Abstract
Key message
Overexpression of CpbHLH1 in Arabidopsis and tobacco resulted in a dramatic decrease in anthocyanin accumulation by repressing the expression of late biosynthesis genes in the flavonoid biosynthesis pathway.
Abstract
Many basic helix–loop–helix (bHLH) transcription factors (TFs) of subgroup IIIf have been characterized as anthocyanin-associated activators in higher plants, but information regarding bHLH TFs that inhibit anthocyanin accumulation remains scarce. In this study, the subgroup IIIf bHLH TF CpbHLH1 from Chimonanthus praecox (L.) was identified as a negative regulator of anthocyanin accumulation. Our results showed that overexpression of CpbHLH1 in model plant species, Arabidopsis and tobacco, resulted in a dramatic decrease in anthocyanin content, whereas the content of proanthocyanidin was little affected. Quantitative RT-PCR (qRT-PCR) assays of the structural genes in the flavonoid biosynthesis pathway revealed that CpbHLH1 inhibits anthocyanin accumulation mainly through repressing the expression of late biosynthesis genes (LBGs). Interactions between CpbHLH1 protein and AtPAP1/NtAN2 protein were detected via yeast two-hybrid (Y2H) and bimolecular fluorescence complementation (BiFC) assays. This is the first bHLH repressor of anthocyanin biosynthesis identified in dicotyledons. These results can help us better understand the anthocyanin regulatory network in plants and may provide insights into the diverse functions of bHLH proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In nature, flavonoids, carotenoids and betalains provide natural color to flowers and fruits of plants. The first two categories are widely distributed in plants, while betalains exist only in several species of the Caryophyllales (Mol et al. 1998). Flavonoids constitute one of the main products of secondary polyphenol metabolism in plants and can be divided into six categories on the basis of their molecular structure: chalcones, flavones, flavonols, flavanols, anthocyanins and procyanidins (also called condensed tannins) (Li 2014). Anthocyanins have many important functions in plants, such as coloring for flowers and fruits, protecting plants from ultraviolet damage, promoting resistance to low-temperature stress, and attracting pollinators and seed dispersers (Holton and Cornish 1995; Mol et al. 1998; Ilk et al. 2015). Besides, owing to their strong antioxidant activity and cardiovascular protective effects, anthocyanins are used as healthcare products (Tsuda 2012).
The synthesis and regulation of flavonoids have been characterized in many important species, including maize (Zea mays), petunia (Petunia hybrida), Arabidopsis thaliana and snapdragon (Cone et al. 1986; Chandler et al. 1989; Ludwig et al. 1989; Goff et al. 1992; Spelt et al. 2000; Schwinn et al. 2006; Gonzalez et al. 2008; Albert et al. 2014). Flavonoid synthesis in higher plants occurs via the same enzymatic steps consisting of a series of enzymes, including chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3ß-hydroxylase (F3H), dihydroflavonol 4-reductase (DFR) and anthocyanidin synthase (ANS). CHS, CHI and F3H are classified as early biosynthesis genes (EBGs), and DFR and ANS are categorized as late biosynthesis genes (LBGs). The EBGs in Arabidopsis are controlled by MYB11, MYB12 and MYB 111, which are R2R3-MYB TFs that promote flavonol biosynthesis with tissue specificity (Mehrtens et al. 2005; Stracke et al. 2007, 2010). Research on the regulation of LBGs has been increasing in recent years. The LBGs of flavonoid biosynthesis are mediated by a MBW complex that consists of an R2R3-MYB TF, a subgroup IIIf bHLH TF and a WD40 repeat protein. In Arabidopsis, members of the MBW transcriptional activator complex include R2R3-MYB TFs (PAP1, PAP2, MYB113 or MYB114), bHLH TFs (TT8, GL3 or EGL3) and the WD40 protein TTG1 (Gonzalez et al. 2008). In petunia, the complex mainly consists of the R2R3-MYB protein AN2, the bHLH protein AN1 and the WD40 protein AN11 (Farcy and Cornu 1979; Gerats et al. 1984; Spelt et al. 2000; Albert et al. 2014). In apple, the MBW transcriptional activator complex consists of MdMYB10, MdbHLH3 and MdTTG1 (Espley et al. 2007; An et al. 2012; Xie et al. 2012). In general, in the MBW complex, the bHLH protein can interact with the MYB protein and the WD40 protein, while the MYB protein cannot interact with the WD40 protein. The bHLH protein is considered an important component in the MBW complex, serving as a bridge between the MYB protein and the WD40 protein.
Transcriptional regulation of genes involved in anthocyanin biosynthesis is controlled by both activators and inhibitors simultaneously. MYB TFs that repress anthocyanin synthesis have recently been characterized in many plant species (Chen et al. 2019). All anthocyanin-related MYB repressors in plants share the same region that binds bHLH proteins and can be divided into two categories: R2R3-MYBs and R3-MYBs. The inhibitory effects of anthocyanin-related MYB repressors may be achieved by the alteration of the transcriptional activation ability of the MBW complex or by the direct binding of the promoters of LBGs and inhibition of their expression. To date, research on TFs that inhibit anthocyanin synthesis has focused on MYB, bHLH TFs that inhibit anthocyanin accumulation have seldom been reported. However, one bHLH TF belonging to subgroup IIIf (Heim et al. 2003) has been reported to inhibit anthocyanin biosynthesis—intensifier 1 (IN1) from maize (Burr et al. 1996). Recently, the sheepgrass protein LcbHLH92, a homolog of Arabidopsis bHLH92 belonging to subgroup IVd (Zhao et al. 2019), has been proven to be an inhibitor of anthocyanin/proanthocyanidin (PA) accumulation. To our knowledge, no other anthocyanin-related bHLH repressors have been reported.
Wintersweet [Chimonanthus praecox (L.)], a traditional Chinese flowering shrub, has been cultivated as an ornamental plant species for more than 1000 years (Zhao et al. 2007). It has yellow middle tepals and red inner tepals, making it a good material for studying the mechanism involved in pigment synthesis and regulation. Research on the mechanisms underlying the biosynthesis and regulation of floral fragrance substances, floral development, and response to cold stress in wintersweet has been performed (Xiang et al. 2010; Tian et al. 2019; Zhang et al. 2012; Sui et al. 2012). However, only a few studies on the flower pigmentation of wintersweet have been carried out. According to our previous study, CpANS1 is the key gene that contributes to the pigmentation differences between the yellow middle tepals and the red inner tepals in wintersweet (Yang et al. 2018). TFs involved in regulating anthocyanin accumulation in wintersweet have not been reported yet.
To gain insights into the regulatory mechanisms underlying pigment formation in wintersweet, we conducted a study of TFs that regulate anthocyanin accumulation, and the subgroup IIIf bHLH TF CpbHLH1 was identified as a negative regulator of anthocyanin accumulation. We generated 35S::CpbHLH1 lines of model plant species, including Arabidopsis and tobacco. The phenotypes of the transgenic plants revealed that CpbHLH1 has a negative impact on anthocyanin accumulation. We further detected the expression of genes that regulate anthocyanin synthesis via qRT-PCR. The results revealed that CpbHLH1 inhibits anthocyanin accumulation mainly by repressing LBGs involved in the flavonoid biosynthesis pathway. We also carried out a transcriptional activity analysis, and the results showed that CpbHLH1 was a transcriptional inhibitor. This is the first bHLH repressor of anthocyanin biosynthesis to be identified in dicotyledons. The present results help provide a better understanding of the complicated regulatory network of anthocyanin biosynthesis and may provide insights into the diverse functions of bHLH proteins.
