscular disease (after muscular dystrophy) and the second most common autosomal recessive disease (after cystic fibrosis), with an incidence of at least 1 in 10,000 live births and a carrier frequency of 1 in 35 to 1 in 50 [25, 78].
Since the initial description of SMA at the end of nineteenth century by Guido Werdnig, it has been postulated that pathological changes in SMA consist of four major features (the so-called “neuropathologic tetrad”): (1) loss of anterior horn cells (α-motoneurons as well as γ-motoneurons and interneurons), (2) empty cell beds (at locations of neuron loss), (3) glial cell bundles in the ventral spinal roots, and (4) heterotopic motoneurons (HMN) [85]. About a hundred years later, Melki and collaborators revealed that recessive forms of proximal SMA are caused by reduced survival of motor neuron (SMN) protein due to deletions or mutations of the SMN1 gene on chromosome 5 [46]. However, it is still not clear why, and how, reduced levels of the SMN protein cause such neuropathological changes. The answer to this question is obviously related to our understanding of the normal function(s) of the SMN protein and particularities of ventral horn neurons development and function. Here, an overview of novel findings regarding the cellular functions of SMN protein in motoneurons are put in perspective together with a recent detailed description of the neuropathological aspects of SMA [76]. Finally, these data are used to propose a possible pathogenetic model. It is hoped that this will help elucidating the mechanisms responsible for SMA genotype-phenotype relationships and serve for future development of possible treatment and prevention stategies.
Clinical presentation and diagnosis
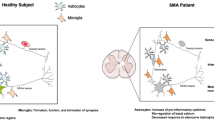
Due to the symmetrical loss of spinal cord anterior horn neurons (Fig. 1a, b), SMA is characterized by progressive denervation of skeletal muscles. The weakness and atrophy are usually first noted for the proximal voluntary muscles of the extremities. During disease progression the distal voluntary muscles of the extremities and eventually entire trunk are also affected. Diagnosis is confirmed by muscle biopsy, which seems to be very useful particularly in case of chronic infantile forms (Fig. 1c, d), electromyography (showing typical “denervation pattern”: abnormal spontaneous activity with fibrillations, positive sharp waves and increased mean duration and amplitude of motor unit action potentials), MRI of the spine and DNA genotyping.
a Normal control spinal cord b SMA spinal cord showing loss of anterior horn cells. Cresyl-violet stain. (c) and (d) Muscle biopsy of biceps brachii muscle from SMA-1 subject showing neurogenic muscle atrophy (c, d) Large groups of circular atrophic muscle fibers (arrowheads) mixed with fascicles of hypertrophied fibers (arrow). Haematoxylin-eosin stain. Scale bars: (a, b) 100μm, (c) 20× magnification, (d) 100× magnification. Photomicrographs (a) and (b) are reproduced from Simic et al. [75] with the permission of Wolters Kluwer Health (Lippincott Williams and Wilkins)
Based on age at onset and severity, SMA is classified in several types that form the clinical spectrum [59, 93]. The clinical symptoms in all types of SMA include hypotonia, symmetrical muscle weakness and atrophy (predominantly of the proximal muscles of the shoulder and pelvic girdle), absence of deep tendon reflexes, tremor of fingers and hands, fasciculation of the tongue muscles and hyporeflexia with contractures of some muscle groups (the diaphragm and extraocular muscles remain unaffected until late stages of the disease and there is little or no impairment of sensory systems) [35]. These symptoms are thought to be due to selective loss of motoneurons in the spinal cord and cranial motor nuclei leading to progressive muscle weakness, atrophy and paralysis [32]. In its most severe type (SMA-1, also called Werdnig-Hoffmann disease or acute SMA, MIM# 253300), onset is usually before 9 months. Most of the children are completely asymptomatic at birth, with symptoms emerging at around 2–4 months after birth. Affected infants are hypotonic (“floppy”) with weak, thin muscles, and display feeding and breathing problems, fail to achieve early motor milestones and are never able to sit. Death occurs within the first 2 years of life, usually due to respiratory failure. If symptoms can be seen already at birth, some researchers tend to call this form as SMA-0 type. Infants with SMA-2 (the intermediate or chronic infantile form, MIM# 253550) have an onset around 3–15 months with less severe symptoms, but they become progressively weaker with time. Since legs are usually weaker than arms, children with SMA-2 may sit but do not learn to ambulate. Survival time is longer, but many patients die generally while still in childhood. The third type (SMA-3 or Kugelberg-Welander disease, MIM# 253400) has an onset from 4 to 15 years. Disease onset before the age of 3 years is classified as type IIIa, whereas an age of onset beyond 3 years is classified as type IIIb SMA [93]. Weakness is often first noted in the proximal leg and shoulder muscles. These children are able to achieve walking and generally live into adulthood (44% of the type IIIa and 90% of type IIIb individuals are able to walk by the age of 20 years) [93]. SMA-4 is a rare adult form (MIM# 271150) with onset after 30 years of age. This milder form of the disease may be inherited in either an autosomal dominant or rarely autosomal recessive manner. Individuals with this form of the disease have a normal life expectancy and may sometimes be difficult to separate from long-duration forms of pure lower motor variant of amyotrophic lateral sclerosis [86].
SMN genes and mutations
Nearly all of SMA patients display homozygous deletions, rearrangements or other mutations in the telomeric copy of the survival of motor neuron (SMN) gene on chromosome 5q13 [46]. Telomeric (SMN1, MIM# 600354) and centromeric (SMN2, MIM# 601627) genes differ by 5 single nucleotide changes, two of which are in exons 7 and 8 [66]. These differences do not alter the sequence of amino acids encoded by these two genes. However, the exon 7 C-to-T transition at codon 280 has been shown to be necessary and sufficient for skipping of exon 7 during alternative splicing of the SMN2 gene [49]. Consequently, about 80–90% of SMN2 transcripts lack exon 7 and their truncated transcript encodes for a dysfunctional protein that lacks the last 16 residues at the C-terminal and is insufficient to compensate for the loss of SMN1 protein [56]. In other words, the full-length transcripts are predominantly produced by SMN1 and not by SMN2 genes. Therefore, mutations in the SMN2 gene do not lead to disease. However, since SMN2 gene does give rise to a small number of the full-length SMN transcripts, higher numbers of SMN2 copies have been linked to the less severe SMA types [25, 38, 52]. In SMA-1, often two or sometimes three copies of the SMN2 gene exist and the amount of full-length SMN protein is, on average, around 9% of normal levels. In SMA-2, usually three or more SMN2 copies are found together with 14% amount of full-length SMN protein. In SMA-3, four to eight SMN2 copies are responsible for an average of 18% of normal SMN protein amount. When SMN protein levels approach about 23% function of motor neurons is normal. Carriers with one functional copy of SMN1 gene who are generally having about 45–55% of normal amount of SMN protein are, therefore, asymptomatic. Different copy number of the SMN2 gene is thought to occur by random duplication events. However, in a small population of SMA patients, the SMN1 gene is converted to the SMN2 gene by replacing exons 7 and 8 (gene conversion). In SMA-2 and SMA-3 missense point mutations are far more common than in SMA-1 type.
