Abstract
Increasing crop production to meet the demands of a growing population depends largely on crop improvement through new plant-breeding techniques (NPBT) such as genome editing. CRISPR/Cas systems are NPBTs that enable efficient target-specific gene editing in crops, which is supposed to accelerate crop breeding in a way that is different from genetically modified (GM) technology. Herein, we review the applications of CRISPR/Cas systems in crop breeding focusing on crop domestication, heterosis, haploid induction, and synthetic biology, and summarize the screening methods of CRISPR/Cas-induced mutations in crops. We highlight the importance of molecular characterization of CRISPR/Cas-edited crops, and pay special attentions to emerging highly specific genome-editing tools such as base editors and prime editors. We also discuss future improvements of CRISPR/Cas systems for crop improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Feeding a growing population in a sustainable way is a great challenge to current crop-breeding efforts (Schaart et al. 2016). Traditional breeding technology based on crossing and selection without any knowledge about genetics, even aided by marker-assisted selection, is a very labor-intensive and time-consuming process, which also shows drawbacks with complex genetic outputs (Schaart et al. 2016). Mutation breeding technology based on chemical and physical genotoxins dramatically increases the mutation rate above natural levels; however, artificially induced changes are unquestionably uncontrolled, requiring complex and expensive screening and selection procedures (Pacher and Puchta 2017). Due to the random modification in the crop genome, outcomes of both natural mutation and mutational breeding are unpredictable. The transgenic breeding technique transfers desired trait-coding genes via an exogenous T-DNA cassette into the elite cultivars; its outcome is relatively predictable. However, time and expenses for research and development of a genetically modified (GM) crop with desirable traits free of unexpected insertions are huge to meet demands from safety regulations and social acceptance (Araki and Ishii 2015; Schaeffer and Nakata 2015).
In contrast, site-directed nucleases (SDNs)-based genome-editing technologies increase significantly the precision of gene modification in crop systems (Jaganathan et al. 2018). SDNs include zinc finger nucleases (ZFNs), transcription-activator-like effector nucleases (TALENs), and clustered regulatory interspaced short palindromic repeats associated protein 9 (CRISPR/Cas9) (Lusser et al. 2012; Zhu et al. 2017). They precisely cut the genomic DNA at the targeted loci to generate double-strand breaks (DSBs), which triggers specialized repair pathways, either homologous recombination (HR) or non-homologous end-joining (NHEJ), and results in indel mutations (Jinek et al. 2012). Besides, simultaneous introduction of several DSBs by genome editing allows to break genetic linkages, to reshuffle entire chromosome orientations, to create inversions, and to permit reciprocal chromosomal translocations or chromosome fragment exchanges (Pacher and Puchta 2017). Thus, SDN-based genome editing holds a great potential for precise crop improvement.
Each SDN has its particular characteristics. ZFNs are artificially engineered chimeric restriction enzymes composed of site-specific DNA binding zinc finger proteins (ZFPs) fused with the non-specific DNA cleavage domain of the FokI restriction enzyme (Guo et al. 2010). Typically, ZFNs consist of three-four lower-affinity ZFPs and two tail-to-tail ZFP-binding sites separated by a 5- to 7-bp spacer sequence, therefore, not all crop sequences can be efficiently targeted (Li et al. 2019b). ZFNs have been reported to induce efficient site-directed mutagenesis in several crop species, including maize (Ainley et al. 2013) and soybean (Curtin et al. 2011). However, due to limitations such as low target specificity, time-consuming, and narrow available target sites (Chen and Gao 2013), ZFNs have given way to other SDNs. TALENs have similar principles to ZFNs but harbor different site-specific DNA binding proteins, named transcription activator-like effectors (TALEs). Each effector domain recognizes a single nucleotide pair in TALEN, therefore, compared with ZFNs, TALENs show higher target specificity (Baltes and Voytas 2015). TALENs have been successfully applied to edit several crop species including tobacco (Zhang et al. 2013), rice (Li et al. 2012), and maize (Liang et al. 2014). Despite its advantages over ZFNs, using of TALENs as genome-editing tools still needs the assembly of complex tandem repeats to bind targeted DNA sequences. In addition, large size and tedious nature make the transfer of the TALEN system to plant cell a challenge (Baltes and Voytas 2015).
CRISPR/Cas9 is a RNA-guided nuclease, and its specificity is single-guide RNA (sgRNA) dependent. Theoretically, CRISPR/Cas9 can bind to any DNA sequence that contains a protospacer adjacent motif (PAM) site when sgRNA is present for identifying the target (Jinek et al. 2012). Unlike protein-guided nucleases such as ZFNs and TALENs, CRISPR/Cas9 introduces a blunt DSB, in the case of SpCas9, the cleavage occurs at a site three nucleotides upstream of the PAM (Jinek et al. 2012). Due to its high target specificity, simplicity, ease of use, CRISPR/Cas9 technique has been widely used as a dominant technique of SDNs for gene editing in crops, humans, and animals (Doudna and Charpentier 2014). In crops, CRISPR/Cas9-induced DSBs are repaired mainly via NHEJ; as a result, indel genetic variations are generated (Zhu et al. 2017). So far, CRISPR/Cas9 system has been widely used for crop improvement in rice, sorghum, wheat, maize, soybean, tomato, potato, apple, and banana (Osakabe and Osakabe 2015; Jaganathan et al. 2018; Tripathi et al. 2020). With advancements in the improvement of CRISPR/Cas systems (Anzalone et al. 2019), such as CRISPR/Cpf1 (Chen et al. 2019; Kim et al. 2017) and nucleotide substitutions tools (Shimatani et al. 2017; Zong et al. 2017), CRISPR systems are becoming to be more efficient for crop improvement (Gao 2021; Hong et al, 2021; Miladinovic et al. 2021). Its applications in crops range from boosting yield, resisting against pests and diseases (Wang et al. 2014), to improving nutritive value (Table 1) (Li et al. 2016, 2018b; Ma et al. 2016; Do et al. 2019; Dong et al. 2020). The objective of this review is to summarize the current developments and applications of CRISPR/Cas systems in crop improvement, discuss the regulatory landscape of genome-edited crops, and to propose future prospects.
Current applications of CRISPR/Cas systems in crop breeding
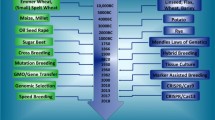
Providing the world with diverse, abundant, nutritive plants and plant-derived products in a sustainable manner cannot be achieved without better understanding of plant biology under both normal and stressful conditions. CRISPR/Cas-mediated genome editing not only revolutionizes crop biology but also providing means for crop improvement (Chen et al. 2019; Kumar et al. 2020a; El-Mounadi et al. 2020). Current applications of CRISPR/Cas systems in crop improvement regarding yield, quality, biotic and abiotic stress tolerance, and herbicide resistance are summarized in Table 1, albeit the fact that increasing excellent reviews are emerging (Gao 2021; Miladinovic et al. 2021). Outputs of CRISPR/Cas9-edited crops, whether targeting a single gene or multiple genes, include mainly small indels or single nucleotide base substitutions; however, high frequency of large deletions and/or reorganizations are also reported (Zhu et al. 2017; Li et al. 2019b; Biswas et al. 2020a). Nevertheless, all mutations can pass faithfully to subsequent generations without any novel modifications (Zhu et al. 2017; Biswas et al. 2020a). Thus, CRISPR/Cas9 finds its way with high potential to be widely adopted for crop breeding (Fig. 1), because it is blurring the boundaries in the GM regulations (Araki and Ishii 2015).
Crop domestication
Compared with their wild ancestors, currently cultivated crops have reduced genetic diversity and resilience to biotic and abiotic stresses (Doebley et al. 2006). On the other hand, compared with cultivated crops, wild crops and/or orphan crops could harbor desired high yield/nutrition traits and favorable resilience that could readily to adapt to changing climates. Therefore, domestication of wild species/orphan crops could be a fascinating way to secure food supply. Conventional domestication is a time-consuming and laborious process (Fernie and Yan 2019; Yu et al. 2021), and the CRISPR/Cas9 system, with its precise, accurate and multiplex genome modification capacity, could accelerate the process of crop domestication. In tomato, targeting six genes of agronomic importance present in cultivated tomato by CRISPR/Cas9 system resulted in successful de novo domestication of wild tomato Solanum pimpinellifolium, fruit size and fruit numbers in edited wild tomato were significantly increased and lycopene accumulation in edited was remarkably enhanced (Zsögön et al. 2018). Taking advantage of multiple targeting capacity of CRISPR/Cas9 (targeting SP5G, SP, SlCLV3, and SlGGP1), several desirable traits were created into four stress-tolerant wild tomatoes; Cas9-free CRISPR/Cas9-edited tomato plants showed domesticated desirable traits (early harvest, determinate shoot architecture, large fruit size, and improved nutritional benefits) while maintained bacterial spot disease and salt tolerance (Li et al. 2018a). Targeting known domestication loci in African landrace Kabre with superior endemic pest resistance and drought and nutrient deficiency tolerance resulted in the reduction of plant stature (targeting HTD1) and increase of yields (targeting GS3, GW2 and GN1A) in rice (Lacchini et al. 2020). Targeting two domestication-related genes (qSH1 for shattering and An-1 for awn length) and two agronomic trait associated genes (SD1 for height and GS3 for grain length) through CRISPR/Cas9 demonstrated successful rapid domestication of wild rice polyploidy rice 1 (PPR1), a wild allotetraploid rice Oryza alta (CCDD) (Yu et al., 2021). Similarly, simultaneously targeting Ghd7 and DTH7 using a multiplex CRISPR/Cas9 editing approach in PPR1 significantly altered their heading date (Yu et al. 2021). Those pioneer studies not only proved the concept but also paved the way to utilize the genetic diversity hidden in wild crops for molecular based breeding to achieve rapid de novo domestication of wild crops into staple foods.
