Abstract
Key message
The transcription factor (TF) IbERF71 forms a novel complex, IbERF71-IbMYB340-IbbHLH2, to coregulate anthocyanin biosynthesis by binding to the IbANS1 promoter in purple-fleshed sweet potatoes.
Abstract
Purple-fleshed sweet potato (Ipomoea batatas L.) is very popular because of its abundant anthocyanins, which are natural pigments with multiple physiological functions. TFs involved in regulating anthocyanin biosynthesis have been identified in many plants. However, the molecular mechanism of anthocyanin biosynthesis in purple-fleshed sweet potatoes has rarely been examined. In this study, TF IbERF71 and its partners were screened by bioinformatics and RT-qPCR analysis. The results showed that the expression levels of IbERF71 and partners IbMYB340 and IbbHLH2 were higher in purple-fleshed sweet potatoes than in other colors and that the expression levels positively correlated with anthocyanin contents. Moreover, transient expression assays showed that cotransformation of IbMYB340+IbbHLH2 resulted in anthocyanin accumulation in tobacco leaves and strawberry receptacles, and additional IbERF71 significantly increased visual aspects. Furthermore, the combination of the three TFs significantly increased the expression levels of FvANS and FvGST, which are involved in anthocyanin biosynthesis and transport of strawberry receptacles. The dual-luciferase reporter system verified that cotransformation of the three TFs enhanced the transcription activity of IbANS1. In addition, yeast two-hybrid and firefly luciferase complementation assays revealed that IbMYB340 interacted with IbbHLH2 and IbERF71 but IbERF71 could not interact with IbbHLH2 in vitro. In summary, our findings provide novel evidence that IbERF71 and IbMYB340-IbbHLH2 form the regulatory complex IbERF71-IbMYB340-IbbHLH2 that coregulates anthocyanin accumulation by binding to the IbANS1 promoter in purple-fleshed sweet potatoes. Thus, the present study provides a new regulatory network of anthocyanin biosynthesis and strong insight into the color development of purple-fleshed sweet potatoes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Purple-fleshed sweet potatoes (Ipomoea batatas L.) are very popular because of their attractiveness and abundant anthocyanins. Anthocyanin is a natural pigment and has many functions related to nutrition and health (Choi et al. 2010; Kwak et al. 2019). Indeed, these compounds could act as important natural antioxidants to scavenge free radicals (Hwang et al. 2011; Zhang et al. 2013).
Anthocyanin biosynthesis is carried out by structural genes encoding a series of catalytic enzymes, including the upstream enzymes phenylalanine ammonia lyase (PAL), chalcone synthase (CHS), chalcone isomerase (CHI), and flavanone 3-hydroxylase (F3H) and downstream enzymes dihydroflavonol 4-reductase (DFR), anthocyanin synthase (ANS) and glycosyltransferase (UFGT). The anthocyanin biosynthesis pathway has been widely identified in different plants (Holton and Cornish 1995). In several plants, the process is regulated by the MYB-bHLH complex or the MYB-bHLH-WD40 complex (Gonzalez et al. 2008; Liu et al. 2013). Among them, central R2R3-MYB TFs have been identified as activators in several crops, such as AtMYB75 and AtMYB90 in Arabidopsis, MdMYB10, MdMYB1, and MdMYB110a in apple, PpMYB10.1 in peach, AN1, MYBA1, and MYB113 in potatoes, and IbMYB1 and IbMYB340 in sweet potatoes (Mano et al. 2007; Chagné et al. 2013; Liu et al. 2016; Kim et al. 2016; Wei et al. 2020). As a partner, bHLH TFs can interact with MYBs to regulate anthocyanin biosynthesis. For example, MYBA1 and MYB113 interact with bHLH TFs to regulate anthocyanin biosynthesis in potato (Liu et al. 2016), and the MrMYB1-MrbHLH1 complex promotes anthocyanin biosynthesis in Chinese bayberry (Liu et al. 2013). In addition to the MBW complex, other TFs also affect anthocyanin biosynthesis, such as PyERF3 and PyWRKW26 in pear, PpNAC1 in peach, IbNAC26 in sweet potato, and a SQUAMOSA MADS-box in bilberry (Jaakola et al. 2010; Zhou et al. 2015; Yao et al. 2017; Li et al. 2020; Wei et al. 2020). They directly or indirectly interact with the MBW complex to regulate anthocyanin biosynthesis.
AP2/ERFs play multiple roles in regulating fruit ripening, participating in secondary metabolism and tolerance to multiple abiotic stresses. AP2/ERFs positively or negatively regulate fruit ripening. SlERF2 is involved in the ethylene response in fruit ripening in tomato (Julien et al. 2006) and AdERF9 suppresses the activity of the AdXET5 promoter to regulate fruit ripening in kiwifruit (Yin et al. 2010). It has also been reported that AP2/ERFs are involved in anthocyanin biosynthesis. In Arabidopsis, mutations in AtERF6 result in anthocyanin accumulation (Sewelam et al. 2013). In apple, MdERF1B regulates anthocyanin and proanthocyanidin biosynthesis, whereas MdERF17 degrades chlorophyll, resulting in a red color (Han et al. 2008; Zhang et al. 2018). PyERF3 with PybHLH3 and PyMYB114 coregulate red-skinned pear color formation (Yao et al. 2017), and MdERF3 with MdEIN3-LIKE1 and MdMYB1 synergistically regulated ethylene synthesis and anthocyanin accumulation (An et al. 2018). Liu et al. (2015) screened 17 differentially expressed genes (DEGs) of AP2/ERF TFs by comparing purple with white potatoes via transcriptome profiling, and the results suggested that AP2/ERF TFs are related to anthocyanin biosynthesis in purple-fleshed potatoes.
