Abstract
Biological soil crusts (BSCs) are key components of polar ecosystems. These complex communities are important for terrestrial polar habitats as they include major primary producers that fix nitrogen, prevent soil erosion and can be regarded as indicators for climate change. To study the genus richness of microalgae and Cyanobacteria in BSCs, two different methodologies were employed and the outcomes were compared: morphological identification using light microscopy and the annotation of ribosomal sequences taken from metatranscriptomes. The analyzed samples were collected from Ny-Ålesund, Svalbard, Norway, and the Juan Carlos I Antarctic Base, Livingston Island, Antarctica. This study focused on the following taxonomic groups: Klebsormidiophyceae, Chlorophyceae, Trebouxiophyceae, Xanthophyceae and Cyanobacteria. In total, combining both approaches, 143 and 103 genera were identified in the Arctic and Antarctic samples, respectively. Furthermore, both techniques concordantly determined 15 taxa in the Arctic and 7 taxa in the Antarctic BSC. In general, the molecular analysis indicated a higher microalgal and cyanobacterial genus richness (about 11 times higher) than the morphological approach. In terms of eukaryotic algae, the two sampling sites displayed comparable genus counts while the cyanobacterial genus richness was much higher in the BSC from Ny-Ålesund. For the first time, the presence of the genera Chloroidium, Ankistrodesmus and Dunaliella in polar regions was determined by the metatranscriptomic analysis. Overall, these findings illustrate that only the combination of morphological and molecular techniques, in contrast to one single approach, reveals higher genus richness for complex communities such as polar BSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Arctic region and Antarctica are extreme environments defined by low temperatures year-round, low water availability, and large seasonal fluctuations with long periods of darkness during winter versus continuous irradiance in summer (Thomas et al. 2008a). Thus, plant cover is sparse and biodiversity is generally low, e.g. only two autochthonous flowering plants occur in Antarctica and these are mainly found in a few suitable areas along the coast (Thomas et al. 2008b; Pointing et al. 2015). These ecosystems are instead dominated by biological soil crusts (BSCs), which are the main terrestrial primary producers and also important ecosystem engineers (Evans and Johansen 1999; Yoshitake et al. 2010; Pointing and Belnap 2012; Williams et al. 2017). BSCs are communities of eukaryotic algae, Cyanobacteria, lichens, bryophytes, as well as heterotrophic fungi and bacteria which colonize the surface and top millimeters of the soil (Belnap et al. 2001a; Belnap 2006; Büdel et al. 2016). Typically occurring phyla of eukaryotic microalgae are Streptophyta (Klebsormidiophyceae, Zygnematophyceae), Chlorophyta (Chlorophyceae, Trebouxiophyceae, Ulvophyceae) and Ochrophyta (Bacillariophyceae, Eustigmatophyceae, Xanthophyceae) (Büdel et al. 2016). Certain crust organisms, such as Cyanobacteria and eukaryotic algae, form a matrix which aggregates the living cells and soil particles (Belnap 2006; Breen and Lévesque 2008). Hence, BSCs provide protection against soil erosion and cryoturbation (Evans and Johansen 1999). Moreover, BSCs enhance water and nutrient retention and the crust-associated Cyanobacteria fix nitrogen which in turn creates improved conditions (fertilization) for seed germination and plant growth (Belnap et al. 2001b; Breen and Lévesque 2008). This accumulation of biomass forms the nutritional basis for higher trophic levels such as the Svalbard reindeer (Rangifer tarandus platyrhynchus; Elster et al. 1999; Cooper and Wookey 2001). In general, BSCs are essential components of sparse ecosystems such as the polar deserts. These communities are valuable biological indicators for abiotic factors, ecological health and climate change (Belnap et al. 2001b; Pushkareva et al. 2016). Due to global warming which particularly affects the Arctic and the Antarctic Peninsula, an invasion by alien species is likely, which in turn might alter the BSC composition (Frenot et al. 2005; Chown et al. 2012; Pushkareva et al. 2016; Lee et al. 2017). Hence, studying individual BSC organisms, as well as the whole community, is important to fully address biodiversity, which is a prerequisite for monitoring changes, as well as for predictions of future climate change (Pushkareva et al. 2016).
The identification of autotrophic microorganisms in BSCs, such as eukaryotic algae and Cyanobacteria, is traditionally performed using light microscopy to examine BSCs in situ, or through established cultures of selected crust organisms (Skinner 1932; Bischoff and Bold 1963; Waterbury and Stanier 1978; Büdel et al. 2009). The taxa of interest are typically identified using morphological traits such as color, cell size and shape, or motility (Prescott 1964; Lind and Brook 1980; Cox 1996; John et al. 2002). The utilization of light microscopy for identification is rather quick and inexpensive compared to molecular techniques (Misawa 1999), but requires expert knowledge for many taxa as morphological features are often difficult to recognize and distinguish (Manoylov 2014). In addition, morphological features are not always stable as they can change in response to environmental factors (Albrecht et al. 2017). As an alternative, a number of molecular markers, so-called barcodes, can be used if unialgal cultures are available (Vieira et al. 2016). Typical barcode sequences are the 16S/18S rRNA gene and the internal transcribed spacer (ITS) rDNA, the RuBisCO large subunit (rbcL), the plastid elongation factor tufA, the cytochrome oxidase I (COX I), as well as internal simple sequence repeats (ISSR) of microsatellite regions (Buchheim and Chapman 1991; Wilmotte 1994; Doyle et al. 1997; An et al. 1999; Evans et al. 2007; Shen 2008; Sherwood et al. 2008; Hall et al. 2010). A huge pitfall is that the majority of microorganisms present in environmental samples, such as BSCs, are unculturable (Ward et al. 1990; Schloss and Handelsman 2005; Shi et al. 2009; Massana et al. 2014). To overcome the limitation of cultivation, biodiversity assessments can be performed by metabarcoding (Taberlet et al. 2012; Yoon et al. 2016; Elferink et al. 2017). For this purpose, total DNA is extracted from an environmental sample (eDNA) and used as a template to generate an amplicon mixture from a barcoding gene (Taberlet et al. 2012). Subsequently, the generated PCR products are sequenced with a high-throughput sequencing (HTS) technique, such as Roche 454 or Illumina (Margulies et al. 2005; Bentley et al. 2008; Taberlet et al. 2012). The identification of taxa is carried out by annotating the obtained sequence reads against an adequate database and sequence counts provide information about taxonomic abundance in the sample (Pawlowski et al. 2011; Taberlet et al. 2012). However, metabarcoding also exhibits various pitfalls such as the introduction of sequence errors during PCR, the design of suitable metabarcoding primers, covering all taxa of interest, and, again, the need for appropriate reference databases (Taberlet et al. 2012). Fortunately, PCR-dependent bias and the dependency on single barcodes can be avoided by exploiting shotgun metagenomics and metatranscriptomics (Urich et al. 2008, 2014). Similar to metabarcoding, total nucleic acids are isolated from an environmental sample but the amplification step is omitted (Urich et al. 2008, 2014). Instead, total DNA or cDNA is directly applied to HTS and the resulting sequences are assembled for any desired gene or transcript of interest, e.g. the small ribosomal subunit RNA (SSU; Geisen et al. 2015). This powerful approach enables a more reliable taxonomic identification compared to metabarcoding but relies on the availability of correctly determined sequence data (Urich et al. 2008; Klimke et al. 2011). Additionally, the isolation of high-quality nucleic acids from soil samples, suitable for HTS, can be difficult due to the presence of humic substances and exocellular RNase activity (Greaves and Wilson 1970; Rippin et al. 2016).
The present study focused on a methodological comparison of morphological and molecular approaches to analyze the microalgal and cyanobacterial genus richness of Arctic and Antarctic BSCs. Studying polar BSCs improves the knowledge on these microecosystems, their biodiversity and ecological structure. In this study, the identification of various groups of eukaryotic algae (Klebsormidiophyceae, Chlorophyceae, Trebouxiophyceae, Xanthophyceae) and Cyanobacteria in one Arctic and one Antarctic BSC sample was carried out. Eukaryotic and prokaryotic algae were identified using light microscopy in combination with suitable taxonomic keys. Furthermore, active species in the crust were targeted by applying a metatranscriptomic approach and algal as well as cyanobacterial taxa were identified using fully assembled SSUs. The final results were combined and compared to assess the comprehensiveness of both techniques regarding the genus richness.