In addition, using one-step CRISPR/Cas9 targeting regulator genes of stem length (SlER), rapid flowering (SP5G) and precocious growth termination (SP), vine-like tomato was reconstructed to compact and early yielding plants suitable for urban agriculture (Kwon et al. 2020). This could be a novel application direction of CRISPR/Cas9 for horticultural crop research and development.
Heterosis
Heterosis (hybrid vigor) is the genetic phenomenon in which hybrid offsprings display better performance (in yields/nutrition; stress tolerance, or adaptability) than their parents (Birchler et al. 2010). Heterosis has been widely utilized in modern agriculture, contributing remarkably to food supply worldwide (Schnable and Springer 2013). Heterosis can be maintained only in F1 generation. Farmers have to buy hybrid seeds every year; on the other hand, producing hybrid seeds is time-consuming, laborious, and costly. Thus, fixing desirable hybrid traits is the most challenging neck bottle for using heterosis in crop breeding.
CRISPR/Cas systems show particular promising applications in this area. If the heterozygosity of F1 hybrid can be fixed while haploid seeds can be produced, heterosis can be maintained via self-propagation through such seeds. Genome editing of three meiotic genes (REC8, PAIR1 and OSD1) using multiplex CRISPR–Cas9 system produced clonal diploid gametes and tetraploid seeds while editing a fertilization gene (MATRILINEAL, MTL) produced haploid seeds in hybrid rice. Thus, simultaneously editing of all four genes (REC8, PAIR1, OSD1, and MTL) in hybrid rice through CRISPR–Cas9 fixed favorite F1 traits (Wang et al. 2019). Overexpressing of BBM1 (a sperm cell-specific expressed AP2 transcription factor) in egg cells in a triple knockout mutant simultaneously targeting three meiotic genes (REC8, PAIR1 and OSD1) has successfully replaced meiosis by mitosis, which resulted in asexual propagation of hybrid rice through seeds (Khanday et al. 2019). These studies demonstrated the feasibility to maintain hybrid rice clonally through seed propagation with the help of CRISPR/Cas systems.
Haploid induction
Haploid induction (HI), the first step of doubled haploid technology, aims to regenerate haploid or spontaneous doubled haploid plants based on intraspecific crossing. Traditional HI, depending on species, is achieved by several approaches, such as androgenesis, gynogenesis or parthenogenesis. It takes time, needs substantial personnel and equipment, and always has unavoidable variability in efficiency (Hooghvorst and Nogués 2020). Targeting genes involved in natural fertilization of female gametic cells using CRISPR/Cas9 impeded fertilization and resulted in haploid embryos generated through egg cells (Hooghvorst and Nogués 20200). CRISPR–Cas9-based HI systems avoided wide ranging adaptation of protocols to different genotypes. Currently, genes involved in chromosome segregation (MATL, CENH3, and DMP) are well-known targets for HI in both monocots and dicots. Targeting MALT by CRISPR/Cas9 in wheat obtained 18.9% haploid progeny (Liu et al. 2019a; b). Similarly, targeting DMP in maize by CRISPR/Cas9 achieved maternal haploids with the efficacy of 0.1–0.3% (Zhong et al. 2019a, b). To obtain doubled haploid homozygous lines, traditional HI needs six to eight generations, genome-editing mediated HI needs only 1 year (Hooghvorst and Nogués 2020), thus, CRISPR/Cas9 accelerates crop breeding via haploid induction.
Another HI-editing technology (HI-Edit) has been developed to directly edit elite inbred lines of diverse monocot and dicot species by a single cross. In MALT-based maternal HI system, the cross between CRISPR/Cas9-edited sperm cells and elite line egg cells leads to successful fertilization off egg cells and edited elite doubled haploid whose chromosomes are exclusively derived from the female parent. In CENH3-based paternal HI system, the cross between elite line (pollen donor) with CRISPR/Cas9-edited line results in female genome elimination and doubled haploid (Kelliher et al. 2019). Compared with existed HI systems, HI-Edit avoids the delay and high cost of introgression due to its faster and more effective delivery of edits to advanced breeding materials. HI-Edit provides transgene-free edited inbred lines lacking haploid-inducer parental DNA and the editing machinery as well (Kelliher et al. 2019).
Synthetic biology
Plant synthetic biology integrates engineering principles with biology to design and produce novel biological devices or systems (Wurtzel et al. 2019; Tian et al. 2020). Genome-editing technology could be used for targeted metabolic engineering to produce desirable products through direct knocking out or overexpressing of specific genes, or through the introduction of a combination of existing enzymes. Actually, genome-editing technology could play essential roles in plant synthetic biology to introduce new reactions/pathways that are not present in nature through de novo design, and to renovate endogenous signaling pathways (Tian et al. 2020; Zhang et al. 2020a). Currently, several redesigned/or synthesized novel biological devices or systems have been reported. For example, the rubisco subunits with RAF1 to enhance photosynthesis in maize (Salesse-Smith et al, 2018), a synthetic CETCH cycle constituting a reaction network of 17 enzymes from 9 different organisms of all 3 domains of life to continuously fix CO2 (Schwander et al. 2016), a synthetic glycolate metabolic pathway to increase C3 crop yield (South et al. 2019). However, none of them was introduced into plant systems using genome-editing approach. One reason is that CRISP/Cas system has been almost exclusively employed for gene knocking out and deletion but not for gene insertion. A recent study successfully re-oriented a 75.5-Mb-long targeted chromosomal region in maize using CRISPR/Cas9 approach (Schwartz et al. 2020), showing a great potential for the application of CRISPR/Cas9 in synthetic biology for chromosomal engineering to introduce large synthetic device or systems to plants, particularly in the case of overexpression of multiple stacking traits.
Molecular characteristics of genome-edited crops
The outcomes of CRISPR/Cas9 system in crops are affected by various factors, including Cas9 activity, gRNA expression, transformation procedure, callus culture time, and gRNA protospacer sequence (Mikami et al. 2015; Doench et al. 2016; Zhu et al. 2017). While most of the studies focus mainly on transient or early stable transformants (Feng et al. 2014; Wang et al. 2014; Zhang et al. 2014; Zhou et al. 2014; Zhu et al. 2017), very few pay attention to molecular characteristics in the consecutive generations. So far, mutation patterns and inheritability have been investigated mainly in Arabidopsis and rice (Feng et al. 2014; Zhang et al. 2014; Čermák et al. 2015). In Arabidopsis, a small incidence of homozygous mutation can be identified in the T1 generation, and rarely off-target mutation is reported (Fauser et al. 2014; Feng et al. 2014; Ma et al. 2015). On the other hand, in rice, homozygous and biallelic mutations appear even in T0 plants (Ma et al. 2015), and pass stably to next generations following laws of Mendel inheritance (Zhang et al. 2014).
The isolating transformants and identifying expected mutations are not the only task for molecular characterization of genome-edited crops, particularly for breeding purposes. Generally, exogenous T-DNA elements and Cas9 could be eliminated as early as in T1 generation (Xu et al. 2015; Zhou et al., 2014). However, if they are not eliminated intentionally at earlier stages, the presence of them in T2 generation is still high (Biswas et al. 2020a). The presence of exogenous element Cas9 in genome-edited plants could make new mutations in every subsequent generation, making it difficult for mutations transmission analysis and hampering inherit stability. Moreover, it may potentially cause off-target mutations. On the other hand, the presence of other exogenous elements rather than Cas9 in the genome-edited plants could be facing regulatory issues, making it difficult for moving forward to commercialization, because transgene-free is a prerequisite for regulatory approval of commercial utilization of genome-edited plants (He et al. 2018). There are close connections between the presence of exogenous elements and the presence of Cas9 (Biswas et al. 2020a), therefore, screening for the absence of Cas9 in T1 could help to eliminate the exogenous elements. Nevertheless, the screening for Cas9 cannot replace the screen for other exogenous elements in T1 or T2 generation (Biswas et al. 2020a). A potential commercialized product could be heritable, exogenous genetic elements-free, target-modified, and with expected traits that can be influenced by various factors that need to be characterized accordingly (Fig. 2).
Methods for the screening of CRISPR/Cas9-induced mutations in crops
The development of efficient, reliable, and inexpensive methods to effective screening for on-target and off-target genome-edited mutations from a pool of mutants in the early stages helps to speed up further basic and applied studies. To date, many different methods for the screening of genome editing-induced, specifically CRISPR/Cas9-induced indels, have been developed and applied in crops (Table 2).
These methods mentioned in Table 2 are generally PCR based, which are reported to be effective under certain circumstance, and thus, each has its own intrinsic limitations. For example, all of them can reveal mutated genotypes (insertion, deletion or substitution) but cannot reveal the exact nucleotide changes (which nucleotide is inserted, deleted or substituted) without Sanger sequencing. In contrast, many methods based on targeted deep sequencing, such as AGEseq, Cas-Analyzer, CRISPR-GA, CRISPResso and Hi-TOM, have been developed to identify simultaneously mutated genotypes and exact nucleotide changes with high accuracy and sensitivity (see review in Liu et al. 2019a, b). Obviously, as compared with targeted deep sequencing-based methods, PCR-based methods are cheaper and more suitable for screening purpose. Combined with Sanger sequencing, these abovementioned screening methods can also be used for identification purpose. In this case, desirable mutation genotypes (homozygous insertion, deletion or substitution mutants) are first screened from plenty of lines by these PCR-based methods and then confirmed with Sanger sequencing. Among methods mentioned in Table 2, multiplex ligation-dependent probe amplification (MLPA)-based method, when combined with Sanger sequencing, could be the most suitable and accurate approach to screen mutants. It has multiplex capabilities (about 60 different target sites in a single assay), high sensitivity (down to ± 1 bp and single nucleotide replacement) and reliability (suitable for different targets), and multiple functions (on- and off-target detection simultaneously) (Biswas et al. 2020b).
