Abstract
It is generally accepted that bacterial diversity in a community confers resistance to environmental perturbation. Communities with high bacterial diversity are less likely to be impacted by environmental changes such as warming. As such, hydrocarbon-contaminated Antarctic soil that are typically characterised by low bacterial diversity and highly selective taxonomic composition are expected to be more sensitive to changes in temperature than uncontaminated Antarctic soil. To test this hypothesis, we evaluated the response of bacterial community structure to warming of hydrocarbon-contaminated and uncontaminated soil collected from Casey Station, Windmill Island, East Antarctica by using microcosms incubated at 5, 10 and 15 °C over a period of 12 weeks. Our results showed that shifts occurred in the bacterial community in relation to the incubation temperatures in both the hydrocarbon-contaminated and uncontaminated soil, with a stronger response observed in the contaminated soil. Taxa referred as comprising hydrocarbon-degrading genera such as Rhodococcus, was the most prevalent genus in the contaminated soil after incubation at 15 °C, accounting for approximately 32–50% of the total detected genera. However, there were no significant differences in the selected functional genes, potentially suggesting high levels of metabolic plasticity in the studied soil bacterial communities. Overall, we showed that hydrocarbon contamination in soil might lead to lower bacterial community stability against environmental perturbation such as temperature variation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Human activities in Antarctica are heavily reliant on petroleum hydrocarbon and synthetic oil products for transport, aircraft, ships, power generation, and heating. Despite the heavy usage, guidelines for proper hydrocarbon disposal were only introduced after the implementation of The Protocol on Environmental Protection to the Antarctic Treaty in 1998. Nearly 20 years after the Protocol was signed, the regulation of fuel management and contaminants is highly dependent on self-regulation by national operators, and their interpretation of the protocol (Aislabie et al. 2004; Bennett et al. 2015). Due to the weakness in past disposal practice and management, and a lack of independent regulation, the occurrence of accidental spills and contamination of soils and sediments with anthropogenic-related hydrocarbons remain common in Antarctica (Filler et al. 2008; Tin et al. 2009; Fryirs et al. 2015; McWatters et al. 2016).
Light (jet oil) and heavy fuel (diesel) are able to move across and into unfrozen soil layers, resulting in the reduction of soil pH and indirectly affect soil nitrite through microbial activities (Aislabie et al. 2004). Hydrocarbons also increase soil surface temperature due to decreasing soil albedo, and they increase the local soil carbon pool (Aislabie et al. 2004; Saul et al. 2005; Vazquez et al. 2009). Anthropogenic hydrocarbons can disrupt soil stoichiometry, causing imbalance in C:N:P ratio (Dias et al. 2015). These alterations in soil properties can exert a strong selection pressure to the bacterial community by preferentially selecting for heterotrophic hydrocarbon-degraders such as Pseudomonas, Rhodococcus and Sphingomonas (Aislabie et al. 2001; Saul et al. 2005; Vázquez et al. 2013; Muangchinda et al. 2015). Furthermore, hydrocarbon-contaminated soils were frequently found to harbour lower bacterial diversity in comparison to the uncontaminated soil, due probably to the selection of microorganisms that are able to utilise hydrocarbons as well as those affected by hydrocarbon toxicity (Aislabie et al. 2004; Saul et al. 2005; Labud et al. 2007; Chong et al. 2009; Cury et al. 2015).
Over the past two decades, a strong warming trend was recorded in the Antarctic Peninsula and West Antarctica (Turner et al. 2014; Ludescher et al. 2016), although a recent report suggested that the warming at the Antarctic Peninsula is slowly attenuated (Turner et al. 2016). Bacterial communities showed rapid and significant responses in structure, activity and diversity to the changes of climate (Yergeau et al. 2012). For instance, Rinnan et al. (2009) showed that the adaptation of Antarctic soil community to warming is highly dependent on external factors such as location and presence of vegetation. In an in situ soil warming experiment on the Antarctic Peninsula, soil bacterial communities exhibited significant shifts and significantly lower functional gene diversity (Yergeau et al. 2012). Separately, under a laboratory-based soil microcosm setting, vegetated soil incubated at 15 °C showed elevated active bacterial diversity (RNA) in comparison to microcosm soils incubated in lower temperatures (3–7 °C) after 6 months of incubation (Yergeau and Kowalchuk 2008).
Previous reports showed that community changes in response to warming were also detectable in Antarctic hydrocarbon-contaminated soil (Coulon et al. 2005; Ferguson et al. 2008). It was found that warming generally established a positive effect to hydrocarbon remediation in Antarctic soil and sediments (Ferguson et al. 2008; McDonald and Knox 2014). Interestingly, however, few studies have compared community and functional stability between hydrocarbon-contaminated and uncontaminated soil during warming. Specifically, it is unclear whether the microbial community assembly in hydrocarbon-contaminated soil is more or is less sensitive to warming than uncontaminated soil. We postulate that the reduced diversity and specialised population dominated by hydrocarbon-degraders in the hydrocarbon-contaminated soil is less resistant to warming than the uncontaminated soil. To test our hypothesis, we compared the response of bacterial communities from both hydrocarbon-contaminated and uncontaminated Antarctic soil using microcosms incubated at 5, 10 and 15 °C over a period of 12 weeks.
Materials and methods
Description of study site
Soil samples used for the microcosm study were collected in the vicinity of Casey Station, Windmill Island, East Antarctica (Fig. 1). Soil around the station is composed of mainly coarse and sandy Antarctic mineral soil. The hydrocarbon-contaminated (HC) site (S66°16′54.305″ E110°31′27.124″) is located where an oil spill occurred in 2012 near the Emergency Power House. The soil was excavated to a depth of approximately 0.2–1.25 m below ground surface, mixed and placed in biopile treatment cells for remediation. The same soil used for the biopile construction was collected (~ 5 kg) and frozen until the microcosm setup. The initial total petroleum hydrocarbon content (TPH) was 2180 ± 118 mg/kg, (carbon 9–40, dry mass basis) and was generally considered to be within the medium contamination range (401–5000 mg/kg) (van Dorst et al. 2014, 2016). The uncontaminated (UC) site (S66°16′54.644″ E110°31′30.374″) is located approximately 50 m east of the spill site and has negligible amount of TPH. The TPH content of the HC and UC soil was supplied by the Australian Antarctic Division.