Materials and methods
Sampling
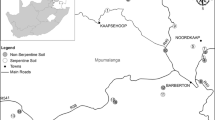
BSC samples were collected during expeditions to the Arctic and Antarctica in August 2014 and February 2015, respectively. The Arctic BSC (NÅ; cf. Rippin et al. 2016; Borchhardt et al. 2017; Williams et al. 2017, Jung et al. (in preparation)) was sampled in close vicinity to Ny-Ålesund, Svalbard, Norway (78°55′26.33″N 11°55′23.84″E; Fig. 1a). The research station Ny-Ålesund is located at Kongsfjorden and its climate is classified as polar tundra (ET) according to the Köppen-Geiger system (Peel et al. 2007; Vogel et al. 2012). The temperature is low year-round with an annual average of − 4.5 °C, the highest monthly temperature is in July with 5.8 °C and the lowest in March with −12 °C (all temperatures are averaged from August 1993 to July 2011; Maturilli et al. 2013). The mean annual precipitation is 433 mm (Førland et al. 2011). Williams et al. (2017) found 90% of the area to be covered by BSCs. The Antarctic BSC (Gr1; cf. Williams et al. 2017) was collected close by the Spanish Juan Carlos I Antarctic Base at Livingston Island, South Shetland Islands (62°39′55.44″S 60°23′42.76″W; Fig. 1b). Similar to Svalbard, the South Shetland Islands can be regarded as polar tundra (ET) according to the Köppen-Geiger system (Pereira et al. 2006; Michel et al. 2014). The temperature at the Juan Carlos I Antarctic Base averages − 1 °C annually, with the highest mean temperature (2.1 °C) in austral summer and the lowest mean temperature (− 4.6 °C) in winter (all temperatures are averaged from December 2001 to April 2003 and January 2007 to February 2011; Bañón et al. 2013). The site receives on average 444.5 mm of precipitation annually (Bañón et al. 2013). The BSC coverage is approximately 43% with skeletal soils being predominant (Williams et al. 2017). BSC samples for metatranscriptomic analysis (1 g) were preserved using the LifeGuard™ Soil Preservation Solution (MO BIO Laboratories, Carlsbad, CA, USA).
Overview map of sampling sites in the Arctic and Antarctica. a Svalbard in the Arctic. Samples were taken at Ny-Ålesund (NÅ). b Livingston Island which is part of the South Shetland Islands. Isolates were collected in close vicinity of the Juan Carlos I Antarctic Base (Gr1). Maps were created using snazzymaps.com
Cultivation and microscopy
Enrichment cultures of eukaryotic algae were established by transferring the sampled BSCs (NÅ, Gr1) onto solid 1.5% Difco™ Agar (Becton Dickonson, Heidelberg, Germany) containing Bold’s Basal Medium modified according to Starr and Jeffrey (1993). The cultures were kept at 15 °C, 30 μmol photons m−2 s−1 under a 16:8-h light–dark regime. A detailed outline on the procedure is given in Borchhardt et al. (2017b). The identification of genera or species was carried out by light microscopy (BX51, Olympus, Tokyo, Japan) with oil-immersion and a 1000-fold magnification. Relevant identification keys are listed in Borchhardt et al. (2017b). A complete list of the identified Arctic algae (NÅ) was published by Borchhardt et al. (2017a) and is used for comparison in the present study.
Cyanobacterial genera, present in the Arctic (NÅ) and Antarctic (Gr1) BSC were directly determined using light microscopy with oil immersion and a 630-fold magnification using identification keys (Geitler 1932; Komárek and Anagnostidis 1998, 2005). Cyanobacteria were additionally pre-cultivated on BG-11 medium at 15–17 °C, 20–50 µm photons m−2 s−1 under a 14:10-h light–dark cycle and identified using suitable taxonomic keys as described by Williams et al. (2016). The data of Jung et al. (in preparation), which examined the cyanobacterial diversity of both sites, were compared to the molecular dataset.
RNA isolation and sequencing
The Arctic sample (NÅ) was processed according to Rippin et al. (2016) using the CTAB protocol, DNase I (Thermo Fisher Scientific, Waltham, MA, USA) and the RNeasy MinElute Cleanup Kit (Qiagen, Hilden, Germany). Due to low RNA yields, a total of six biological replicates were extracted and combined to obtain three pooled replicates. RNA from three biological replicates, collected at Livingston Island (Gr1), was isolated using the Spectrum™ Plant Total RNA Kit (Sigma-Aldrich, St. Louis, MO, USA), treated with DNase I (Thermo Fisher Scientific, Waltham, MA, USA) and purified using the RNeasy MinElute Cleanup Kit (Qiagen, Hilden, Germany) as described by Rippin et al. (2016). Single samples yielded sufficient amounts of RNA.
All RNA samples were further processed by Eurofins Genomics (Ebersberg, Germany). The processing included quality control utilizing the BioAnalyzer (Agilent Technologies, Santa Clara, CA, USA) and library preparation for both triplicates. Eukaryotic mRNA was enriched using oligo-(dT) beads, fragmented and, subsequently, cDNA was synthesized using random hexamers. Finally, Illumina compatible adapters were ligated. The libraries of the individual samples NÅ and Gr1 were applied to an Illumina HiSeq 2500, all triplicates multiplexed on one lane, using 125 bp paired-end and single-end mode, respectively. For sample NÅ, the HiSeq Control Software 2.2.58, RTA 1.18.64 and bcl2fastq-1.8.4 were used while the detected signals from sample Gr1 were processed by operating the HiSeq Control Software 2.2.38, RTA 1.18.61 and bcl2fastq-1.8.4 (Illumina, San Diego, CA, USA).
Bioinformatics
The obtained raw reads (SRA bioproject PRJNA421095) were quality trimmed using Trimmomatics 0.35 (Bolger et al. 2014) and rRNA gene reads were filtered out using SortMeRNA 2.1 (Kopylova et al. 2012) and the SILVA SSU NR Ref 119. All rRNA SSU reads were assembled using EMIRGE 0.61.0 (Miller et al. 2011) combined with the SILVA SSU NR Ref 128. The script emirge_amplicon.py for single-end reads with the flag phred33 was run. Only successfully assembled SSUs were considered. The assigned genera for Klebsormidiophyceae, Chlorophyceae, Trebouxiophyceae, Xanthophyceae and Cyanobacteria were extracted and compiled with an in-house R-script (Version 3.3.2).
Results
The genus richness of phototrophic organisms in BSCs, isolated from Svalbard (NÅ) and Livingston Island (Gr1), was assessed using morphological features and annotated environmental rRNA SSU sequences. We obtained 18,041,893, 28,446,946 and 116,112,134 forward raw reads for sample NÅ and 62,235,763, 70,163,318 and 48,605,923 for Gr1. After quality trimming and rRNA SSU filtering we ended up with 8936, 7818 and 38,320 sequences for NÅ and 706,549, 1081,090 and 8646 for Gr1. This study focused on genera belonging to the Klebsormidiophyceae, Chlorophyceae, Trebouxiophyceae, Xanthophyceae or Cyanobacteria. All the taxa that were identified are included in the supplementary information (Online Resource 1). Additional taxonomic groups, for example Fungi, that were identified in the molecular analysis were not further investigated in this study.
The Arctic BSC
The Arctic sample contained 26 genera of eukaryotic algae and Cyanobacteria identified by light microscopy while 132 were assigned by metatranscriptomic analysis. The average number of genera in the metatranscriptomic triplicates was 71.7 ± 36.1 (SD). In total, 15 genera were determined by both methodologies, compared to 117 and 11 genera which were solely found by molecular and morphological identification, respectively (Fig. 2a). The utilized 18S reference database contained sequences for 5 of the 11 genera identified by morphology. Figures 3a and 4a display all genera identified with 1, 44, 25, 6 and 67 taxa belonging to Klebsormidiophyceae, Chlorophyceae, Trebouxiophyceae, Xanthophyceae and Cyanobacteria, respectively. The genera Coccomyxa, Elliptochloris, Stichococcus (Trebouxiophyceae), Leptolyngbya, Microcoleus, Nostoc and Phormidium (Cyanobacteria) were found in the culture isolates and in all three metatranscriptomes while Klebsormidium (Klebsormidiophyceae), Chlorococcum, Coelastrella, Desmodesmus, Lobochlamys (Chlorophyceae), Botrydiopsis, Heterococcus (Xanthophyceae) and Stigonema (Cyanobacteria) were identified in at least one of the three replicates and by light microscopy. All three metatranscriptomes per site included rRNA SSU sequences of the following genera, which could not be cultivated: Chloromonas, Dictyococcus, Oedogonium, Scenedesmus (Chlorophyceae), Chlorella, Chloroidium, Dictyochloropsis (Trebouxiophyceae), Anabaena, Aphanizonemon, Arthronema, Calothrix, Chroococcidiopsis, Crinalium, Cyanothece, Cylindrospermum, Gloeobacter, Nodularia, Tolypothrix and Trichormus (Cyanobacteria). The morphology-based method identified the eukaryotic genera Gloeocystis*, Macrochloris*, Neocystis*, Tetracystis (Chlorophyceae), Desmococcus*, Koliella, Lobosphaera (Trebouxiophyceae), Pleurochloris*, Pleurogaster*, Tribonema and Xanthonema (Xanthophyceae), of which six were not represented by 18S rRNA gene sequences in the reference database (asterisk).