Newly emerging genome-editing tools in crops
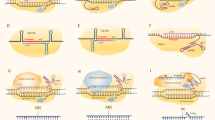
During past few years, several new CRISPR systems have been developed to improve the specificity and overcome the bottlenecks of the CRISPR/Cas9 system for more effective genome editing (Fig. 3), which continue to drive major advances in crop sciences and breeding. These emerging technologies could be essential tools for molecular crop-breeding purposes are discussed in below.
Genome-editing tools in crop improvement. a Cas9 protein forms a crRNA-tracrRNA complex with the sgRNA to bind upstream of the G/C-rich PAM sequence and initiates DSB through its RuvC and HNH domains. b Cpf1 and crRNA form a complex to bind the downstream of the A/T-rich PAM sequence and introduces DSB via a RuvC-like nuclease domain. c Cas13 targets a single strand RNA (ssRNA) molecule at the outer nuclease surface guided by crRNA in the absence of a PAM recognition site. d, e Cytosine base editor and adenine base editor, respectively. The complex of nickase Cas9 (nCas9-D10A) fused with cytidine deaminase/deoxyadenosine deaminase converts cytosine (C) or adenine (A) in the target site to uracil (U) or inosine (I), resulting C-T and A-G replacement, respectively. f Prime editor. The nCas9-RT complex formed between Cas9 nickase and engineered reverse transcriptase domains targets to editing sites by engineered prime editing guide RNAs (pegRNAs), and nicks the PAM-containing DNA strand by Cas9 nuclease. HEPN higher eukaryotes and prokaryotes nucleotide-binding domains; HNH His-Asn-His; pegRNA prime editing guide RNA; sgRNA single-guide RNA; UGI uracil glycosylase inhibitor
Cas12a/Cpf1 nuclease
The class 2 type V-A Cas protein Cpf1, also known as Cas12a with RNA-guided DNA endonuclease activity, has been widely applied in genome editing (Chen et al. 2019; Kim et al. 2017). Cpf1 uses a T-rich PAM sequence to recognize the target site in genomic DNA, which prolongs the editing sites behind those of G-rich PAM preferred by Cas9. The guide RNA of Cpf1 is shorter (about 43 bp) than sgRNA of Cas9 (about 100 bp), and the Cpf1 target site is positioned distal and downstream of the PAM sequence (Chen et al. 2019; Kim et al. 2017). Cpf1 produces staggered-ended DSBs at the distal location of a PAM, which provides further benefits than Cas9 due to knock-in strategies and enhances efficiency for the NHEJ-based gene insertion (Kim et al. 2017; Moon et al. 2018). Genome editing using Cpf1 system in crops has been reported in rice and soybean (Kim et al. 2017; Xu et al. 2017). Whole-genome sequencing analysis results indicated that neither Cas9 nor Cpf1 generates bona fide off-target mutations due to continued expression of Cas9 or Cpf1 in T1 rice (Tang et al. 2018). Notably, in vitro studies show that Cpf1 has robust non-specific activated nicking activities, which may lead to off-target editing (Murugan et al. 2020). Further investigations are needed to evaluate the specificity of Cpf1 in vivo in other crops and to improve current Cas12a-based applications (Schindele and Puchta 2020).
Cas12b/C2c1 nuclease
CRISPR-associated Cas12b, a class 2 type V-B nuclease, prefers T-rich PAM, creates staggered ends of DNA DSBs, and requires a crRNA and a trans-activating crRNA (combined as a sgRNA). In addition, Cas12b protein is smaller than Cas9 or Cas12a, which has been regarded as the promising CRISPR system for genome editing in crops. In rice, it recognizes VTTV PAMs, more preferring ATTV and GTTG PAMs. The successful establishment of a compelling Cas12b transcriptional activation system in rice indicated that Cas12b is more adaptable for versatile guide RNA engineering (Ming et al. 2020). Cas12b/C2c1 has been successfully used to induce mutations including to create large deletions at multiple loci, and to perform multiplex genome editing in Arabidopsis, which does not show any mutations at potential off-target sites (Wu et al. 2020). Nevertheless, Cas12b requires higher temperature for optimal activity (Teng et al. 2018), which needs to be modified to make it more practically for crop applications.
Cas13/C2c2 nuclease
Cas13, also known as C2c2, is a newly identified CRISPR effector, specifically cutting single-stranded RNA in eukaryotic cells (Wolter and Puchta 2018). Cas13 protein is assigned into class 2 type VI, which acts solely on RNA because of its unique HEPN (higher eukaryotes and prokaryotes nucleotide-binding) domains are exclusively associated with RNase activity. Notably, there is no strict requirement for PAM sequence for some Cas13 orthologues (Wolter and Puchta 2018). To date, three different Cas13 protein classes, such as Cas13a, Cas13b, and Cas13d, have been applied for RNA editing in plants (Schindele et al. 2019), mainly to target RNA for cleavage, for combating RNA viruses (Aman et al. 2018; Wolter and Puchta 2018). Combining Cas13 with other DNA-directed Cas nucleases opens new opportunities for crop breeding by targeting at both DNA and RNA levels.
Cas14/Cas12f nucleases
Cas14a, a highly compact class 2 type V nuclease, is an RNA-guided DNA nuclease that can be utilized for target-specific single-stranded DNA (ssDNA) cleavage (Harrington et al. 2018; Khan et al. 2019b). Cas14a does not require restrictive sequence to target and cleave ssDNA, which is different from other known class 2 systems (Harrington et al. 2018). Cas14a is the smallest functional CRISPR system to date, which is only one-third size of Cas9 (Harrington et al. 2018). The CRISPR/Cas14a system shows potential application in crops in defense against ssDNA viruses or mobile genetic elements. It has been used to create resistance against ssDNA viruses, including Geminiviridae and Nanoviridae families, in crops (Khan et al. 2019b).
Altogether, these abovementioned several nucleases (Table 3) enable a wide range of genome-editing applications in crops, with their unique characteristics either at DNA or RNA level. It is worthy to note that except Cas14a (Harrington et al. 2018), LwaCas13a and PspCas13b (Wolter and Puchta 2018), other nucleases require specific PAM sequences for their functions. Two recent studies demonstrate that using SpRY, a modified SpCAS9, does not need all those specific PAM sequences, and realizes PAM-less genome editing in rice (Ren et al. 2021; Xu et al. 2021), which greatly extends the application of CRISPR system in crop genome, and facilitates moving genome-edited crops towards commercialization. Nevertheless, the identification of novel CRISPR–Cas genome-editing systems free of off-target editing activity while maintaining robust on-target editing efficiency and compatibility with crop genomes continues to be a challenge in the future.
Base editors
Different from abovementioned nucleases, base editors (BEs) precisely generate targeted mutations without requirement of DSBs or donor DNA, and independent on homology-directed repair (HDR), providing efficient, simple, well-accepted techniques for specific base replacement at the target site (Chen et al. 2019). BEs are extremely useful when base editing of interested protein-coding genes is needed to generate genetic variants with improved agronomic traits (Li et al. 2020c). Currently, there are two classes of BEs: cytosine base editor (CBE) and adenine base editor (ABE). CBE converts of C-G base pair to T-A base pair while ABE converts of A-T base pair to G-C base pairs (Komor et al. 2016; Gaudelli et al. 2017). CBEs use cytidine deaminases to convert cytosine to uracil (Komor et al. 2016) while ABEs use TadA deoxyadenosine deaminases to convert adenosines to inosines (Gaudelli et al. 2020). Activities of both BEs depend largely on PAM availability, because they both use CRISPR–Cas DNA binding proteins to allow the targeted deamination of single nucleotides at the targeted sites. BEs install transition point mutations but cannot install transversion point mutations, precise insertions or deletions. Both CBE and ABE have been tested in rice, and genome-wide sequencing data indicated that CBE not ABE induces substantial genome-wide off-target mutations, highlighting needs to optimize fidelities of CBEs (Jin et al. 2019). Progresses have been made to increase genome-targeting scope and fidelity of BEs (Anzalone et al. 2020; Yan et al. 2021); hopefully, BEs will play more roles in both random mutagenesis and targeted random mutagenesis (Li et al. 2020c) in crop breeding.
Prime editors
Prime editing, an emerging genome-editing tool, can precisely introduce all possible types of point mutations, and small insertions/deletions without donor DNA or DSBs (Anzalone et al. 2020). Prime editors (PEs) are fused proteins of Cas9 nickase domains with engineered reverse transcriptase domains. PEs target to editing sites by engineered prime editing guide RNAs (pegRNAs), nick the PAM-containing DNA strand by Cas9 nuclease, and prime reverse transcriptions using extensions in the pegRNAs as templates (Anzalone et al. 2020). Plant PEs have been successfully developed and applied to precisely edit several endogenous genes in rice and wheat protoplasts (Lin et al. 2020; Tang et al. 2020), to achieve stable edited lines with desired edits in both exogenous and endogenous genes (Butt et al., 2020); Li et al., 2020e). Nevertheless, this new technology is still at the experimental stages, more studies are needed to apply PEs in crop for different trait improvement.