Location of Casey Station, Antarctica. Source: Australian Antarctic Division Data Centre (adapted from: https://data.aad.gov.au/aadc/mapcat/)
Soil incubation
The soil samples were homogenised by sieving through a 2-mm mesh sieve prior to setting up the microcosms. The microcosms were set up using sterile falcon tubes (diameter = 3 cm, capacity = 50 mL). The bottom of each tube was filled with sterile glass beads up to 2 cm in height to allow drainage to avoid waterlogging of the soil. Water usually exist only transiently in liquid form in Antarctic soil (Adams et al. 2006; Cary et al. 2010). Approximately 20 g of soil (HC or UC) was put into each sterile falcon tube. The top of the tubes had a headspace of approximately 5 cm and they were loosely plugged with sterilised rolled cotton wool to trap dust and allow aeration during incubation. The tubes of soil were incubated at three different temperatures (5, 10 and 15). 500 µL sterile distilled water was added to each tube every 2 days to prevent dehydration.
Fifteen tubes each of HC and UC soil were incubated at each temperature. Therefore, for three temperature treatments, 45 tubes of HC soil and 45 tubes of UC soil were incubated. Five tubes each of HC and UC soil were removed and analysed at each of the three incubation periods: 4, 8 and 12 weeks. Separately, five tubes each of untreated HC and UC soil were analysed as control.
Soil chemical analyses
All samples collected were dried at 70 °C until a constant weight was obtained (Chong et al. 2012). Soil water content was determined based on percentage of water lost from drying. Dried samples were used for subsequent soil chemical analysis. Soil pH was measured in 1:2 (w/v) suspensions of dried soil in distilled water while electrical conductivity (µS/cm) was measured in 1:5 (w/v) suspensions of dried soil in distilled water (Chong et al. 2010). Nitrate and nitrite in the soil were extracted by addition of calcium chloride and activated charcoal, while phosphate was analysed by digestion of the soil with concentrated sulphuric acids and hydrogen peroxide. Nitrate, nitrite and phosphate contents in the soil (mg/kg) were determined photometrically using spectroquant photometer (MERCK, USA) at International Medical University, Malaysia.
Bacterial community analyses
Bacterial DNA extraction and Terminal Restriction Fragment Length Polymorphism (T-RFLP) analysis
PowerSoil® DNA Isolation Kit (MoBio Inc., USA) was used to extract DNA from the soil samples, according to the manufacturer’s instructions. Soil DNA was eluted in 50 μL TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0). T-RFLP was conducted as described in Chong et al. (2012). Briefly, template DNA from each soil sample was amplified using the bacteria-specific 16S primers, 27F (5′- FAM- GAGTTTGATCMTGGCTCAG-3′) and 1492R (5′- HEX-GGYTACCTTGTTACGACTT-3′), to produce amplicons of ~ 1500 base pairs (bp). The amplified DNA was then purified with MEGAquick-spin™ PCR Product Purification Kit (iNtRON Biotechnology, Korea) and digested with Msp-I (Fermentas, USA). Electrophoretic separation of restriction fragments was analysed by FirstBase Laboratories (Selangor, Malaysia) using ABI 3100 and ABI 3730 XL DNA analyser (Applied Biosystems Inc.) resulting in a unique genotype profile for each sample. The profiles were ran along with GeneScan™-500HD ROX™ marker (Applied Biosystems, USA), which permits determination of terminal restriction fragments (T-RF) up to 500 bp in length. These terminal fragments were analysed using the web-based programme T-Rex (Culman et al. 2009). The fluorescence intensity of each peak indicates the relative abundance of a taxon which is expressed as either peak height or peak area. Statistical determination of the threshold was carried out (Abdo et al. 2006) to distinguish true peaks from noise in the range of 50–500 bp. T-RFs that are < 50 or > 500 bp were eliminated from all datasets. In addition, to reduce run-to-run variability, fingerprints were aligned using T-Align (Smith et al. 2005). In this method, the smallest T-RF present among all replicate profiles was identified and marked. Peaks within this range of 0.5 bp were considered identical and binned into a single T-RF. The next smallest peak within the same range (± 0.5 bp) not falling into the first T-RF, was binned into the second T-RF.
Statistical analyses of the T-RFLP derived community patterns
Multivariate data analysis was conducted with PRIMER v6 with PERMANOVA add-on programme package (Anderson 2001; Anderson et al. 2008). Permutational multivariate analysis of variance (PERMANOVA) based on Bray–Curtis similarity coefficient was performed to test for significant difference in bacterial assemblage pattern across treatments. Constraint ordination in the form of canonical analysis of principal coordinates (CAP) was conducted to visualise the distribution of bacterial assemblage pattern by selecting axes from the multidimensional data cloud that showed the strongest difference between treatments. Distance-based linear model (DISTLM) routine was also performed to analyse and model the correlation between the bacterial assemblages and environmental factors. DISTLM was constructed using the step-wise selection procedure under adjusted Akaike information criterion with a correction for finite sample sizes (AICc) selection criterion to enable the fitting of the best explanatory environmental variables in the model.
Roche 454 pyrosequencing
The HC and UC microcosms which showed the highest diversity (based on T-RFLP analysis) under different temperatures and incubation durations were further analysed using Roche 454 pyrosequencing which was commissioned to ScienceVision Sdn. Bhd (Selangor, Malaysia). Sequencing was conducted using PCR primer pair (5′ GAGTTTGATCMTGGCTCAG-3′ and 5′WTTACCGCGGCTGCTGG-3′) targeting V3-V4 regions of the 16S rRNA gene. A total of 111,902 reads were obtained with an average sequence length of ~ 280 bp. The raw data were noise-filtered, screened for chimeras and aligned to SILVA alignment database, Release 128. The resulting output consisted of 65,972 sequences with an average sequence length of ~ 245 bp. Sequences were subsequently clustered into operational taxonomic unit (OTU) at 3% evolutionary distance and different levels of phylogenetic taxonomic unit based on SILVA reference taxonomy Release 128 with a cut-off of 80% homology. Alpha diversity of the bacterial assemblages at OTU level was assessed based on observed richness (Sobs) and Inverse Simpson Index (Invsimpson), community evenness using Simpson evenness index (Simpsoneven) while the sequencing coverage was estimated based on Good’s coverage. Analysis of molecular variance (AMOVA), as implemented in mothur, was used to determine if there were statistically significant differences in the Sorensen and Hellinger similarity coefficient between HC and UC soil samples. Differentially abundant features in genus and phylum level were identified based on linear discriminant analysis (LDA) effect size (LEfSe) method. The alpha diversity indices including Invsimpson and Simpsoneven, as well as AMOVA and LEFse, were calculated using the subsampled data (n = 1565 bp).