Moreover, since it has been shown that even diseased siblings with the same number of SMN2 genes can have different SMA phenotypes, many researchers thought that the SMN locus alone cannot explain the whole genetic basis for phenotypic variability of SMA [18, 71]. To identify those putative SMA modifier genes, a transcriptome-wide differential expression analysis using total RNA from the lymphoblastoid cell lines derived from both unaffected and affected SMN1-deleted siblings was recently carried out by Wirth and collaborators [63]. In total, 18 transcripts showed a greater than threefold difference in expression, but only plastin 3 (PLS3, MIM# 300131) exhibited statistically significantly higher expression in fully asymptomatic siblings of all six SMA-discordant families analyzed [63]. All unaffected SMN1-deleted subjects were females and interestingly also the PST3 gene is localized on the sex-determining X chromosome. The authors concluded that this is the first modifier gene able to fully protect from developing SMA, but despite extensive efforts, the exact cause of this gender-specific protection remains largely unknown. Subsequent experiments showed that overexpression of PLS3 rescued the axon length and outgrowth defects associated with SMN downregulation in motor neurons of SMA mouse embryos and in zebrafish [63]. Since it has also been found that plastin 3 is highly expressed during neuronal differentiation in spinal cord, associates with SMN protein and increases levels of F-actin, which is important for axonal outgrowth and guidance, these findings support the view that the involvement of SMN in axonal outgrowth and pathfinding may be the major pathogenetic defect in SMA [63].
It may be concluded that genotype–phenotype relationship in autosomal recessive SMA is very complex. Namely, although nearly all of the affected patients are having the same homozygous absence of SMN1 gene, the different copy numbers of SMN2, together with the effect(s) of PLASTIN3 and possibly other, yet unknown genes, gives rise to extremely variable phenotypes: from congenital SMA (type 0), through SMA types 1–4, to subjects with minimal symptoms or even totally unaffected ones [63, 82].
Spinal muscular atrophy without mutations in the SMN1 gene has also been associated with mtDNA depletion due to mutations of nuclear thymidine kinase 2 gene (TK2, MIM# 188250) that controls mtDNA replication or more often with mutations of the cytochrome-c oxidase assembly gene that controls respiratory chain complexes (SCO2, MIM# 604377) [36, 81].
While SMN1 gene is highly conserved through evolution, the SMN2 gene is unique to humans: most other organisms possess a single copy of the SMN gene. Yeast, worm, fly, zebrafish and mouse models with no functional SMN protein have a uniformly embryonic lethal phenotype [72]. Out of several surrogate mouse models developed, the strategy of generating SMA mice which express the human SMN2 gene in the presence of Smn gene knockout was shown to be quite promising. Mice expressing one copy of SMN2 died one day after birth and mice with two SMN2 copies survived for 6 days, showing reduced number of motor neurons, while mice expressing 8–16 SMN2 copies were completely rescued from the SMA phenotype [37]. Most recently, a line of mice that express high levels of a SMN transcript lacking exon 7 (SMNΔ7) in the SMN2+/+, Smn−/− background was generated [45]. These mice confirm the susceptibility of motoneurons to SMN deficiency and live 5–14 days, due to complexing of the SMNΔ7 protein with the full-length protein [45]. This model is now most commonly being used to test potential therapeutics. Unfortunately, an in vitro motor neuronal model of SMA is missing to date. Currently, much effort is given to develop such a model using embryonic stem cells differentiating into motoneurons [48].
SMN protein and its functions
Since the functional SMN protein seems to be critical for survival of α-motoneurons, it would be of great importance to know its exact cellular function(s). So far, it is known that the SMN protein is an unique molecule with many binding partners and possible functions (Table 2). The SMN protein has several identified motifs, such as a lysine-rich basic region encoded by exon 2, a Tudor domain important for RNA processing encoded by exon 3, a polyproline region encoded by exons 4 and 5, and region with several tyrosine–glycine pairs encoded by exon 6 [73]. Identification of missense mutations in these regions indicates that each of these domains may be functionally important [79].
The SMN protein is found in the nucleus and cytoplasm of all cells, but most abundantly in α-motoneurons [5, 7, 91]. Within the nucleus, the SMN protein forms heteromeric complexes and seems to play an important role in small ribonucleoproteins (snRNPs) biogenesis (such as hnRNP U protein and the small RNA binding protein) and pre-mRNA processing (splicing) [26, 96]. It is localized in the nuclear domains called Cajal bodies (CBs), which are also involved in the biogenesis and recycling of splicing snRNPs [23]. In SMA, low SMN protein levels result in altered CBs composition and a notable separation of SMN protein into distinct nuclear bodies called gems. In severe cases of SMA, a dramatic reduction in nuclear gems can be observed [21].
The first suggestion that SMN protein might have other important functions than snRNP assembly came from electron-microscopical analysis of mouse spinal cord that revealed SMN protein present in dendrites and axons [64]. Further analysis of the subcellular localization of SMN during retinoic-acid-induced neuronal differentiation of mouse teratocarcinoma P19 cells demonstrated SMN accumulation in growth cone and filopodia in both neuronal- and glial-like cells [24]. Moreover, SMN was present at the leading edge of neurite outgrowths, suggesting its specific role in axono- and dendrogenesis. Then, SMN protein was found to be present at axon branching points and growth cones in cultured motoneurons [24, 39]. In a more detailed analysis that Rossoll and collaborators performed in n PC12 cells, is was found that Smn colocalizes with hnRNP-R in cell bodies and neurite-like proceses [68]. In contrast, hnRNP-R lacking the Smn interaction domain localized primarily to the nucleus indicating that hnRNP-R is targeted to cell processes by Smn where it mediates processes other than pre-mRNA splicing. Furthermore, it was also demonstrated that hnRNP-R is present in significantly reduced levels in neurites and growth cones of motor neurons from Smn−/−, SMN2 mice compared to Smnt/t;SMN2 motor neurons [68]. The interaction between Smn and hnRNP-R was functionally relevant since upregulation of either Smn or hnRNP-R resulted in enhanced outgrowth of neurites, which could be suppressed by hnRNP-R lacking the Smn interaction domain [68]. By expressing truncated proteins, the same authors also showed that this positive effect on neurite outgrowth depends on functional Smn as well as on the RNA-binding domain of hnRNP-R. Accordingly, Smn mutant motor neurons showed reduced growth cone size.