Safety regulations of genome-edited crops
There is an ongoing argument whether a genome-edited organism obtained by the CRISPR technology is or is not regarded as a genetically modified organism (GMO), and regulated or not regulated as a GMO (Eş et al. 2019). In fact, similar to GMO, genome-edited crops are regulated globally in either technology-based or final product-based manner (Table 4) (Eckerstorfer et al. 2019; Van Vu et al. 2019). Some countries, including European Union (EU), New Zealand, and India, recognize and regulate genome-edited crops as GMOs based on technologies used to generate them (Jouanin et al. 2018; Fritsche et al. 2018; Friedrichs et al. 2019). On the other hand, Argentina, Australia, Brazil, Canada, Chile, Japan, and the USA, recognize and regulate genome-edited crops based on the final products, and if they are free of transgene, they are regarded as non-GMOs (Lema 2019; Eckerstorfer et al. 2019; Eriksson et al. 2019; Smyth 2017; Van Vu et al. 2019; Razzaq et al. 2019). Several countries like Nigeria and Kenya are in their way of developing regulatory policy on genome-editing crops, but many countries have not yet confirmed their positions (Tripathi et al. 2020; Eckerstorfer et al. 2019). In China, according to the current law, GE plants fall in the regulation scope of GMO, specific laws regarding genome-edited products are not yet announced (Gao et al. 2018).
Different countries take different initiative policies regarding regulatory landscapes on genome-edited crops, resulting in an inconsistent global regulatory system, which somehow hinders commercial utilization of genome-edited crops and pragmatic technological improvement. There is a need to establish a more optimistic and more realistic regulatory system regarding genome-edited plants globally, bring the world under the one safety regulation umbrella.
Conclusions and future directions
In addition to applications in basic researches in crops sciences, CRISPR/Cas systems find ways in many aspects of crop breeding. With advances in the development of novel CRISPR/Cas systems that are more specific, accurate, efficient and feasible, CRISPR/Cas systems will play more roles in securing global food supply in a sustainable manner.
However, to be fully applied to crop improvement, further improvements on these versatile tools are needed. These include: (1) the fidelity, where the incidence of off-target effect should be null; (2) the applicability, where the activity is independent on PAM and the system is free of donor DNA; (3) the compatibility, where the delivery into crop cells is independent on species; and (4) the traceability, where any modifications in the genome should be traceable. Molecular characterization of crops generated from any genome-editing tools should be performed before any filed trials. Last but not the least, development of a pragmatic product-based global regulatory policy on genome-edited crops is necessary for speeding up the applications of these tools in crop breeding.
References
Ainley WM, Dent SL, Welter ME, Murray MG, Zeitler B, Amora R, Corbin DR, Miles RR, Arnold NL, Strange TL, Simpson MA (2013) Trait stacking via targeted genome editing. Plant Biotechnol J 11:1126–1134. https://doi.org/10.1111/pbi.12107
Ali Z, Shami A, Sedeek K, Kamel R, Alhabsi A, Tehseen M, Hassan N, Butt H, Kababji A, Hamdan SM, Mahfouz MM (2020) Fusion of the Cas9 endonuclease and the VirD2relaxase facilitates homology-directed repair forprecise genome engineering in rice. Commun Biol 3(1):44. https://doi.org/10.1038/s42003-020-0768
Aman R, Ali Z, Butt H, Mahas A, Aljedaani F, Khan MZ, Ding S, Mahfouz M (2018) RNA virus interference via CRISPR/Cas13a system in plants. Genome Biol 19:1. https://doi.org/10.1186/s13059-017-1381-1
Anzalone AZ, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, Chen PJ, Wilson C, Newby GA, Raguram A, Liu DR (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157. https://doi.org/10.1038/s41586-019-1711-4
Anzalone AZ, Koblan LW, Liu DR (2020) Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat Biotechnol 38:824–844. https://doi.org/10.1038/s41587-020-0561-9
Araki M, Ishii T (2015) Towards social acceptance of plant breeding by genome editing. Trends Plant Sci 20:145–149. https://doi.org/10.1016/j.tplants.2015.01.010
Arndell T, Sharma N, Langridge P, Baumann U, Watson-Haigh NS, Whitford R (2019) gRNA validation for wheat genome editing with the CRISPR-Cas9 system. BMC Biotechnol 19:71. https://doi.org/10.1186/s12896-019-0565-z
Baltes NJ, Voytas DF (2015) Enabling plant synthetic biology through genome engineering. Trends Biotechnol 33:120–131. https://doi.org/10.1016/j.tibtech.2014.11.008
Birchler JA, Yao H, Chudalayandi S, Vaiman D, Veitia RA (2010) Heterosis. Plant Cell 22:2105–2122. https://doi.org/10.1105/tpc.110.076133
Biswas S, Li R, Yuan Z, Zhang D, Zhao X, Shi J (2019) Development of methods for effective identification of CRISPR/Cas9- induced indels in rice. Plant Cell Rep 38(4):503–510. https://doi.org/10.1007/s00299-019-02392-3
Biswas S, Li R, Hong J, Zhao X, Yuan Z, Zhang D, Shi J (2020a) Effective identification of CRISPR/Cas9-induced and naturally occurred mutations in rice using a multiplex ligation-dependent probe amplification-based method. Theor Appl Genet 133:2323–2334. https://doi.org/10.1007/s00122-020-03600-5
Biswas S, Tian J, Li R, Chen X, Luo Z, Chen M, Zhao X, Zhang D, Persson S, Yuan Z, Shi J (2020b) Investigation of CRISPR/Cas9-induced SD1 rice mutants highlights the importance of molecular characterization in plant molecular breeding. J Genet Genom. https://doi.org/10.1016/j.jgg.2020.04.004
Brauer EK, Balcerzak M, Rocheleau H, Leung W, Schernthaner J, Subramaniam R, Ouellet T (2020) Genome editing of a deoxynivalenol-induced transcription factor confers resistance to Fusarium graminearum in wheat. Mol Plant Microbe Interact 33(3):553–560. https://doi.org/10.1094/MPMI-11-19-0332-R
Butt H, Eid A, Momin AA, Bazin J, Crespi M, Arold ST, Mahfouz MM (2019) CRISPR directed evolution of the spliceosome for resistance to splicing inhibitors. Genome Biol 20:73. https://doi.org/10.1186/s13059-019-1680-9
Butt H, Rao GS, Sedeek K, Aman R, Mahfouz M (2020) Engineering herbicide resistance via prime editing in rice. Plant Biotechnol J 18:2370–2372. https://doi.org/10.1111/pbi.13399
Čermák T, Baltes NJ, Čegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol 16:232. https://doi.org/10.1186/s13059-015-0796-9
Chen K, Gao C (2013) TALENs: Customizable molecular DNA scissors for genome engineering of plants. J Genet Genomics 40:271–279. https://doi.org/10.1016/j.jgg.2013.03.009
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697. https://doi.org/10.1146/annurev-arplant-050718-100049
Curtin SJ, Zhang F, Sander JD, Haun WJ, Starker C, Baltes NJ, Reyon D, Dahlborg EJ, Goodwin MJ, Coffman AP, Dobbs D, Joung JK, Voytas DF, Stupar RM (2011) Targeted mutagenesis of duplicated genes in soybean with zinc-finger nucleases. Plant Physiol 156:466–473. https://doi.org/10.1104/pp.111.172981
Curtin SJ, Xiong Y, Michno JM, Campbell BW, Stec AO, Cermak T, Starker C, Voytas DF, Eamens AL, Stupar RM (2018) CRISPR/Cas9 and TALENs generate heritable mutations for genes involved in small RNA processing of Glycine max and Medicago truncatula. Plant Biotechnol J 16:1125–1137. https://doi.org/10.1111/pbi.12857
Do PT, Nguyen CX, Bui HT, Tran LTN, Stacey G, Gillman JD, Zhang ZJ, Stacey MG (2019) Demonstration of highly efficient dual gRNA CRISPR/Cas9 editing of the homeologous GmFAD2–1A and GmFAD2–1B genes to yield a high oleic, low linoleic and α-linolenic acid phenotype in soybean. BMC Plant Biol 19:311. https://doi.org/10.1186/s12870-019-1906-8
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321. https://doi.org/10.1016/j.cell.2006.12.006
Doench JG, Fusi N, Sullender M, Hegde M, Vaimberg EW, Donovan KF, Smith I, Tothova Z, Wilen C, Orcherd R, Virgin HW, Listgarten J, Root DE (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34:184–191. https://doi.org/10.1038/nbt.3437