Materials and methods
Plant materials and growth conditions
Chimonanthus praecox variety ‘H29’ was used in the study, which has been described by Yang et al. (2018). The flowers were collected at each developmental stage (S1–S5). The S1 samples were collected on November 10, 2018, and the subsequent samples were collected every 20 days until the S4 stage. The S5 samples were collected 10 days after the S4 stage. The young leaves, stems, and fruits were also collected for further analysis. All samples were flash frozen with liquid nitrogen and stored at – 80 °C until use.
The Arabidopsis thaliana ecotypes Col-0 and pap1-D (stock name CS3884) and Nicotiana tabacum ‘Samsun’ were used to produce genetically modified plants. The growth conditions of the Arabidopsis plants were the same as described by Zhang et al. (2019). Tobacco was planted in a greenhouse under normal sunlight. The seeds of the transgenic A. thaliana ecotype Col-0 plants and tobacco plants were screened on Murashige and Skoog (MS) (1962) salt media that contained 50 mg/L kanamycin, and the seeds of the transgenic pap1-D plants were screened on MS (1962) salt media that contained 25 mg/L hygromycin B. The seeds of all the transgenic plants were sterilized in a vacuum dryer with two beakers containing 50 mL of chlorine-containing disinfectant and 1.5 mL of hydrochloric acid for 4–6 h.
Cloning and sequence analysis of the CpbHLH1 gene in wintersweet
CpbHLH1 was obtained from a wintersweet flower transcriptome database (Yang et al. 2018). The sequences of CpbHLH1 and anthocyanin-related bHLHs from other plant species were used to assess the completeness and homology via the BLASTX algorithm. Total RNA extraction and cDNA synthesis were conducted according to the methods of Yang et al. (2018). The complete coding sequence of CpbHLH1 was cloned via PCR with Phanta® EVO Super-Fidelity DNA Polymerase (Vazyme Biotech). The complete coding sequence was confirmed via ORF Finder and the BLAST network service. The primers used to amplify CpbHLH1 are presented in Supplementary Table S1.
Construction of overexpression vectors and generation of transgenic plants
To transform wild-type Arabidopsis (Col-0) and tobacco plants, the entire coding sequence of CpbHLH1 was inserted into a modified pCAMBIA 2300S vector (Munis et al. 2010). To transform A. thaliana pap1-D plants, a pCAMBIA1302 plasmid containing CpbHLH1 was constructed. All of the overexpression vectors were then transferred into Agrobacterium tumefaciens strain GV3101 via electroporation and used to transform Arabidopsis and tobacco plants via the floral dip method (Clough and Bent 1998) and the leaf disc method (Horsch et al. 1985), respectively. PCR amplification was applied for the confirmation of the transgenic plants. All primers used in the overexpression vector construction are presented in Supplementary Table S1.
Measurement of total anthocyanin and flavonol contents
The total anthocyanin content in Arabidopsis seedlings and tobacco flowers (whose sepals were removed) was determined according to the methods of Zhang et al. (2009). The flavonol content was measured according to the methods of Luo et al. (2016); quercetin (Sigma, USA) equivalents were used to determine the flavonol content. Three independent biological replicates were made.
Quantitative RT-PCR (qRT-PCR) analysis
RNA extraction from the samples of Arabidopsis plants and the fully open flowers of tobacco plants, along with subsequent cDNA synthesis, was performed as described above. In addition, wintersweet H29 flowers at different developmental stages and different tissues of H29 were selected for gene expression analysis. qRT-PCR analysis was conducted using the method described by Yang et al. (2018). ACT2 (NM_112764) and UBI (Pandey et al. 2014) were used as housekeeping genes to analyze gene expression levels in Arabidopsis and tobacco, respectively. RPL8 (Yang et al. 2018) was selected as a housekeeping gene in wintersweet. The Primer 5 program was used to design the primers used in the qRT-PCR analysis, which are presented in Supplementary Table S2.
Stress conditions
For sucrose stress, seeds were germinated on MS medium containing 6% sucrose for 7 days, and in another treatment, seedlings were first grown on MS medium for 7 days and then transferred to MS medium containing 6% sucrose and cultured for another 2 weeks. For salt stress, seedlings were first grown on MS medium for 7 days and then transferred to MS medium containing 150 mM NaCl and cultured for another 2 weeks. For nitrogen stress, seeds were germinated on MS medium without nitrogen (MS − N media) for 7 days, and in another treatment, 14-day-old seedlings grown on MS medium were transferred to MS − N medium for 7 days.
Yeast two-hybrid (Y2H) assays
The coding sequences of NtAn2, AtPAP1, and CpbHLH1 were inserted into a pGADT7 vector (Clontech, USA) and a pGBKT7 vector (Clontech, USA), respectively. The resulting pGADT7-NtAn2/AtPAP1/CpbHLH1 and pGBKT7-NtAn1a/AtPAP1/CpbHLH1 plasmids were cotransformed into yeast strain AH109 (Clontech, USA) using the method described by Zhang et al. (2019). To detect whether the cotransformation was successful, synthetic defined media lacking leucine and tryptophan (SD/-Leu-Trp) were used. To detect whether the proteins interact, quadruple-selection synthetic defined media lacking adenine, histidine, leucine and tryptophan (SD/-Ade-His-Leu-Trp) and SD/-Ade-His-Leu-Trp media plus 25 mg/L X-α-Gal were used. All primers used in the Y2H experiments are listed in Supplementary Table S1.
CpbHLH1 protein subcellular localization analysis and bimolecular fluorescence complementation (BiFC) assays
The coding sequence of CpbHLH1 without the stop codon was amplified and recombined into a pMDC83 plasmid with a green fluorescent protein (GFP) gene controlled by the 35S promoter. The fusion constructs and the GFP control vector were then introduced into Nicotiana benthamiana via Agrobacterium strain GV3101. With respect to the BiFC assays, the complete cDNA sequences of NtAn2/AtPAP1 and CpbHLH1 were fused into pSPYNE-35S and pSPYCE-35S vectors, respectively. The coding sequences of the N and C termini of yellow fluorescent proteins (NYFP and CYFP) are present in pSPYNE-35S and pSPYCE-35S, respectively. The resulting vectors, NtAn2-NYFP, AtPAP1-NYFP, and CpbHLH1-CYFP, were introduced into A. tumefaciens GV3101. Injection of 5-week N. benthamiana leaves was conducted using the methods described by Espley et al. (2007). A nucleolar marker vector containing the Arabidopsis thaliana fibrin 2 (FIB2) gene fused to the red fluorescent protein (RFP) gene was used to locate the position of the nucleus. A confocal microscope was used to detect the GFP fluorescence 3 days after injection. All primers used in the subcellular localization analysis and BiFC assays are presented in Supplementary Table S1.