Further experiments confirmed actin as a most probable downstream mediator of the Smn–hnRNP–R complex [87]. Actin is a major component of outgrowing axons responsible for growth cone motility and transport of its mRNA into distal axons and growth cones can be stimulated by neurotrophin treatment [10, 87]. In addition to interactions with hnRNP-R and Q [69], SMN has also been shown to interact with profilin II, an actin-binding protein [31]. Profilin II is highly expressed in brain and spinal cord, where it localizes predominantly to anterior horn neurons [31].
The asymmetric SMN staining demonstrated in the germinative neuroepithelium further supported a role for SMN in neuronal migration and differentiation [23]. Finally, new approaches to study SMN protein function, particularly the use of antisense morpholino and synthetic RNA injections to knockdown SMN levels in the zebrafish model of SMA, have confirmed the hypothesis that, in addition to its role in snRNP biogenesis, SMN protein has an additional and independent function which is critical for motor neuron axon outgrowth [10, 11, 30, 54, 74]. Moreover, the severity of motor axon defects was shown to correlate with decreased survival of the zebrafish, suggesting that SMN protein is not only needed for motor neuron outgrowth, but perhaps also in its maintenance of, and interactions with, muscle [11], which is further supported by the finding of increased vulnerability of the neuromuscular juntions in mouse SMA models [60]. One of the most recent exciting new findings has been the identification of an SMN transcript which encodes the axonal-SMN protein which is found to be selectively expressed in developing spinal cord motoneurons and mainly localized in axons [74]. The pre` of diverse types of SMN complexes that have functions in neurons, other than its well characterized role to assemble snRNPs was recently further confirmed using high-resolution fluorescence of an SMN-Gemin complex in neuronal processes and growth cones of embryonic stem cell-derived motor neurons [95]. An attractive hypothesis driven from these findings is that the SMN complexes play a role in the assembly of localized mRNP complexes needed for axonal growth and synaptogenesis.
Neuropathological changes in SMA and proposed pathogenetic model
Based on the published neuropathological data [15, 75, 76], it can be stated that four different morphological types of motoneurons exist in the spinal cord of SMA subjects, each probably having a different cell fate:
-
(1)
Morphologically normal motoneurons in their normal position: on average 10–20% of motoneurons seen during postmortem analysis of SMA-1 subjects. It is supposed that the number of these neurons correlates with the amount of SMN protein produced.
-
(2)
Morphologically abnormal motoneurons in their normal position, i.e., chromatolytic neurons within anterior spinal grey matter. These neurons die slowly by necrotic cell death and comprise about 60–85% of the remaining motoneurons seen at autopsy.
-
(3)
Apoptotic motoneurons within the ventral horns. Although the number of these neurons is relatively low (on average 2–3% of the remaining neurons seen in SMA-1 subjects), since they die very rapidly and disappear even faster due to heterophagic elimination by activated microglial cells, their cumulative number is probably largely underestimated.
-
(4)
Heterotopic (migratory, displaced) motoneurons (HMN). Unlike controls, SMA patients with confirmed homozygous deletion of exon 7 in the SMN1 gene display a significant number of HMN at all levels of the spinal cord [76]. These heterotopic motoneurons are situated all along the ventral outflow of the white matter and within the ventral roots, have no axon or dendrites, appear hyperchromatic in Nissl-stained sections and their number positively correlates with clinical severity [76]. It was shown that HMN have no synapses, rarely activate microglial cells, and eventually die slowly by necrosis [76].
Taken all the findings together, an integrated model of SMA pathogenesis can be proposed. The presumptive series of pathological events during the development of SMA according to this model is given in Figs. 2 and 3 and follow the sequence below:
The proposed pathogenetic mechanism of SMA. A sequence of events during normal motoneuron development is given in a and b for comparison with SMA (c, d). Schemes a and c represent presumed events during early prenatal stage, while b and d represent late prenatal stage. The numbers on the scheme correspond to numbers of the proposed pathogenetic steps in the text (1–10). A anterior corticospinal tract fibers (green), L lateral corticospinal tract fibers (green), blue dots astrocytes, blue lines glial bundles, red dots microglial cells, SMA spinal muscular atrophy
Illustrative examples for Fig. 2. Letters a–j correspond to pathogenetic events 1–10, respectively. a PCR analysis of SMN exon 7, b Western blot analysis of SMN protein, c impaired axono- and dendrogenesis of heterotopic spinal motoneurons in SMA d spinal motoneuron of SMA patient without synapses (electron microscopy) e heterotopic (migratory) motoneurons (arrows) in SMA (cresyl-violet stain) f apoptotic motoneurons (arrowheads) in SMA (cresyl-violet stain) g heterotopic (migratory) motoneurons (mmn) in the ventral roots (VR) of SMA patient (cresyl-violet stain) h glial bundles of ventral roots (GB) (cresyl-violet stain) i anterior horn of a SMA subject: an apoptotic motoneuron engulfed by an ISEL-positive microglial cell (large arrowhead); arrow shows weakly ISEL-positive neuron probably representing an early stage of the apoptotic process; small arrowheads show ISEL-positive nuclei of microglial cells that appear to be involved in neuronophagia of chromatolytic neurons (in-situ end labeling) j neuronophagia of a chromatolytic neuron (cresyl-violet stain). Scale bars: c, e, g, h = 20 μm, (d) = 1 μm, (f) = 5 μm, (i, j) = 10 μm. Photomicrographs (c), (d), (e), (g) and (h) are reproduced from Simic et al. [76]. Photomicrographs f, i and j are reproduced from Simic et al. [75], with the permission of Wolters Kluwer Health (Lippincott Williams and Wilkins)
- (1)
-
(2)
Increased SMN mRNA decay [8] and reduction in functional, full-length SMN protein production [47] lead to decreased intracellular concentration of oligomerization-competent SMN proteins proportional to disease severity [50, 92].
-
(3)
Impaired spinal motoneuron axono- and dendrogenesis [76].