Dong OX, Yu S, Jain R, Zhang N, Duong PQ, Butler C, Li Y, Lipzen A, Martin JA, Barry KW, Schmutz J, Tian L, Ronald PC (2020) Marker-free carotenoid-enriched rice generated through targeted gene insertion using CRISPR-Cas9. Nat Commun 11:1178. https://doi.org/10.1038/s41467-020-14981-y
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346:1258096. https://doi.org/10.1126/science.1258096
Eckerstorfer MF, Engelhard M, Heissenberger A, Simon S, Teichmann H (2019) Plants developed by new genetic modification techniques-Comparison of existing regulatory frameworks in the EU and Non-EU countries. Front Bioeng Biotechnol 7:26. https://doi.org/10.3389/fbioe.2019.00026
El-Mounadi K, Morales-Floriano ML, Garcia-Ruiz H (2020) Principles, applications, and biosafety of plant genome editing using CRISPR-Cas9. Front Plant Sci 11:56. https://doi.org/10.3389/fpls.2020.00056
Eriksson D, Kershen D, Nepomuceno A, Pogson BJ, Prieto H, Purnhagen K, Smyth S, Wesseler J, Whelan A (2019) A comparison of the EU regulatory approach to directed mutagenesis with that of other jurisdictions, consequences for international trade and potential steps forward. New Phytol 222:1673–1684. https://doi.org/10.1111/nph.15627
Eş I, Gavahian M, Marti-Quijal FJ, Lorenzo JM, Khaneghah AM, Tsatsanis C, Kampranis SC, Barba FJ (2019) The application of the CRISPR-Cas9 genome editing machinery in food and agricultural science: Current status, future perspectives, and associated challenges. Biotechnol Adv 37:410–421. https://doi.org/10.1016/j.biotechadv.2019.02.006
Fauser F, Schiml S, Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J 79:348–359. https://doi.org/10.1111/tpj.12554
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL, Wang Z, Zhang Z, Zheng R, Yang L, Zeng L, Liu X, Zhu ZK (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc Natl Acad Sci U S A 111:4632–4637. https://doi.org/10.1073/pnas.1400822111
Fernie AR, Yan J (2019) De novo domestication: an alternative route toward new crops for the future. Mol Plant 5:615–631. https://doi.org/10.1016/j.molp.2019.03.016
Friedrichs S, Takasu Y, Kearns P, Dagalliar B, Oshima R, Schofield J, Moreddu C (2019) Meeting report of the OECD conference on “genome editing: applications in agriculture-implications for health, environment and regulation.” Transgenic Res 28:419–463. https://doi.org/10.1007/s11248-019-00154-1
Fritsche S, Poovaiah C, MacRae E, Thorlby G (2018) A New Zealand perspective on the application and regulation of gene editing. Front Plant Sci 9:1323. https://doi.org/10.3389/fpls.2018.01323
Gao C (2021) Genome engineering for crop improvement and future agriculture. Cell 184:1621–1635. https://doi.org/10.1016/j.cell.2021.01.005
Gao W, Xu WT, Huang KL, Guo MZ, Luo YB (2018) Risk analysis for genome editing-derived food safety in China. Food Control 84:128–137. https://doi.org/10.1016/j.foodcont.2017.07.032
Gao H, Gadlage MJ, Lafitte HR, Lenderts B, Yang M, Schroder M, Farrell J, Snopek K, Peterson D, Feigenbutz L, Jones S, Clair GST, Rahe M, Sanyour-Doyel N, Peng C, Wang L, Young JK, Beatty M, Dahlke B, Hazebroek J, Greene TW, Cigan AM, Chilcoat ND, Meeley RB (2020) Superior field performance of waxy corn engineered using CRISPR–Cas9. Nat Biotechnol 38:579–581. https://doi.org/10.1038/s41587-020-0444-0
Garcia-Doval C, Jinek M (2017) Molecular architectures and mechanisms of Class 2 CRISPR-associated nucleases. Curr Opin Struc Biol 47:157–166. https://doi.org/10.1016/j.sbi.2017.10.015
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, Liu DR (2017) Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551:464–471. https://doi.org/10.1038/nature24644
Gaudelli NM, Lam DK, Rees HA, Solá-Esteves NM, Barrera LA, Born DA, Edwards A, Gehrke JM, Lee SJ, Liquori AJ, Murray R, Packer MS, Rinaldi C, Slaymaker IM, Yen J, Young LE, Ciaramella G (2020) Directed evolution of adenine base editors with increased activity and therapeutic application. Nature Biotechnol 38:892–900. https://doi.org/10.1038/s41587-020-0491-6
Gomez MA, Lin ZD, Moll T, Chauhan RD, Hayden L, Renninger K, Beyene G, Taylor NJ, Carrington JC, Staskawicz BJ, Bart BS (2019) Simultaneous CRISPR/Cas9-mediated editing of cassava eIF4E isoforms nCBP-1 and nCBP-2 reduces cassava brown streak disease symptom severity and incidence. Plant Biotechnol J 17:421–434. https://doi.org/10.1111/pbi.12987
Guo J, Gaj T, Barbas CF (2010) Directed evolution of an enhanced and highly efficient FokI cleavage domain for zinc finger nucleases. J Mol Biol 400:96–107. https://doi.org/10.1016/s13007-018-0305-8
Guo J, Li K, Jin L, Xu R, Miao K, Yang F, Qi C, Zhang L, Botella JR, Wang R, Miao Y (2018) A simple and cost-effective method for screening of CRISPR/Cas9-induced homozygous/biallelic mutants. Plant Methods 14:40. https://doi.org/10.1186/s13007-018-0305-8
Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, White IP, Cofsky JC, Kyrpides NC, Banfield JF, Doudna JA (2018) Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362:839–842. https://doi.org/10.1126/science.aav4294
He Y, Zhu M, Wang L, Wu J, Wang Q, Wang R, Zhao Y (2018) Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol Plant 11:1210–1213. https://doi.org/10.1016/j.molp.2018.05.005
Holubova K, Hensel G, Vojta P, Tarkowski P, Bergougnoux V, Galuszka P (2018) Modification of barley plant productivity through regulation of cytokinin content by reverse-genetics approaches. Front Plant Sci 9:1676. https://doi.org/10.3389/fpls.2018.01676
Hong J, Shi Q, Biswas S, Jiang SC, Shi J (2021) Moving genome edited crops forward from the laboratory bench to the kitchen table. Food Cont 122:107790. https://doi.org/10.1016/j.foodcont.2020.107790
Hooghvorst I, Nogués S (2020) Chromosome doubling methods in doubled haploid and haploid inducer-mediated genome-editing systems in major crops. Plant Cell Rep. https://doi.org/10.1007/s00299-020-02605-0
Hua Y, Wang C, Huang J, Wang K (2017) A simple and efficient method for CRISPR/Cas9-induced mutant screening. J Genet Genomics 44:207–213. https://doi.org/10.1016/j.jgg.2017.03.005
Jaganathan D, Ramasamy K, Sellamuthu G, Jayabalan S, Venkataraman G (2018) CRISPR for crop improvement: an update review. Front Plant Sci 9:985. https://doi.org/10.3389/fpls.2018.00985
Jin S, Zong Y, Gao Q, Zhu Z, Wang Y, Qin P, Liang C, Wang D, Qiu JL, Zhang F, Gao C (2019) Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364:292–295. https://doi.org/10.1126/science.aaw7166
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821. https://doi.org/10.1126/science.1225829
Jouanin A, Boyd LA, Visser RGF, Smulders MJ (2018) Development of wheat with hypoimmunogenic gluten obstructed by the gene editing policy in Europe. Front Plant Sci 9:1523. https://doi.org/10.3389/fpls.2018.01523
Karunarathna NL, Wang H, Harloff HJ, Jiang L, Jung C (2020) Elevating seed oil content in a polyploid crop by induced mutations in SEED FATTY ACID REDUCER genes. Plant Biotechnol J 18:2251–2266. https://doi.org/10.1111/pbi.13381
Kelliher T, Starr D, Su X, Tang G, Chen Z, Carter J, Wittich PE, Song S, Green J, Burch E, McCuiston J, Gu W, Sun Y, Strebe T, Roberts J, Bate NJ, Que Q (2019) One-step genome editing of elite crop germplasm during haploid induction. Nat Biotechnol 37:287–292. https://doi.org/10.1038/s41587-019-0038-x
Khan MSS, Basnet R, Islam SA, Shu Q (2019a) Mutational analysis of OsPLDα1 reveals its involvement in phytic acid biosynthesis in rice grains. J Agric Food Chem 41:11436–11443. https://doi.org/10.1021/acs.jafc.9b05052
Khan MZ, Haider S, Mansoor S, Amin I (2019b) Targeting plant ssDNA viruses with engineered miniature CRISPR-Cas14a. Trends Biotech 37:800–804. https://doi.org/10.1016/j.tibtech.2019.03.015
Khanday I, Skinner D, Yang B, Mercier R, Sundaresan V (2019) A male-expressed rice embryogenic trigger redirected for asexual propagation through seeds. Nature 565:91–95. https://doi.org/10.1038/s41586-018-0785-8
Kim H, Kim ST, Ryu J, Kang BC, Kim JS, Kim SG (2017) CRISPR/Cpf1-mediated DNA-free plant genome editing. Nat Commun 8:14406. https://doi.org/10.1038/ncomms14406