Dual-luciferase (LUC) transient expression assay
The coding sequences of full-length and truncated CpbHLH1 proteins were fused into a pBD vector as effectors; the double-reporter vector included a firefly LUC gene controlled by the 35S minipromoter and a Renilla (REN) gene controlled by the 35S promoter (Fu et al. 2017). Injection of five-week-old N. benthamiana leaves was conducted using the methods described by Espley et al. (2007). All primers used in this experiment are presented in Supplementary Table S1.
For the promoter activity analysis, the promoters of AtANS and NtAN1a were isolated from Arabidopsis (Col-0) and tobacco (N. tabacum) genomic DNA, respectively, and fused into a pGreen0800LUC dual-LUC plasmid. The coding sequences of AtPAP1, AtTT8, NtAN2 and NtAN1a were recombined into a modified pCAMBIA 2300S vector (Munis et al. 2010). The experiments were performed in N. benthamiana leaves using the methods described by Espley et al. (2009). The sequences of the AtANS and NtAN1a promoters were acquired from The Arabidopsis Information Resource (https://www.arabidopsis.org/) and the article by Bai et al. (2011), respectively. All primers used in this experiment are presented in Supplementary Table S1.
A dual-LUC assay kit (Promega) and a Promega GloMax 20/20 luminometer were used to measure the LUC and REN activities. The results are represented as the ratio of LUC to REN. Each pair was measured at least three times.
Bioinformatics and statistical analyses
MEGA version 7 was used to construct a phylogenetic tree. ProtComp version 9.0 was used to predict the subcellular localization of CpbHLH1 (https://www.softberry.com/berry.phtml?topic=protcomppl&group=help&subgroup=proloc).
The Student’s t test was performed to determine the anthocyanin content and gene expression differences between CpbHLH1 transgenic plants and controls. To analyze the gene expression results for CpbHLH1 and CpANS1 in wintersweet, the one-way ANOVA LSD test was conducted using the method described by Yang et al. (2018).
Results
CpbHLH1 is a subgroup IIIf bHLH TF that is localized in the nucleus
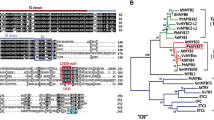
A BLAST search of all bHLH TFs obtained from the wintersweet flower transcriptome database was conducted. The BLAST results suggested that the CpbHLH1 protein might be involved in flavonoid synthesis. The open reading frame (ORF) of CpbHLH1 consists of 2025 bp encoding 674 amino acids, and the sequence from the 473rd amino acid to the 528th amino acid constitutes the specific bHLH domain. The N-terminal MYB-interacting region (MIR) (Pattanaik et al. 2008) and conserved bHLH domain were revealed by comparing the protein sequence of CpbHLH1 with the protein sequences of well-known anthocyanin-associated bHLH TFs in other species, including MdbHLH3, PhAN1, IpIVS, AtTT8, ZmIN1 and AmDELILA (Fig. 1). The MIR in subgroup IIIf bHLH proteins is critical for binding to MYB-type TFs (Pattanaik et al. 2008). We constructed a phylogenetic tree of CpbHLH1 and the Arabidopsis bHLH family (Fig. 2). The results suggested that CpbHLH1 belonged to subgroup IIIf. According to previous studies, most proteins in this subgroup are involved in the synthesis and regulation of anthocyanins.
Complete protein sequence alignment of CpbHLH1 and other subgroup IIIf proteins: Malus domestica bHLH3 (HM122458), Petunia hybrida AN1 (AAG25928), the Ipomoea purpurea Ivory seed bHLH protein (BAD18982), Arabidopsis thaliana TT8 (CAC14865), Zea mays IN1 (AAB03841) and Antirrhinum majus DELILA (AAA32663). Identical residues and conserved residues are marked in black and in dark gray, respectively. Conserved MIR and bHLH domains are labeled
Phylogenetic tree of CpbHLH1 and all bHLH TFs from Arabidopsis. All bHLH protein sequences were obtained from Plant Transcription Factor Database (https://planttfdb.cbi.pku.edu.cn/). The analysis was performed with MEGA version 7 via the neighbor-joining method, with 1000 bootstrap replicates. The box indicates subgroup IIIf
CpbHLH1 was predicted to be localized in the nucleus. Furthermore, its nuclear localization was verified by a subcellular localization assay. Agrobacterium strains containing the 35S-GFP::CpbHLH1 plasmid as well as those containing the 35S-GFP control vector were introduced into N. benthamiana leaf epidermal cells, and the fluorescence was visualized by laser confocal microscopy, revealing nuclear localization of CpbHLH1 (Fig. 3).
Expression profiling of CpbHLH1 in wintersweet
To determine whether CpbHLH1 expression was correlated with anthocyanin biosynthesis during the wintersweet flower developmental stages (Fig. 4a), the expression levels of CpbHLH1 and CpANS1 were analyzed by qRT-PCR. The expression pattern of CpbHLH1 was negatively correlated with that of CpANS1 expression. In S3, the expression level of CpbHLH1 was lowest but CpANS1 was highest (Fig. 4b, c). The expression level of CpbHLH1 in the middle (yellow) tepals was twice as high as that in the inner (red) tepals (Fig. 4d). Among different tissues of wintersweet, expression level of CpbHLH1 was highest in the leaves, followed by the stems and fruits, and was lowest in the flowers (Fig. 4e). We also measured the anthocyanin content in different tissues of wintersweet. It was found that the anthocyanin content in the flower was much higher than that in vegetative tissues (Supplementary Figure S3). These results suggested a negative correlation between the expression level of CpbHLH1 and anthocyanin biosynthesis in wintersweet.
Developmental and tissue-specific analyses of CpbHLH1 expression via qRT-PCR. a Five developmental stages of wintersweet H29 flowers. Bar, 1 cm. b–c Expression of CpbHLH1 and CpANS1 during the five wintersweet flower developmental stages. d Expression of CpbHLH1 in the middle (yellow) and inner (red) tepals of fully open wintersweet flowers. e Tissue-specific analysis of CpbHLH1 expression in wintersweet. Different letters above the columns indicate a significant difference at p < 0.05. All the data are presented as the means of three replicates, with the error bars indicating ± standard deviations (SDs)
CpbHLH1 represses anthocyanin accumulation in Arabidopsis
As genetic transformation is still very difficult in wintersweet, we generated CpbHLH1 overexpression Arabidopsis (Col-0) plants to confirm whether CpbHLH1 could regulate anthocyanin accumulation. The positive lines with the greatest expression level of CpbHLH1 were screened for the next generation, and the T3 generation was used in the study.
Under high-light (1000 lx) conditions, 35S::CpbHLH1 Arabidopsis lines exhibited phenotypes with reduced anthocyanin accumulation in young stems (Fig. 5a), and the total anthocyanin content was significantly reduced in transgenic plants compared with WT plants (Fig. 5c).