All genera identified belonging to either Klebsormidiophyceae (Klebsormidium), Chlorophyceae, Trebouxiophyceae or Xanthophyceae. The metatranscriptome (blue and violet shades) is separated into three replicates while the culture isolates are represented by a single category (red, orange). Chords connect the methodologies with the identified genera. Genera displayed in gray were absent from the database used for the metatranscriptomic analyses. a NÅ; b Gr1
The Antarctic BSC
A total of 15 genera, present in the Antarctic BSC, were determined based on morphology while molecular identification yielded 95 taxa. The genera mean value of the three molecular replicates was 51.3 ± 5.7 (SD). Both techniques found seven genera while 8 and 88 genera were solely determined by morphology or sequence homology search, respectively (Fig. 2b). For three genera, identified from culture isolates, rRNA reference sequences were present in the database. Regarding the different taxonomic groups, several genera were determined: one genus of Klebsormidiophyceae, 53 genera of Chlorophyceae, 28 genera of Trebouxiophyceae, six genera of Xanthophyceae and 16 genera of Cyanobacteria (Figs. 3b and 4b). The genera Coccomyxa (Trebouxiophyceae), Leptolyngbya, Nostoc and Phormidium (Cyanobacteria) could be determined morphologically and were identified in all replicates used for the molecular approach. In addition, the genera Elliptochloris, Prasiola (Trebouxiophyceae), Heterococcus (Xanthophyceae) were determined by morphology and present in at least one out of three metatranscriptomic replicates. All three replicates prepared for molecular identification included the genera Ankistrodesmus, Bracteacoccus, Chlamydomonas, Chloromonas, Dunaliella, Hemiflagellochloris, Monoraphidium, Oedogonium, Scenedesmus (Chlorophyceae), Botryococcus, Chlorella, Chloroidium, Heterochlorella, Nannochloris, Stichococcus (Trebouxiophyceae), Botrydiopsis (Xanthophyceae) and Chamaesiphon (Cyanobacteria); however, cultivation failed for these organisms. In contrast, cultures were established for the genera Chlorococcum, Heterotetracystis, Mychonastes, Neocystis* (Chlorophyceae), Desmococcus*, Muriella*, Myrmecia* (Trebouxiophyceae) and Pleurochloris* (Xanthophyceae) which are missing in the rRNA dataset. Five of these genera were not represented by 18S sequences in the used database (asterisk).
Site comparison
By comparing the Arctic and Antarctic crust samples we found 63 genera that were shared between both locations (Fig. 5). However, the BSC collected from Svalbard, included 80 microalgae and Cyanobacteria which were not shared with the Antarctic sample, although the sample from Livingston Island contained 40 additional taxa. The genus Klebsormidium occurred at both sites. The biggest portion of chlorophycean (28) and trebouxiophycean (16) genera was found in both samples. Regarding Xanthophyceae, both BSCs contained 6 genera of which 3 were shared between sites. The analyses of the Arctic and Antarctic samples revealed 67 and 16 cyanobacterial genera to be present, respectively. An overlap of 15 cyanobacterial genera was detected.
Discussion
Overall, the molecular survey of the BSC samples revealed a higher degree of genera richness for Chlorophyceae, Trebouxiophyceae and Cyanobacteria compared to the culture and morphology-based methodology. However, the rRNA SSU identification did not recover all genera determined by light microscopy. Regarding Cyanobacteria, the Arctic crust sample exhibited a much higher variety compared to the sample collected at Livingston Island.
Assessment of microalgal and cyanobacterial genera richness
The genus Klebsormidium (Klebsormidiophyceae) is typically found in BSC communities and has been previously found to colonize soils of both Livingston Island and Svalbard (Büdel 2001; Kastovská et al. 2005; Zidarova 2008; Borchhardt et al. 2017b). Our metatranscriptomic analyses confirmed the presence of this streptophytic taxon in the polar regions, although Klebsormidium was not identified by light microscopy in the sample collected from Livingston Island.
Several chlorophycean genera were identified in both the Arctic and the Antarctic samples by metatranscriptomic analysis: Ankistrodesmus, Chlamydomonas, Chloromonas, Coelastrella, Desmodesmus, Dictyococcus, Dunaliella, Lobochlamys, Monoraphidium, Oedogonium and Scenedesmus. Coelastrella, Desmodesmus and Lobochlamys were also found in Arctic culture isolates. All the aforementioned genera are known to occupy terrestrial habitats from temperate to cold climates (Büdel 2001; Borie and Ibraheem 2003; Lürling 2003; Tschaikner et al. 2007; Matuła et al. 2007; Thorn and Lynch 2007; Uzunov et al. 2008; Zidarova 2008; de Wever et al. 2009; Wu et al. 2016; Büdel et al. 2016; Pushkareva et al. 2016; Schulz et al. 2016; Borchhardt et al. 2017b). For Chlamydomonas, Chloromonas, Coelastrella and Monoraphidium, earlier records exist confirming their occurrence in both polar regions; however, Dictyococcus, Oedogonium and Scenedesmus have only been identified from Arctic BSCs (Matuła et al. 2007; Zidarova 2008; Pushkareva et al. 2016; Borchhardt et al. 2017b). The genera Chlamydomonas, Coelastrella and Scenedesmus were previously identified from Svalbard and Monoraphidium from Livingston Island (Matuła et al. 2007; Zidarova 2008; Pushkareva et al. 2016). Previous research describing the genera Ankistrodesmus and Dunaliella in polar habitats was not found; therefore, this is the first study which confirms their presence in Svalbard and Livingston Island BSCs. Species of Chlorococcum were found in culture isolates prepared from both the Arctic and Antarctic BSCs as well as in the metatranscriptomic dataset generated from the NÅ sample. Kastovská et al. (2005) also found Chlorococcum in close vicinity to Ny-Ålesund and Borchhardt et al. (2017b) identified this genus from samples collected from Ardley and King George Island, Antarctica. The genus Tetracystis was identified from both BSC communities. Based on morphology, Tetracystis was found in the Arctic sample and metatranscriptomics confirmed its presence in the BSC from Livingston Island. Evidence exists that suggests Tetracystis colonizes the soil of Arctic and Antarctica ecosystems (Broady 1986; Giełwanowska and Olech 2012; Patova et al. 2015; Pushkareva et al. 2016). The following eukaryotic algae were only identified in culture isolates: Gloeocystis and Macrochloris were present in isolates from the Arctic while Heterotetracystis and Mychonastes grew in the Antarctic enrichment cultures. The genus Neocystis was found in both samples. For almost five genera their presence in polar and alpine ecosystems has been reported previously (Matuła et al. 2007; Zidarova 2008; Lukešová et al. 2010; Andreyeva and Kurbatova 2014; Patova et al. 2015; Pushkareva et al. 2016; Borchhardt et al. 2017b).