-
(4)
Failure of motoneurons to form synapses onto the corticospinal fibers of the upper motoneurons [15].
-
(5)
Due to the disturbed differentiation (i.e., impaired dendrite and axon outgrowth) and lack of synapse formation many motoneurons continue to migrate along the ventral outflow in the anterior direction towards the ventral roots of spinal cord (heterotopic motoneurons) [76].
-
(6)
In some of these abnormally migrating undifferentiated neurons, apoptotic mode of programmed cell death program is activated and they rapidly die (because they are immediately being eliminated by nearby microglial cells, this leaves no biochemical or morphological trace). It seems that apoptosis is inappropriately prolonged [3, 38, 42, 65, 70, 75, 84, 92], because during the midgestational period this is the normal process by which 40–70% of the embryonic motor neurons in the spinal cord undergo naturally occurring programmed cell death immediately following the arrival of their axons in muscle cells [16, 62, 90].
-
(7)
The larger part of motoneurons continues its abnormal migration, some being stopped by the anterior rim of the spinal cord, others migrating even outside the anterior white matter and entering the ventral roots [76].
-
(8)
As a secondary phenomenon, attracted glial cells follow abnormally migrating HMN thus forming glial bundles within the ventral roots [14, 29].
-
(9)
Impaired axonal transport of actin causes remaining motoneurons to become chromatolytic. It is the author’s speculation that glial bundles of ventral roots may further compromise axoplasmic flow of these neurons causing a so-called axonal reaction of the central type.
-
(10)
In the end-stage of the disease, all apoptotic, heterotopic and chromatolytic neurons [27] eventually die. Apoptotic cells do so more rapidly and predominantly during earlier stages of SMA, while heterotopic and chromatolytic cells die more slowly by necrosis during the later stages of SMA. It is hoped that future studies will use unbiased stereological methods, to precisely quantify the magnitude of motoneuron cell loss due to each type of cell death as well as with respect to each particular SMA genotype.
Within the framework of this model, it is supposed that individually variable levels of SMN protein together with influences of other phenotype modifier genes and their products cause different types of SMA through differential degree of motoneuron functional loss.
SMA treatment and prevention strategies
Treatment and prevention strategies for SMA can be divided in at least five broad groups: (1) use of compounds that drive SMN2 promoter activity; (2) use of drugs that modulate SMN2 splicing; (3) use drugs that stabilize SMN2 mRNA or SMN protein; (4) gene therapy; and (5) stem cell therapy [79].
Compounds that drive SMN2 promoter activity and activate SMN2 gene expression
As noted previously, there is a strong inverse correlation between SMN protein production from the SMN2 gene and disease severity, both in SMA patients [47] and SMA mouse models [28, 58]. For the purpose of quantification of SMN protein levels, a cell immunoassay [43] and a two site enzyme-linked immunosorbent assay (ELISA) have been developed to measure accurately drug-induced SMN elevation in model systems, such as skin fibroblast cultures from type I SMA patients [53]. It is of importance that the SMN genotypes producing less SMN protein have also been shown to increase susceptibility to and severity of sporadic amyotrophic lateral sclerosis [83].
The SMN2 gene is regulated by a promoter that is nearly identical in sequence and activity to the SMN1 promoter [22, 57]. The level of histone acetylation is determined by the balance of activities of histone acetyltransferases, which acetylate histones, and histone deacetylase (HDAC), which deacetylate histones. Control of the acetylation state of histones is an important epigenetic mechanism regulating gene expression. When the NH2-terminus of core histones is acetylated, corresponding chromatine region is more transcriptionally active due to increased accessibility of DNA. It has been demonstrated that HDAC 1 and 2 proteins may modulate the histone acetylation state at the SMN promoter and determine SMN2 gene expression [41]. One of the earliest discovered HDAC inhibitors was sodium butyrate, which was shown to increase full-length SMN2 transcript levels and SMN protein levels in lymphoblastoid cell lines derived from SMA-1 patients [13]. Subsequent studies showed that the same effect can be achived by several other HDAC inhibitors, such as phenylbutyrate [2], valproate [8, 80], trichostatin A [4] and suberoyl anilide hydroxamic acid [9, 40, 41, 80]. Other drugs that have been proposed to work by activating SMN2 gene expression are hydroxyurea [33] and indoprofen [51]. Most of the drugs mentioned are currently being studied in clinical trials worldwide.
Recently, a screening of 550,000 different small molecules was performed to identify those that activate SMN2 promoter present in a motoneuron-like NSC34 cells [40]. None of the 17 different compounds were acting as HDAC inhibitors. Two compounds, one indole and one quinazoline were confirmed to increase full-length SMN transcript and protein in SMA-derived cells.
Drugs that modulate SMN2 splicing
The mechanisms that direct splicing of the SMN gene transcripts showed that full-length SMN transcript is encoded by exons 1, 2a, 2b and 3–8, while exons 1–7 are being translated into SMN protein. It seems that there are many exonic and intronic splice enhancer and silencer motifs that play role in SMN transcript splicing. After a screen in patient fibroblasts, a chemotherapeutic drug aclarubicin was shown to stimulate exon 7 inclusion in SMN2 gene transcripts [2] therefore yielding a higher proportion of full-length SMN protein [25]. Unfortunately, the toxicity profile of this drug prohibits its long-term use.
Another strategy to enhance exon 7 inclusion is the use of synthetic antisense oligonucleotides that bind to SMN2-derived transcripts and promote exon 7 inclusion during splicing. Although it has been shown that this strategy may work well in vitro [12, 77], achieving efficient delivery of these oligonucleotides to motor neurons remains to be an unresolved problem.
Drugs that stabilize SMN2 mRNA or SMN protein
As it has been shown that in a subset of SMA patients intragenic SMN1 mutations render SMN1 transcripts susceptible to nonsense-mediated SMN mRNA decay (resulting in mRNA degradation), future treatment strategies may also be directed towards increasing SMN mRNA stability [8]. One group of the candidate compounds may be aminoglycosides, which stabilize SMN2 mRNA by triggering stop codon read through [89].
Gene therapy for SMA
Since the SMN2 gene is present in the same region of chromosome 5 and is similar in sequence to SMN1 except for a T at position +6 of exon 7 that is likely the predominant functional difference between the two genes, an approach has been developed to use single-stranded oligonucleotides (ODN) to repair genes within the context of the native chromosome. Indeed, using SMN2-sequence-specific ODN to direct the exchange of a T to a C in an SMA skin fibroblast cell line from a type 1 patient was shown to increase production of full-length SMN mRNA, measured by qRT-PCR, and SMN protein, measured by Western blotting [19]. However, the technical difficulty of gene delivery, together with problems related to random insertion of the therapeutic gene, still represent the major obstacles for its use in treatment of SMA patients.