Sweet potato is a hexaploid (2n = 6× = 90) and highly polymorphic plant, and its genome is very complex and resulting in the regulatory mechanism of pigment formation in sweet potatoes lags behind. Previous studies mainly focused on the metabolomics of flavonoids in sweet potatoes with different flesh colors (Wang et al. 2018), and 240 different flesh color sweet potatoes were also identified by molecular variance analysis and found that there were small but with significant differences between the white- and orange-fleshed sweet potatoes (Zhang et al. 2014). Recently, with the completion of the hexaploid sweet potato genome sequencing, the molecular mechanism of pigment formation has developed rapidly. A novel glutathione S-transferase gene involved in anthocyanin sequestration was identified in sweet potatoes (Kou et al. 2019), Yang et al. (2020) found that TF WRKY75 and fructose-bisphosphate aldolase 2 genes are involved in the regulation of anthocyanin metabolism in Sushu8 and its mutant Zhengshu20 by transcript profiling. However, it is unclear how AP2/ERFs regulate anthocyanin biosynthesis in purple-fleshed sweet potatoes.
In this study, we screened the AP2/ERF family TF IbERF71 and its partners IbMYB340 and IbbHLH2 by bioinformatics and RT-qPCR analysis. In addition, correlation between the expression level of candidate genes and anthocyanin contents was assessed in sweet potatoes of different colors. Furthermore, transient cotransformation of IbMYB340 and IbbHLH2 in tobacco leaves and strawberry receptacles resulted in visible anthocyanin accumulation, and additional IbERF71 increased anthocyanin levels. Interaction between IbMYB340 with IbbHLH2 and IbERF71 was confirmed by yeast two-hybrid (Y2H) and firefly luciferase complementation (FLC) assays. In addition, the transactivation activity of the complex IbMYB340-IbbHLH2-IbERF71 to the IbANS1 promoter was tested by a dual-luciferase reporter system. The results provide new insight into the regulatory network of anthocyanin biosynthesis in purple-fleshed sweet potato roots and demonstrate the interaction of different TFs in regulating anthocyanin biosynthesis.
Materials and methods
Plant materials
Sweet potatoes including ‘Hanzi’, ‘Xuzi No. 8’, ‘Zhezi No. 3’, ‘Zhezi No. 4’, ‘Guangshu No. 87’ and ‘Xushu No. 18’ were provided by the National Sweet Potato Improvement Center (Xuzhou, Jiangsu Province, China). Storage roots were harvested approximately 120 days after planting in the field in October 2018. Nine uninjured roots were selected from each sweet potato cultivar and randomly divided into three groups, each of which was quickly chopped and frozen in liquid nitrogen and then stored at – 80 °C for later anthocyanin content measurement and RNA extraction.
Nicotiana tabacum was grown in the glasshouse of Hefei University of Technology (Hefei, Anhui province, China), and the temperature was controlled at 25 °C under natural light (daylight, 16 h). When the plant grew six leaves and the young leaves were more than 1 cm long, they were used for transient transformation experiments, FLC assays and dual-luciferase reporter assays.
Diploid strawberry (Fragaria vesca) ‘Yellow Wonder’ 5AF7 (YW5AF7) is the seventh generation inbred line of the F. vesca variety Yellow Wonder. The yellow–white fruit is the phenotype of the mutant of transcription factor FvMYB10, it was planted in an environment-controlled glasshouse, where the temperature was controlled at 23 °C under natural light; daylight was extended to 14 h (Slovin et al. 2009; Hawkins et al. 2016). Receptacles at 2 to 3 weeks after flowering were used for transient transformation experiments.
Sequence alignment
To screen candidate AP2/ERF TFs involved in anthocyanin biosynthesis in sweet potato, 264 amino acid sequences obtained by BLASTP searches using AP2/ERF proteins from A. thaliana as queries were matched as candidate AP2/ERF genes across the entire sweet potato genome (https://sweetpotato.plantbiology.msu.edu/index.shtml). Furthermore, a phylogenetic tree was constructed from the amino acid sequences of 264 AP2/ERFs and some AP2/ERF TFs from other species known to regulate anthocyanin biosynthesis using MEGA 7.0, including PyERF3 (ASY06613.1), PyERF27 (XP_009371279.1) and PyERF73 (XP_009378746.1) in pear (Yao et al. 2017) and MdERF17 (AVP27531.1) and MdERF1B (XP_008341120.2) in apple. All protein sequences are listed in Table S1. The reliability of the phylogenetic tree was evaluated by the bootstrap test with 1000 replicates. Five AP2/ERFs related to anthocyanin synthesis and 9 candidate AP2/ERF amino acid sequences were aligned using ClustalW and DNAMAN.
Total RNA extraction and RT-qPCR analysis
Samples of sweet potato roots were lyophilized, and approximately 0.1 g of every fully mixed sample was used to extract total RNA with a Plant Total RNA Isolation Kit Plus (Foregene). First-strand cDNA synthesis was performed using Prime ScriptTM RT Master Mix (Takara). For RT-qPCR analysis, SYBR® Premix ExTaq™ II (Takara) was used according to the manufacturer’s instructions. The RT-qPCR program was 95 °C for 20 s, followed by 35 cycles of 95 °C for 15 s, 55 °C for 25 s, and 72 °C for 15 s. The primers used are listed in Table S2. Transcript data were analyzed using the 2−ΔΔCt method (Livak et al. 2001). All analyses were repeated three times as biological replicates for error bars.
Measurement of anthocyanins in sweet potato, tobacco leaves and strawberry receptacles
The method of extracting anthocyanins was described by Yao et al. (2017). In summary, sample of 0.5 g (strawberry receptacles) or approximately 0.2 g (tobacco leaves) or 1 g (sweet potato roots) was ground and added to 0.5% hydrochloric acid in methanol at 4 °C for 24 h. Supernatants were collected at 13,000×g for 10 min, and the absorbance of the solution was measured at 530, 620 and 650 nm using a spectrophotometer (UV-1800; MAPADA). The results were processed according to the following formula: optical density (OD) = (A530 – A620) – [0.1 × (A650 – A620)]. Three independent samples as three biological replicates were processed for each of the sweet potato cultivars.
Transient expression assays in tobacco and strawberry receptacles
The full-length CDS of IbMYB340 (MN602041), IbbHLH2 (JQ337863.1) and IbERF71 (MN602040) was isolated and then inserted into the pSAK277 vector under the control of the 35S promoter using EcoRI and XbaI sites (Hellen et al. 2005). The constructs were transformed into Agrobacterium tumefaciens strain GV3101 via transient transformation according to Yao et al. (2017). Approximately, 300 μl of the above agrobacterium mixture was injected into tobacco leaves or strawberry receptacles, and samples were collected approximately 1 week after transformation to measure anthocyanin contents and extract RNA. The primers used for cloning are listed in Table S2.
