Abstract
Key message
A gene not only control tiller and plant height, but also regulate panicle structure by QTL dissection in rice.
Abstract
An ideal panicle structure is important for improvement of plant architecture and rice yield. In this study, using recombinant inbred lines (RILs) of PA64s and 93-11, we identified a quantitative trait locus (QTL), designated qPPB3 for primary panicle branch number. With a BC3F2 population derived from a backcross between a resequenced RIL carrying PA64s allele and 93-11, qPPB3 was fine mapped to a 34.6-kb genomic region. Gene prediction analysis identified four putative genes, among which Os03g0203200, a previously reported gene for plant height and tiller number, Dwarf 88 (D88)/Dwarf 14 (D14), had three nucleotide substitutions in 93-11 compared with PA64s. The T to G substitution resulted in one amino acid change from valine in 93-11 to glycine in PA64s. Real-time PCR analysis showed expression level of D88 was higher in 93-11 than PA64s. The expression of APO1 and IPA1 increased, while GN1a and DST decreased in 93-11 compared with PA64s. Therefore, D88/D14 is not only a key regulator for branching, but also affects panicle structure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant architecture is very important for improving rice yield, grain quality, resistance to multiple biotic and abiotic stresses, and nutrient utilization efficiency. Panicle structure, belonging to plant architecture, is one of the important factors for rice yield. It includes four components: panicle length, primary panicle branch number, secondary panicle branch number, and spikelet number. Several genes that regulate the development of rice panicle branches have been cloned, such as MOC1 (Li et al. 2003), LAX1 (Komatsu et al. 2003), LAX2 (Tabuchi et al. 2011), GN1a/OsCKX2 (Ashikari et al. 2005), DST (Li et al. 2013), DEP1 (Huang et al. 2009), SP1 (Li et al. 2009), and APO1 (Ikeda et al. 2007).
Panicle branches are lateral organs emerged at reproductive stage, and their number affects spikelet number. In some cases, panicle branches are regulated by common mechanisms shared with tiller formation and elongation. MOC1 and LAX1 were genes that regulate both tillers and panicle branches. Reduction in tiller number and panicle branch number was discovered in both mutants. Mutation in LAX2 led to decrease in tiller number. However, the primary panicle branch number remained unchanged (Tabuchi et al. 2011). The near isogenic lines (NILs) carrying ipa1 displayed reduced tillers and increased panicle branches (Jiao et al. 2010). DEP1 (Huang et al. 2009) was a major dominant QTL that control panicle branches. The NIL-dep1 showed more panicle branches, which may result from cell proliferation through GN1a (Huang et al. 2009). APO1, a pivotal gene to regulate primary panicle branch, controlled the vascular bundle formation and could increase the harvest index and grain yield in rice (Terao et al. 2010).
In this study, QTL analysis of panicle branches was performed with a RIL population derived from an indica/javonica cross, and a major QTL (qPPB3) for primary panicle branch number was detected on chromosome 3. The region of qPPB3 was narrowed by map-based cloning strategy with BC3F2 population derived from a chromosome segment substitution line (CSSL). Based on sequencing and expression analysis, the predicted ORF for qPPB3 encoding a protein of the α/β-fold hydrolase superfamily (D88/D14), was proved to be not only a key regulator for plant shoot branching (Jiang et al. 2013; Arite et al. 2009; Gao et al. 2009), but also effected primary panicle branching.
Materials and methods
Mapping population and phenotyping
The core mapping population of 132 LYP9 RILs was randomly chosen from 1,841 RILs derived by single-seed descend from a cross between an elite paternal inbred Oryza sativa. indica cv. 93-11 and the maternal inbred Oryza sativa ssp. javonica cv. PA64s, a photo-thermo-sensitive male sterile line with a mixed genetic background of indica, javanica, and japonica. The population was developed in the experimental fields at China National Rice Research Institute in Hangzhou, Zhejiang Province, and in Sanya, Hainan Province, China. After 12 generations of self-fertilization, genomic DNA samples of the F13 RILs were isolated for genotyping (Gao et al. 2013). Phenotyping was conducted according to Gao et al. (2013).
Data analysis and QTL detection
All statistical analyses were completed using the SAS (Statistical Analysis System) V8.01. QTL analysis was performed with the MultiQTL1.6 package (http://www.multiqtl.com) using maximum likelihood interval mapping (MIM) based on a permutation test (1,000 permutation, P = 0.05) for each dataset to confirm the LOD threshold. It was considered as a major effect QTL when its LOD was larger than 2.5. QTLs were named according to McCouch (Mccouch et al. 1997).
Acquisition of CSSL involving qPPB3
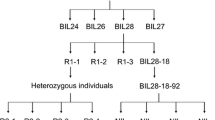
To develop a CSSL containing qPPB3, a line of RILs with PA64s’ genotype in the qPPB3 region was selected to backcross with recurrent parent 93-11. Two STS markers P1 and P2 (Table 1) were used for marker-assisted selection (MAS) of each generation in the segregating progenies. As a result, a BC3F1 line GH18, with 93-11 genetic background exhibiting heterozygous across the entire qPPB3 region, was constructed. After self-crossing, we acquired a BC3F2 population for fine mapping of qPPB3.
Design of fine mapping markers
Primers were designed around qPPB3 on chromosome 3 using insertions/deletions (InDels) identified between 93-11 and PA64s (Gao et al. 2013) by two software, Primer Premier 5.0 and Oligo 7 (Table 1).