Stem cell therapy for SMA
Embryonic stem (ES) cells are omnipotent cells that can be directed to differentiate into motor neurons. Transplantation of such differentiated ES into the spinal cord of rats with induced motoneuron injury showed promising results because these cells survived and produced axons that were able to grow into the ventral root [34]. With the growing understanding of ES cell biology and SMA pathogenesis, ES cells-based therapy is moving closer to clinical application [17, 48, 61, 67].
Two hypotheses on SMA pathogenesis
Based on the data available in the literature two main hypotheses about the pathogenesis of SMA prevail. The first one is that severe SMN deficiency causes widespread pre-mRNA splicing defects in numerous mouse transcripts of diverse genes, preferentially those containing a large number of introns [96]. Since only a large degree of SMN decrease (>80%) is required to cause a significant change in the levels of snRNAs or cause cell death in cultured cells, this suggests that cells normally contain a large excess capacity of SMN complex to maintain their normal inventory of snRNAs. According to this hypothesis, SMA is a general splicing disease not restricted to motor neurons. However, the degree of snRNAs reduction is not uniform across different tissues and cell types. Within this framework it is speculated that the affected motor neurons, being large and high energy requiring cells, would have a lower tolerance for depleted SMN levels and are uniquely sensitive.
The alternative hypothesis on pathogenesis of SMA is that the disease is a consequence of a motor neuron specific function of the SMN protein. This hypothesis stems from observations demonstrating the accumulation of the SMN protein in the axons and growth cones of motor neuron like cells in vitro and anterior horn cells in vivo [6, 7, 10, 11, 24, 30, 54, 64, 68, 74, 94], and its interaction with the heterogeneous nuclear riboprotein Q and R (hnRNP-R) and β-actin mRNA, targeting them into the growth cones [68].
According to the pathogenetic model proposed in this review, the selective motoneuron axonopathy appears to be far more important than ubiquitous nuclear splicing deficit.
Conclusion
In conclusion, recent molecular and neuropathological findings suggest that impaired outgrowth of motoneurons’ axon and dendrites with subsequent abnormal migration may represent the early key events of SMA pathogenesis and are responsible for all of the subsequent pathological changes observed in these cases. It is supposed that individually variable levels of SMN protein, together with influences of other phenotype modifier genes and their products, cause the clinical SMA spectrum through differential degree of motoneuron functional loss. The dynamics of the described sequence of interrelated events still needs to be fully elucidated. Hopefully the present review will help building more efficient treatment and prevention strategies for SMA.
References
Andreassi C, Jarecki J, Zhou J, Coovert DD, Monani UR, Chen X, Whitney M, Pollok B, Zhang M, Androphy E, Burghes AH (2001) Aclarubicin treatment restores SMN levels to cells derived from type I spinal muscular atrophy patients. Hum Mol Genet 10:2841–2849
Andreassi C, Angelozzi C, Tiziano FD, Vitali T, De Vincenzi E, Boninsegna A, Villanova M, Bertini E, Pini A, Neri G, Brahe C (2004) Phenylbutyrate increases SMN expression in vitro: relevance for treatment of spinal muscular atrophy. Eur J Hum Genet 12:59–65
Araki S, Hayashi M, Tamagawa K, Saito M, Kato S, Komori T, Sakakihara Y, Mizutani T, Oda M (2003) Neuropathological analysis in spinal muscular atrophy type II. Acta Neuropathol 106:441–448
Avila AM, Burnett BG, Taye AA, Gabanella F, Knight MA, Hartenstein P, Cizman Z, Di Prospero NA, Pellizzoni L, Fischbeck KH, Sumner CJ (2007) Trichostatin A increases SMN expression and survival in a mouse model of spinal muscular atrophy. J Clin Invest 117:659–671
Battaglia G, Princivalle A, Forti F, Lizier C, Zeviani M (1997) Expression of the SMN gene, the spinal muscular atrophy determining gene, in the mammalian central nervous system. Hum Mol Genet 6:1961–1971
Beattie CE, Carrel TE, McWhorter ML (2007) Fishing for a mechanism: using zebrafish to understand spinal muscular atrophy. J Child Neurol 22:995–1003
Béchade C, Rostaing P, Cisterni C, Kalisch R, La Bella V, Pettmann B, Triller A (1999) Subcellular distribution of SMN protein: possible involvement in nucleoplasmic and dendritic transport. Eur J Neurosci 11:293–304
Brichta L, Hofmann Y, Hahnen E, Siebzehnrubl FA, Raschke H, Blumcke I, Eyupoglu IY, Wirth B (2003) Valproic acid increases the SMN2 protein level: a well-known drug as a potential therapy for spinal muscular atrophy. Hum Mol Genet 12:2481–2489
Brichta L, Garbes L, Jedrzejowska M, Grellscheid SN, Holker I, Zimmermann K, Wirth B (2008) Nonsense-mediated messenger RNA decay of survival motor neuron 1 causes spinal muscular atrophy. Hum Genet 123:141–153
Briese M, Esmaeili B, Sattelle DB (2005) Is spinal muscular atrophy the result of defects in motor neuron processes? BioEssays 27:946–957
Carrel TL, McWhorter ML, Workman E, Zhang H, Wolstencroft EC, Lorson C, Bassell GJ, Burghes AH, Beattie CE (2006) Survival motor neuron function in motor axons is independent of functions required for small nuclear ribonucleoprotein biogenesis. J Neurosci 26:11014–11022
Cartegni L, Krainer AR (2003) Correction of disease-associated exon skipping by synthetic exon-specific activators. Nat Struct Biol 10:120–125