CpbHLH1 represses anthocyanin synthesis in WT (Col-0) and pap1-D mutant Arabidopsis plants. a Anthocyanin accumulation in the young stems (2–3 cm) of WT Arabidopsis plants and three 35S::CpbHLH1 Arabidopsis lines under high-light conditions. Images of approximately 1-month-old plants were collected. b Relative expression levels of CpbHLH1 in three 35S::CpbHLH1 Arabidopsis lines in the WT background. c Total anthocyanin content in the young stems (2–3 cm) of WT Arabidopsis plants and three 35S::CpbHLH1 Arabidopsis lines under high-light conditions. FW, fresh weight. d Anthocyanin accumulation in pap1-D Arabidopsis plants and three 35S::CpbHLH1 lines in the pap1-D background. Images of 24-day-old plants were collected. e Relative expression levels of CpbHLH1 in three CpbHLH1-overexpressing lines in the pap1-D background. f Total anthocyanin content in 24-day-old seedlings of pap1-D Arabidopsis plant and three 35S::CpbHLH1 lines in the pap1-D background. FW, fresh weight. The asterisks represent significant differences (**p < 0.01) between Arabidopsis control plants and the CpbHLH1 transgenic Arabidopsis lines according to the Student's t test. All the data are presented as the means of three replicates, with the error bars indicating ± SDs
To further confirm that CpbHLH1 reduced anthocyanin accumulation in Arabidopsis, we generated three CpbHLH1-overexpressing Arabidopsis lines in the pap1-D background, which represents an activation-tagged Arabidopsis line with increased anthocyanin accumulation (Borevitz et al. 2000). The phenotypes of the transgenic plants and quantitative analysis of anthocyanin levels confirmed that CpbHLH1 was a TF that negatively regulated anthocyanin accumulation (Fig. 5d, f).
A series of stress experiments were also conducted to verify the anthocyanin repressor function of CpbHLH1 in Arabidopsis. Seedlings of 35S::CpbHLH1 Arabidopsis and WT (Col-0) were grown under various stress conditions, and then the anthocyanin contents were measured. The results showed that 35S::CpbHLH1 Arabidopsis seedlings produced fewer anthocyanins than WT seedlings under sucrose (Fig. 6a, b, g), salt (Fig. 6e, g), and nitrogen stress conditions (Fig. 6c, d, g).
Anthocyanin accumulation in 35S::CpbHLH1 Arabidopsis (Col-0) lines and WT plants under various stress conditions. a and c Anthocyanin accumulation in WT (Col-0) Arabidopsis seedlings and three 35S::CpbHLH1 Arabidopsis lines in the presence of 6% sucrose and on MS−N media. Images of 7-day-old seedlings were collected. b Twenty-one-day-old seedlings subjected to sucrose stress. d Twenty-day-old seedlings subjected to nitrogen stress. e Twenty-one-day-old seedlings subjected to salt stress. f Twenty-day-old seedlings grown on control media (MS media). g Determination of anthocyanin content in 21-day-old seedlings grown on control media and subjected to sucrose stress, nitrogen stress, and salt stress. FW, fresh weight. The asterisks represent significant differences (**p < 0.01) between WT (Col-0) and 35S::CpbHLH1 Arabidopsis seedlings according to the Student’s t test. All the data are presented as the means of three replicates, with the error bars indicating ± SDs
Therefore, we considered CpbHLH1 to be a bHLH TF that negatively regulates anthocyanin accumulation in Arabidopsis.
Overexpression of CpbHLH1 inhibits anthocyanin accumulation in tobacco
To evaluate whether CpbHLH1 inhibits pigmentation in other plant species, CpbHLH1-overexpressing tobacco plants that were stably transformed were generated via the leaf disc method. Fourteen transgenic tobacco lines were produced, three of which exhibited a phenotype of white flowers (Fig. 7a). A high expression level of CpbHLH1 was confirmed in these three lines by qRT-PCR. These three lines were used to produce the T2 generation.
Overexpression of CpbHLH1 inhibits anthocyanin synthesis in tobacco. a Color alterations of the flowers of three 35S::CpbHLH1 tobacco lines compared with WT plants. b Relative expression levels of CpbHLH1 in three 35S::CpbHLH1 tobacco lines. c Total flavonol content in the petals of WT plants and three 35S::CpbHLH1 tobacco lines. FW, fresh weight. d Determination of anthocyanin content in the petals of WT plants and three 35S::CpbHLH1 tobacco lines. FW, fresh weight. The asterisks represent significant differences (*0.01 < p < 0.05; **p < 0.01) between the WT plants and 35S::CpbHLH1 tobacco lines according to the Student’s t test. All the data are presented as the means of three replicates, with the error bars indicating ± SDs
Because enhanced flavonol accumulation can cause a white-flower phenotype in tobacco plants, the flavonol extracts from flowers of 35S::CpbHLH1 tobacco T2 lines and WT plants were analyzed by HPLC. Compared with the WT tobacco plants, the CpbHLH1-overexpressing tobacco lines exhibited largely unchanged flavonol accumulation levels in their flowers (Fig. 7c). We thus excluded the possibility that the white-flower phenotype resulted from an excessive accumulation of flavonols. The anthocyanin content decreased sharply in flowers of 35S::CpbHLH1 tobacco plants but not in WT plants (Fig. 7d). Determination of the anthocyanin content in flowers confirmed that the white-flower phenotype originated from the reduction in anthocyanin content in tobacco flowers. The results revealed that CpbHLH1 inhibited pigmentation in tobacco.
CpbHLH1 inhibits the expression of regulatory and structural genes associated with anthocyanin biosynthesis in both Arabidopsis and tobacco
To investigate the means by which CpbHLH1 inhibits anthocyanin accumulation, the relative transcript levels of the structural genes responsible for anthocyanin synthesis in young stems of Arabidopsis (Col-0) exposed to high-light conditions and flowers of tobacco were measured. The expression levels of AtDFR/AtANS and NtDFR/NtANS decreased in the three 35S::CpbHLH1 Arabidopsis and tobacco lines, while no consistent change trend was noticed for the expression levels of EBGs such as CHS and CHI, among others (Fig. 8a, c). The results suggested that CpbHLH1 inhibited anthocyanin synthesis mainly by inhibiting LBGs in the flavonoid biosynthesis pathway.
Relative expression of genes involved in anthocyanin biosynthesis in 35S::CpbHLH1 Arabidopsis (Col-0) and tobacco lines via qRT-PCR. a and c Expression analysis of the structural genes in the flavonoid biosynthesis pathway in Arabidopsis (Col-0) and tobacco. b and d Expression analysis of TFs that regulate anthocyanin biosynthesis in Arabidopsis (Col-0) and tobacco. Young stems (2–3 cm) of Arabidopsis (Col-0) under high-light conditions and flowers of tobacco were used for RNA extraction. The asterisks represent significant differences (*0.01 < p < 0.05; **p < 0.01) between the WT and the 35S::CpbHLH1 transgenic plants according to the Student's t test. All the data are presented as the means of three replicates, with the error bars indicating ± SDs
We further measured the expression levels of TFs that regulate anthocyanin synthesis in Arabidopsis and tobacco. The expression levels of AtTT8 and NtAN1a/NtAN1b decreased in the three 35S::CpbHLH1 Arabidopsis and tobacco lines, respectively, while no consistent change trend was detected for other TFs such as AtPAP1 and AtTTG1 (Fig. 8b, d). These results suggested that CpbHLH1 did not affect the expression of MYB and WD40 in the MBW complex but inhibited the expression of the bHLH activator.