Quantitative evaluation of functional genes involved in nitrogen and carbon metabolism
Quantitative PCR (qPCR) was conducted to measure the abundance of nifH (nitrogen fixation), nirS (denitrification), nosZ (denitrification), Chitinase GA (carbon degradation), amo-A (nitrification) and nirK (nitrate reductase) genes in the soil samples. Each 10 µl reaction contained 1 µl of the target DNA extract (10 ng), 5 µl of IQ SYBR Green Supermix, 1.25 µl of each primer (10 µm) and 1.5 µl of sterile distilled water. Thermal cycling conditions used were different for all six primers tested (Online Resource 1). Fluorescence signal (520 nm) was detected at the end of the elongation phase for each cycle. To evaluate amplification specificity, melt curve analysis was performed at the end of each PCR run. A melting curve profile was obtained by heating the mixture to 95 °C, cooling to 60 °C for each 10 s. To quantify unknown concentrations of each target gene, a standard curve was generated by the amplification of a tenfold dilution series of target gene in the presence of DNA extracted from Pseudomonas fluorescens isolates (control). The coefficient of correlation (R2) between the cycle threshold value (Ct) and target DNA concentration was between 0.990 and 0.997 while the PCR efficiency was between 95 and 98%. Subsequently, the gene copy number was calculated using the formula: number of copies = (DNA concentration × 6.022 × 1023)/(product length in bp × 1 × 109 × 650) (Aghamollaei et al. 2015).
Results
Responses of bacterial community pattern to temperature and incubation durations
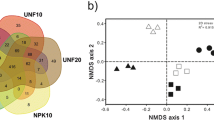
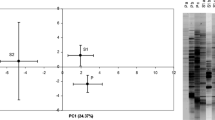
CAP ordination based on all samples suggested that the bacterial community pattern of the temperature-treated UC soil was more similar to the untreated UC soil compared to the bacterial community pattern of temperature-treated HC soil and untreated HC soil (Fig. 2a). Separately, the HC soil showed stronger sample dispersion than UC soil across treatments (Fig. 2b), with calculated PERMDISP average distance to centroid of 39.25 ± 1.08 for HC soil and 27.54 ± 1.93 for UC soil (pairwise T = 5.33, P = 0.001). Significant separations according to temperatures and incubation durations were detected in both HC and UC soil (P < 0.05; Table 1). Among them, greater effects of temperature and incubation duration were observed for bacterial composition in HC soil (pseudo-T for temperature = 3.77, pseudo-T for incubation duration = 44.18) compared to UC soil (pseudo-T for temperature = 2.96, pseudo-T for incubation temperature = 10.86) (Table 1). In addition, the influence of incubation duration was found to be stronger than temperature on bacterial community composition. For instance, clear clustering based on the duration of incubation was apparent in the CAP ordination plot for both HC and UC soil (Fig. 3). A follow-up pairwise PERMANOVA (Online Resource 2) comparing the effect of temperature across weeks of incubation suggests that no significant community changes had occurred at Week 4 for both the HC and UC soil. However, at Week 8 and Week 12, the HC soil bacterial community patterns in the 15 °C microcosms showed strong separation compared to the 5 and 10 °C microcosms. On the other hand, the UC soil bacterial assemblage from the 5 and 15 °C microcosms remained similar in the Week 8 and Week 12 samples, while both were significantly different to the 10 °C microcosms.
CAP ordination based on bacterial community composition inferred using TRFLP (UC open symbols, HC closed symbols). a Ordination with complete sample set including untreated soil; b ordination excluding untreated soil. Symbol: (open triangle) Week 4, (open inverted triangle) Week 8, (open square) Week 12, blue = 5 °C, green = 10 °C, red = 15 °C. (Color figure online)
CAP ordination of bacterial community composition in a hydrocarbon-contaminated Antarctic soil; b uncontaminated soil. Symbol: (open triangle) Week 4, (open inverted triangle) Week 8, (open square) Week 12, blue = 5 °C, green = 10 °C, red = 15 °C. Note that when compared with Fig. 2, the separation for each treatment is amplified when the ordination was conducted separately for HC and UC. (Color figure online)
Taxonomic distribution based on 454 pyrosequencing of the 16S rDNA
From the 454 pyrosequencing, 36,281 processed reads were obtained from the HC soil, and 42,281 processed reads were obtained from the UC soil. The sequencing coverage was between 91 and 97%. Based on the calculated α-diversity indices, HC soil was generally less diverse than the UC soil (Table 2) and the bacterial richness in both soils were generally positively correlated with the incubation temperature. At the highest phylogenetic rank, sixteen phyla were identified across the two soils (Fig. 4a), with two phyla specific only to UC soil and three to HC soil. Overall, both soils were dominated by Proteobacteria and Actinobacteria. Nevertheless, taxonomic shift in accordance to the incubation temperature was apparent. For instance, at 5 and 10 °C, HC soils were over-represented by Proteobacteria. However, the proportion of Proteobacteria was reduced and was overtaken by Actinobacteria at the highest incubation temperature (15 °C) at all incubation durations. UC soil, on the other hand, showed lower proportions of Acidobacteria and Bacteroidetes at 5 °C than in 10 and 15 °C.
The distribution pattern was highly variable when the comparison was made at genus level (Fig. 4b). In HC soil, Rhodococcus was the most prevalent bacterial genus. The dominance of Rhodococcus was particularly discernible at 15 °C, accounting for approximately 32–50% of the total detected genera. It is noteworthy that at genus level, the taxonomic distribution of bacteria in the HC soil microcosms at 15 °C was distinctly different from the untreated HC soil. In the UC soil, a high prevalence of unclassified taxa under Betaproteobacteria group was detected.