Both the Arctic and Antarctic metatranscriptomic datasets and culture isolates contained the trebouxiophycean genus Coccomyxa. Previous studies also found that this genus is an inhabitant of Svalbard and Livingston Island ecosystems (Matuła et al. 2007; Zidarova 2008; Pushkareva et al. 2016). Coccomyxa has a ubiquitous distribution and exhibits an extraordinary versatility, it occurs in terrestrial mats, in associations with soil, Fungi and bryophytes, and is also planktonic in freshwater systems (Schmidle 1901; Vogel 1955; Peveling and Galun 1976; Büdel 2001; Darienko et al. 2015). An ecophysiological study on three Coccomyxa strains isolated from Antarctic BSC samples points to pronounced cold and drought tolerance (Pfaff et al. 2016). Rivasseau et al. (2016) even found a new species of Coccomyxa, C. actinabiotis, in the cooling pool of a nuclear power plant. Thus, it is not surprising that members of this genus can colonize the extreme habitats which prevail in the polar regions. The green alga Elliptochloris generally occupies terrestrial habitats and associates with Fungi to form lichens (Ettl and Gärtner 2014). Furthermore, this genus was found in the Canadian Arctic and King George Island, Antarctica (Pushkareva et al. 2016; Borchhardt et al. 2017b). Both methodologies that were used in this study confirm that Elliptochloris is present in Svalbard and Livingston Island BSCs. In contrast, the genus Stichococcus was found in both the Arctic and the Antarctic metatranscriptomic data and in isolates from Livingston Island but not from Svalbard. This alga has often been identified as a member of BSC communities (Kastovská et al. 2005). Different species of Stichococcus have been previously found to colonize the soils of both Livingston Island and Svalbard (Kastovská et al. 2005; Zidarova 2008). The green alga Prasiola was identified through both the metatranscriptomic and microscopic identification for Livingston Island terrestrial habitats. However, only the Arctic metatranscriptomes showed evidence for Prasiola species. These findings are supported by a previous survey on species richness which reported Prasiola from both Livingston Island and Svalbard (Matuła et al. 2007; Zidarova 2008). Based solely on the metatranscriptomic analysis, the genera Chlorella, Chloroidium and Nannochloris were identified in both the Arctic and Antarctic BSC samples. Different species of Chlorella have been previously isolated from Svalbard and Livingston Island (Matuła et al. 2007; Zidarova 2008; Pushkareva et al. 2016). Vishnivetskaya (2009) identified Nannochloris in Siberian permafrost using molecular techniques, which suggests that this alga can survive in extreme conditions. The genus Chloroidium has been previously identified in BSCs; however, it has not been reported from polar habitats so far (Schulz et al. 2016; Williams et al. 2016). Therefore, this study provides the first molecular record of Chloroidium in Arctic and Antarctic ecosystems. The presence of Lobosphaera in the Arctic and Antarctic BSC samples was supported by the morphological and metatranscriptomic identification. Additionally, Giełwanowska and Olech (2012) identified Lobosphaera as a photobiont of Antarctic lichens. Based on morphological identification, the genus Desmococcus was found in BSC samples from both sites, Koliella was identified in isolates from the Arctic, while Muriella and Myrmecia grew in the Antarctic enrichment cultures. These findings are in accordance with previous studies reporting the presence of these genera in polar and alpine habitats (Kawecka 1986; Takeuchi 2001; Zidarova 2008; Lukešová et al. 2010; Andreyeva and Kurbatova 2014; Czerwik-Marcinkowska et al. 2015; Patova et al. 2015; Pushkareva et al. 2016; Borchhardt et al. 2017b).
The xanthophyte Heterococcus was identified by both techniques from the Svalbard and Livingston Island samples. Pushkareva et al. (2016) and Borchhardt et al. (2017b) also identified this genus in BSCs collected in the Canadian Arctic and King George Island, Antarctica. The genus Botrydiopsis is also typically found in BSCs and was previously isolated from other Arctic and Antarctic locations (Broady 1976; Büdel 2001; Pushkareva et al. 2016). Our metatranscriptomic analyses support the presence of this xanthophycean alga in the polar regions. However, it was identified by light microscopy only in the sample collected from Livingston Island. On the other hand, the genera Pleurogaster, Tribonema and Xanthonema were only present in isolates from the Arctic, while Pleurochloris was identified in both samples by means of morphological methods. Pleurochloris, Tribonema and Xanthonema have been previously found in polar and alpine ecosystems (Matuła et al. 2007; Zidarova 2008; Patova et al. 2015; Pushkareva et al. 2016; Borchhardt et al. 2017b). However, the genus Pleurogaster had previously only been isolated from terrestrial habitats outside polar regions (Stoyneva 2000).
All metatranscriptomes and morphological analyses identified the cyanobacterial genera Leptolyngbya, Nostoc and Phormidium from Svalbard and Livingston Island samples, which was also reported by Pushkareva et al. (2016) and Zidarova (2008), respectively. All genera occur in various climates and in aquatic and terrestrial habitats (Rippka et al. 1979; Büdel 2001; Casamatta et al. 2005; Dojani et al. 2014). The cyanobacterium Microcoleus also has a ubiquitous distribution, is often associated with BSCs and has been found in both Arctic and Antarctic environments (Büdel 2001; Pushkareva et al. 2015). All metatranscriptomic datasets contained evidence of Microcoleus; however, it was only identified from Svalbard through the morphological investigation. The genus Stigonema was identified from Svalbard through both metatranscriptomic and morphological methods, which is in agreement with other reports (Matuła et al. 2007; Pushkareva et al. 2015). However, in contrast to our results Stigonema can also be found in Antarctic ecosystems (Büdel 2001). The cyanobacterial genera Calothrix, Chamaesiphon, Chroococcidiopsis, Nodularia and Tolypothrix, typically found in soil biocoenoses (Büdel 2001; Wang et al. 2015), were identified in the Arctic and the Antarctic sample using metatranscriptomics. Earlier studies, assessing the species richness of aeroterrestrial algae from Svalbard, recorded all of these genera (Matuła et al. 2007; Pushkareva et al. 2015, 2016).
Interestingly, the Arctic crust community exhibited higher cyanobacterial genus richness than the BSC from Livingston Island, while the genus richness of eukaryotic algae was similar. Our morphological and molecular analyses found only 16 different genera of Cyanobacteria in the Antarctic sample compared to 67 in the Arctic. BSCs develop over time and undergo changes in species composition (Büdel et al. 2009). Early successional stages of BSCs are usually dominated by Cyanobacteria; however, bryophytes, lichens and/or liverworts may become the dominant groups in many ecosystems (Büdel et al. 2009). Thus, differences in cyanobacterial genus richness may be explained by the different developmental stages of the communities. Furthermore, Williams et al. (2017) reported a lower abundance of cyanobacterial crusts on Livingston Island than at Ny-Ålesund, Svalbard. Our observation might also be linked to the reduced occurrence of Cyanobacteria in these soil communities which was previously reported by Colesie et al. (2014a, b).
Methodological differences and difficulties
The identification of microalgae through metatranscriptomics yielded a higher number of genera compared to the morphological determination of cultivated and directly observed organisms. This is an indication for the lack of culturability of most microorganisms associated with BSCs (Ward et al. 1990; Schloss and Handelsman 2005; Shi et al. 2009; Massana et al. 2014). The successful cultivation of microalgae is highly dependent on the applied growth conditions including temperature, irradiance and culture medium (Bold 1942; Hoham 1975; Singh and Singh 2015; Wu et al. 2016). For example, Hoham (1975) tested various cryophilic algae, isolated from alpine habitats in Washington, USA, and discovered that Chloromonas pichinchae reached the highest growth rate at 1 °C while Cylindrocystis brébissonii grew fastest at 10 °C. Therefore, the genus Chloromonas might have been inhibited during our culturing procedure due to unsuitable temperatures. Regarding the growth medium, one single solution, that is suitable for all algae and Cyanobacteria, does not exist (Bold 1942; Lee et al. 2014). Even though the medium, used to establish the enrichment cultures, was optimized for a broad range of algae, certain strains may not exhibit growth due to individual nutrient requirements. Some species of the genus Dunaliella, for example, prefer higher salt concentrations compared to other algae (Oren 2005). The salt concentrations of Bold’s Basal Medium, used to establish the enrichment cultures, could have been too low to enable the growth of Dunaliella. Similar reports exist for the cyanobacterium Chroococcidiopsis. Different strains of Chroococcidiopsis, isolated from different habitats, exhibited growth only if the medium was supplied with the appropriate amount of sea salt (Cumbers and Rothschild 2014). However, growth parameters and culture medium determine the culturability of microorganisms only to a certain extent. For many organisms, especially algae, the co-cultivation of certain helper organisms (other algae, bacteria, etc.) is essential for growth and development (Jones et al. 1973; Sanders et al. 2001; Vartoukian et al. 2010). Some Cyanobacteria, which form symbioses with eukaryotic algae, fail to grow in the absence of their partner (Thompson et al. 2012). It is likely that quorum sensing plays an important role for these biotic interactions as certain signal molecules from one organism influence growth and development of another (Poonguzhali et al. 2007; Williams 2007). Chi et al. (2017) studied the marine bacterium Ponticoccus sp. PD-2 and how it controls the growth of the dinoflagellates Alexandrium tamarense, Prorocentrum donghaiense and the haptophyte Phaeocystis globosa. Although this clear indication of microalgae interacting with other microbes exists, hardly anything is known about the underlying mechanisms. BSCs are complex biocoenoses with many dependencies and co-dependencies which are not yet fully understood.