CpbHLH1 is a transcriptional repressor
By overexpressing CpbHLH1 from wintersweet in Arabidopsis and tobacco, we found that this protein was a TF that negatively regulates anthocyanin accumulation. To explore the transcriptional activity of CpbHLH1, we carried out dual-LUC transient expression assays in N. benthamiana. The vector containing the transcriptional activator VP16 control had a significantly greater relative LUC/REN ratio than did the pBD control vector; however, compared with that of the control vector pBD, the relative LUC/REN ratio was greatly decreased for CpbHLH1 (Fig. 9b). These results confirmed that CpbHLH1 acted as a transcriptional repressor.
Analysis of transcriptional repression ability of CpbHLH1 protein. a Schematic of the vectors used in the transient expression assay. b The transcriptional repression ability of CpbHLH1 is suggested by the relative LUC/REN ratio. pBD and pBD-VP16 were used as a negative control vector and a positive control vector, respectively. **represents significant differences (**p < 0.01) between pBD and pBD-VP16 or pBD-CpbHLH1 according to Student’s t test. c Transcriptional repression ability of CpbHLH1 truncation mutants. MIR, MYB-interacting region. Different letters above the columns indicate a significant difference at p < 0.05. All the data are presented as the means of three replicates, with the error bars indicating ± SDs
To explore the domain(s) responsible for the repressive activity of CpbHLH1, truncation mutants harboring the GAL4BD–CpbHLH1 fusion protein were generated. The results of the transient expression assays with the complete and truncated CpbHLH1 fusion protein suggested that the removal of the C-terminal region of CpbHLH1 did not influence CpbHLH1-mediated repression (Fig. 9c). Deletion of both the C-terminal and the bHLH domains of CpbHLH1 slightly relieved the CpbHLH1-mediated repression (Fig. 9c). When only the MIR remained, the suppression effect of CpbHLH1 was removed, indicating that the region between the MIR and the bHLH domain was critical for the repressive function.
CpbHLH1 interacts with NtAN2 and AtPAP1, which are positive R2R3-MYB regulators of anthocyanin biosynthesis in tobacco and Arabidopsis
The MIR motif plays a critical role in binding MYB TFs, and as it was identified in the CpbHLH1 sequence, we carried out Y2H assays to investigate the possible interaction between CpbHLH1 and MYB TFs. PAP1 from Arabidopsis and AN2 from tobacco were selected in the assays. As shown in Fig. 10, all the transformed colonies grew well on SD/-Leu-Trp media, while only the colonies cotransformed with CpbHLH1 and AtPAP1/NtAN2 survived on SD/-Ade-His-Leu-Trp media (Fig. 10). Moreover, only the colonies cotransformed with CpbHLH1 and AtPAP1/NtAN2 survived and turned blue on quadruple-selection SD media that contained 25 mg/L X-α-Gal (Fig. 10). These results suggested that CpbHLH1 could interact with both PAP1 from Arabidopsis and AN2 from tobacco.
Interactions between AtPAP1, NtAN2 and CpbHLH1 were determined by Y2H assays. All cotransformed yeast cells were screened on SD/-Leu-Trp media, and protein interactions were determined via SD/-Ade–His–Leu-Trp and SD/-Ade-His-Leu-Trp plus 25 mg/L X-α-Gal media. AD, GAL4 activation domain; BD, GAL4 DNA-binding domain
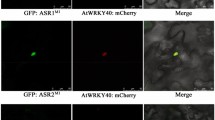
To further confirm the interaction of CpbHLH1 with AtPAP1 and NtAN2, BiFC assays were conducted. Obvious yellow fluorescence was observed in the nucleus of cells cotransformed with CpbHLH1-CYFP and either AtPAP1-NYFP or NtAN2-NYFP (Fig. 11), whereas no fluorescence was visible in the leaves injected with A. tumefaciens GV3101 harboring CpbHLH1-CYFP plus an NYFP empty vector or a CYFP empty vector plus AtPAP1-NYFP or NtAN2-NYFP, indicating that CpbHLH1 could interact with both AtPAP1 and NtAN2 in vivo.
Discussion
CpbHLH1 from wintersweet belonging to subgroup IIIf-2 is a TF that inhibits anthocyanin accumulation
bHLH TF members constitute one of the main eukaryotic TF families. In Arabidopsis, bHLH proteins participate in many biological functions, such as flavonoid biosynthesis, epidermal cell development, and responses to various stresses. All bHLH TFs in Arabidopsis can be divided into 25 subgroups according to their regions other than their bHLH-conserved domain (Heim et al. 2003). CpbHLH1 and AtGL3, AtEGL3 and AtTT8 in Arabidopsis belong to subgroup IIIf; these three proteins participate in the regulation of anthocyanin synthesis. Subgroup IIIf can be further divided into two categories (Supplementary Figure S1): the bHLH proteins AtTT8, PhAN1, and IpIVS cluster into subgroup IIIf-1, whereas CpbHLH1 clusters with AtEGL3, AtGL3, PhJAF13, and MdbHLH33 into subgroup IIIf-2. Subgroup IIIf-1 proteins promote anthocyanin as well as PA synthesis (Nesi et al. 2000; Spelt et al. 2000; Park et al. 2007). However, subgroup IIIf-2 proteins promote only anthocyanin synthesis and have no effect on PA synthesis (Ramsay et al. 2003; Ramsay and Glover 2005; Espley et al. 2007; Zhou et al. 2012). In our study, 35S::CpbHLH1 Arabidopsis and tobacco plants presented a clear reduction in anthocyanin accumulation; the seed coat color of the transgenic lines was the same as that of the WT plants, which means that CpbHLH1 did not affect PA accumulation. The result that CpbHLH1 only regulates anthocyanin accumulation is consistent with its clustering prediction.
Surprisingly, unlike other genes in subgroup IIIf-2, CpbHLH1 is an inhibitor of anthocyanin accumulation. The pattern of CpbHLH1 expression during wintersweet flower development was negatively correlated with that of CpANS1, which is the key structural gene controlling anthocyanin biosynthesis in wintersweet. The CpbHLH1 expression level in the middle (yellow) tepals of wintersweet flowers was twice of that in the inner (red) tepals. Besides, the expression level of CpbHLH1 in the flowers tissues of wintersweet was lower than that of vegetative tissues that do not synthesize anthocyanins. These results showed that the expression level of CpbHLH1 was negatively related to anthocyanin biosynthesis in wintersweet. We also measured the anthocyanin content during the flower developmental stages in wintersweet. The results revealed that the flower anthocyanin content increased continuously from S1 to S5 (Supplementary Figure S2), which might have been due to the synthesis of anthocyanin in the flower of wintersweet rather than its degradation via anthocyanin synthase activity from S1 to S5. These results indicated that the expression level of CpbHLH1 was negatively related to anthocyanin biosynthesis rather than accumulation in wintersweet.
In model species, CpbHLH1 showed a similar effect of inhibiting anthocyanin accumulation. The total anthocyanin content was greatly reduced in 35S::CpbHLH1 plants in both WT (Col-0) and pap1-D mutant backgrounds. Similar results were obtained when CpbHLH1-overexpressing Arabidopsis (Col-0) plants were exposed to various stress conditions. The CpbHLH1-overexpressing tobacco plants exhibited a white-flower phenotype due to the decreased anthocyanin content.