Bacterial community comparison between HC and UC soil
Significant differences in phylum-based and genus-based bacterial composition were detected based on AMOVA (P value < 0.001 for both Sorensen-distance and Hellinger distance matrices). A followed-up LEFse suggests that at phylum level, Actinobacteria and Saccharibacteria were enriched in the HC soil while Acidobacteria, Bacteroidetes, Chloroflexi, Gemmatimonadetes and members of unclassified phylum were more abundant in UC soil (LDA > 3.5, P < 0.01). In contrast, at genus level, Brevundimonas, Rhodococcus, Rhodopseudomonas, Sphingomonas, Novosphingobium, unclassified genus under Sphingomonadaceae, and unclassified genus under Gammaproteobacteria were greater in HC soil while Methylotenera, Porphyrobacter, Sulfuritalea, Thermomonas and unclassified genera under Actinobacteria, Comamonadaceae and Oxalobacteraceae were higher in UC soil (LDA ≥ 4, P < 0.01).
Soil chemical properties of microcosms under different temperatures
Soil chemical parameters were significantly different between HC and UC soil (PERMANOVA pseudo-F = 25.21, PMC = 0.001). For example, nitrate, nitrite, phosphate and the electrical conductivity were lower in UC soil compared to HC soil (Table 3). Significant differences in soil chemical properties in relation to temperature were detected in HC soil (PERMANOVA pseudo-F = 54.39, PMC = 0.001) but not in UC soil (pseudo-F = 2.27, PMC = 0.057). However, significant differences in the soil chemical parameters were observed for both soil when the data were compared across incubation durations (HC pseudo-F = 54.40, PMC = 0.001; UC pseudo-F = 8.18, PMC = 0.001). While the pH of both soils was weakly acidic, the pH of the HC soil decreased with incubation duration. Greater decrease in pH occurred at higher incubation temperatures (Table 3). A similar trend was also seen for the nitrate content in HC soil.
Changes in soil chemical properties in relation to bacterial assemblage pattern were modelled using DISTLM. There were no significant correlations between the six measured soil parameters (pH, electrical conductivity, water, nitrate, nitrite and phosphate contents) and bacterial community patterns in the HC soil. However, water content significantly correlated with the bacterial assemblage pattern in the UC soil (R2 = 9.0%, P = 0.003).
Association between the abundance of functional genes and bacterial assemblage pattern
The overall functional gene abundance is given in Online Resource 3. HC and UC soils showed significant differences in the abundance of functional genes (PERMANOVA pseudo-T = 3.8045, PMC = 0.006). There were no significant differences in functional genes in relation to temperature, whereas significant effect of incubation duration on the functional genes was detected in UC soil (PERMANOVA pseudo-T = 2.0755, P = 0.049) although not in HC soil. We subsequently modelled the bacterial assemblage pattern of the two soils based on functional genes abundance. Among the genes tested, chitinase GA, nirK and nosZ showed significant correlation to the overall bacterial community patterns in the HC and UC soil (cumulative R2 = 33.64%, P = 0.022). However, when we repeated the modelling separately using only HC soil alone, a significant correlation was only established between amoA gene and bacterial composition pattern (R2 = 5%, P = 0.018), while no correlation was detected between each functional gene and the bacterial community in the UC soil when considered alone.
Discussion
In the current study, we evaluated the effect of warming on bacterial community in hydrocarbon-contaminated soil and uncontaminated soil from Antarctica. The two soils were morphologically similar and were collected from closely located sites (> 50 m apart) near Casey Station, Windmill Island, East Antarctica. Soil microcosms were set up and incubated at 5, 10 and 15 °C. The temperatures used are within the range of normal surface soil temperature in Antarctica (0 to + 22 °C) (Balks et al. 2002) and comparable to similar studies (Ferguson et al. 2008; Yergeau and Kowalchuk 2008). In summer, soil temperature in Antarctica may be subjected to large fluctuations (Cary et al. 2010). In our current study, our soil microcosms were incubated in three different temperatures for up to 12 weeks each. This was to evaluate the effect of prolonged warming on the soil bacterial community in HC and UC soil.
Distinct bacterial community structure in HC and UC soil
Low bacterial richness is a general characteristic of HC Antarctic soil (Saul et al. 2005; Chong et al. 2009; Cury et al. 2015). Our results support this notion as the HC soil was found to harbour lower bacterial richness in comparison to the UC soil despite their close proximity and similar environmental conditions. The two soils had different bacterial compositions. The untreated HC soil had high proportions of taxa previously reported as hydrocarbon-degrading, such as Rhodococcus, Sphingomonas and Brevundimonas (Saul et al. 2005; Aislabie et al. 2006; Adriaenssens et al. 2014; van Dorst et al. 2016). In contrast, untreated UC soil had higher proportions of proteobacteria genera including Thermomonas were detected in the untreated UC soil (Fig. 4). The dominance of alkane and aromatic hydrocarbon-degraders in petroleum-contaminated Antarctic soil is common and is related to the selective pressure from the hydrocarbon contamination, and occurs at the expense of bacterial richness (Powell et al. 2006; Ferguson et al. 2008; Vázquez et al. 2013; Yang et al. 2016). Higher Acidobacteria, Bacteroidetes, and Gemmatimonadetes were also found in UC soil compared to HC soil. The lower representation of Acidobacteria in HC soil might be attributed to its preference for a low nutrient environment (Fierer et al. 2007; Ward et al. 2009), as well as its sensitivity to the hydrocarbon contamination (Saul et al. 2005; Vazquez et al. 2017).
Response of bacterial community composition to warming
Allison and Martiny (2008) defined resistance as “the degree to which microbial composition remains unchanged in the face of a disturbance” and resilience as “the rate at which microbial composition returns to its original composition after being disturbed”. It is hypothesised that communities with lower richness have less resistance to environmental perturbation (Allison and Martiny 2008; Chong et al. 2015; Isbell et al. 2015). In line with this, we showed that the bacterial community in HC soil was more sensitive to the incubation conditions (temperature and incubation duration) than the UC soil. For instance, there was a greater separation in bacterial community composition in the treated soil relative to the untreated soil in the HC than the UC soils (Fig. 2a). In addition, higher overall community variation was detected in the HC soil while the UC soil remained relatively homogenous throughout the experiment (Fig. 2b). The HC soil also had stronger response to temperature treatments especially at Week 12 under 15 °C, compared to the UC soil (Table 1, Fig. 3a), specifically, a sharp increase in Rhodococcus (Gram-positive) (Fig. 4). However, there was a decrease in 16S signatures belonging to an unclassified genus under Betaproteobacteria (Gram-negative) in UC soil at 15 °C (Fig. 4).