Another potential problem when comparing morphological and molecular methodologies is the false identification of taxa due to ambiguous morphological traits (Wu et al. 2001; Hodač et al. 2016). Some genera, such as Chlorella, are difficult to identify as their morphological characters are limited and changeable depending on environmental conditions (Bock et al. 2011; Hodač et al. 2016). This might be one possible explanation why the metatranscriptomic data confirmed the presence of Chlorella in the Arctic and the Antarctic sample but the morphology-based identification did not. However, it is more likely that the spatial heterogeneity of the sample replicates collected for molecular and light microscopic analysis caused some of the differences (Concostrina-Zubiri et al. 2013; Kim et al. 2015). A clear indication of heterogeneity is the differing results observed for the replicates of the same site in the molecular analysis. For example, the genera Coelastrella, Chlorococcum and Chloromonas were found in one, two and three replicates of the Arctic metatranscriptome, respectively. Furthermore, the high standard deviation of the averaged genera number for the molecular dataset of the Arctic BSC suggests sample heterogeneity. Differences in taxonomic determination may also arise due to a lack of consensus in the algal taxonomy. Many species concepts are still being revised based on molecular analyses rendering an unambiguous morphological identification difficult (Stoyanov et al. 2014; Champenois et al. 2015; Kawasaki et al. 2015; Škaloud et al. 2016). The genus Leptolyngbya, for example, was revealed to be heterogeneous and only recently split up into 15 new genera (Komárek 2016).
Finally, we came across a number of genera which could be identified by light microscopy but not by the analysis of the rRNA SSU sequences. Regarding the genera Gloeocystis, Macrochloris, Neocystis, Desmococcus, Muriella, Myrmecia, Pleurochloris and Pleurogaster, the corresponding 18S sequences were missing from the database utilized in the course of identification. Incomplete and wrongly annotated references are a major problem when performing molecular analyses of any kind and impair the correct determination of genera in environmental samples (Klimke et al. 2011; Taberlet et al. 2012). The microbial dark matter, meaning the total of unculturable microorganisms, complicates this matter even more (Solden et al. 2016). However, by studying metagenomics, metatranscriptomics and single cell genomics researchers can overcome the limit of culturability and supply the databases with novel sequences (Solden et al. 2016). A number of algae, such as Chlorococcum, Heterotetracystis, Mychonastes, Tetracystis, Koliella, Lobosphaera, Tribonema and Xanthonema, were isolated from the BSC samples and were included in the reference database, used for this study. However, they were not detected in the metatranscriptomes. One possible explanation is the spatial heterogeneity of the sample replicates (Concostrina-Zubiri et al. 2013; Kim et al. 2015). Furthermore, these organisms may have been inactive during sampling and were, therefore, undetectable during total RNA analysis (Geisen et al. 2015). Many algae are known to develop resting cells to survive unfavorable conditions such as drought (Holzinger and Karsten 2013; Li et al. 2015). Li et al. (2015) identified 35 different algae in resting stages, among them Chlorococcum which surrounds resting cells with a thin layer of mucus. A species of Mychonastes, M. desiccatus, may produce desiccation-resistant cysts if dried out (Margulis et al. 1988). Massalski et al. (1994) isolated Lobosphaera from King George Island, Antarctica, and studied its ultrastructure. The alga, colonizing volcanic rocks, was able to produce resting cysts with thickened cell walls to outlast detrimental conditions. Similar observations were made for Tribonema bombycinum which produced akinetes in response to starvation (Nagao et al. 1999). Our crust samples were collected in rather extreme habitats exhibiting low temperatures, high irradiance or complete darkness and low water availability throughout the year resulting in a hostile environment (Thomas et al. 2008a). Thus, the absence of certain genera in the molecular datasets might be attributed to the occurrence of resting stages, which likely occur under the conditions prevailing at Svalbard and Livingston Island.
Conclusion
BSCs represent an important part of polar ecosystems and serve as indicators for ecological health and climate change. Thus, these complex communities need to be closely examined to gain insights into not only taxonomy but also physiological plasticity and functionality. This study focused on comparing methodologies to analyze the genus richness of these interesting biocoenoses. Cultivating microorganisms, present in the BSC, combined with light microscopy offers individual analysis of certain organisms while metatranscriptomics enables investigation of the whole genera richness within the crust. The selection of one or the other technique should be based on the individual research focus as both come with certain benefits, as well as costs. The isolation of microorganisms and establishment of cultures enable detailed investigations and follow-up experiments on the isolated algae. However, unculturable genera will be neglected. In contrast, molecular surveys are always dependent on appropriate and complete reference datasets as it is difficult to correctly annotate taxa which are absent from, or misidentified in the database. Moreover, PCR biases and sequencing errors may falsify the results. In our opinion, the combination of both approaches will generate reliable results of higher quality. Thus, we recommend conjoined studies to validate findings and maximize outcome.
References
Albrecht M, Pröschold T, Schumann R (2017) Identification of Cyanobacteria in a eutrophic coastal lagoon on the Southern Baltic coast. Front Microbiol 8:923. https://doi.org/10.3389/fmicb.2017.00923
An SS, Friedel T, Hegewald E (1999) Phylogenetic relationships of Scenedesmus and Scenedesmus-like coccoid green algae as inferred from ITS-2 rDNA sequence comparison. Plant Biol 1:418–428. https://doi.org/10.1111/j.1438-8677.1999.tb00724.x
Andreyeva V, Kurbatova L (2014) Terrestrial and aerophilic nonmotile green microalgae (Chlorophyta) from regions of investigation of Russian Antarctic expedition. Nov Sist Nizsh Rast 46:12–26
Bañón M, Justel A, Velázquez D, Quesada A (2013) Regional weather survey on Byers Peninsula, Livingston Island, South Shetland Islands, Antarctica. Antarct Sci 25:146–156. https://doi.org/10.1017/S0954102012001046
Belnap J (2006) The potential roles of biological soil crusts in dryland hydrologic cycles. Hydrol Process 20:3159–3178. https://doi.org/10.1002/hyp.6325
Belnap J, Büdel B, Lange OL (2001a) Biological soil crusts: characteristics and distribution. In: Belnap J, Lange OL (eds) Biological soil crusts: structure, function, and management. Springer, Berlin, pp 3–30
Belnap J, Rosentreter R, Leonard S et al (2001b) Biological soil crusts: Ecology and management. US Department of the Interior, Bureau of Land Management, National Science and Technology Center, Denver
Bentley DR, Balasubramanian S, Swerdlow HP et al (2008) Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456:53–59. https://doi.org/10.1038/nature07517.Accurate
Bischoff H, Bold H (1963) Some soil algae from enchanted rock and related algal species. Univiversity of Texas Publications, Austin
Bock C, Krienitz L, Pröschold T (2011) Taxonomic reassessment of the genus Chlorella (Trebouxiophyceae) using molecular signatures (barcodes), including description of seven new species. Fottea 11:293–312. https://doi.org/10.5507/fot.2011.028
Bold HC (1942) The cultivation of algae. Bot Rev 8:69–138
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Borchhardt N, Baum C, Mikhailyuk T, Karsten U (2017a) Biological soil crusts of Arctic Svalbard—Water availability as potential controlling factor for microalgal biodiversity. Front Microbiol 8:1485. https://doi.org/10.3389/fmicb.2017.01485