CpbHLH1 is the first bHLH TF found to inhibit anthocyanin synthesis in dicotyledons. In a previous study, IN1 from maize was the first bHLH TF reported to inhibit anthocyanin accumulation. Although CpbHLH1 and IN1 belong to the same subgroup IIIf, they share low similarity except for the MIR and bHLH domains. Another reported TF that inhibits anthocyanin accumulation is LcbHLH92 from sheepgrass. CpbHLH1 and LcbHLH92 differ greatly in amino acid sequence. The ORF of CpbHLH1 consists of 2025 bp encoding 674 amino acids, while the complete ORF of LcbHLH92 consists of 873 bp encoding 290 amino acids (Zhao et al. 2019). According to the classification system established by Heim et al. (2003), CpbHLH1 belongs to the subgroup IIIf, while LcbHLH92 belongs to the subgroup IVd and does not contain a MIR domain. In a previous study, the inhibitory effects of LcbHLH92 were achieved by activating JAZ genes to repress the expression of TT8; the expression of DFR and ANS was subsequently downregulated (Zhao et al. 2019). The mechanisms underlying the inhibitory effects of IN1, however, remain unknown.
CpbHLH1 downregulates the expression of both structural genes and endogenous TFs in model plant species potentially by inhibiting activity of the MYB–bHLH complex
We evaluated the relative transcript levels of the structural genes that regulate anthocyanin synthesis in Arabidopsis and tobacco. Although most EBGs in the flavonoid biosynthesis pathway exhibited a different trend in the transgenic lines compared with the WT plants, the expression levels of LBGs, AtDFR, AtANS, NtDFR, and NtANS, dramatically decreased in all transgenic lines. As transformation can sometimes lead to disorders, the different expression trends of EBGs in transgenic lines might be due to the ectopic expression of CpbHLH1. In addition, overexpression of CpbHLH1 resulted in the downregulation of endogenous TFs in transgenic model plants, including AtTT8, NtAN1a, and NtAN1b. Recent studies have shown that the bHLH activator can form a complex with the MYB activator to modulate the expression of bHLH itself (Baudry et al. 2006; Albert et al. 2014). The decrease in bHLH activator expression in transgenic plants might also be due to the effect of CpbHLH1 on the MBW complex.
Using Y2H and BiFC assays, we found that CpbHLH1 could interact with MYB activators of anthocyanin synthesis in Arabidopsis and tobacco, suggesting the possibility that CpbHLH1 could directly affect the MBW complex. The results of the dual-LUC transient expression assay confirmed that CpbHLH1 was a transcriptional repressor (Fig. 9b). In apple, MdMYB16 belonging to anthocyanin-related R2R3-MYB repressor can directly bind to the promoters of the ANS and UFGT genes to suppress their expression (Xu et al. 2017). It remains unclear whether CpbHLH1 directly affects the MBW complex or binds to the promoters of the LBGs to suppress their expression. To test these assumptions, dual-LUC transient expression experiments were performed. The results confirmed that CpbHLH1 could not repress genes involved in anthocyanin biosynthesis on its own (Supplementary Figure S5, Supplementary Figure S6) and that CpbHLH1 had a negative impact on both the TT8–PAP1 complex from Arabidopsis (Supplementary Figure S5) and the AN1–AN2 complex from tobacco (Supplementary Figure S6). The reduction of endogenous gene expression in transgenic plants might have been due to the effect of CpbHLH1 on the MBW complex.
The repressive motif(s) in CpbHLH1 protein may be located within the domain between the MIR and the bHLH domain
Transcriptional inhibition domains have recently been found within many negative regulatory plant proteins. These domains include the EAR motif L/FDLNL/F(x)P that was first identified in class II ERF transcriptional negative regulatory proteins (Ohta et al. 2001), the LxLxL motif within domain I of Aux/IAA proteins (Tiwari et al. 2004), the TLLLFR motif within AtMYBL2 (Matsui et al. 2008) and the R/KLFGV motif within numerous B3 DNA-binding domain transcriptional regulators (Ikeda and Ohme-Takagi 2009). CpbHLH1 is a TF that negatively regulates anthocyanin accumulation, but bioinformatics analysis revealed no known repressive motif within the CpbHLH1 protein. We inadvertently found an “LKLTL” sequence in the C-terminal region of the CpbHLH1 protein. To evaluate whether the “LKLTL” sequence controls the repressive function of CpbHLH1, deletion of the CpbHLH1 protein was performed such that only the amino acid upstream of the “LKLTL” sequence remained. Deletion of the “LKLTL” sequence only slightly relieved the repression by CpbHLH1 (Supplementary Figure S4), indicating that the “LKLTL” sequence may not control the repressive function of CpbHLH1. The active repressive motif LxLxL may exist only within domain I of Aux/IAA proteins.
To further investigate the domain(s) responsible for the repressive activity of CpbHLH1, we tested the transcriptional repression ability of truncated mutants of CpbHLH1. Deletion of the C-terminal region of CpbHLH1 had no obvious influence on the repression by CpbHLH1, but removal of both the C-terminal and the bHLH domain of CpbHLH1 slightly relieved the repression by CpbHLH1. These results suggest that the bHLH domain also mediates the repressive ability. A similar result in which the bHLH domain was shown to mediate the repression of a bHLH protein was also found for the bHLH protein SHARP-1 (Garriga-Canut et al. 2001). Further deletion mutation of the CpbHLH1 protein such that only the MIR was left relieved the repressive activity (Fig. 9c), suggesting that the repressive motif(s) might be located in the region between the MIR and the bHLH domain.
Conclusion
We identified that overexpression of CpbHLH1 from wintersweet greatly reduced anthocyanin content in transgenic Arabidopsis and tobacco plants, and the results suggested that CpbHLH1 is a transcription factor that inhibits anthocyanin accumulation. This is the first bHLH repressor of anthocyanin biosynthesis to be identified in dicotyledons. The present results can help us better understand the anthocyanin regulatory network in plants and may provide insights into the diverse functions of bHLH proteins.