Dennis et al. (2013) observed increase in Gram-positive-to-Gram-negative bacteria ratio in desert soil from Mars Oasis, Alexander Island after simple nutrient supplementation and/or warming under open top chamber (OTC) for 10–12 months. However, they did not see the same response in the more northerly and more diverse location at Wynn Knolls, Signy Island. While the report by Dennis et al. (2013) established that slow community responses towards low temperature and nutrient supplement is possible at locations with low bacterial diversity, our data further suggest that significant shifts in bacterial community can occur within a short period (e.g., 3 months) for habitats under strong selective pressures, such as through warming of hydrocarbon-contaminated soil.
In a parallel microcosm study using soil from McMurdo Dry Valleys, de Scally et al. (2016) found no significant influence of warming to the soil microbial composition and extracellular enzymatic activities over an incubation period of 40 days. The difference from our study might be due to the higher microbial diversity in their soil and relatively lower bacterial richness in our soil (H′ = 4.6–5.3 in de Scalley et al. (2016) as opposed to H′ = 2.8–5.1 in our study), as well as longer incubation duration used in our experiment. It has been shown that some Antarctic soil communities are resistant to low magnitudes of warming (< 1 °C) (Bokhorst et al. 2007; Yergeau and Kowalchuk 2008). However, Rinnan et al. (2009) described significant positive correlation between mean soil temperature and minimum temperature for bacterial growth, and suggested that “every 1 °C rise in soil temperature will increase minimum growth temperature for Antarctic bacteria by 0.24–0.38 °C”.
In our current study, under all three incubation temperatures (5, 10 and 15 °C), the bacterial community composition in HC and UC soil microcosms was highly homogenous at Week 4, and compositional changes became apparent at Week 8 and Week 12 (Fig. 3, Table S2). In comparison, in another study of this group using tropical soil microcosms incubated at 25, 30 and 35 °C for durations of 2, 4 and 8 weeks, a dramatic bacterial community shift was observed after 2 weeks of warming, followed by adaptation/stabilisation at Week 4 (Supramaniam et al. 2016). The difference in bacterial community response to warming between tropical and Antarctic soil could be due to microbial density, diversity and stronger microbial resilience in tropical soil, while the low metabolic rate of cold-adapted bacteria in Antarctic soil might explain the delayed response to warming. Previous open top chamber (OTC) experiments across a latitudinal gradient in Antarctica suggested that Antarctic soil bacterial communities continue to change over 3 years under experimental warming at 0.5–2 °C (Yergeau et al. 2012).
In this study, the final hydrocarbon concentration in the microcosms was not measured because hydrocarbon degradation was not an objective. Therefore, it is unclear whether the differences in bacterial community shift influenced the level of hydrocarbon degradation in the microcosms. However, Aislabie et al. (2006) reported that substantial hydrocarbon degradation was achievable at higher temperatures (e.g., 23 °C) compared to lower temperatures such as those used here (5–15 °C). Similarly, under microcosm conditions, Ferguson et al. (2008) observed little difference in hydrocarbon mineralisation between sample incubated at 4 and 10 °C but a drastically higher degradation was observed at 42 °C. The low water content measured in our samples may further impede the degradation of the hydrocarbon. Low water content (< 5%) is a general characteristics of Antarctic mineral soil (Fell et al. 2006; Chong et al. 2009, 2012). However, the extremely low water content measured in the untreated (control) samples in this study (< 1%) might be due to the water loss during the sieving process. Nonetheless, the association of bacterial composition with water content was only established in the UC soil but not the HC soil. The difference may suggest that the impact of hydrocarbon in the soil outweighed the influence of the water content in the HC soil.
Responses of functional genes abundance and soil parameters to warming
Using soil from King George Island, maritime Antarctica, Han et al. (2013) recorded elevated nifH and amoA genes in microcosms incubated at 8 °C for 14 days as opposed to microcosms incubated at 5 °C. However, we did not find any significant difference in functional gene abundance with temperature or incubation duration. The non-response of C-cycling and N-cycling genes to temperature is consistent with other warming experiments conducted in Antarctica (Yergeau and Kowalchuk 2008; Yergeau et al. 2012; de Scally et al. 2016). Using lab-based microcosms, Wertz et al. (2007) demonstrated that microbial functional groups were resistant to diversity decline. Similarly, using Canadian High Arctic soil, Lamb et al. (2011) found that fertilisation and temperature treatments did not affect soil nitrogen fixation gene markers such as nosZ, pmoA and amoA. It is thus suggested that ecological function may be more resistant and resilient than community structure (Bissett et al. 2011). However, this is perhaps more relevant to highly diverse soils which support ecosystems with high functional redundancy (Chong et al. 2015). The lack of response of functional genes in relatively simpler ecosystems such as Antarctic soils is less expected especially for HC soil where the diversity is further reduced (see Awasthi et al. 2014). One possible explanation is the presence of high functional plasticity among the bacterial taxa supporting high functional redundancy in the soil (de Scally et al. 2016). By comparing hydrocarbon-contaminated and non-contaminated subtidal sediments from Polar and subpolar coastal environments, Espínola et al. (2018) showed that the metabolic potentials of the sediments are remarkably similar despite the distinct difference in community structure across sites.
In our study, the difference in bacterial composition and richness between HC and UC soil is not reflected in the functional gene abundance. For instance, only amoA gene showed correlation to the changes in bacterial community pattern in the HC soil while no correlation between functional gene abundance and bacterial community composition was detected in the UC soil. The lack of significance might also reflect that the functional gene number is highly heterogeneous across Antarctic soil replicates (Yergeau and Kowalchuk 2008).