Borchhardt N, Schiefelbein U, Abarca N et al (2017b) Diversity of algae and lichens in biological soil crusts of Ardley and King George Island, Antarctica. Antarct Sci 29:229–237
Borie I, Ibraheem M (2003) Preliminary survey of microalgal soil crusts in a xeric habitats (Wadi-Araba, eastern desert, Egypt). Egypt J Phycol 4:17–33
Breen K, Lévesque E (2008) The influence of biological soil crusts on soil characteristics along a high Arctic glacier foreland, Nunavut, Canada. Arctic Antarct Alp Res 40:287–297. https://doi.org/10.1657/1523-0430(06-098)
Broady PA (1976) Six new species of terrestrial algae from Signy Island, South Orkney Islands, Antarctica. Br Phycol J 11:387–405. https://doi.org/10.1080/00071617700650031
Broady P (1986) Ecology and taxonomy of the terrestrial algae of the Vestfold Hills. In: Pickard J (ed) Antarctic oasis: terrestrial environments and history of the Vestfold Hills. Academic Press, North Ryde, pp 165–202
Buchheim MA, Chapman RL (1991) Phylogeny of the colonial green flagellates: a study of 18S and 26S rRNA sequence data. BioSystems 25:85–100. https://doi.org/10.1016/0303-2647(91)90015-D
Büdel B (2001) Synopsis: comparative biogeography of soil-crust biota. In: Belnap J, Lange OL (eds) Biological soil crusts: structure, function, and management. Springer, Berlin, pp 141–152
Büdel B, Darienko T, Deutschewitz K et al (2009) Southern African biological soil crusts are ubiquitous and highly diverse in drylands, being restricted by rainfall frequency. Microb Ecol 57:229–247. https://doi.org/10.1007/s00248-008-9449-9
Büdel B, Dulic T, Darienko T et al (2016) Cyanobacteria and algae of biological soil crusts. In: Weber B, Büdel B, Belnap J (eds) Biological soil crusts: an organizing principle in drylands. Springer, Switzerland, pp 55–80
Casamatta DA, Johansen JR, Vis ML, Broadwater ST (2005) Molecular and morphological characterization of ten polar and near-polar strains within the Oscillatoriales (Cyanobacteria). J Phycol 41:421–438. https://doi.org/10.1111/j.1529-8817.2005.04062.x
Champenois J, Marfaing H, Pierre R (2015) Review of the taxonomic revision of Chlorella and consequences for its food uses in Europe. J Appl Phycol 27:1845–1851. https://doi.org/10.1007/s10811-014-0431-2
Chi W, Zheng L, He C et al (2017) Quorum sensing of microalgae associated marine Ponticoccus sp. PD-2 and its algicidal function regulation. AMB Express 7:59. https://doi.org/10.1186/s13568-017-0357-6
Chown S, Huiskes A, Gremmen N et al (2012) Continent-wide risk assessment for the establishment of nonindigenous species in Antarctica. PNAS 109:4938–4943
Colesie C, Gommeaux M, Green TGA, Büdel B (2014a) Biological soil crusts in continental Antarctica: garwood Valley, southern Victoria Land, and Diamond Hill, Darwin Mountains region. Antarct Sci 26:115–123. https://doi.org/10.1017/S0954102013000291
Colesie C, Green TGA, Türk R et al (2014b) Terrestrial biodiversity along the Ross Sea coastline, Antarctica: lack of a latitudinal gradient and potential limits of bioclimatic modeling. Polar Biol 37:1197–1208. https://doi.org/10.1007/s00300-014-1513-y
Concostrina-Zubiri L, Huber-Sannwald E, Martínez I et al (2013) Biological soil crusts greatly contribute to small-scale soil heterogeneity along a grazing gradient. Soil Biol Biochem 64:28–36. https://doi.org/10.1016/j.soilbio.2013.03.029
Cooper E, Wookey P (2001) Field measurements of the growth rates of forage lichens, and the implications of grazing by Svalbard reindeer. Symbiosis 31:173–186
Cox EJ (1996) Identification of freshwater diatoms from live material. Chapman & Hall, London
Cumbers J, Rothschild LJ (2014) Salt tolerance and polyphyly in the cyanobacterium Chroococcidiopsis (Pleurocapsales). J Phycol 50:472–482. https://doi.org/10.1111/jpy.12169
Czerwik-Marcinkowska J, Massalski A, Olech M, Wojciechowska A (2015) Morphology, ultrastructure and ecology of Muriella decolor (Chlorophyta) from subaerial habitats in Poland and the Antarctic. Polish Polar Res 36:163–174. https://doi.org/10.1515/popore
Darienko T, Gustavs L, Eggert A et al (2015) Evaluating the species boundaries of green microalgae (Coccomyxa, Trebouxiophyceae, Chlorophyta) using integrative taxonomy and DNA barcoding with further implications for the species identification in environmental samples. PLoS ONE 10:e0127838. https://doi.org/10.1371/journal.pone.0127838
de Wever A, Leliaert F, Verleyen E et al (2009) Hidden levels of phylodiversity in Antarctic green algae: further evidence for the existence of glacial refugia. Proc R Soc B Biol Sci 276:3591–3599. https://doi.org/10.1098/rspb.2009.0994
Dojani S, Kauff F, Weber B, Büdel B (2014) Genotypic and phenotypic diversity of Cyanobacteria in biological soil crusts of the succulent Karoo and Nama Karoo of Southern Africa. Microb Ecol 67:286–301. https://doi.org/10.1007/s00248-013-0301-5
Doyle JJ, Doyle JL, Ballenger JA et al (1997) A phylogeny of the chloroplast gene rbcL in the Leguminosae: taxonomic correlations and insights into the evolution of nodulation. Am J Bot 84:541–554. https://doi.org/10.2307/2446030
Elferink S, Neuhaus S, Wohlrab S et al (2017) Molecular diversity patterns among various phytoplankton size-fractions in West Greenland in late summer. Deep Res Part I Oceanogr Res Pap 121:54–69. https://doi.org/10.1016/j.dsr.2016.11.002
Elster J, Lukesova A, Svoboda J et al (1999) Diversity and abundance of soil algae in the polar desert, Sverdrup Pass, central Ellesmere Island. Polar Rec 35:231–254 (Gr Brit)
Ettl H, Gärtner G (2014) Syllabus der Boden-, Luft- und Flechtenalgen
Evans RD, Johansen JR (1999) Microbiotic crusts and ecosystem processes. CRC Crit Rev Plant Sci 18:183–225. https://doi.org/10.1016/S0735-2689(99)00384-6
Evans KM, Wortley AH, Mann DG (2007) An assessment of potential diatom “barcode” genes (cox1, rbcL, 18S and ITS rDNA) and their effectiveness in determining relationships in Sellaphora (Bacillariophyta). Protist 158:349–364. https://doi.org/10.1016/j.protis.2007.04.001
Førland EJ, Benestad R, Hanssen-Bauer I et al (2011) Temperature and precipitation development at Svalbard 1900–2100. Adv Meteorol. https://doi.org/10.1155/2011/893790
Frenot Y, Chown SL, Whinam J et al (2005) Biological invasions in the Antarctic: extent, impacts and implications. Biol Rev 80:45–72. https://doi.org/10.1017/S1464793104006542
Geisen S, Tveit AT, Clark IM et al (2015) Metatranscriptomic census of active protists in soils. ISME J 9:2178–2190. https://doi.org/10.1038/ismej.2015.30
Geitler L (1932) Cyanophyceae. Akadademische Verlagsgesellschaft M.B.H, Leipzig
Giełwanowska I, Olech M (2012) New ultrastructural and physiological features of the thallus in Antarctic lichens. Acta Biol Cracoviensia Ser Bot 54:40–52. https://doi.org/10.2478/v10182-012-0004-0
Greaves MP, Wilson MJ (1970) The degradation of nucleic acids and montmorillonite-nucleic-acid complexes by soil microorganisms. Soil Biol Biochem 2:257–268
Hall JD, Fucikova K, Lo C et al (2010) An assessment of proposed DNA barcodes in freshwater green algae. Cryptogam Algol 31:529–555
Hodač L, Hallmann C, Spitzer K et al (2016) Widespread green algae Chlorella and Stichococcus exhibit polar-temperate and tropical-temperate biogeography. FEMS Microbiol Ecol 92:1–16. https://doi.org/10.1093/femsec/fiw122
Hoham R (1975) Optimum temperatures and temperature ranges for growth of snow algae. Arct Alp Res 7:13–24
Holzinger A, Karsten U (2013) Desiccation stress and tolerance in green algae: consequences for ultrastructure, physiological, and molecular mechanisms. Front Plant Sci 4:327. https://doi.org/10.3389/fpls.2013.00327
John DM, Whitton BA, Brook AJ (2002) The freshwater algal flora of the British Isles: an identification guide to freshwater and terrestrial algae. Cambridge University Press, New York
Jones AK, Rhodes ME, Evans SC (1973) The use of antibiotics to obtain axenic cultures of algae. Br Phycol J 8:185–196. https://doi.org/10.1080/00071617300650211