Chang JG, Hsieh-Li HM, Jong YJ, Wang NM, Tsai CH, Li H (2001) Treatment of spinal muscular atrophy by sodium butyrate. Proc Natl Acad Sci USA 98:9808–9813
Chou SM, Nonaka I (1978) Werdnig-Hoffmann disease: proposal of a pathogenetic mechanism. Acta Neuropathol 41:45–54
Chou SM, Wang HS (1997) Aberrant glycosylation/phoshorylation in chromatolytic motoneurons of Werdnig–Hoffmann disease. J Neurol Sci 152:198–209
Clarke PGH (1994) Neuronal death in the development of the vertebrate central nervous system. Sem Neurosci 6:291–297
Coutts M, Keirstead HS (2008) Stem cells for the treatment of spinal cord injury. Exp Neurol 209:368–377
Cuscò I, Barceló MJ, Rojas-García R, Illa I, Gamez J, Cervera C, Pou A, Izquierdo G, Baiget M, Tizzano EF (2006) SMN copy number predicts acute or chronic spinal muscular atrophy but does not account for intrafamilial variability in siblings. J Neurol 253:21–25
DiMatteo D, Callahan S, Kmiec EB (2008) Genetic conversion of an SMN2 gene to SMN1: a novel approach to the treatment of spinal muscular atrophy. Exp Cell Res 314:878–886
Dubowitz V, Sewry CA (2007) Muscle biopsy: a practical approach. Saunders Elsevier, Amsterdam
Dundr M, Hebert MD, Karpova TS, Stanek D, Xu H, Shpargel KB, Meier UT, Neugebauer KM, Matera AG, Misteli T (2004) In vivo kinetics of Cajal body components. J Cell Biol 164:831–42
Echaniz-Laguna A, Miniou P, Bartholdi D, Melki J (1999) The promoters of the survival motor neuron gene (SMN) and its copy (SMNc) share common regulatory elements. Am J Hum Genet 64:1365–1370
Eggert C, Chari A, Laggerbauer B, Fischer U (2006) Spinal muscular atrophy: the RNP connection. Trends Mol Med 12:113–121
Fan L, Simard LR (2002) Survival motor neuron (SMN) protein: role in neurite outgrowth and neuromuscular maturation during neuronal differentiation and development. Hum Mol Genet 11:1605–1614
Feldkötter M, Schwarzer V, Wirth R, Wienker TF, Wirth B (2002) Quantitative analyses of SMN1 and SMN2 based on realtime light-cycler PCR: fast and highly reliable carrier testing and prediction of severity of spinal muscular atrophy. Am J Hum Genet 70:358–368
Fischer U, Liu Q, Dreyfuss G (1997) The SMN-SIP1 complex has an essential role in spliceosome biogenesis. Cell 90:1023–1029
Galluzzi L, Maiuri MC, Vitale I, Zischka H, Castedo M, Zitvoogel L, Kroemer G (2007) Cell death modalities: classification and pathophysiological implications. Cell Death Diff 14:1237–1243
Gavrilina TO, McGovern VL, Workman E, Crawford TO, Gogliotti RG, DiDonato CJ, Monani UR, Morris GE, Burghes AH (2008) Neuronal SMN expression corrects spinal muscular atrophy in severe SMA mice while muscle-specific SMN expression has no phenotypic effect. Hum Mol Genet 17:1063–1075
Ghatak NR (1978) Spinal roots in Werdnig-Hoffmann disease. Acta Neuropathol 41:1–7
Giavazzi A, Setola V, Simonati A, Battaglia G (2006) Neuronal-specific roles of the survival motor neuron protein: evidence from survival motor neuron expression patterns in the developing human central nervous system. J Neuropathol Exp Neurol 65:267–277
Giesemann T, Rathke-Hartlieb S, Rothkegel M, Barthsch JW, Buchmeier S, Jockusch BM, Jockusch H (1999) A role for polyproline motifs in the spinal muscular atrophy protein SMN. Profilins bind to and colocalize with smn in nuclear gems. J Biol Chem 274:37908–37914
Greenfield JC, Stern RO (1927) The anatomical identity of the Werdnig-Hoffmann and Oppenheim forms of infantile muscular atrophy. Brain 50:652–686
Grzeschik SM, Ganta M, Prior TW, Heavlin WD, Wang CH (2005) Hydroxyurea enhances SMN2 gene expression in spinal muscular atrophy cells. Ann Neurol 58:194–202
Harper JM, Krishnan C, Darman JS, Deshpande DM, Peck S, Shats I, Backovic S, Rothstein JD, Kerr DA (2004) Axonal growth of embryonic stem cell-derived motoneurons in vitro and in motoneuron-injured adult rats. Proc Natl Acad Sci USA 101:7123–7128
Hausmanowa-Petrusiewicz I, Zaremba J (2000) Proximal spinal muscular atrophy of childhood. In: Deymeer F (ed) Neuromuscular diseases: From basic mechanisms to clinical menagement. Basel: Karger, Monogr Clin Neurosci 18:163–176
Hirano M, Angelini C, Montagna P, Hays AP, Tanji K, Mitsumoto H, Gordon PH, Naini AB, DiMauro S, Rowland LP (2008) Amyotrophic lateral sclerosis with ragged-red fibers. Arch Neurol 65:403–406
Hsieh-Li HM, Chang JG, Jong YJ, Wu MH, Wang NM, Tsai CH, Li H (2000) A mouse model for spinal muscular atrophy. Nat Genet 24:66–70
Iwahashi H, Eguchi Y, Yasuhara N, Hanafusa T, Matsuzawa Y, Tsujimoto Y (1997) Synergistic antiapoptotic activity between bcl-2 and SMN implicated in spinal muscular atrophy. Nature 390:413–417
Jablonka S, Bandilla M, Wiese S, Buhler D, Wirth B, Sendtner M, Fischer U (2001) Co-regulation of survival of motor neuron (SMN) protein and its interactor SIP1 during development and in spinal muscular atrophy. Hum Mol Genet 10:497–505
Jarecki J, Chen X, Bernardino A, Coovert DD, Whitney M, Burghes A, Stack J, Pollok BA (2005) Diverse small-molecule modulators of SMN expression found by high-throughput compound screening: early leads towards a therapeutic for spinal muscular atrophy. Hum Mol Genet 14:2003–2018
Kernochan LE, Russo ML, Woodling NS, Huynh TN, Avila AM, Fischbeck KH, Sumner CJ (2005) The role of histone acetylation in SMN gene expression. Hum Mol Genet 14:1171–1182
Kerr DA, Nery JP, Traystman RJ, Chau BN, Hardwick JM (2000) Survival motor neuron protein modulates neuron-specific apoptosis. Proc Natl Acad Sci USA 97:13312–13317
Kolb SJ, Gubitz AK, Olszewski RF Jr, Ottinger E, Sumner CJ, Fischbeck KH, Dreyfuss G (2006) A novel cell immunoassay to measure survival of motor neurons protein in blood cells. BMC Neurology 6:6