In conclusion, our data suggest that warming can induce significant community shifts in Antarctic soil within 3 months. The effect of warming was stronger in the hydrocarbon-contaminated soil which is characterised by lower bacterial richness and a more highly specialised community than the uncontaminated soil. The bacterial community in the UC soil was more resistant to the temperatures tested than the HC soil and remained relatively homogenous throughout the experiment. It is noteworthy that a relatively simple methodology was employed in this study to infer the community assemblage pattern (i.e., TRFLP) and taxonomic identity (16S 454 pyrosequencing). The whole genome sequencing approach used in Espínola et al. (2018) could be used to further explore the direct relationship between the changes in bacterial community pattern and the overall metabolic potential of the sample. Nonetheless, rates of natural degradation in Antarctica are low (Bargagli 2008), and heating was found to be an effective strategy to increase bioremediation of hydrocarbon in Antarctica, especially where nutrient limitations are overcome with supplemental amendments (Ferguson et al. 2008; Delille et al. 2009; McDonald and Knox 2014). Our data showed that higher temperature (15 °C) selected for hydrocarbon-degraders that could potentially improve the hydrocarbon degradation rate. However, the effect might occur at the expense of community stability. Long term monitoring is therefore warranted to understand if the soil community composition is able to recover from the community change induced by both hydrocarbon contamination and warming.
References
Abdo Z, Schüette UME, Bent SJ, Williams CJ, Forney LJ, Joyce P (2006) Statistical methods for characterizing diversity of microbial communities by analysis of terminal restriction fragment length polymorphisms of 16S rRNA genes. Environ Microbiol 8:929–938. https://doi.org/10.1111/j.1462-2920.2005.00959.x
Adams BJ, Bardgett RD, Ayres E, Wall DH, Aislabie J, Bamforth S, Bargagli R, Cary C, Cavacini P, Connell L, Convey P, Fell JW, Frati F, Hogg ID, Newsham KK, O’Donnell A, Russell N, Seppelt RD, Stevens MI (2006) Diversity and distribution of Victoria Land biota. Soil Biol Biochem 38:3003–3018
Adriaenssens EM, Guerrero LD, Makhalanyane TP, Aislabie JM, Cowan DA (2014) Draft genome sequence of the aromatic hydrocarbon-degrading bacterium Sphingobium sp. Strain Ant17, isolated from Antarctic soil. Genome Announc. https://doi.org/10.1128/genomea.00212-14
Aghamollaei H, Moghaddam MM, Kooshki H, Heiat M, Mirnejad R, Barzi NS (2015) Detection of Pseudomonas aeruginosa by a triplex polymerase chain reaction assay based on lasI R and gyrB genes. J Infect Public Health 8:314–322. https://doi.org/10.1016/j.jiph.2015.03.003
Aislabie JM, Fraser R, Duncan S, Farrell RL (2001) Effects of oil spills on microbial heterotrophs in Antarctic soils. Polar Biol 24:308–313
Aislabie JM, Balks MR, Foght JM, Waterhouse EJ (2004) Hydrocarbon spills on Antarctic soils: effects and management. Environ Sci Technol 38:1265–1274. https://doi.org/10.1021/es0305149
Aislabie JM, Saul DJ, Foght JM (2006) Bioremediation of hydrocarbon-contaminated polar soils. Extremophiles 10:171–179. https://doi.org/10.1007/s00792-005-0498-4
Allison SD, Martiny JBH (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 105:11512–11519. https://doi.org/10.1073/pnas.0801925105
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46
Anderson MJ, Gorley RN, Clarke KR (2008) PERMANOVA+ for PRIMER: guide to software and statistical methods. PRIMER-E, Plymouth
Awasthi A, Singh M, Soni SK, Singh R, Kalra A (2014) Biodiversity acts as insurance of productivity of bacterial communities under abiotic perturbations. ISME J 8:2445–2452. https://doi.org/10.1038/ismej.2014.91
Balks MR, Paetzold RF, Kimble JM, Aislabie JM, Campbell IB (2002) Effects of hydrocarbon spills on the temperature and moisture regimes of Cryosols in the Ross Sea region. Antarctic Sci 14:319–326
Bargagli R (2008) Environmental contamination in Antarctic ecosystems. Sci Total Environ 400:212–226. https://doi.org/10.1016/j.scitotenv.2008.06.062
Bennett JR, Shaw JD, Terauds A, Smol JP, Aerts R, Bergstrom DM, Blais JM, Cheung WWL, Chown SL, Lea MA, Nielsen UN, Pauly D, Reimer KJ, Riddle MJ, Snape I, Stark JS, Tulloch VJ, Possingham HP (2015) Polar lessons learned: long-term management based on shared threats in Arctic and Antarctic environments. Front Ecol Environ 13:316–324. https://doi.org/10.1890/140315
Bissett A, Richardson AE, Baker G, Thrall PH (2011) Long-term land use effects on soil microbial community structure and function. Appl Soil Ecol 51:66–78. https://doi.org/10.1016/j.apsoil.2011.08.010
Bokhorst S, Huiskes A, Convey P, Aerts R (2007) Climate change effects on organic matter decomposition rates in ecosystems from the Maritime Antarctic and Falkland Islands. Glob Change Biol 13:2642–2653. https://doi.org/10.1111/j.1365-2486.2007.01468.x
Cary SC, McDonald IR, Barrett JE, Cowan DA (2010) On the rocks: the microbiology of Antarctic Dry Valley soils. Nat Rev Microbiol 8:129–138
Chong CW, Tan GYA, Wong RCS, Riddle MJ, Tan IKP (2009) DGGE fingerprinting of bacteria in soils from eight ecologically different sites around Casey Station, Antarctica. Polar Biol 32:853–860. https://doi.org/10.1007/s00300-009-0585-6
Chong CW, Pearce DA, Convey P, Tan GYA, Wong RCS, Tan IKP (2010) High levels of spatial heterogeneity in the biodiversity of soil prokaryotes on Signy Island, Antarctica. Soil Biol Biochem 42:601–610. https://doi.org/10.1016/j.soilbio.2009.12.009