Kastovská K, Elster J, Stibal M, Santrůcková H (2005) Microbial assemblages in soil microbial succession after glacial retreat in Svalbard (High Arctic). Microb Ecol 50:396–407. https://doi.org/10.1007/s00248-005-0246-4
Kawasaki Y, Nakada T, Tomita M (2015) Taxonomic revision of oil-producing green algae, Chlorococcum oleofaciens (Volvocales, Chlorophyceae), and its relatives. J Phycol 51:1000–1016. https://doi.org/10.1111/jpy.12343
Kawecka B (1986) Ecology of snow algae. Polish Polar Res 7:407–415
Kim M, Cho A, Lim HS et al (2015) Highly heterogeneous soil bacterial communities around Terra Nova Bay of Northern Victoria Land, Antarctica. PLoS ONE 10:e0119966. https://doi.org/10.1371/journal.pone.0119966
Klimke W, O’Donovan C, White O et al (2011) Solving the problem: genome annotation standards before the data deluge. Stand Genom Sci 5:168–193. https://doi.org/10.4056/sigs.2084864
Komárek J (2016) Review of the cyanobacterial genera implying planktic species after recent taxonomic revisions according to polyphasic methods: state as of 2014. Hydrobiologia 764:259–270. https://doi.org/10.1007/s10750-015-2242-0
Komárek J, Anagnostidis K (1998) Freshwater flora of Central Europe—Cyanoprokaryota, vol 1. Chroococcales. Springer, Berlin
Komárek J, Anagnostidis K (2005) Freshwater flora of Central Europe—Cyanoprokaryota, vol 2. Oscillatorialles. Springer, Berlin
Kopylova E, Noé L, Touzet H (2012) SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28:3211–3217. https://doi.org/10.1093/bioinformatics/bts611
Lee K, Eisterhold ML, Rindi F et al (2014) Isolation and screening of microalgae from natural habitats in the midwestern United States of America for biomass and biodiesel sources. J Nat Sci Biol Med 5:333. https://doi.org/10.4103/0976-9668.136178
Lee JR, Raymond B, Bracegirdle TJ et al (2017) Climate change drives expansion of Antarctic ice-free habitat. Nature 547:49–54. https://doi.org/10.1038/nature22996
Li Z, Shin HH, Lee T, Han M-S (2015) Resting stages of freshwater algae from surface sediments in Paldang Dam Lake, Korea. Nov Hedwigia 101:475–500. https://doi.org/10.1127/nova
Lind EM, Brook AJ (1980) A key to the commoner desmids of the English Lake District. Freshwater Biological Association
Lukešová A, Kociánová M, Váňa J et al (2010) Mud boils of the Giant Mts and Abisko Mts tundra—preliminary comparative study. Opera Corcon 47:55–82
Lürling M (2003) Phenotypic plasticity in the green algae Desmodesmus and Scenedesmus with special reference to the induction of defensive morphology. Ann Limnol 39:85–101. https://doi.org/10.1051/limn/2003014
Manoylov KM (2014) Taxonomic identification of algae (Morphological and molecular): species concepts, methodologies, and their implications for ecological bioassessment. J Phycol 50:409–424. https://doi.org/10.1111/jpy.12183
Margulies M, Egholm M, Altman WE et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–381. https://doi.org/10.1038/nature03959
Margulis L, Hinkle G, McKhann H, Moynihan B (1988) Mychonastes desiccatus Brown sp. nova (Chlorococcales, Chlorophyta)—an intertidal alga forming achlorophyllous desiccation-resistant cysts. Arch Hydrobiol Suppl Algol Stud 78:425–446
Massalski A, Mrozinska T, Olech M (1994) Ultrastructure of Lobosphaera reniformis (Watanabe) Komárek et fott (= Chlorellales) from King George Island, South Shetland Islands, Antarctica. Acta Soc Bot Pol 63:205–210
Massana R, del Campo J, Sieracki ME et al (2014) Exploring the uncultured microeukaryote majority in the oceans: reevaluation of ribogroups within stramenopiles. ISME J 8:854–866. https://doi.org/10.1038/ismej.2013.204
Matuła J, Pietryka M, Richter D, Wojtuń B (2007) Cyanoprokaryota and algae of Arctic terrestrial ecosystems in the Hornsund area, Spitsbergen. Polish Polar Res 28:283–315
Maturilli M, Herber A, König-Langlo G (2013) Climatology and time series of surface meteorology in Ny-Ålesund, Svalbard. Earth Syst Sci Data 5:155–163. https://doi.org/10.5194/essd-5-155-2013
Michel RFM, Schaefer CEGR, Simas FMB et al (2014) Active-layer thermal monitoring on the Fildes Peninsula, King George Island, maritime Antarctica. Solid Earth 5:1361–1374. https://doi.org/10.5194/se-5-1361-2014
Miller CS, Baker BJ, Thomas BC et al (2011) EMIRGE: reconstruction of full-length ribosomal genes from microbial community short read sequencing data. Genome Biol 12:R44. https://doi.org/10.1186/gb-2011-12-5-r44
Misawa S (1999) Rapid diagnosis of infectious diseases; features and limitations of the microscopic examination of clinical specimens. J Assoc Rapid Method Autom Microbiol 10:121–131
Nagao M, Arakawa K, Takezawa D et al (1999) Akinete formation in Tribonema bombycinum Derbes et Solier (Xanthophyceae) in relation to freezing tolerance. J Plant Res 112:163–174
Oren A (2005) A hundred years of Dunaliella research: 1905-2005. Saline Syst 1:2. https://doi.org/10.1186/1746-1448-1-2
Patova E, Davydov D, Vera A (2015) Cyanoprokaryotes and algae. In: Matveyeva M (ed) Plants and fungi of the polar deserts in the northern hemisphere. MAPAФOH, pp 133–164
Pawlowski J, Christen R, Lecroq B et al (2011) Eukaryotic richness in the abyss: insights from pyrotag sequencing. PLoS ONE 6:e18169. https://doi.org/10.1371/journal.pone.0018169
Peel MC, Finlayson BL, McMahon TA (2007) World map of the Köppen-Geiger climate classification updated. Hydrol Earth Syst Sci 11:1633–1644. https://doi.org/10.1127/0941-2948/2006/0130
Pereira EB, Evangelista H, Pereira KCD et al (2006) Apportionment of black carbon in the South Shetland Islands, Antartic Peninsula. J Geophys Res 111:D03303. https://doi.org/10.1029/2005JD006086
Peveling E, Galun M (1976) Electron-microscopical studies on the phycobiont Coccomyxa Schmidle. New Phytol 77:713–718. https://doi.org/10.1111/j.1469-8137.1976.tb04665.x
Pfaff S, Borchardt N, Boy J et al (2016) Desiccation tolerance and growth-temperature requirements of Coccomyxa (Trebouxiophyceae, Chlorophyta) strains from Antarctic biological soil crusts. Arch Hydrobiol Suppl Algol Stud 151:3–19. https://doi.org/10.1127/algol
Pointing SB, Belnap J (2012) Microbial colonization and controls in dryland systems. Nat Rev Microbiol 10:551–562. https://doi.org/10.1038/nrmicro2831
Pointing SB, Büdel B, Convey P et al (2015) Biogeography of photoautotrophs in the high polar biome. Front Plant Sci 6:692. https://doi.org/10.3389/fpls.2015.00692
Poonguzhali S, Madhaiyan M, Sa T (2007) Quorum-sensing signals produced by plant-growth promoting Burkholderia strains under in vitro and in planta conditions. Res Microbiol 158:287–294. https://doi.org/10.1016/j.resmic.2006.11.013
Prescott GW (1964) How to know the fresh-water algae. Plenum Press, New York
Pushkareva E, Pessi IS, Wilmotte A, Elster J (2015) Cyanobacterial community composition in Arctic soil crusts at different stages of development. FEMS Microbiol Ecol 91:fiv143. https://doi.org/10.1093/femsec/fiv143
Pushkareva E, Johansen JR, Elster J (2016) A review of the ecology, ecophysiology and biodiversity of microalgae in Arctic soil crusts. Polar Biol. https://doi.org/10.1007/s00300-016-1902-5
Rippin M, Komsic-Buchmann K, Becker B (2016) RNA isolation from biological soil crusts: methodological aspects. Arch Hydrobiol Suppl Algol Stud 151:21–37. https://doi.org/10.1127/algol
Rippka R, Deruelles J, Waterbury JB et al (1979) Generic assignments, strain histories and properties of pure cultures of Cyanobacteria. J Gen Microbiol 111:1–61. https://doi.org/10.1099/00221287-111-1-1
Rivasseau C, Farhi E, Compagnon E et al (2016) Coccomyxa actinabiotis sp. nov. (Trebouxiophyceae, Chlorophyta), a new green microalga living in the spent fuel cooling pool of a nuclear reactor. J Phycol 52:689–703. https://doi.org/10.1111/jpy.12442
Sanders RW, Caron DA, Davidson JM et al (2001) Nutrient acquisition and population growth of a mixotrophic alga in axenic and bacterized cultures. Microb Ecol 42:513–523. https://doi.org/10.1007/s00248-001-1024-6
Schloss PD, Handelsman J (2005) Metagenomics for studying unculturable microorganisms: cutting the Gordian knot. Genome Biol 6:229. https://doi.org/10.1186/gb-2005-6-8-229