La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck KH (1991) Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352:77–79
Le TT, Pham LT, Butchbach ME, Zhang HL, Monani UR, Coovert DD, Gavrilina TO, Xing L, Bassell GJ, Burghes AH (2005) SMNDelta7, the major product of the centromeric survival motor neuron (SMN2) gene, extends survival in mice with spinal muscular atrophy and associates with full-length SMN. Hum Mol Genet 14:845–857
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L, Benichou B, Cruaud C, Millasseau P, Zeviani M, Le Paslier D, Frézal J, Cohen D, Weissenbach J, Munnich A, Melki J (1995) Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80:155–165
Lefebvre S, Burlet P, Liu Q, Bertrandy S, Clermont O, Munnich A, Dreyfuss G, Melki J (1997) Correlation between severity and SMN protein level in spinal muscular atrophy. Nat Genet 16:265–269
Li XJ, Hu BY, Jones SA, Zhang YS, Lavaute T, Du ZW, Zhang SC (2008) Directed differentiation of ventral progenitors and motor neurons from human embryonic stem cells by small molecules. Stem Cells 26:886–893
Lorson CL, Strasswimmer J, Yao J-M, Baleja JD, Hahnen E, Wirth B, Le T, Burghes AH, Androphy EJ (1998) SMN oligomerization defect correlates with SMA severity. Nat Genet 19:63–66
Lorson CL, Hahnen E, Androphy EJ, Wirth B (1999) A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc Natl Acad Sci 96:6307–6311
Lunn MR, Root DE, Martino AM, Flaherty SP, Kelley BP, Coovert DD, Burghes AH, Man NT, Morris GE, Zhou J, Androphy EJ, Sumner CJ, Stockwell BR (2004) Indoprofen upregulates the survival motor neuron protein through a cyclooxygenase-independent mechanism. Chem Biol 11:1489–1493
Mailman MD, Heinz JW, Papp AC, Snyder PJ, Sedra MS, Wirth B, Burghes AH, Prior TW (2002) Molecular analysis of spinal muscular atrophy and modification of the phenotype by SMN2. Genet Med 4:20–26
Man NT, Humphrey E, Le Thanh L, Fuller HR, Lynch TA, Sewry CA, Goodwin PR, MacKenzie AE, Morris GE (2008) A two-site ELISA can quantify up-regulation of SMN protein by drugs for spinal muscular atrophy. Neurology (accepted)
McWhorter ML, Monani UR, Burghes AH, Beattie CE (2003) Knockdown of the survival motor neuron (Smn) protein in zebrafish causes defects in motor axon outgrowth and pathfinding. J Cell Biol 162:919–931
Monani UR (2005) Spinal muscular atrophy: A deficiency in a ubiqutous protein; a motor neuron specific disease. Neuron 48:885–896
Monani UR, Lorson CL, Parsons DW, Prior TW, Androphy EJ, Burghes AH, McPherson JD (1999) A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum Mol Genet 8:1177–1183
Monani UR, McPherson JD, Burghes AH (1999) Promoter analysis of the human centromeric and telomeric survival motor neuron genes (SMNC and SMNT). Biochim Biophys Acta 1445:330–336
Monani UR, Sendtner M, Coovert DD, Parsons DW, Andreassi C, Le TT, Jablonka S, Schrank B, Rossoll W, Prior TW, Morris GE, Burghes AH (2000) The human centromeric survival motor neuron gene (SMN2) rescues embryonic lethality in Smn(–/–) mice and results in a mouse with spinal muscular atrophy. Hum Mol Genet 9:333–339
Munsat TL, Davies KE (1992) International SMA consortium meeting. Neuromuscul Disord 2:423–428
Murray LM, Comley LH, Thomson D, Parkinson N, Talbot K, Gillingwater TH (2008) Selective vulnerability of motor neurons and disociation of pre- and post-synaptic pathology at the neuromuscular junction in mouse models of spinal muscular atrophy. Hum Mol Genet 17:949–962
Nayak MS, Kim Y-S, Goldman M, Keirstead HS, Kerr DA (2006) Cellular therapies in motor neuron diseases. Biochim Biophys Acta 1762:1128–1138
Oppenheim RW (1991) Cell death during development of the nervous system. Annu Rev Neurosci 14:453–501
Oprea GE, Kröber S, McWhorter ML, Rossoll W, Müller S, Krawcak M, Bassell GJ, Beattie CE, Wirth B (2008) Plastin 3 is a protective modifier of autosomal recessive spinal muscular atrophy. Science 320:524–527
Pagliardini S, Giavazzi A, Setola V, Lizier C, Di Luca M, DeBiasi S, Battaglia G (2000) Subcellular localization and axonal transport of the survival motor neuron (SMN) in the developing rat spinal cord. Hum Mol Genet 9:47–56
Parker GC, Li X, Anguelov RA, Toth G, Cristescu A, Acsadi G (2008) Survival motor neuron protein regulates apoptosis in an in vitro model of spinal muscular atrophy. Neurotox Res 13:39–48
Parsons DW, McAndrew PE, Iannaccone ST, Mendell JR, Burghes AH, Prior TW (1998) Intragenic telSMN mutations: frequency, distribution, evidence of a founder effect, and modification of the spinal muscular atrophy phenotype by cenSMN copy number. Am J Hum Genet 63:1712–1723
Renoncourt Y, Carroll P, Filippi P, Arce V, Alonso S (1998) Neurons derived in vitro from ES cells express homeoproteins characteristic of motoneurons and interneurons. Mech Dev 79:185–197
Rossoll W, Kroning AK, Ohndorf UM, Steegborn C, Jablonka S, Sendtner M (2002) Specific interaction of Smn, the spinal muscular atrophy determining gene product, with hnRNP-R and gry-rbp/hnRNPQ: a role for Smn in RNA processing in motor axons? Hum Mol Genet 11:93–105
Rossoll W, Jablonka S, Andreassi C, Kroning AK, Karle K, Monani UR, Sendtner M (2003) Smn, the spinal muscular atrophy-determining gene product, modulates axon growth and localization of beta-actin mRNA in growth cones of motoneurons. J Cell Biol 163:801–812
Sato K, Eguchi Y, Kodama TS, Tsujimoto Y (2000) Regions essential for the interaction between Bcl-2 and SMN, the spinal muscular atrophy disease gene product. Cell Death Differ 7:374–383