Chong CW, Pearce DA, Convey P, Tan IKP (2012) The identification of environmental parameters which could influence soil bacterial community composition on the Antarctic Peninsula—a statistical approach. Antarct Sci 24:249–258. https://doi.org/10.1017/S0954102012000028
Chong CW, Pearce DA, Convey P (2015) Emerging spatial patterns in Antarctic prokaryotes. Front Microbiol 6:1058
Coulon F, Pelletier E, Gourhant L, Delille D (2005) Effects of nutrient and temperature on degradation of petroleum hydrocarbons in contaminated sub-Antarctic soil. Chemosphere 58:1439–1448. https://doi.org/10.1016/j.chemosphere.2004.10.007
Culman SW, Bukowski R, Gauch HG, Cadillo-Quiroz H, Buckley DH (2009) T-REX: software for the processing and analysis of T-RFLP data. BMC Bioinform 10:171. https://doi.org/10.1186/1471-2105-10-171
Cury JC, Jurelevicius DA, Villela HDM, Jesus HE, Peixoto RS, Schaefer CEGR, Bícego MC, Seldin L, Rosado AS (2015) Microbial diversity and hydrocarbon depletion in low and high diesel-polluted soil samples from Keller Peninsula, South Shetland Islands. Antarct Sci 27:263–273
de Scally SZ, Makhalanyane TP, Frossard A, Hogg ID, Cowan DA (2016) Antarctic microbial communities are functionally redundant, adapted and resistant to short term temperature perturbations. Soil Biol Biochem 103:160–170. https://doi.org/10.1016/j.soilbio.2016.08.013
Delille D, Pelletier E, Rodriguez-Blanco A, Ghiglione J-F (2009) Effects of nutrient and temperature on degradation of petroleum hydrocarbons in sub-Antarctic coastal seawater. Polar Biol 32:1521–1528. https://doi.org/10.1007/s00300-009-0652-z
Dennis PG, Newsham KK, Rushton SP, Ord VJ, O’Donnell AG, Hopkins DW (2013) Warming constrains bacterial community responses to nutrient inputs in a southern, but not northern, maritime Antarctic soil. Soil Biol Biochem 57:248–255. https://doi.org/10.1016/j.soilbio.2012.07.009
Dias RL, Ruberto L, Calabró A, Balbo AL, Del Panno MT, Mac Cormack WP (2015) Hydrocarbon removal and bacterial community structure in on-site biostimulated biopile systems designed for bioremediation of diesel-contaminated Antarctic soil. Polar Biol 38:677–687. https://doi.org/10.1007/s00300-014-1630-7
Espínola F, Dionisi HM, Borglin S, Brislawn CJ, Jansson JK, Mac Cormack WP, Carroll J, Sjöling S, Lozada M (2018) Metagenomic analysis of subtidal sediments from polar and subpolar coastal environments highlights the relevance of anaerobic hydrocarbon degradation processes. Microb Ecol 75:123–139
Fell JW, Scorzetti G, Connell L, Craig S (2006) Biodiversity of micro-eukaryotes in Antarctic Dry Valley soils with < 5% soil moisture. Soil Biol Biochem 38:3107–3119
Ferguson SH, Powell SM, Snape I, Gibson JAE, Franzmann PD (2008) Effect of temperature on the microbial ecology of a hydrocarbon-contaminated Antarctic soil: implications for high temperature remediation. Cold Reg Sci Technol 53:115–129. https://doi.org/10.1016/j.coldregions.2007.04.006
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Filler DM, Snape I, Barnes DL (2008) Bioremediation of petroleum hydrocarbons in cold regions. Cambridge University Press, Cambridge
Fryirs KA, Hafsteinsdóttir EG, Stark SC, Gore DB (2015) Metal and petroleum hydrocarbon contamination at Wilkes Station, East Antarctica. Antarct Sci 27:118–133
Han J, Jung J, Park M, Hyun S, Park W (2013) Short-term effect of elevated temperature on the abundance and diversity of bacterial and archaeal amoA genes in Antarctic soils. J Microbiol Biotechnol 23:1187–1196
Isbell F, Craven D, Connolly J, Loreau M, Schmid B, Beierkuhnlein C, Bezemer TM, Bonin C, Bruelheide H, de Luca E, Ebeling A, Griffin JN, Guo Q, Hautier Y, Hector A, Jentsch A, Kreyling J, Lanta V, Manning P, Meyer ST, Mori AS, Naeem S, Niklaus PA, Polley HW, Reich PB, Roscher C, Seabloom EW, Smith MD, Thakur MP, Tilman D, Tracy BF, van der Putten WH, van Ruijven J, Weigelt A, Weisser WW, Wilsey B, Eisenhauer N (2015) Biodiversity increases the resistance of ecosystem productivity to climate extremes. Nature 526:574–577. https://doi.org/10.1038/nature15374
Labud V, Garcia C, Hernandez T (2007) Effect of hydrocarbon pollution on the microbial properties of a sandy and a clay soil. Chemosphere 66:1863–1871. https://doi.org/10.1016/j.chemosphere.2006.08.021
Lamb EG, Han S, Lanoil BD, Henry GHR, Brummell ME, Banerjee S, Siciliano SD (2011) A High Arctic soil ecosystem resists long-term environmental manipulations. Glob Change Biol 17:3187–3194. https://doi.org/10.1111/j.1365-2486.2011.02431.x
Ludescher J, Bunde A, Franzke CLE, Schellnhuber HJ (2016) Long-term persistence enhances uncertainty about anthropogenic warming of Antarctica. Clim Dyn 46:263–271. https://doi.org/10.1007/s00382-015-2582-5
McDonald R, Knox OGG (2014) Cold region bioremediation of hydrocarbon contaminated soils: do we know enough? Environ Sci Technol 48:9980–9981. https://doi.org/10.1021/es5036738
McWatters RS, Wilkins D, Spedding T, Hince G, Raymond B, Lagerewskij G, Terry D, Wise L, Snape I (2016) On site remediation of a fuel spill and soil reuse in Antarctica. Sci Total Environ 571:963–973. https://doi.org/10.1016/j.scitotenv.2016.07.084
Muangchinda C, Chavanich S, Viyakarn V, Watanabe K, Imura S, Vangnai AS, Pinyakong O (2015) Abundance and diversity of functional genes involved in the degradation of aromatic hydrocarbons in Antarctic soils and sediments around Syowa Station. Environ Sci Pollut Res 22:4725–4735. https://doi.org/10.1007/s11356-014-3721-y