Dual-luciferase reporter assays
The 1228-bp IbANS1 promoter sequence was amplified and inserted into the pGreenII 0800-LUC vector. The constructs were subsequently transformed into A. tumefaciens strain GV3101 (pMP90) with the helper vector pSoup by the freeze–thaw method, and cells were plated on Lennox agar with 100 μg ml-1 rifampicin and 50 μg ml-1 kanamycin at 28 ℃ for 48 h. A 10-μl loop sample of confluent bacteria at OD600 of 1.0–1.2 was added to 10 ml of infiltration buffer [10 mM MES, 10 mM MgCl2 and 80 μM acetosyringone, pH 5.5] followed by infiltration after incubating at 25 ℃ with shaking for 3 h. The infiltration was performed with the IbANS1 promoter reconstructs and the above recombinant TFs IbMYB340, IbERF71 and IbbHLH2 at a ratio of 1:9 (Voinnet et al. 2003). The tobacco (N. benthamiana) leaves were injected and then placed in an environment-controlled glasshouse at 25 ℃ for 3 days. A Dual-Luciferase Reporter Assay System (E1910, Promega) was used to assess LUC and Renilla luciferase activities.
Y2H assay
The CDS sequence of IbMYB340 was isolated and inserted into the pGBKT7 vector; the CDS sequences of IbERF71 and IbbHLH2 were isolated and inserted into the pGADT7 vector (Clontech). For the Y2H assay, IbMYB340 was transformed with IbERF71 or IbbHLH2 into the Y2HGold strain using the LiCl-PEG method according to the manufacturer’s manual (Clontech). SD/–Leu/–Trp medium was used to select transformants, and SD/–Leu/–Trp/–His/–Ade medium was used to examine interactions. The cloning vector pGADT7-T with pGBKT7-Lam or pGBKT7-53 was cotransformed as a negative or positive control, respectively.
FLC assay
FLC assays were performed according to a previous report (Zhou et al. 2018). The full-length coding sequence of IbMYB340 (without a stop codon) was cloned and inserted into the binary vector pCAMBIA1300-nLuc; the full-length coding sequences of IbERF71 and IbbHLH2 were isolated and inserted into the binary vector pCAMBIA1300-cLuc. Agrobacterium harboring nLuc or cLuc was cultivated and infiltrated using the same protocol as described above for the dual-luciferase reporter assay. Luciferase activity at the infiltration site was measured using a Steady-Glo Luciferase Assay System (Promega).
Statistical analysis
SPSS version 18.0 was used for data analysis. One-way analysis of variance (ANOVA) followed by pairwise comparisons was performed with Tukey’s honestly significant difference (HSD) post hoc tests, with the significance level at P< 0.05 or P< 0.01.
Results
Screening of IbERF71, IbbHLH2 and IbMYB340 as anthocyanin-related TFs in purple-fleshed sweet potato
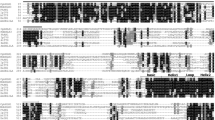
To screen candidate AP2/ERF genes regulating anthocyanin biosynthesis in sweet potato, a genome-wide phylogenetic tree was constructed. The results revealed evolutionary relationships between the 264 AP2/ERF TFs in sweet potato and some anthocyanin-related AP2/ERF genes from different species, such as MdERF1B, MdERF17, PyERF3, PyERF73, and PyERF27 (Han et al. 2008; Yao et al. 2017; Zhang et al. 2018) (Fig. 1a). Among them, MdERF1B and itf04 g07800.t1 clustered together on a branch; MdERF17 with itf06 g05430.t1, itf01 g14500.t1, and itf15 g20250.t1 also grouped together on a branch, as did PyERF3 with itf06 g25270.t1, PyERF73 and itf14 g19640.t1 (named as IbERF71), PyERF27 with itf03 g12870.t1, itf08 g00420.t1, and itf13 g03580.t1. Therefore, these genes were considered candidate genes for regulating purple-fleshed anthocyanin biosynthesis. Alignment analysis of nine AP2/ERF amino acid sequences revealed that all genes contained an AP2/ERF domain. The 14th and 19th amino acids of the AP2 domain of IbERF71 (itf14 g19640.t1), itf04 g07800.t1 and itf06 g25270.t1 are alanine and aspartic acid residues (Fig. 1b). Hence, they belong to the ERF family, as described by Sakuma (Sakuma et al. 2002).
Bioinformatics analysis of AP2/ERF TFs in different species and the transcript abundance of candidate AP2/ERFs in different sweet potato roots. a Phylogenetic tree of 264 AP2/ERFs in sweet potato and other AP2/ERFs related to the synthesis of anthocyanins from other species. b Protein sequence alignment of 14 candidate AP2/ERF TFs from the phylogenetic tree. The accession numbers are as follows: IbERF71, MN602040; PyERF3, ASY06613.1; PyERF27, XP_009371279.1; PyERF73, XP_009378746.1; MdERF17, AVP27531.1; and MdERF1B, XP_008341120.2. The black frame is the AP2/ERF domain, the blue highlighting represents conserved amino acid residues in all the AP2/ERFs, and the orange represents conserved amino acid residues in most of the AP2/ERF domain
To investigate the potential relationship of AP2/ERF TFs with anthocyanin biosynthesis, we examined the total anthocyanin contents and expression levels of ERFs in six sweet potato roots with different colors. The results showed the higher total anthocyanin contents in ‘Hanzi’ and ‘Xuzi No.8’ (Fig. 2a). Moreover, the correlation between anthocyanin content and the expression levels of the candidate genes in different color sweet potato cultivars, the result showed that itf14 g19640.t1 (IbERF71) and itf06 g05430.t1 were considerably positive correlation with anthocyanin contents, and the correlation coefficients were 0.9537 and 0.8686 in Fig. 2b. In addition, the pheatmap analysis showed that fold difference of transcript abundance of the candidate genes itf14 g19640.t1 (IbERF71) in purple-/yellow-fleshed sweet potatoes was significantly higher than other color sweet potatoes in Fig. 2c. Thus, IbERF71 is as a candidate gene for regulating anthocyanin biosynthesis. To screen partner genes of coregulating anthocyanin biosynthesis in sweet potato, AmMYB340 (P81396) in Antirrhinum (another Convolvulaceae plant) and AtTT8 (NM_117050.3) in A. thaliana were queried via BLASTP against the sweet potato genomic database (https://sweetpotato.plantbiology.msu.edu/ index.shtml), and ten highly homologous genes were selected to analyze expression levels by RT-qPCR. For ten homologous MYB genes, itf14 g19640.t1 (IbMYB340) displayed the highest transcript abundance (P<0.01) (Fig. 2d), and four bHLHs (itf14 g18730.t1, itf14 g18730.t3, itf2 g16510.t1 and JQ337863.1) had relatively higher transcript abundance than other bHLHs in purple-fleshed sweet potato ‘Hanzi’ (P<0.01) (Fig. 2e). Recently, JQ337863.1 (named IbbHLH2) was also reported to regulate anthocyanin biosynthesis in purple-fleshed sweet potatoes (Wei et al. 2020). Thus, IbMYB340 and IbbHLH2 were further studied as candidate partners of IbERF71.