Schmidle W (1901) Ueber drei Algengenera. Berichte der Dtsch Bot Gessellschaft 19:10–24
Schulz K, Mikhailyuk T, Dreßler M et al (2016) Biological soil crusts from coastal dunes at the Baltic Sea: Cyanobacterial and algal biodiversity and related soil properties. Microb Ecol 71:178–193. https://doi.org/10.1007/s00248-015-0691-7
Shen S (2008) Genetic diversity analysis with ISSR PCR on green algae Chlorella vulgaris and Chlorella pyrenoidosa. Chin J Oceanol Limnol 26:380–384. https://doi.org/10.1007/s00343-008-0380-1
Sherwood AR, Vis ML, Entwisle TJ et al (2008) Contrasting intra versus interspecies DNA sequence variation for representatives of the Batrachospermales (Rhodophyta): insights from a DNA barcoding approach. Phycol Res 56:269–279. https://doi.org/10.1111/j.1440-1835.2008.00508.x
Shi XL, Marie D, Jardillier L et al (2009) Groups without cultured representatives dominate eukaryotic picophytoplankton in the oligotrophic South East Pacific Ocean. PLoS ONE 4:e7657. https://doi.org/10.1371/journal.pone.0007657
Singh SP, Singh P (2015) Effect of temperature and light on the growth of algae species: a review. Renew Sustain Energy Rev 50:431–444. https://doi.org/10.1016/j.rser.2015.05.024
Škaloud P, Friedl T, Hallmann C et al (2016) Taxonomic revision and species delimitation of coccoid green algae currently assigned to the genus Dictyochloropsis (Trebouxiophyceae, Chlorophyta). J Phycol 52:599–617. https://doi.org/10.1111/jpy.12422
Skinner CE (1932) Isolation in pure culture of green algae from soil by a simple technique. Plant Physiol 7:533–537. https://doi.org/10.1104/pp.7.3.533
Solden L, Lloyd K, Wrighton K (2016) The bright side of microbial dark matter: lessons learned from the uncultivated majority. Curr Opin Microbiol 31:217–226. https://doi.org/10.1016/j.mib.2016.04.020
Starr RC, Jeffrey AZ (1993) UTEX—the culture collection of algae at the University of Texas at Austin 1993 list of cultures. J Phycol 29:1–106
Stoyanov P, Moten D, Mladenov R et al (2014) Phylogenetic relationships of some filamentous cyanoprokaryotic species. Evol Bioinform 10:39–49. https://doi.org/10.4137/EBo.s13748
Stoyneva M (2000) Soil algae in museum samples from some Southwest Asia sites I. Hist Nat Bulg 12:129–146
Taberlet P, Coissac E, Pompanon F et al (2012) Towards next-generation biodiversity assessment using DNA metabarcoding. Mol Ecol 21:2045–2050. https://doi.org/10.1111/j.1365-294X.2012.05470.x
Takeuchi N (2001) Seasonal and altitudinal variations in snow algal communities on an Alaskan glacier (Gulkana glacier in the Alaska range). Hydrol Process 15:3447–3459. https://doi.org/10.1088/1748-9326/8/3/035002
Thomas DN, Fogg T, Convey P et al (2008a) Introduction to the polar regions. In: Thomas DN (ed) The biology of polar regions. University Press, Oxford, pp 1–27
Thomas DN, Fogg T, Convey P et al (2008b) Periglacial and terrestrial habitats in polar regions. In: Thomas DN (ed) The biology of polar regions. University Press, Oxford, pp 53–100
Thompson AW, Foster RA, Krupke A et al (2012) Unicellular cyanobacterium symbiotic with a single-celled eukaryotic alga. Science 337:1546–1550
Thorn R, Lynch M (2007) Fungi and eukaryotic algae. In: Paul EA (ed) Soil microbiology, ecology, and biochemistry. Elsevier, Oxford, pp 145–162
Tschaikner A, Ingolic E, Gärtner G (2007) Observations in a new isolate of Coelastrella terrestris (Reisigl) Hegewald & Hanagata (Chlorophyta, Scenedesmaceae) from Alpine Soil (Tyrol, Austria). Phyton (B Aires) 46:237–245
Urich T, Lanzén A, Qi J et al (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS ONE 3:e2527. https://doi.org/10.1371/journal.pone.0002527
Urich T, Lanzén A, Stokke R et al (2014) Microbial community structure and functioning in marine sediments associated with diffuse hydrothermal venting assessed by integrated meta-omics. Environ Microbiol 16:2699–2710. https://doi.org/10.1111/1462-2920.12283
Uzunov BA, Stoyneva MP, Gärtner G, Koefler W (2008) First record of Coelastrella species (Chlorophyta: Scenedesmaceae) in Bulgaria. Berichte des naturwissenschaftlichen-medizinischen Verein Innsbruck 95:27–34
Vartoukian SR, Palmer RM, Wade WG (2010) Strategies for culture of “unculturable” bacteria. FEMS Microbiol Lett 309:1–7. https://doi.org/10.1111/j.1574-6968.2010.02000.x
Vieira HH, Bagatini IL, Guinart CM, Vieira AAH (2016) tufA gene as molecular marker for freshwater Chlorophyceae. Algae 31:155–165. https://doi.org/10.4490/algae.2016.31.4.14
Vishnivetskaya TA (2009) Viable Cyanobacteria and green algae from the permafrost darkness. Permafrost soils. Springer, Heidelberg, pp 73–84
Vogel S (1955) Niedere “Fensterpflanzen” in der Südafrikanischen Wüste: Eine ökologische Schilderung. Duncker & Humblot
Vogel S, Eckerstorfer M, Christiansen HH (2012) Cornice dynamics and meteorological control at Gruvefjellet, Central Svalbard. Cryosph 6:157–171. https://doi.org/10.5194/tc-6-157-2012
Wang NF, Zhang T, Zhang F et al (2015) Diversity and structure of soil bacterial communities in the Fildes Region (maritime Antarctica) as revealed by 454 pyrosequencing. Front Microbiol 6:1188. https://doi.org/10.3389/fmicb.2015.01188
Ward DM, Weller R, Bateson MM (1990) 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature 345:63–65. https://doi.org/10.1038/346183a0
Waterbury JB, Stanier RY (1978) Patterns of growth and development in pleurocapsalean Cyanobacteria. Microbiol Rev 42:2–44
Williams P (2007) Quorum sensing, communication and cross-kingdom signalling in the bacterial world. Microbiology 153:3923–3938. https://doi.org/10.1099/mic.0.2007/012856-0
Williams L, Loewen-Schneider K, Maier S, Büdel B (2016) Cyanobacterial diversity of western European biological soil crusts along a latitudinal gradient. FEMS Microbiol Ecol 92:fiw157. https://doi.org/10.1093/femsec/fiw157
Williams L, Borchhardt N, Colesie C et al (2017) Biological soil crusts of Arctic Svalbard and of Livingston Island, Antarctica. Polar Biol 40:399–411. https://doi.org/10.1007/s00300-016-1967-1
Wilmotte A (1994) Molecular evolution and taxonomy of the Cyanobacteria. In: Bryant D (ed) The molecular biology of cyanobacteria. Springer, Dordrecht, pp 1–25
Wu HL, Hseu RS, Lin LP (2001) Identification of Chlorella spp. isolates using ribosomal DNA sequences. Bot Bull Acad Sin 42:115–121
Wu Z, Duangmanee P, Zhao P et al (2016) The effects of light, temperature, and nutrition on growth and pigment accumulation of three Dunaliella salina strains isolated from saline soil. Jundishapur J Microbiol 9:e26732. https://doi.org/10.5812/jjm.26732
Yoon T-H, Kang H-E, Kang C-K et al (2016) Development of a cost-effective metabarcoding strategy for analysis of the marine phytoplankton community. PeerJ 4:e2115. https://doi.org/10.7717/peerj.2115
Yoshitake S, Uchida M, Koizumi H et al (2010) Production of biological soil crusts in the early stage of primary succession on a high Arctic glacier foreland. New Phytol 186:451–460. https://doi.org/10.1111/j.1469-8137.2010.03180.x
Zidarova R (2008) Algae from Livingston Island (S Shetland Islands): a checklist. Phytol Balc 14:19–35
Acknowledgements
This study was funded by the Deutsche Forschungsgemeinschaft (DFG) within the project ‘Polarcrust’ (BE1779/18-1, KA899/23-1, BU666/17-1) which is part of the Priority Program 1158 ‘Antarctic Research’. We also thank the AWIPEW station, the Instituto Antártico Chileno, the Spanish Antarctic Committee and the Juan Carlos I Antarctic Base for their logistic support. Sampling and research activities were approved by the German authorities (Umwelt Bundesamt: Biological soil crust algae from the polar regions; 24.09.2014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rippin, M., Borchhardt, N., Williams, L. et al. Genus richness of microalgae and Cyanobacteria in biological soil crusts from Svalbard and Livingston Island: morphological versus molecular approaches. Polar Biol 41, 909–923 (2018). https://doi.org/10.1007/s00300-018-2252-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-018-2252-2