Kim D, Alptekin B, Budak H (2018) CRISPR/Cas9 genome editing in wheat. Funct Integr Genomics 18:31–41. https://doi.org/10.1007/s10142-017-0572-x
Kis A, Hamar E, Tholt G, Bán R, Havelda Z (2019) Creating highly efficient resistance against wheat dwarf virus in barley by employing CRISPR/Cas9 system. Plant Biotechnol J 17(6):1004–1006. https://doi.org/10.1111/pbi.13077
Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR (2016) Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533:420–424. https://doi.org/10.1038/nature17946
Kuang Y, Li S, Ren B, Yan F, Spetz C, Li X, Zhou X, Zhou H (2020) Base-editing-mediated artificial evolution of OsALS1 in planta to develop novel herbicide-tolerant rice germplasms. Mol Plant 13:565–572. https://doi.org/10.1016/j.molp.2020-01-010
Kumar N, Galli M, Ordon J, Stuttmann J, Kogel K-H, Imani J (2018) Further analysis of barley MORC1 using a highly efficient RNA-guided Cas9 gene-editing system. Plant Biotechnol j 16(11):1892–1903. https://doi.org/10.1111/pbi.12924
Kumar P, Alok A, Kumar J (2020a) Expanding the potential of CRISPR-Cas9 technology for crops improvement. In: Singh V (ed) Advances in Synthetic Biology. Springer, Singapore. https://doi.org/10.1007/978-981-15-0081-7_15
Kumar VVS, Verma RK, Yadav RK, Yadav P, Watts A, Rao MV, Chinnusamy V (2020b) CRISPR-Cas9 mediated genome editing of drought and salt tolerance (OsDST) gene in indica mega rice cultivar MTU1010. Physiol Mol Biol Plants 26:1099–1110. https://doi.org/10.1007/s12298-020-00819-w
Kwon CT, Heo J, Lemmon ZH, Capua Y, Hutton SF, Eck JV, Park SJ, Lippman ZB (2020) Rapid customization of Solanaceae fruit crops for urban agriculture. Nat Biotechnol 38:182–188. https://doi.org/10.1038/s41587-019-0361-2
Lacchini E, Kiegle E, Castellani M, Adam H, Jouannic S, Gregis V, Kater MM (2020) CRISPR-mediated accelerated domestication of African rice landraces. PLoS ONE 15(3):e0229782. https://doi.org/10.1371/journal.pone.0229782
Lema MA (2019) Regulatory aspects of gene editing in Argentina. Transgenic Res 28:147–150. https://doi.org/10.1007/s11248-019-00145-2
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390–392. https://doi.org/10.1038/nbt.2199
Li M, Li X, Zhou Z, Wu P, Fang M, Pan X, Lin Q, Luo W, Wu G, Li H (2016) Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci 7:377. https://doi.org/10.3389/fpls.2016.00377
Li T, Yang X, Yu Y, Si X, Zhai X, Zhang H, Dong W, Gao C, Xu C (2018a) Domestication of wild tomato is accelerated by genome editing. Nature Biotechnol 36:1160–1163. https://doi.org/10.1038/nbt.4273
Li X, Wang Y, Chen S, Tian H, Fu D, Zhu B, Luo Y, Zhu H (2018b) Lycopene is enriched in tomato fruit by CRISPR/Cas9-mediated multiplex genome editing. Front Plant Sci 9:559. https://doi.org/10.3389/fpls.2018.00559
Li J, Li Y, Ma L (2019a) CRISPR/Cas9-based genome editing and its applications for functional genomic analyses in plants. Small Methods 3:1800473. https://doi.org/10.1002/smtd.201800473
Li R, Liu C, Zhao R, Wang L, Chen L, Yu W, Zhang S, Sheng J, Shen L (2019b) CRISPR/Cas9-Mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol 19:38. https://doi.org/10.1186/s12870-018-1627-4
Li C, Zhang R, Meng X, Chen S, Zong Y, Lu C, Qiu J-L, Chen Y-H, Li J, Gao C (2020a) Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors. Nat Biotechnol 38:875–882. https://doi.org/10.1038/s41587-019-0393-7
Li H, Li J, Chen J, Yan L, Xia L (2020b) Precise modifications of both exogenous and endogenous genes in rice by prime editing. Mol Plant 13:671–674. https://doi.org/10.1016/j.molp.2020.03.011
Li H, Li X, Xu Y, Liu H, He M, Tian X, Wang Z, Wu X, Bu Q, Yang J (2020c) High-efficiency reduction of rice amylose content via CRISPR/Cas9-mediated base editing. Rice Sci 27(6):445–448. https://doi.org/10.1016/j.rsci.2020.09.001
Li Q, Wu G, Zhao Y, Wang B, Zhao B, Kong D, Wei H, Chen C, Wang H (2020d) CRISPR/Cas9-mediated knockout and overexpression studies reveal a role of maize phytochrome C in regulating flowering time and plant height. Plant Biotechnol J. https://doi.org/10.1111/pbi.13429
Li R, Ba Y, Song Y, Cui J, Zhang X, Zhang D, Yuan Z, Yang L (2020e) Rapid and sensitive screening and identification of CRISPR/Cas9 edited rice plants using quantitative real-time PCR coupled with high resolution melting analysis. Food Cont 112:107088. https://doi.org/10.1016/j.foodcont.2020.107088
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted mutagenesis in Zea mays using TALENs and the CRISPR/Cas system. J Genet Genom 41:63–68. https://doi.org/10.1016/j.jgg.2013.12.001
Liao S, Qin X, Luo L, Han Y, Wang X, Usman B, Nawaz G, Zhao N, Liu Y, Li R (2019) CRISPR/Cas9-induced mutagenesis of semi-rolled leaf1,2 confers curled leaf phenotype and drought tolerance by influencing protein expression patterns and ROS scavenging in rice (Oryza sativa L.). Agrnomoy 9:728. https://doi.org/10.3390/agronomy9110728
Lin Q, Zong Y, Xue C, Wang S, Jin S, Zhu Z, Wang Y, Anzalone AV, Raguram A, Domam JL, Liu DR, Gao C (2020) Prime genome editing in rice and wheat. Nat Biotechnol 38:582–585. https://doi.org/10.1038/s41587-020-0455-x
Liu H, Wang K, Jia Z, Gong Q, Lin Z, Du L, Pei X, Ye X (2019a) Efficient induction of haploid plants in wheat by editing of TaMTL using an optimized Agrobacterium-mediated CRISPR system. J Exp Bot 71:1337–1349. https://doi.org/10.1093/jxb/erz529
Liu Y, Li G, Zhang Y, Chen L (2019b) Current advances on CRISPR/Cas genome editing technologies in plants. J South China Agric Univ 40:38–49. https://doi.org/10.7671/j.issn.1001-411X.201905058
Liu L, Kuang Y, Yan F, Li S, Ren B, Gosavi G, Spetz C, Li X, Wang X, Zhou X, Zhou H (2020a) Developing a novel artificial rice germplasm for dinitroaniline herbicide resistance by base editing of OsTubA2. Plant Biotechnol J. https://doi.org/10.1111/pbi.13430
Liu X, Qin R, Li J, Liao S, Shan T, Xu R, Wu D, Wei P (2020b) A CRISPR-Cas9-mediated domain-specific base-editing screen enables functional assessment of ACCase variants in rice. Plant Biotechnol J 18:1845–1847. https://doi.org/10.1111/pbi.13348
Lusser M, Parisi C, Plan D, Rodríguez-Cerezo E, Rodríguez-Cerezo E (2012) Deployment of new biotechnologies in plant breeding. Nat Biotechnol 30:231–239. https://doi.org/10.1038/nbt.2142
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Dong Z, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8:1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Ma X, Zhu Q, Chen Y, Liu YG (2016) CRISPR/Cas9 platforms for genome editing in plants: developments and applications. Mol Plant 9:961–974. https://doi.org/10.1016/j.molp.2016.04.009
Mikami M, Toki S, Engdo M (2015) Parameters affecting frequency of CRISPR/Cas9 mediated targeted mutagenesis in rice. Plant Cell Rep 34:1807–1815. https://doi.org/10.1007/s00299-015-1826-5
Miladinovic D, Antunes D, Yildirim K, Bakhsh A, Cvejić S, Kondić-Špika A, Jeromela AM, Opsahl-Sorteberg HG, Zambounis A, Hilioti Z (2021) Targeted plant improvement through genome editing: from laboratory to field. Plant Cell Rep. https://doi.org/10.1007/s00299-020-02655-4
Ming M, Ren Q, Pan C, He Y, Zhang Y, Liu S, Zhong Z, Wang J, Malzahn AA, Wu J, Zheng X, Zhang Y, Qi Y (2020) Crispr–Cas12b enables efficient plant genome engineering. Nat Plants 6:202–208. https://doi.org/10.1038/s41477-020-0614-6
Moon SB, Lee JM, Kang JG, Lee NE, Ha DI, Kim DY, Kim SH, Yoo K, Kim D, Ko JH, Kim YS (2018) Highly efficient genome editing by CRISPR-Cpf1 using CRISPR RNA with a uridinylate-rich 3’-overhang. Nature Commun 9:3651. https://doi.org/10.1038/s41467-018-06129-w
Moon SB, Kim DY, Ko JH, Kim YS (2019) Recent advances in the CRISPR genome editing tool set. Exp Mol Med 51:1–11. https://doi.org/10.1038/s12276-019-0339-7
Murugan K, Seetharam AS, Severin AJ, Sashital DG (2020) CRISPR-Cas12a has widespread off-target and dsDNA-nicking effects. J Biol Chem 295:5538–5553. https://doi.org/10.1074/jbc.RA120.012933
Nekrasov V, Staskawicz B, Weigel D, Jones JDG, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31:691–693. https://doi.org/10.1038/nbt.2655