Measurement of total anthocyanin content and screening of candidate TFs in different sweet potato roots. a Total anthocyanin contents were measured in six different sweet potato cultivars (‘Hanzi’, ‘Xuzi No. 8’, ‘Zhezi No. 3’, ‘Zhezi No. 4’, ‘Guangshu No. 87’, and ‘Xushu No. 18’). b The correlation analysis between anthocyanin contents and the transcript abundance of nine candidate AP2/ERF genes in sweet potatoes. c The pheatmap analysis of fold difference of the transcript abundance of the candidate AP2/ERF TFs in different color sweet potatoes. d Relative expression levels of ten candidate IbMYB TFs in the purple-fleshed sweet potato ‘Hanzi’. e Relative expression levels of ten candidate IbbHLH TFs in the purple-fleshed sweet potato ‘Hanzi’. The error bars show the means (±SD) of three biological replicates. The lowercase letters represent significant differences at P < 0.05; the capital letters represent highly significant differences at P < 0.01
Heterologous overexpression of IbERF71, IbMYB340 and IbbHLH2 induces anthocyanin accumulation in tobacco leaves and strawberry receptacles
To explore the function of IbERF71 and its partners IbMYB340 and IbbHLH2 in anthocyanin biosynthesis of purple-fleshed sweet potatoes, the sequences were transformed into tobacco leaves. No anthocyanin was observed when transforming the empty vector (pSAK277) or IbERF71 or IbbHLH2 alone. However, when transforming IbMYB340 alone, some pigmentation was visible at the injection site of tobacco leaves; when TF IbbHLH2 or IbERF71 was cotransformed with IbMYB340, significant pigmentation was observed. Moreover, we noticed deeper pigmentation after the addition of IbERF71 with IbMYB340+IbbHLH2 (Fig. 3a). Furthermore, anthocyanin content analysis indicated that cotransformation of IbERF71+IbMYB340+IbbHLH2 resulted in higher anthocyanin contents than when cotransforming IbMYB340 with IbbHLH2 or IbERF71, and the ratio of a*/b* was similar (P<0.01) (Fig. 3b, c).
Transient expression of IbERF71 and other TFs in tobacco leaves. a Phenotypic changes at 5 days after infiltration in tobacco leaves. (a) pSAK277; (b) IbbHLH2; (c) IbERF71; (d) IbMYB340; (e) IbERF71+IbbHLH2; (f) IbMYB340+IbbHLH2; (g) IbMYB340+IbERF71; (h) IbMYB340+IbbHLH2+IbERF71; b Colored regions were measured using a Minolta Chroma Meter, and the results are indicated as the a*/b* ratio. Among them, “a” means red-green, and an a* value from negative to positive means the color changes from greenish to reddish; “b” means yellow-blue, and b* value from negative to positive means the color changes from blueish to yellowish. The value of a*/b* changed from negative to positive, indicating that the tobacco leaf color changed from green to red. The statistical significance means six color regions were randomly selected around six injection sites to evaluate anthocyanin accumulation, and the average value was used as the ratio of a*/b*. c The total anthocyanin content was measured in tobacco leaves with induced coloration. The error bars show the means (±SDs) of three biological replicates. Significant differences are indicated by lowercase letters, with P < 0.05; significant differences are indicated by capital letters, with P < 0.01
To further verify the function of IbERF71 and other TFs, a transient expression assay was performed in strawberry receptacles, and the observed changes in appearance were similar to those in tobacco leaves. No pigmentation was detected with the control (pSAK277) or IbERF71 or IbbHLH2 alone (Fig. 4a). Pink pigmentation was observed when IbMYB340 was transformed alone, and the anthocyanin content reached 1.048 mg g-1. The strawberry receptacles became redder in color when IbMYB340 was cotransformed with IbbHLH2, and the addition of IbERF71 with IbMYB340+IbbHLH2 resulted in crimson-colored strawberry receptacles, with the pigment content reaching 3.805 mg g-1. Meanwhile, the ratio of a*/b* was higher in the three cotransformed-TF group than in the other groups (P<0.01), and the total anthocyanin content coincided with the above observations (Fig. 4b, c). These results are consistent with the tobacco leaf phenotype.