References
Albert NW, Davies KM, Lewis DH, Zhang H, Montefiori M, Brendolise C, Boase MR, Ngo H, Jameson PE, Schwinn KE (2014) A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant Cell 26(3):962–980. https://doi.org/10.1105/tpc.113.122069
An XH, Tian Y, Chen KQ, Wang XF, Hao YJ (2012) The apple WD40 protein MdTTG1 interacts with bHLH but not MYB proteins to regulate anthocyanin accumulation. J Plant Physiol 169(7):710–717. https://doi.org/10.1016/j.jplph.2012.01.015
Bai Y, Pattanaik S, Patra B, Werkman JR, Xie CH, Yuan L (2011) Flavonoid-related basic helix-loop-helix regulators, NtAn1a and NtAn1b, of tobacco have originated from two ancestors and are functionally active. Planta 234(2):363–375. https://doi.org/10.1007/s00425-011-1407-y
Baudry A, Caboche M, Lepiniec L (2006) TT8 controls its own expression in a feedback regulation involving TTG1 and homologous MYB and bHLH factors, allowing a strong and cell-specific accumulation of flavonoids in Arabidopsis thaliana. Plant J 46(5):768–779. https://doi.org/10.1111/j.1365-313X.2006.02733.x
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12(12):2383–2394. https://doi.org/10.1105/tpc.12.12.2383
Burr FA, Burr B, Scheffler BE, Blewitt M, Wienand U, Matz EC (1996) The maize repressor-like gene intensifier1 shares homology with the r1/b1 multigene family of transcription factors and exhibits missplicing. Plant Cell 8(8):1249–1259. https://doi.org/10.1105/tpc.8.8.1249
Chandler VL, Radicella JP, Robbins TP, Chen J, Turks D (1989) Two regulatory genes of the maize anthocyanin pathway are homologous: isolation of B utilizing R genomic sequences. Plant Cell 1(12):1175–1183. https://doi.org/10.1105/tpc.1.12.1175
Chen L, Hu B, Qin Y, Hu G, Zhao J (2019) Advance of the negative regulation of anthocyanin biosynthesis by MYB transcription factors. Plant Physiol Biochem 136:178–187. https://doi.org/10.1016/j.plaphy.2019.01.024
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Cone KC, Burr FA, Burr B (1986) Molecular analysis of the maize anthocyanin regulatory locus C1. P Natl Acad Sci USA 83(24):9631–9635. https://doi.org/10.1073/pnas.83.24.9631
Espley RV, Hellens RP, Putterill J, Stevenson DE, Kutty-Amma S, Allan AC (2007) Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J 49(3):414–427. https://doi.org/10.1111/j.1365-313X.2006.02964.x
Espley RV, Brendolise C, Chagne D, Kutty-Amma S, Green S, Volz R, Putterill J, Schouten HJ, Gardiner SE, Hellens RP, Allan AC (2009) Multiple repeats of a promoter segment causes transcription factor autoregulation in red apples. Plant Cell 21(1):168–183. https://doi.org/10.1105/tpc.108.059329
Farcy E, Cornu A (1979) Isolation and characterization of anthocyanin variants originating from the unstable system an2–1 in Petunia hybrida (Hort.). Theor Appl Genet 55(6):273–278. https://doi.org/10.1007/BF00265365
Fu CC, Han YC, Kuang JF, Chen JY, Lu WJ (2017) Papaya CpEIN3a and CpNAC2 co-operatively regulate carotenoid biosynthesis-related genes CpPDS2/4, CpLCY-e and CpCHY-b during fruit ripening. Plant Cell Physiol 58(12):2155–2165. https://doi.org/10.1093/pcp/pcx149
Garriga-Canut M, Roopra A, Buckley NJ (2001) The basic helix-loop-helix protein, sharp-1, represses transcription by a histone deacetylase-dependent and histone deacetylase-independent mechanism. J Biol Chem 276(18):14821–14828. https://doi.org/10.1074/jbc.M011619200
Gerats AGM, Farcy E, Wallroth M, Groot SPC, Schram A (1984) Control of anthocyanin synthesis in Petunia-hybrida by multiple allelic series of the genes An1 and An2. Genetics 106(3):501–508
Goff SA, Cone KC, Chandler VL (1992) Functional analysis of the transcriptional activator encoded by the maize B gene: evidence for a direct functional interaction between two classes of regulatory proteins. Genes Dev 6(5):864–875. https://doi.org/10.1101/gad.6.5.864
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53(5):814–827. https://doi.org/10.1111/j.1365-313X.2007.03373.x
Heim MA, Jakoby M, Werber M, Martin C, Weisshaar B, Bailey PC (2003) The basic helix-loop-helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 20(5):735–747. https://doi.org/10.1093/molbev/msg088
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7(7):1071–1083. https://doi.org/10.1105/tpc.7.7.1071
Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SG, Fraley RT (1985) A simple and general-method for transferring genes into plants. Science 227(4691):1229–1231
Ikeda M, Ohme-Takagi M (2009) A novel group of transcriptional repressors in Arabidopsis. Plant Cell Physiol 50(5):970–975. https://doi.org/10.1093/pcp/pcp048
Ilk N, Ding J, Ihnatowicz A, Koornneef M, Reymond M (2015) Natural variation for anthocyanin accumulation under high-light and low-temperature stress is attributable to the ENHANCER OF AG-4 2 (HUA2) locus in combination with PRODUCTION OF ANTHOCYANIN PIGMENT1 (PAP1) and PAP2. New Phytol 206(1):422–435. https://doi.org/10.1111/nph.13177
Li S (2014) Transcriptional control of flavonoid biosynthesis: fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal Behav 9(1):e27522. https://doi.org/10.4161/psb.27522
Ludwig SR, Habera LF, Dellaporta SL, Wessler SR (1989) Lc, a member of the maize R gene family responsible for tissue-specific anthocyanin production, encodes a protein similar to transcriptional activators and contains the myc-homology region. Proc Natl Acad Sci USA 86(18):7092–7096. https://doi.org/10.1073/pnas.86.18.7092
Luo P, Ning GG, Wang Z, Shen YX, Jin HA, Li PH, Huang SS, Zhao J, Bao MZ (2016) Disequilibrium of flavonol synthase and dihydroflavonol-4-reductase expression associated tightly to white vs red color flower formation in plants. Front Plant Sci 6:1257. https://doi.org/10.3389/fpls.2015.01257
Matsui K, Umemura Y, Ohme-Takagi M (2008) AtMYBL2, a protein with a single MYB domain, acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J 55(6):954–967. https://doi.org/10.1111/j.1365-313X.2008.03565.x
Mehrtens F, Kranz H, Bednarek P, Weisshaar B (2005) The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol 138(2):1083–1096. https://doi.org/10.1104/pp.104.058032
Mol J, Grotewold E, Koes R (1998) How genes paint flowers and seeds. Trends Plant Sci 3(6):212–217. https://doi.org/10.1016/S1360-1385(98)01242-4
Munis MF, Tu L, Deng F, Tan J, Xu L, Xu S, Long L, Zhang X (2010) A thaumatin-like protein gene involved in cotton fiber secondary cell wall development enhances resistance against Verticillium dahliae and other stresses in transgenic tobacco. Biochem Biophys Res Commun 393(1):38–44. https://doi.org/10.1016/j.bbrc.2010.01.069
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15(3):473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nesi N, Debeaujon I, Jond C, Pelletier G, Caboche M, Lepiniec L (2000) The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 12(10):1863–1878. https://doi.org/10.1105/tpc.12.10.1863
Ohta M, Matsui K, Hiratsu K, Shinshi H, Ohme-Takagi M (2001) Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13(8):1959–1968. https://doi.org/10.1105/tpc.13.8.1959