Ortigosa A, Gimenez-lbanez S, Leonhardt N, Solano R (2019) Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol J 17:665–673. https://doi.org/10.1111/pbi.13006
Osakabe Y, Osakabe K (2015) Genome editing with engineered nucleases in plants. Plant Cell Physiol 56:389–400. https://doi.org/10.1093/pcp/pcu170
Pacher M, Puchta H (2017) From classical mutagenesis to nuclease-based breeding-directing natural DNA repair for a natural end-product. Plant J 90:819–833. https://doi.org/10.1111/tpj.13469
Peng C, Wang H, Xu X, Wang X, Chen X, Wei W, Lai Y, Liu G, Godwin ID, Li J, Zhang L, Xu J (2018) High-throughput detection and screening of plants modified by gene editing using quantitative real-time polymerase chain reaction. Plant J 95:557–567. https://doi.org/10.1111/tpj.13961
Pröbsting M, Schenke D, Hossain R, Häder C, Thurao T et al (2020) Loss of function of CRT1a (calreticulin) reduces plant susceptibility to Verticillium longisporum in both Arabidopsis thaliana and oilseed rape (Brassica napus). Plant Biotechnol j 18:2328–2344. https://doi.org/10.1111/pbi.13394
Razzaq A, Saleem F, Kanwal M, Mustafa G, Yousaf S, Arshad HMI, Hameed MK, Khan MS, Joyia FA (2019) Modern trends in plant genome editing: an inclusive review of the CRISPR/Cas9 toolbox. Int J Mol Sci 20:4045. https://doi.org/10.3390/ijms20164045
Ren Q, Sretenovic S, Liu S, Tang X, Huang L, He Y, Liu L, Guo Y, Zhong Z, Liu G, Cheng Y, Zheng X, Pan C, Yin D, Zhang Y, Li W, Qi L, Li C, Qi Y, Zhang Y (2021) PAM-less plant genome editing using a CRISPR–SpRY toolbox. Nat Plants. https://doi.org/10.1038/s41477-020-00827-4
Safari F, Zare K, Negahdaripour M, Barekati-Mowahed M, Ghasemi Y (2019) CRISPR- Cpf1 proteins: structure, function and implications for genome editing. Cell Biosci 9:36. https://doi.org/10.1186/s13578-019-0298-7
Salesse-Smith CE et al (2018) Overexpression of Rubisco subunits with RAF1 increases Rubisco content in maize. Nat Plants 4:802–810. https://doi.org/10.1038/s41477-018-0252-4
Sánchez-León S, Gil-Humanes J, Ozuna CV, Giménez MJ, Sousa C, Voytas DF, Barro F (2018) Low-gluten, nontransgenic wheat engineered with CRISPR/Cas9. Plant Biotechnol J 16:902–910. https://doi.org/10.1111/pbi.12837
Sashidhar N, Harloff HJ, Potgieter L, Jung C (2020) Gene editing of three BnITPK genes in tetraploid oilseed rape leads to significant reduction of phytic acid in seeds. Plant Biotechnol J 18:2241–2250. https://doi.org/10.1111/pbi.13380
Schaart JG, van de Wiel CCM, Lotz LAP, Smulders MJM (2016) Opportunities for products of new plant breeding techniques. Trends Plant Sci 21:438–449. https://doi.org/10.1016/j.tplants.2015.11.006
Schaeffer SM, Nakata PA (2015) CRISPR/Cas9-mediated genome editing and gene replacement in plants: Transitioning from lab to field. Plant Sci 240:130–142. https://doi.org/10.1016/j.plantsci.2015.09.011
Schindele P, Puchta H (2020) Engineering CRISPR/LbCas12a for highly efficient, temperature-tolerant plant gene editing. Plant Biotechnol J 8:1118–1120. https://doi.org/10.1111/pbi.13275
Schindele A, Dorn A, Puchta H (2019) CRISPR/Cas brings plant biology and breeding into the fast lane. Curr Opin Biotechnol 61:7–14. https://doi.org/10.1016/j.copbio.2019.08.006
Schnable PS, Springer NM (2013) Progress toward understanding heterosis in crop plants. Annu Rev Plant Biol 64:71–88. https://doi.org/10.1141/annurev-arplant-042110-103827
Schwander T, Schada von Borzyskowski L, Burgener S, Cortina NS, Erb TJ (2016) A synthetic pathway for the fixation of carbon diox-ide in vitro. Science 354:900–904. https://doi.org/10.1126/science.aah5237
Schwartz C, Lenderts B, Feigenbutz L, Barone P, Llaca V, Fengler K, Svitashev S (2020) CRISPR–Cas9-mediated 75.5-Mb inversion in maize. Nat Plants 6:1427–1431. https://doi.org/10.1038/s41477-020-00817-6
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9(10):2395–2410. https://doi.org/10.1038/nprot.2014.157
Shimatani Z, Kashojiya S, Takayama M, Terada R, Arazoe T, Ishii H, Teramura H, Yamamoto T, Komatsu H, Miura K, Ezura H, Nishida K, Ariizumi T, Kondo A (2017) Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat Biotechnol 35:441–443. https://doi.org/10.1038/nbt.3833
Smyth SJ (2017) Canadian regulatory perspectives on genome engineered crops. GM Crop Food 8:35–43. https://doi.org/10.1080/21645698.2016.1257468
South PF, Cavanagh AP, Liu HW, Ort DR (2019) Synthetic glycolate metabolism pathways stimulate crop growth and productivity in the field. Science 363:9077. https://doi.org/10.1126/science.aat9077
Tang X, Liu G, Zhou J, Ren Q, You Q, Tian L, Xin X, Zhong Z, Liu B, Zheng X, Zhang D, Malzahn A, Gong Z, Qi Y, Zhang T, Zhang Y (2018) A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 (Cas12a) nucleases in rice. Genome Biol 19:84. https://doi.org/10.1186/s13059-018-1458-5
Tang X, Sretenovic S, Ren Q, Jia X, Li M et al (2020) Plant prime editors enable precise gene editing in rice cells. Mol Plant 13:667–670. https://doi.org/10.1016/j.molp.2020.03.010
Teng F, Cui T, Feng G, Guo L, Xu K, Gao Q, Li T, Li J, Zhou Q, Li W (2018) Repurposing CRISPR-Cas12b for mammalian genome engineering. Cell Discov 4:63. https://doi.org/10.1038/s41421-018-0069-3
Tian S, Jiang L, Cui X, Zhang J, Guo S, Li M, Zhang H, Ren Y, Gong G, Zong M, Liu F, Chen Q, Xu Y (2018) Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep 37:1353–1356. https://doi.org/10.1007/s00299-018-2299-0
Tian X, Wang JW, Li J, Han B (2020) Designing future crops: challenges and strategies forsustainable agriculture. Plant J 105:1165–1178. https://doi.org/10.1111/tpj.15107
Tripathi L, Ntui VO, Tripathi JN (2020) CRISPR/Cas9-based genome editing of banana for disease resistance. Curr Opin Plant Biol 56:118–126. https://doi.org/10.1016/j.pbi.2020.05.003
Tuncel A, Corbin KR, Ahn-Jarvis J, Harris S, Hawkins E et al (2019) Cas9-mediated mutagenesis of potato starch-branching enzymes generates a range of tuber starch phenotypes. Plant Biotechnol j 17(12):2259–2271. https://doi.org/10.1111/pbi.13137
Van Vu T, Sung YW, Kim J, Doan DTH, Tran MT, Kim JY (2019) Challenges and perspectives in homology-directed gene targeting in monocot plants. Rice 12:95. https://doi.org/10.1186/s12284-019-0355-1
Veillet F, Chauvin L, Kermarrec MP, Sevestre F, Merrer M et al (2019a) The Solanum tuberosum GBSSI gene: a target for assessing gene and base editing in tetraploid potato. Plant Cell Rep 38:1065–1080. https://doi.org/10.1007/s00299-019-02426-w
Veillet F, Perrot L, Chauvin L, Kermarrec MP, Guyon-Debast A, Chauvin JE, Nogue F, Mazier M (2019b) Transgene-free genome editing in tomato and potato plants using agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int j Mol Sci 20:402. https://doi.org/10.3390/ijms20020402
Vu TV, Sivankalyani V, Kim EJ, Doan DTH, Tran MT, Kim J, Sung YW, Park M, Kang YJ, Kim JY (2020) Highly efficient homology-directed repair using CRISPR/Cpf1-geminiviral replicon in tomato. Plant Biotechnol J 18:2133–2143. https://doi.org/10.1111/pbi.13373
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947–951. https://doi.org/10.1038/nbt.2969
Wang K, Mei DY, Liu QN, Xiao XH, Ruan WM, Huang T, Cao GS (2015) Research of methods to detect genomic mutations induced by CRISPR/Cas systems. J Biotechnol 214:128–132. https://doi.org/10.1016/j.jbiotec.2015.09.029
Wang C, Liu Q, Shen Y, Hua Y, Wang J, Lin J, Wu M, Sun T, Cheng Z, Mercier R, Wang K (2019) Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes. Nat Biotechnol 37:283–286. https://doi.org/10.1038/s41587-018-0003-0
Wang C, Wang G, Gao Y, Lu G, Habben JE, Mao G, Chen G, Wang J, Yang F, Zhao X, Zhang J, Mo H, Qu P, Liu J, Greene TW (2020a) A cytokinin-activation enzyme-like gene improves grain yield under various field conditions in rice. Plant Mol Biol 102:373–388. https://doi.org/10.1007/s11103-019-00952-5
Wang S, Yang Y, Guo M, Zhong C, Yan C, Sun S (2020b) Targeted mutagenesis of amino acid transporter genes for rice quality improvement using the CRISPR/Cas9 system. Crop J 8:457–464. https://doi.org/10.1016/j.cj.2020.02.005
Wolter F, Puchta H (2018) The CRISPR/Cas revolution reaches the RNA world: Cas13, a new Swiss Army Knife for plant biologists. Plant J 94:767–775. https://doi.org/10.1111/tpj.13899