Transient expression of IbERF71 and other TFs to observe the appearance in strawberry receptacles at 5 days after injection. a The phenotype of strawberry receptacles at 5 days after infiltration. (a) pSAK277; (b) IbbHLH2; (c) IbERF71; (d) IbMYB340; (e) IbERF71+IbbHLH2; (f) IbMYB340+IbbHLH2; (g) IbMYB340+IbERF71; (h) IbMYB340+IbbHLH2+IbERF71; b Colored regions were measured using a Minolta Chroma Meter, and the results are indicated as the ratio of a*/b*. The value of a*/b* changed from negative to positive, indicating strawberry receptacle color changes from green to red. c Quantification of anthocyanin contents in the samples shown in a–h. d Expression levels of anthocyanin biosynthesis-related genes were analyzed in strawberry receptacles. RT-qPCR was used to analyze the expression patterns of FvPAL, FvCHS, FvCHI, FvF3H, FvDFR, FvANS, FvUFGT, FvGST and FvMYB10 in Yellow Wonder ‘5AF7’ strawberry receptacles. The error bars show the means (±SDs) of three biological replicates. Significant differences are indicated by lowercase letters, with P < 0.05, and capital letters, with P < 0.01
Expression analysis of anthocyanin biosynthesis-related genes in strawberry receptacles
Expression levels of anthocyanin biosynthesis-related genes in strawberry receptacles were analyzed, and the results revealed changes in FvPAL, FvCHS, FvCHI, FvF3H, FvDFR, FvANS, FvUFGT, FvGST and FvMYB10 (Fig. 4d). When IbERF71 was cotransformed with IbMYB340 and IbbHLH2, the expression levels of FvPAL, FvCHS, FvCHI, FvF3H, FvDFR, FvANS, FvUFGT and FvGST were higher to varying degrees than that cotransformed with IbMYB340+IbbHLH2 or IbMYB340 alone. For FvANS in particular, the expression level of the three cotransformed-TF group was three times higher than that of the IbMYB340+IbbHLH2 group (Fig. 4d). In contrast, the expression level of FvMYB10 was not markedly different when the three TFs were cotransformed compared with two TFs or IbMYB340 alone, and the expression level was much higher than that of the empty vector. Therefore, it is suggested that exogenous genes rarely activate the expression of the endogenous gene FvMYB10 in strawberry receptacles.
Verification of the interaction of IbERF71 with other TFs in vivo
To assess the interaction between IbERF71 and IbMYB340 or IbbHLH2, a Y2H assay was performed by cloning the full-length CDS sequence of IbMYB340 into pGBKT7 and the full-length CDS sequences of IbERF71 and IbbHLH2 into pGADT7 to examine interaction with IbMYB340. When IbMYB340 was cotransformed with IbbHLH2, growth on SD/–Leu/–Trp and SD/–Leu/–Trp/–His/–Ade media occurred. Thus, IbMYB340 could interact with IbbHLH2. Cotransformation of IbERF71 and IbMYB340 generated the same results. However, IbERF71 cotransformed with IbbHLH2 only grew on SD/–Leu/–Trp medium, whereas it could not grow on SD/–Leu/–Trp/–His/–Ade medium. Thus, IbERF71 could not interact with IbbHLH2 in vivo (Fig. 5a).
Interaction of IbMYB340 with IbERF71 or IbbHLH2 in vivo. a Verification of the interaction of IbMYB340 and IbERF71 or IbbHLH2 in vivo by a Y2H assay. b Schematic diagram of NLuc, CLuc and NLuc/CLuc constructs. c FLC assay in young Nicotiana tabacum leaves. The error bars show the means (±SD) of three biological replicates. ** indicates significant differences, with P < 0.01
To validate these results, we performed a FLC assay. IbMYB340 was inserted into the N-terminal region of firefly luciferase (NLuc), and IbbHLH2 and IbERF71 were inserted into the C-terminal region of firefly luciferase (CLuc). Cotransformation of IbMYB340-NLuc with IbERF71-CLuc and IbMYB340-nLuc with IbbHLH2-cLuc resulted in the highest level of luciferase enzyme activity. Conversely, no obvious changes were detected for coexpression of IbERF71-CLuc with IbbHLH2-NLuc, IbMYB340-nLuc with CLuc, IbERF71-CLuc with NLuc, and IbERF71-NLuc with CLuc. Thus, IbMYB340 is able to interact with IbERF71 and IbbHLH2, consistent with the Y2H assay.
The complex IbERF71-IbMYB340-IbbHLH2 regulates anthocyanin biosynthesis by binding to the IbANS1 promoter
To further explore the regulation mode of the complex IbERF71-IbMYB340-IbbHLH2 with regard to anthocyanin biosynthesis-related genes, we analyzed the expression level of three IbANS genes (IbANS1, itf13 g04110.t1; IbANS2, itf13 g15560.t1; IbANS3, itf09 g09340.t1), which were homology aligned with FvANS (gene32347) by blastp searches. The expression level of IbANS1 was highest among the three IbANS genes in purple-fleshed sweet potatoes ‘Hanzi’ and ‘Xuzi No. 8’ (P<0.05), but expression of the three IbANS genes was not different in yellow-/white-fleshed sweet potatoes (Fig. 6a). Moreover, transactivation activity at the IbANS1 promoter was analyzed using a dual-luciferase reporter assay in tobacco leaves. Therefore, IbMYB340+IbbHLH2 activated the IbANS1 promoter compared with the empty vector, and the presence of IbERF71+IbMYB340+IbbHLH2 largely increased the activation effect (Fig. 6b). These results provide further evidence that IbERF71 and its partners coregulate anthocyanin accumulation by binding to the IbANS1 promoter to increase pigment accumulation in purple-fleshed sweet potatoes.