Pandey A, Misra P, Bhambhani S, Bhatia C, Trivedi PK (2014) Expression of arabidopsis MYB transcription factor, AtMYB111, in tobacco requires light to modulate flavonol content. Sci Rep 4:5018. https://doi.org/10.1038/srep05018
Park KI, Ishikawa N, Morita Y, Choi JD, Hoshino A, Iida S (2007) A bHLH regulatory gene in the common morning glory, Ipomoea purpurea, controls anthocyanin biosynthesis in flowers, proanthocyanidin and phytomelanin pigmentation in seeds, and seed trichome formation. Plant J 49(4):641–654. https://doi.org/10.1111/j.1365-313X.2006.02988.x
Pattanaik S, Xie CH, Yuan L (2008) The interaction domains of the plant Myc-like bHLH transcription factors can regulate the transactivation strength. Planta 227(3):707–715. https://doi.org/10.1007/s00425-007-0676-y
Ramsay NA, Glover BJ (2005) MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10(2):63–70. https://doi.org/10.1016/j.tplants.2004.12.011
Ramsay NA, Walker AR, Mooney M, Gray JC (2003) Two basic-helix-loop-helix genes (MYC-146 and GL3) from Arabidopsis can activate anthocyanin biosynthesis in a white-flowered Matthiola incana mutant. Plant Mol Biol 52(3):679–688. https://doi.org/10.1023/A:1024852021124
Schwinn K, Venail J, Shang YJ, Mackay S, Alm V, Butelli E, Oyama R, Bailey P, Davies K, Martin C (2006) A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 18(4):831–851. https://doi.org/10.1105/tpc.105.039255
Spelt C, Quattrocchio F, Mol JN, Koes R (2000) anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12(9):1619–1632. https://doi.org/10.1105/tpc.12.9.1619
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50(4):660–677. https://doi.org/10.1111/j.1365-313X.2007.03078.x
Stracke R, Jahns O, Keck M, Tohge T, Niehaus K, Fernie AR, Weisshaar B (2010) Analysis of PRODUCTION OF FLAVONOL GLYCOSIDES-dependent flavonol glycoside accumulation in Arabidopsis thaliana plants reveals MYB11-, MYB12- and MYB111-independent flavonol glycoside accumulation. New Phytol 188(4):985–1000. https://doi.org/10.1111/j.1469-8137.2010.03421.x
Sui S, Luo J, Ma J, Zhu Q, Lei X, Li M (2012) Generation and analysis of expressed sequence tags from Chimonanthus praecox (Wintersweet) flowers for discovering stress-responsive and floral development-related genes. Comp Funct Genomics 2012:134596. https://doi.org/10.1155/2012/134596
Tian JP, Ma ZY, Zhao KG, Zhang J, Xiang L, Chen LQ (2019) Transcriptomic and proteomic approaches to explore the differences in monoterpene and benzenoid biosynthesis between scented and unscented genotypes of wintersweet. Physiol Plantarum 166(2):478–493. https://doi.org/10.1111/ppl.12828
Tiwari SB, Hagen G, Guilfoyle TJ (2004) Aux/IAA proteins contain a potent transcriptional repression domain. Plant Cell 16(2):533–543. https://doi.org/10.1105/tpc.017384
Tsuda T (2012) Dietary anthocyanin-rich plants: biochemical basis and recent progress in health benefits studies. Mol Nutr Food Res 56(1):159–170. https://doi.org/10.1002/mnfr.201100526
Xiang L, Zhao K, Chen L (2010) Molecular cloning and expression of Chimonanthus praecox farnesyl pyrophosphate synthase gene and its possible involvement in the biosynthesis of floral volatile sesquiterpenoids. Plant Physiol Biochem 48(10–11):845–850. https://doi.org/10.1016/j.plaphy.2010.08.015
Xie XB, Li S, Zhang RF, Zhao J, Chen YC, Zhao Q, Yao YX, You CX, Zhang XS, Hao YJ (2012) The bHLH transcription factor MdbHLH3 promotes anthocyanin accumulation and fruit colouration in response to low temperature in apples. Plant Cell Environ 35(11):1884–1897. https://doi.org/10.1111/j.1365-3040.2012.02523.x
Xu H, Wang N, Liu J, Qu C, Wang Y, Jiang S, Lu N, Wang D, Zhang Z, Chen X (2017) The molecular mechanism underlying anthocyanin metabolism in apple using the MdMYB16 and MdbHLH33 genes. Plant Mol Biol 94(1–2):149–165. https://doi.org/10.1007/s11103-017-0601-0
Yang N, Zhao K, Li X, Zhao R, Aslam MZ, Yu L, Chen L (2018) Comprehensive analysis of wintersweet flower reveals key structural genes involved in flavonoid biosynthetic pathway. Gene 676:279–289. https://doi.org/10.1016/j.gene.2018.08.050
Zhang W, Ning G, Lv H, Liao L, Bao M (2009) Single MYB-type transcription factor AtCAPRICE: a new efficient tool to engineer the production of anthocyanin in tobacco. Biochem Biophys Res Commun 388(4):742–747. https://doi.org/10.1016/j.bbrc.2009.08.092
Zhang LH, Jia BL, Zhuo RY, Liu JL, Pan HY, Baldwin TC, Zhang SH (2012) An acyl-acyl carrier protein thioesterase gene isolated from wintersweet (Chimonanthus praecox), CpFATB, enhances drought tolerance in transgenic tobacco (Nicotiana tobaccum). Plant Mol Biol Rep 30(2):433–442. https://doi.org/10.1007/s11105-011-0359-5
Zhang Y, Zhu H, Shao C, Cai F, Zhang J, Bao M (2019) PaMYB82 from Platanus acerifolia regulates trichome development in transgenic Arabidopsis. Plant Sci 287:110177. https://doi.org/10.1016/j.plantsci.2019.110177
Zhao KG, Zhou MQ, Chen LQ, Zhang DL, Robert GW (2007) Genetic diversity and discrimination of Chimonanthus praecox (L.) link germplasm using ISSR and RAPD markers. HortScience 42(5):1144–1148
Zhao PC, Li XX, Jia JT, Yuan GX, Chen SY, Qi DM, Cheng LQ, Liu GS (2019) bHLH92 from sheepgrass acts as a negative regulator of anthocyanin/proanthocyandin accumulation and influences seed dormancy. J Exp Bot 70(1):269–284. https://doi.org/10.1093/jxb/ery335
Zhou LL, Shi MZ, Xie DY (2012) Regulation of anthocyanin biosynthesis by nitrogen in TTG1-GL3/TT8-PAP1-programmed red cells of Arabidopsis thaliana. Planta 236(3):825–837. https://doi.org/10.1007/s00425-012-1674-2
Acknowledgements
This work was funded by the National Natural Science Foundation of China (Nos: 31272207 and 31972442). We thank Professor Wang-jin Lu (State Key Laboratory for Conservation and Utilization of Subtropical Agro-bioresources, College of Horticulture, South China Agricultural University) and Professor Jin-Long Qiu (State Key Laboratory of Plant Genomics, Institute of Microbiology, Chinese Academy of Sciences) for the generous gifts of the pBD vectors and the pap1-D mutant of Arabidopsis, respectively. We are also grateful to Professor Guogui Ning (Key Laboratory of Horticultural Plant Biology, College of Horticulture and Forestry Sciences, Huazhong Agricultural University) for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
RZ, XS and KZ: designed the experiments; RZ: performed the majority of experiments; XS, NY, and LC: supplied the materials; RZ, XS, and KZ: analyzed experimental results. RZ: prepared the initial manuscript draft; LX, XL, and KZ: produced the final version of the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
We declare that we have no conflicts of interest.
Additional information
Communicated by Salim Al-Babili.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, R., Song, X., Yang, N. et al. Expression of the subgroup IIIf bHLH transcription factor CpbHLH1 from Chimonanthus praecox (L.) in transgenic model plants inhibits anthocyanin accumulation. Plant Cell Rep 39, 891–907 (2020). https://doi.org/10.1007/s00299-020-02537-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-020-02537-9