Powell SM, Ferguson SH, Bowman JP, Snape I (2006) Using real-time PCR to assess changes in the hydrocarbon-degrading microbial community in Antarctic soil during bioremediation. Microb Ecol 52:523–532. https://doi.org/10.1007/s00248-006-9131-z
Rinnan R, Rousk J, Yergeau E, Kowalchuk GA, Baath E (2009) Temperature adaptation of soil bacterial communities along an Antarctic climate gradient: predicting responses to climate warming. Glob Change Biol 15:2615–2625. https://doi.org/10.1111/j.1365-2486.2009.01959.x
Saul DJ, Aislabie JM, Brown CE, Harris L, Foght JM (2005) Hydrocarbon contamination changes the bacterial diversity of soil from around Scott Base, Antarctica. FEMS Microbiol Ecol 53:141–155. https://doi.org/10.1016/j.femsec.2004.11.007
Smith CJ, Danilowicz BS, Clear AK, Costello FJ, Wilson B, Meijer WG (2005) T-Align, a web-based tool for comparison of multiple terminal restriction fragment length polymorphism profiles. FEMS Microbiol Ecol 54:375–380. https://doi.org/10.1016/j.femsec.2005.05.002
Supramaniam Y, Chong CW, Silvaraj S, Tan IKP (2016) Effect of short term variation in temperature and water content on the bacterial community in a tropical soil. Appl Soil Ecol 107:279–289. https://doi.org/10.1016/j.apsoil.2016.07.003
Tin T, Fleming ZL, Hughes KA, Ainley DG, Convey P, Moreno CA, Pfeiffer S, Scott J, Snape I (2009) Impacts of local human activities on the Antarctic environment. Antarct Sci 21:3–33. https://doi.org/10.1017/S0954102009001722
Turner J, Barrand NE, Bracegirdle TJ, Convey P, Hodgson DA, Jarvis M, Jenkins A, Marshall G, Meredith MP, Roscoe H (2014) Antarctic climate change and the environment: an update. Polar Rec 50:237–259
Turner J, Lu H, White I, King JC, Phillips T, Hosking JS, Bracegirdle TJ, Marshall GJ, Mulvaney R, Deb P (2016) Absence of 21st century warming on Antarctic Peninsula consistent with natural variability. Nature 535:411–415. https://doi.org/10.1038/nature18645
van Dorst J, Siciliano SD, Winsley T, Snape I, Ferrari BC (2014) Bacterial targets as potential indicators of diesel fuel toxicity in subantarctic soils. Appl Environ Microbiol 80:4021–4033. https://doi.org/10.1128/aem.03939-13
van Dorst JM, Hince G, Snape I, Ferrari BC (2016) Novel culturing techniques select for heterotrophs and hydrocarbon degraders in a subantarctic soil. Sci Rep 6:36724. https://doi.org/10.1038/srep36724
Vazquez S, Monien P, Pepino Minetti R, Jurgens J, Curtosi A, Villalba Primitz J, Frickenhaus S, Abele D, Mac Cormack W, Helmke E (2017) Bacterial communities and chemical parameters in soils and coastal sediments in response to diesel spills at Carlini Station, Antarctica. Sci Total Environ 605–606:26–37
Vázquez S, Nogales B, Ruberto L, Hernandez E, Christie-Oleza J, Lo Balbo A, Bosch R, Lalucat J, Mac Cormack W (2009) Bacterial community dynamics during bioremediation of diesel oil-contaminated Antarctic soil. Microb Ecol 57:598–610. https://doi.org/10.1007/s00248-008-9420-9
Vázquez S, Nogales B, Ruberto L, Mestre C, Christie-Oleza J, Ferrero M, Bosch R, Mac Cormack WP (2013) Characterization of bacterial consortia from diesel-contaminated Antarctic soils: towards the design of tailored formulas for bioaugmentation. Int Biodeterior Biodegrad 77:22–30. https://doi.org/10.1016/j.ibiod.2012.11.002
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM, Bruce D, Creasy T, Daugherty SC, Davidsen TM, DeBoy RT, Detter JC, Dodson RJ, Durkin AS, Ganapathy A, Gwinn-Giglio M, Han CS, Khouri H, Kiss H, Kothari SP, Madupu R, Nelson KE, Nelson WC, Paulsen I, Penn K, Ren Q, Rosovitz MJ, Selengut JD, Shrivastava S, Sullivan SA, Tapia R, Thompson LS, Watkins KL, Yang Q, Yu C, Zafar N, Zhou L, Kuske CR (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056. https://doi.org/10.1128/aem.02294-08
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Guillaumaud N, Le Roux X (2007) Decline of soil microbial diversity does not influence the resistance and resilience of key soil microbial functional groups following a model disturbance. Environ Microbiol 9:2211–2219. https://doi.org/10.1111/j.1462-2920.2007.01335.x
Yang S, Wen X, Shi Y, Liebner S, Jin H, Perfumo A (2016) Hydrocarbon degraders establish at the costs of microbial richness, abundance and keystone taxa after crude oil contamination in permafrost environments. Sci Rep 6:37473. https://doi.org/10.1038/srep37473
Yergeau E, Kowalchuk GA (2008) Responses of Antarctic soil microbial communities and associated functions to temperature and freeze-thaw cycle frequency. Environ Microbiol 10:2223–2235. https://doi.org/10.1111/j.1462-2920.2008.01644.x
Yergeau E, Bokhorst S, Kang S, Zhou J, Greer CW, Aerts R, Kowalchuk GA (2012) Shifts in soil microorganisms in response to warming are consistent across a range of Antarctic environments. ISME J 6:692–702
Acknowledgements
The HC and UC Antarctic soils were kindly provided by the Australian Antarctic Division through Australian Antarctic Science Project #4036 (Remediation of Petroleum Contaminants in the Antarctic and subantarctic). This work was funded by UMRG (RP007-2012B) and YPASM fellowship (IMUR121/12). The authors thank University of Malaya and International Medical University for providing research facility and support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chong, C.W., Silvaraj, S., Supramaniam, Y. et al. Effect of temperature on bacterial community in petroleum hydrocarbon-contaminated and uncontaminated Antarctic soil. Polar Biol 41, 1763–1775 (2018). https://doi.org/10.1007/s00300-018-2316-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-018-2316-3