Identification of IbERF71, IbMYB340 and IbbHLH2 transactivation activity of the IbANS promoter by a dual-luciferase assay. a Relative expression levels of IbANS1(itf13 g04110.t1), IbANS2 (itf13 g15560.t1) and IbANS3 (itf09 g09340.t1) in different sweet potato cultivars. * indicates significant differences, with P < 0.05. b The promoter activity of anthocyanin biosynthesis pathway genes is expressed as the ratio of LUC to REN activities. The error bars show the means (±SD) of three biological replicates. Significant differences are indicated by small letters, with P < 0.05, and capital letters, with P < 0.01
Discussion
IbERF71 interacts with other TFs to coordinately regulate anthocyanin biosynthesis in purple-fleshed sweet potatoes
AP2/ERF TFs play a pivotal role in plant growth, fruit ripening, and biological or abiotic stress responses. In potato, MYB113 interacts with bHLH TFs to induce the synthesis of anthocyanins (Liu et al. 2016). In pear, the PyERF3-PyMYB114-PybHLH3 complex coordinately regulates the synthesis of anthocyanins (Yao et al. 2017). Recently, it was reported that the interaction of Pp4ERF24/Pp12ERF96 with PpMYB114 modulates blue light-induced anthocyanin biosynthesis in pear (Ni et al. 2019). In this study, nine candidate AP2/ERFs in sweet potato were screened by bioinformatics analysis (Fig. 1a), and analysis of the expression pattern suggested that expression level of IbERF71 was consistent with anthocyanin contents (Fig. 2b). However, some other ERF genes (such as itf08 g00420.t1, itf13 g03580.t1 and itf06 g25270.t1) have expression levels that correlate more in some of purple cultivars, suggesting variation among family member regulation depending on cultivars. One of the reasons is that sweet potato cultivars exhibit genetic diversity, and the regulatory mechanisms need further study. When IbERF71 with IbMYB340 and IbbHLH2 was cotransformed, the color of the tobacco leaves and strawberry receptacles was deeper than that when IbMYB340 and IbbHLH2 were cotransformed (Figs. 3a, 4a). Therefore, IbERF71, as a partner with IbMYB340 and IbbHLH2, coregulates anthocyanin accumulation in purple-fleshed sweet potato.
Effect of endogenous gene expression on the IbERF71-IbMYB340-IbbHLH2 complex
In this study, the expression level of FvMYB10 in strawberry receptacles was analyzed by RT-qPCR (Fig. 4d), and the results demonstrated that the level was not significantly different among groups, including IbMYB340, IbMYB340+IbbHLH2, IbMYB340+IbERF71 or IbMYB340+IbbHLH2+IbERF71. This result suggests that the endogenous gene FvMYB10 was not affected by cotransformation of the exogenous factors IbMYB340+IbbHLH2+IbERF71; however, the IbbHLH2+IbERF71 and empty vector pSAK277 groups were slightly affected by expression of exogenous FvMYB10 (Singer et al. 2012; Zhang et al. 2020). This finding provides new evidence that endogenous genes do not interfere with the expression of the regulatory complex IbERF71-IbMYB340-IbbHLH2 in strawberry receptacles.
The IbERF71-IbMYB340-IbbHLH2 (EMB) complex regulates anthocyanin synthesis via key structural genes
PAL, CHI and CHS are the key upstream enzymes in the metabolic pathway of anthocyanin biosynthesis. DFR, ANS and UFGT are key downstream genes in this pathway (Holton and Cornish 1995). In this study, compared with other combinations, MYB340 alone or cotransformed MYB340+bHLH2 or MYB340+bHLH2+ERF71 caused the expression levels of FvPAL, FvDFR, FvANS and FvGST to be significantly increased, even though the expression level of FvUFGT was only slightly upregulated. Among them, the expression level of FvANS was highest. Further analysis by dual-luciferase reporter assays indicated that IbMYB340 activated expression of IbANS1 and that the combined presence of IbERF71-IbMYB340-IbbHLH2 increased the effect (Fig. 6). These results suggest that the IbERF71-IbMYB340-IbbHLH2 complex binds to the IbANS1 promoter to regulate anthocyanin biosynthesis. In addition, glutathione S-transferase (GST), as the principal transporter of anthocyanins, can be modified to alter fruit color in strawberry fruit (Kitamura et al. 2004; Luo et al. 2018). Kou et al. (2019) reported that a novel glutathione S-transferase gene, IbGSTF4, is involved in anthocyanin sequestration in sweet potato. In this study, GST was related to anthocyanin accumulation in purple flesh sweet potatoes, which is consistent with previous reports. Accumulating evidence shows that anthocyanin biosynthesis correlates positively with DFR and UFGT activity in apple, strawberry, litchi and grape (Castellarin and Di 2007; Zhao et al. 2012). This regulatory pattern in purple-fleshed sweet potatoes differs from that in horticultural crops (Zhao et al. 2012; Yao et al. 2017). We speculate that the anthocyanin biosynthesis pathway differs between light-induced horticultural crops and nonlight-induced purple sweet potatoes. In summary, the TF IbERF71 forms a regulatory complex with IbMYB340-IbbHLH2 and binds to the promoter of IbANS1, increasing anthocyanin biosynthesis and transport in purple-fleshed sweet potato roots. Additional scientific research is needed to elucidate the detailed molecular mechanism.
References
An JP, Wang XF, Li YY, Song LQ, Zhao LL, You CX, Hao YJ (2018) EIN3-LIKE1, MYB1, and ETHYLENE RESPONSE FACTOR3 act in a regulatory loop that synergistically modulates ethylene biosynthesis and anthocyanin accumulation. Plant Physiol. 178(2):808–823
Castellarin SD, Di Gaspero G (2007) Transcriptional control of anthocyanin biosynthetic genes in extreme phenotypes for berry pigmentation of naturally occurring grapevines. BMC Plant Biol. 7:46
Chagné D, Lin-Wang K, Espley RV et al (2013) An ancient duplication of apple MYB transcription factors is responsible for novel red fruit-flesh phenotypes. Plant Physiol. 161(1):225–239
Choi JH, Hwang YP, Choi CY, Chung YC, Jeong HG (2010) Anti-fibrotic effects of the anthocyanins isolated from the purple-fleshed sweet potato on hepatic fibrosis induced by dimethylnitrosamine administration in rats. Food Chem. Toxicol. 48(11):3137–3143