Scharf JM, Endrizzi MG, Wetter A, Huang S, Thompson TG, Zerres K, Dietrich WF, Wirth B, Kunkel LM (1998) Identification of a candidate modifying gene for spinal muscular atrophy by comparative genomics. Nat Genet 20:83–86
Schmid A, Di Donato CJ (2007) Animal models of spinal muscular atrophy. J Child Neurol 22:1004–1012
Selenko P, Sprangers R, Stier G, Bühler D, Fischer U, Sattler M (2001) SMN Tudor domain structure and its interaction with Sm proteins. Nature Struct Biol 8:27–31
Setola V, Terao M, Locatelli D, Bassanini S, Garattini E, Battaglia G (2007) Axonal-SMN (a-SMN), a protein isoform of the survival motor neuron gene, is specifically involved in axonogenesis. Proc Natl Acad Sci USA 104:1959–1964
Simic G, Seso-Simic D, Lucassen P, Islam A, Krsnik Z, Cviko A, Jelasic D, Barisic N, Winblad B, Kostovic I, Kruslin B (2000) Ultrastructural analysis and TUNEL demonstrate motor neuron apoptosis in Werdnig-Hoffmann disease. J Neuropathol Exp Neurol 59:398–407
Simic G, Mladinov M, Seso-Simic D, Jovanov-Milosevic N, Islam A, Pajtak A, Barisic N, Sertic J, Lucassen PJ, Hof PR, Kruslin B (2008) Abnormal motoneuron migration, differentiation and axon outgrowth in spinal muscular atrophy. Acta Neuropathol 115:313–326
Skordis LA, Dunckley MG, Yue B, Eperon IC, Muntoni F (2003) Bifunctional antisense oligonucleotides provide a trans-acting splicing enhancer that stimulates SMN2 gene expression in patient fibroblasts. Proc Natl Acad Sci USA 100:4114–4119
Smith M, Calabro V, Chong B, Gardiner N, Cowie S, du Sart D (2007) Population screening and cascade testing for carriers of SMA. Eur J Hum Genet 15:759–766
Sumner CJ (2006) Therapeutics development for spinal muscular atrophy. NeuroRx 3:235–245
Sumner CJ, Huynh TN, Markowitz JA, Perhac JS, Hill B, Coovert DD, Schussler K, Chen X, Jarecki J, Burghes AH, Taylor JP, Fischbeck KH (2003) Valproic acid increases SMN levels in spinal muscular atrophy patient cells. Ann Neurol 54:647–654
Tarnopolsky MA, Bourgeois JM, Fu M-H, Kataeva G, Shah J, Simon DK, Mahoney D, Johns D, MacKay N, Robinson BH (2004) Novel SCO2 mutation (G1521A) presenting as a spinal muscular atrophy type I phenotype. Am J Med Genet 125A:310–314
Tizzano E, Baiget M (2001) Molecular bases of spinal muscular atrophy: the survival motor neuron gene. Contrib Science 2:35–42
Veldink JH, Kalmijn S, Van der Hout AH, Lemmink HH, Groeneveld GJ, Lummen C, Scheffer H, Wokke JH, Van den Berg LH (2005) SMN genotypes producing less SMN protein increase susceptibility to and severity of sporadic ALS. Neurology 65:820–825
Wang W, DiMatteo D, Funanage VL, Scavivna M (2005) Increased susceptibility of spinal muscular atrophy fibroblasts to camptothecin-induced cell death. Mol Genet Metabol 85:38–45
Werdnig G (1891) Zwei frühinfantile hereditäre Fälle von progressiver Muskelatrophie unter dem Bilde der Dystrophie, aber auf neurotischer Grundlage. Arch Psychiatr Nervenkr 22:437–481
Wharton S, Ince PG (2003) Pathology of motor neuron disorders. In: Shaw PJ, Strong MJ (eds) Motor neuron disorders. Blue books of practical neurology, book 28. Butterworth-Heineman, Philadelphia, pp 17–49
Willis D, Li KW, Zheng JQ, Chang JH, Smit A, Kelly T, Merianda TT, Sylvester J, van Minnen J, Twiss JL (2005) Differential transport and local translation of cytoskeletal, injury-response, and neurodegeneration protein mRNAs in axons. J Neurosci 25:778–791
Wirth B, Brichta L, Hahnen E (2006) Spinal muscular atrophy: from gene to therapy. Sem Pediat Neurol 13:121–131
Wolstencroft EC, Mattis V, Bajer AA, Young PJ, Lorson CL (2005) A non-sequencespecific requirement for SMN protein activity: the role of aminoglycosides in inducing elevated SMN protein levels. Hum Mol Genet 14:1199–1210
Yachnis AT, Giovanini MA, Eskin TA, Reier PJ, Anderson DK (1998) Developmental patterns of bcl-2 and bcl-x polypeptide expression in the human spinal cord. Exp Neurol 150:82–97
Young P, Le TT, Dunckley M, Nguyen TM, Burghes AH, Morris GE (2001) Nuclear gems and Cajal (coiled) bodies in fetal tissues: nucleolar distribution of the spinal muscular atrophy protein, SMN. Exp Cell Res 265:252–261
Young PJ, Day PM, Androphy EJ, Morris GE, Lorson CL (2002) A direct interaction between survival motor neuron protein and p53 and its relationship to spinal muscular atrophy. J Biol Chem 277:2852–2859
Zerres K, Rudnik-Schöneborn S (1995) Natural history in proximal spinal muscular atrophy. Clinical analysis of 445 patients and suggestions for a modification of existing classifications. Arch Neurol 52:518–523
Zhang HL, Pan F, Hong D, Shenoy SM, Singer RH, Bassell GJ (2003) Active transport of the survival motor neuron protein and the role of exon-7 in cytoplasmic localization. J Neurosci 23:6627–6637
Zhang HL, Xing L, Rossoll W, Wichterle H, Singer RH, Bassell GJ (2006) Multiprotein complexes of the survival of motor neuron protein SMN with Gemins traffic to neuronal processes and growth cones of motor neurons. J Neurosci 26:8622–8632
Zhang Z, Lotti F, Dittmar K, Younis I, Wan L, Kasim M, Dreyfuss G (2008) SMN deficiency causes tissue-specific perturbations in the repertoire of snRNAs and widespread deficits in splicing. Cell 133:585–600
Acknowledgments
The help of Dr. Patrick R. Hof (Mount Sinai School of Medicine, NY) in the preparation of this manuscript is greatly appreciated. Figure 3a is kindly provided by Jadranka Sertic and 3b by Glenn E. Morris.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Simic, G. Pathogenesis of proximal autosomal recessive spinal muscular atrophy. Acta Neuropathol 116, 223–234 (2008). https://doi.org/10.1007/s00401-008-0411-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-008-0411-1