Wu F, Qiao X, Zhao Y, Zhang Z, Gao Y, Shi L, Du H, Wang L, Zhang YJ, Zhang Y, Liu L, Wang Q, Kong D (2020) Targeted mutagenesis in Arabidopsis thaliana using CRISPR-Cas12b/C2c1. J Integr Plant Biol 11:1653–1658. https://doi.org/10.1111/jipb.12944
Wurtzel ET, Vickers CE, Hanson AD, Millar AH, Cooper M, Voss-Fels KP, Nikel PI, Erb TJ (2019) Revolutionizing agriculture with synthetic biology. Nat Plants 5:1207–1210. https://doi.org/10.1038/s41477-019-0539-0
Xu RF, Li H, Qin RY, Li J, Qiu CH, Yang YC, Ma H, Li L, Wei PC, Yang JB (2015) Generation of inheritable and “transgene clean” targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Sci Rep 5:11491. https://doi.org/10.1038/srep11491
Xu R, Qin R, Li H, Li D, Li L, Wei P, Yang J (2017) Generation of targeted mutant rice using CRISPR-Cpf1 system. Plant Biotechnol J 15:713–717. https://doi.org/10.1111/pbi.12669
Xu Z, Xu X, Gong Q, Li Z, Li Y, Wang S, Yang Y, Ma W, Liu L, Zhu B, Zou L, Chen G (2019) Engineering broad-spectrum bacterial blight resistance by simultaneously disrupting variable TALE-binding elements of multiple susceptibility genes in rice. Mol Plant 12:1434–1446. https://doi.org/10.1016/j.molp.2019.08.006
Xu Y, Lin Q, Li X, Wang F, Chen Z, Wang J, Li W, Fan F, Tao Y, Jiang Y, Wei X, Zhang R, Zhu Q-H, Bu Q, Yang J, Gao C (2020) Fine-tuning the amylose content of rice by precise base-editing of the Wx gene. Plant Biotechnol J 19:11–13. https://doi.org/10.1111/pbi.13433
Xu Z, Kuang Y, Ren B, Yan D, Yan F, Spetz C, Sun W, Wang G, Zhou X, Zhou H (2021) SpRY greatly expands the genome editing scope in rice with highly flexible PAM recognition. Genome Biol 22:6. https://doi.org/10.1186/s13059-020-02231-9
Yan D, Ren B, Liu L, Yan F, Zhou H (2021) High-efficiency and multiplex adenine base editing in plants using new tada variants. Mol Plant. https://doi.org/10.1016/j.molp.2021.02.007
Yang Q, Zhong X, Li Q, Lan J, Tang H et al (2020) Mutation of the d-hordein gene by RNA-guided Cas9 targeted editing reducing the grain size and changing grain compositions in barley. Food Chem 311:125892. https://doi.org/10.1016/j.foodchem.2019.125892
Yoon YZ, Venkatesh J, Lee JH, Kim J, Lee HE, Kim DS, Kang BC (2020) Genome editing of eIF4E1 in tomato confers resistance to pepper mottle virus. Front Plant Sci 11:1098. https://doi.org/10.3398/fpls.2020.01098
Yu H, Lin T, Meng X, Du H, Zhang J et al (2021) A route to de novo domestication of wild allotetraploid rice. Cell 184(5):1156–1170. https://doi.org/10.1016/j.cell.2021.01.013
Yuste-Lisbona FJ, Fernandez-Lozano A, Pineda B, Bretones S, Ortíz-Atienza A et al (2020) ENO regulates tomato fruit size through the floral meristem development network. PNAS 117(14):8187–8195. https://doi.org/10.1073/pnas.1913688117
Zeng Y, Wen J, Zhao W, Wang Q, Huang W (2019) Rational improvement of rice yield and cold tolerance by editing the three genes OsPIN5b, GS3, and OsMYB30 with the CRISPR-Cas9 system. Front Plant Sci 10:1663. https://doi.org/10.3389/fpls.2019.01663
Zeng D, Liu T, Ma X, Wang B, Zheng Z, Zhang Y, Xie X, Yang B, Zhao Z, Zhu Q, Liu Y-G (2020) Quantitative regulation of Waxy expression by CRISPR/Cas9-based promoter and 5’UTR-intron editing improvesgrain quality in rice. Plant Biotechnol J 18:2385–2387. https://doi.org/10.1111/pbi.13427
Zhai Y, Yu K, Cai S, Hu L, Amoo O et al (2020) Targeted mutagenesis of BnTT8 homologs controls yellow seed coat development for effective oil production in Brassica napus L. Plant Biotechnol J 18:1153–1168. https://doi.org/10.1111/pbi.13281
Zhan X, Zhang F, Zhong Z, Chen R, Wang Y, Chang L, Bock R, Nie B, Zhang J (2019) Generation of virus-resistant potato plants by RNA genome targeting. Plant Biotechnol J 17:1814–1822. https://doi.org/10.1111/pbi.13102
Zhang Y, Zhang F, Li X, Baller JA, Qi Y, Starker CG, Bogdanove AJ, Voytas DF (2013) Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiol 161:20–27. https://doi.org/10.1104/pp.112.205179
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z, Mao Y, Yang L, Zhang H, Xu N, Zhu JK (2014) The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol J 12:797–807. https://doi.org/10.1111/pbi.12200
Zhang M, Cao Y, Wang Z, Wang ZQ, Shi J, Liang X, Song W, Chen Q, Lai J, Jiang C (2018a) A retrotransposon in an HKT1 family sodium transporter causesvariation of leaf Na+exclusion and salt tolerance in maize. New Phytol 217:1161–1176. https://doi.org/10.1111/nph.14882
Zhang Y, Li D, Zhang D, Zhao X, Cao X, Dong L, Liu J, Chen K, Zhang H, Gao C, Wang D (2018b) Analysis of the functions of TaGW2 homoeologs in wheat grain weight and protein content traits. Plant J 94:857–866. https://doi.org/10.1111/tpj.13903
Zhang A, Liu Y, Wang F, Li T, Chen Z, Kong D, Bi J, Zhang F, Luo X, Wang J, Tang J, Yu X, Liu G, Luo L (2019a) Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol Breed 39:47. https://doi.org/10.1007/s11032-019-0954-y
Zhang Z, Hua L, Gupta A, Tricoli A, Edwards KJ, Yang B, Li W (2019b) Development of an agrobacterium delivered CRISPR/Cas9 system for wheat genome editing. Plant Biotechnol j 17:1623–1635. https://doi.org/10.1111/pbi.13088
Zhang M, Liu Q, Yang X, Xu J, Liu G, Yao X, Ren R, Xu J, Lou L (2020a) CRISPR/Cas9-mediated mutagenesis of Clpsk1 in watermelon to confer resistance to Fusarium oxysporum f. sp. niveum. Plant Cell Rep 39:589–595. https://doi.org/10.1007/s00299-020-02516-0
Zhang Y, Pribil M, Palmgren M, Gao C (2020b) A CRISPR way for accelerating improvement of food crops. Nat Food 1:200–205. https://doi.org/10.1038/s43016-020-0051-8
Zhong Y, Blennow A, Kofoed-Enevoldsen O, Jiang D, Hebelstrup KH (2019a) Protein Targeting to Starch 1 is essential for starchy endosperm development in barley. J Exp Bot 70(2):485–496. https://doi.org/10.1093/jxb/ery398
Zhong Y, Liu C, Qi X, Jiao Y, Wang D, Wang Y, Liu Z, Chen C, Chen B, Tian X, Li J, Chen M, Dong X, Xu X, Li L, Li W, Liu W, Jin W, Lai J, Chen S (2019b) Mutation of ZmDMP enhances haploid induction in maize. Nat Plant 5:575–580. https://doi.org/10.1038/s41477-019-0443-7
Zhou H, Liu B, Weeks DP, Spalding MH, Yang B (2014) Large chromosomal deletions and heritable small genetic changes induced by CRISPR/Cas9 in rice. Nucleic Acids Res 42:10903–10914. https://doi.org/10.1093/nar/gku806
Zhou J, Xin X, He Y, Chen H, Li Q, Tang X, Zhong Z, Deng K, Zheng X, Akher SA, Cai G, Qi Y, Zhang Y (2019) Multiplex QTL editing of grain-related genes improves yield in elite rice varieties. Plant Cell Rep 38:475–485. https://doi.org/10.1007/s00299-018-2340-3
Zhu C, Bortesi L, Baysal C, Twyman RM, Fischer S, Capell T, Schillberg S, Christou P (2017) Characteristics of genome editing mutations in cereal crops. Trends Plant Sci 22:38–52. https://doi.org/10.1016/j.tplants.2016.08.009
Zong Y, Wang Y, Li C, Zhang R, Chen K, Ran Y, Qiu JL, Wang D, Gao C (2017) Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat Biotechnol 35:438–440. https://doi.org/10.1038/nbt.3811
Zsögön A, Čermák T, Naves ER, Notini MM, Edel KH, Weinl S, Freschi L, Voytas DF, Kudla J, Peres LEP (2018) De novo domestication of wild tomato using genome editing. Nat Biotechnol 36:1211–1216. https://doi.org/10.1038/nbt.4272
Acknowledgements
This work was partially supported by grants from the China National Transgenic Plant Special Fund (2016ZX08012-002, 2017ZX08013-001) and the Programme of Introducing Talents of Discipline to Universities (111 Project, B14016).
Author information
Authors and Affiliations
Contributions
SJ designed the project, supervised the project, and revised the manuscript. BS collected information, did analysis, and wrote the first draft of the manuscript. ZD participated in the discussion.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Editorial Responsibility: Sukhpreet Sandhu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Biswas, S., Zhang, D. & Shi, J. CRISPR/Cas systems: opportunities and challenges for crop breeding. Plant Cell Rep 40, 979–998 (2021). https://doi.org/10.1007/s00299-021-02708-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02708-2