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J. 53(5):814–827
Han Z, Hu Y, Lv Y, Rose JKC, Sun Y, Shen F, Wang Y, Zhang X, Xu X, Wu T, Han Z (2008) Natural variation underlies differences in ETHYLENE RESPONSE FACTOR 17 activity in fruit peel degreening. Plant Physiol. 176(3):2292–2304
Hawkins C, Caruana J, Schiksnis E, Liu Z (2016) Genome-scale DNA variant analysis and functional validation of a SNP underlying yellow fruit color in wild strawberry. Sci. Rep. 6:29017
Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, Karunairetnam S, Gleave AP, Laing WA (2005) Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1:13
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7(7):1071–1083
Hwang YP, Choi JH, Yun HJ, Han EH, Kim HG, Kim JY, Park BH, Khanal T, Choi JM, Chung YC, Jeong HG (2011) Anthocyanins from purple-fleshed sweet potato attenuate dimethylnitrosamine-induced liver injury in rats by inducing Nrf2-mediated antioxidant enzymes and reducing COX-2 and iNOS expression. Food Chem. Toxicol. 49(1):93–99
Jaakola L, Poole M, Jones MO et al (2010) A SQUAMOSA MADS box gene involved in the regulation of anthocyanin accumulation in bilberry fruits. Plant Physiol. 153(4):1619–1629
Julien P, Fabiola JM, Teresa SBM et al (2006) Sl-ERF2, a tomato ethylene response factor involved in ethylene response and seed germination. Plant Cell Physiol. 47(9):1195–1205
Kim CY, Ahn YO, Kim SH, Kim YH, Lee HS, Catanach AS, Jacobs JM, Conner AJ, Kwak SS (2010) The sweet potato IbMYB1 gene as a potential visible marker for sweet potato intragenic vector system. Physiol Plant. 139(3):229–240
Kitamura S, Shikazono N, Tanaka A (2004) TRANSPARENT TESTA 19 is involved in the accumulation of both anthocyanins and proanthocyanidins in Arabidopsis. Plant J. 37(1):104–114
Kou M, Liu YJ, Li ZY, Zhang YG, Tang W, Yan H, Wang X, Chen XG, Su ZX, Mohamed HA, Li Q, Ma DF (2019) A novel glutathione S-transferase gene from sweet potato, IbGSTF4, is involved in anthocyanin sequestration. Plant Physiol. Biochem. 135:395–403
Kwak SS (2019) Biotechnology of the sweetpotato: ensuring global food and nutrition security in the face of climate change. Plant Cell Rep. 38(11):1361–1363
Li C, Wu J, Hu KD, Wei SW, Sun HY, Hu LY, Han Z, Yao GF, Zhang H (2020) PyWRKY26 and PybHLH3 cotargeted the PyMYB114 promoter to regulate anthocyanin biosynthesis and transport in red-skinned pears. Hortic. Res. 7:37
Lin-Wang K, Micheletti D, Palmer J, Volz R, Lozano L, Espley R, Hellens RP, Chagnè D, Rowan DD, Troggio M, Iglesias I, Allan AC (2011) High temperature reduces apple fruit colour via modulation of the anthocyanin regulatory complex. Plant Cell Environ. 34(7):1176–1190
Liu XF, Yin XR, Andrew C, Wang KL, Shi YN, Huang YJ, Ian B, Xu CJ, Chen KS (2013) The role of MrbHLH1 and MrMYB1 in regulating anthocyanin biosynthetic genes in tobacco and Chinese bayberry (Myrica rubra) during anthocyanin biosynthesis. Plant Cell Tiss Org. 115(3):285–298
Liu Y, Jiang H, Zhao Y, Li X, Dai X, Zhuang J, Zhu M, Jiang X, Wang P, Gao L, Xia T (2019) Three Camellia sinensis glutathione S-transferases are involved in the storage of anthocyanins, flavonols, and proanthocyanidins. Planta 250(4):1163–1175
Liu YH, Lin-Wang K, Espley Richard V, Wang L, Yang HY, Yu B, Dare A, Varkonyi-Gasic E, Wang J, Zhang JL, Wang D, Allan AC (2016) Functional diversification of the potato R2R3 MYB anthocyanin activators AN1, MYBA1, and MYB113 and their interaction with basic helix-loop-helix cofactors. J. Exp Bot. 67(8):2159–2176
Liu Y, Lin-Wang K, Deng C, Warran B, Wang L, Yu B, Yang H, Wang J, Espley RV, Zhang J, Wang D, Allan AC (2015) Comparative transcriptome analysis of white and purple potato to identify genes involved in anthocyanin biosynthesis. Plos One 10(6):e0129148
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408
Luo H, Dai C, Li Y, Feng J, Liu Z1, Kang C, (2018) RAP codes for a GST anthocyanin transporter that is essential for the foliage and fruit coloration in strawberry. J. Exp Bot. 69(10):2595–2608
Mano H, Ogasawara F, Sato K, Higo H, Minobe Y (2007) Isolation of a regulatory gene of anthocyanin biosynthesis in tuberous roots of purple-fleshed sweet potato. Plant Physiol. 143(3):1252–1268
Medina-Puche L, Cumplido-Laso G, Amil-Ruiz F, Hoffmann T, Ring L, Rodríguez-Franco A, Caballero JL, Schwab W, Muñoz-Blanco J, Blanco-Portales R (2014) MYB10 plays a major role in the regulation of flavonoid/phenylpropanoid metabolism during ripening of Fragaria×ananassa fruits. J. Exp Bot. 65(2):401–417
Moyano E, Martínez-Garcia JF, Martin C (1996) Apparent redundancy in myb gene function provides gearing for the control of flavonoid biosynthesis in antirrhinum flowers. Plant Cell 8(9):1519–1532
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 140(2):411–432
Ni J, Bai S, Zhao Y, Qian M, Tao R, Yin L, Gao L, Teng Y (2019) Ethylene response factors Pp4ERF24 and Pp12ERF96 regulate blue light-induced anthocyanin biosynthesis in ‘RedZaosu’ pear fruits by interacting with MYB114. Plant Mol. Biol. 99(1–2):67–78
Riechmann JL, Heard J, Martin G et al (2000) Arabidopsis Transcription factors: genome-wide comparative analysis among eukaryotes. Science 290(5499):2105–2110
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-Binding Specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem. Bioph. Res. Co. 290(3):998–1009
Schaart JG, Dubos C, Romero De La Fuente I, van Houwelingen AM, de Vos RC, Jonker HH, Xu W, Routaboul JM, Lepiniec L, Bovy AG (2013) Identification and characterization of MYB-bHLH-WD40 regulatory complexes controlling proanthocyanidin biosynthesis in strawberry (Fragaria × ananassa) fruits. New Phytol. 197(2):454–467
Sewelam N, Kazan K, Thomas-Hall SR, Kidd BN, Manners JM, Schenk PM (2013) Ethylene response factor 6 is a regulator of reactive oxygen species signaling in Arabidopsis. Plos One 8(8):e70289
Slovin JP, Schmitt K, Folta KM. (2009) An inbred line of the diploid strawberry Fragaria vesca f. semperflorens for genomic and molecular genetic studies in the Rosaceae. Plant Methods 5(1):15.
Sparkes IA, Runions J, Kearns A, Hawes C (2006) Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 1(4):2019–2025
Singer SD, Liu Z, Cox KD (2012) Minimizing the unpredictability of transgene expression in plants: the role of genetic insulators. Plant Cell Rep. 31(1):13–25
Voinnet O, Rivas S, Mestre P, Baulcombe D (2003) Retracted: an enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 33:949–956
Wei ZZ, Hu KD, Zhao DL, Tang J, Huang ZQ, Jin P, Li YH, Han Z, Hu LY, Yao GF、Zhang H, (2020) MYB44 competitively inhibits the formation of the MYB340-bHLH2-NAC56 complex to regulate anthocyanin biosynthesis in purple-fleshed sweet potato. BMC Plant Biol. 20:258
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB–bHLH–WDR complexes. Trends Plant Sci. 20(3):176–185
Xu ZS, Yang QQ, Feng K, Xiong AS (2019) Changing Carrot Color: Insertions in DcMYB7 Alter the Regulation of Anthocyanin Biosynthesis and Modification. Plant Physiol. 181(1):195–207
Yang YF, Shi DY, Wang YN, Zhang L, Chen XG, Yang XP, Xiong HZ, Gehendra B, Waltram R, Dotun O, Yang GH, Shi AN (2020) Transcript profiling for regulation of sweet potato skin color in Sushu8 and its mutant Zhengshu20. Plant Physiol. Biochem. 148:1–9
Yao G, Ming M, Allan AC, Gu C, Li L, Wu X, Wang R, Chang Y, Qi K, Zhang S, Wu J (2017) Map-based cloning of the pear gene MYB114 identifies an interaction with other transcription factors to coordinately regulate fruit anthocyanin biosynthesis. Plant J 92(3):437–451
Yin XR, Allan AC, Chen KS, Ferguson IB (2010) Kiwifruit EIL and ERF Genes Involved in Regulating Fruit Ripening. Plant Physiol. 153(3):1280–1292
Zhang J, Xu H, Wang N, Jiang S, Fang H, Zhang Z, Yang G, Wang Y, Su M, Xu L, Chen X (2018) The ethylene response factor MdERF1B regulates anthocyanin and proanthocyanidin biosynthesis in apple. Plant Mol. Biol. 98(3):205–218
Zhang ZF, Lu J, Zheng YL, Wu DM, Hu B, Shan Q, Cheng W, Li MQ, Sun YY (2013) Purple-fleshed sweet potato color attenuates hepatic insulin resistance via blocking oxidative stress and endoplasmic reticulum stress in high-fat-diet-treated mice. J. Nutr. Biochem. 24(6):1008–1018
Zhang JX, Lei YY, Wang BT, Li S, Yu S, Wang Y, Li H, Liu YX, Ma Y, Dai HY, Wang JH, Zhang ZH (2020) The high-quality genome of diploid strawberry (Fragaria nilgerrensis) provides new insights into anthocyanin accumulation. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13351.
Zhang K, Wu ZD, Li YH, Zhang H, Wang LP, Zhou QL, Tang DB, Fu YF, He FF, Jiang YC, Yang H, Wang JC (2014) ISSR-based molecular characterization of an elite germplasm collection of sweet potato (Ipomoea batatas L) in China. J Integr Agric 13(11):2346–2361
Zhao QZ, Zhao SY, Xia GM (2005) Research advances on the mechanism of RNA silencing in plants. Acta Genet. Sin. 32(1):104–110
Zhao ZC, Hu GB, Hu FC, Wang HC, Yang ZY, Lai B (2012) The UDP glucose: flavonoid-3-O-glucosyltransferase (UFGT) gene regulates anthocyanin biosynthesis in litchi (Litchi chinesis Sonn) during fruit coloration. Mol Biol Rep 39(6):6409–6415
Zhou H, Lin-Wang K, Wang H, Gu C, Dare AP, Espley RV, He H, Allan AC, Han Y (2015) Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors. Plant J. 82(1):105–121
Zhou ZY, Bi GZ, Zhou JM (2018) Luciferase complementation assay for protein-protein interactions in plants. Curr Protoc Plant Biol 3:42–50
Acknowledgements
This work was supported by the Earmarked Fund for the China Agriculture Research System (CARS-10-B1), the National Key R&D Program of China (2019YFD1001303, 2019YFD1001300), the National Natural Science Foundation of China (31901993, 31970312, 31970200, 31872078, 31670278), the Natural Science Foundations of Anhui Province (1908085MC72), the Key Research and Development Program of Anhui Province (201904a06020031), the Fundamental Research Funds for the Central Universities (JZ2020YYPY0249, JZ2018HGBZ0160), National Undergraduate Training Programs for Innovation of China (No.202010359064, S202010359221). We thank Dr. Andrew C. Allan, Dr. Lin-Wang Kui and Dr. Richard Espley for the dual-vector pGreen II 0800-LUC in The New Zealand Institute for Plant & Food Research Limited, Auckland, New Zealand.
Author information
Authors and Affiliations
Contributions
N.Z.Y., K.D.H., G.F.Y. and H.Z. conceived and designed the experiments; N.Z.Y., G.F.Y., Z.Z.L., Z.D.L. and D.C. performed the experiments; C.X.Y., L.L.X., W.H., Z.Z.L. and T.J. analyzed the data; N.Z.Y. and G.F.Y. wrote the paper; K.D.H., G.F.Y. and H.Z. interpreted the data and revised the manuscript
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Neal Stewart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ning, Z., Hu, K., Zhou, Z. et al. IbERF71, with IbMYB340 and IbbHLH2, coregulates anthocyanin accumulation by binding to the IbANS1 promoter in purple-fleshed sweet potato (Ipomoea batatas L.). Plant Cell Rep 40, 157–169 (2021). https://doi.org/10.1007/s00299-020-02621-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-020-02621-0