RNA extraction and real-time PCR analysis
Total RNA was isolated from panicles before booting stage with the micro RNA extraction kit (Axygen). DNase treatment, cDNA synthesis, primer design and SYBR Green I Real-time PCR were carried out as described (Vandesompele et al. 2002) using a ReverTra Ace® qPCR-RT kit (TOYOBA, Japan). Real-time PCR amplification mixtures (10 μl) contained 50 ng template cDNA, 2× SYBR Green PCR Master Mix (10 μl) (Applied Biosystems), and 200 nM forward and reverse primers. Reactions were run on an ABI PRISM® 7900HT Sequence Detector (Applied Biosystems). The relative expression level of each transcript was obtained by comparing to the expression of the OsACTIN1 gene. Primers for D88, APO1, IPA1, GN1a, DST, and Actin are listed in Table 1.
Subcellular localization of D88
Full-length cDNA of D88 was isolated from Nipponbare and sequenced. Then it was inserted into the vector pCAMBIA1302. The D88::GFP fusion was made by inflame fusion of the full-length D88 cDNA with GFP. Rice transformation was performed by an Agrobacterium-mediated method (Hiei et al. 1994). The root tips of transgenic plants expressing the GFP fusions were observed directly with a microscope (Nikon 90i).
Results
Phenotypic performance of panicle branches in parents and RILs
Rice panicle branch traits include primary panicle branch number (PPB) and secondary panicle branch number (SPB). And they directly affect spikelet number per panicle (SN), a crucial factor for rice grain yield. Our study showed PPB was significantly related to SPB and SN, with correlation coefficient 0.5204 and 0.4062, respectively. SPB was significantly related to SN (r = 0.8568) (Table 2). The relationships of PPB, SPB, and SN were positive to each other. Therefore, increase in PPB help improve rice grain yield. In parents, spikelet number per panicle, primary panicle branch number and secondary panicle branch number of 93-11 are larger than PA64s. Significant difference existed between parents in spikelet number per panicle and secondary panicle branch number. On the contrary tiller number exhibits less in 93-11 than PA64s. (Fig. 1).
Panicle morphology of indica 93-11 and Javonica PA64s. a Panicle structure of 93-11 (left) and PA64s (right), Bar = 5 cm. b Spikelet number per panicle in two parents. c Primary panicle branch number, secondary panicle branch number, and tillers per plant of 93-11 and PA64s. Values are the mean ± SD with five replicates
QTL analysis for PPB, SPB, and SN
SNP markers, covered all 12 chromosomes, were used to construct a high-resolution genetic linkage map with total genetic distance of 1,382 cM and an average distance of 0.53 cM between two adjacent SNP markers, as described in Gao et al. (2013).
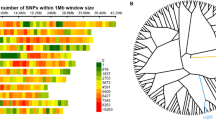
QTL analysis was performed with MultiQTL1.6 using the maximum likelihood interval mapping approach with an LOD threshold 2.5. We detected two QTLs for PPB on chromosomes 3 and 8, named qPPB3 and qPPB8. The qPPB3 explained 9.4 % phenotypic variation with additive effect came from PA64s. And qPPB8 explained 15.3 % phenotypic variation came from 93-11. Two QTLs for SPB were identified. The qSPB1 was mapped on chromosome 1 and explained 10 % phenotypic variation with additive effect came from 93-11. The positive effect of qSPB9 was from 93-11, explained 9.6 % phenotypic variation. PPB and SPB are two key factors for SN determined by panicle architecture. Two QTLs, qSN8 and qSN9 for SN were mapped on chromosomes 8 and 9 with additive effects of 35.5 and 33.2, respectively (Table 3; Fig. 2a). They totally explained 25.4 % phenotypic variation.
a QTLs for panicle traits in the RIL population. SNP markers are shown on the left of chromosomes. QTLs signed on chromosomes were detected in several environments and different years. b QTL peak map of rice chromosome 3. The genetic distances (cM) are shown below the x axis. c Fine mapping of qPPB3. The white, black, and gray bars represent genotypes of PA64s, 93-11, and heterozygote, respectively. All genotypic lines exhibited (G1-G11) were derived from BC3F2 population
Fine mapping of qPPB3
To narrow down the region of qPPB3, we developed seven markers (P1–P7). With 1,522 RIL populations, qPPB3 was fine mapped in the region between markers P1 and P2. Total 1,105 BC3F2 individuals were utilized to further mapping, and 17 lines were selected from BC3F2 progenies derived from the cross between GH18 and 93-11. Parent 93-11 and GH18 were designated as the controls L1 and L2, respectively. Out of ten lines with high PPB similar to L2 (genotype of PA64s), there were five recombinant events between P4 and qPPB3, four between P5 and qPPB3, one between P7 and qPPB3, one between P6 and qPPB3, four between P3 and qPPB3. On the other hand, among seven lines with low PPB similar to L1 (genotype of 93-11), there were two recombinant events between P4 and qPPB3, one between P5 and qPPB3, one between P3 and qPPB3, and P6 and P7 co-segregated with qPPB3. Finally, qPPB3 was delimited to an interval of 34.6-kb region between markers P6 and P7 on the BAC clone AC146702 (Fig. 2b, c).
A candidate gene for qPPB3
To screen candidate genes in the critical 34.6-kb genomic region of Nipponbare genome (http://rapdb.dna.affrc.go.jp/), and four corresponding genes (Os03g0203100, Os03g0203200, Os03g0203700, Os03g0203800) were found. After comparing sequence of the region between parents, only two genes, Os03g0203200 and Os03g0203700, had three SNPs and one InDel between two parents, respectively. However, product of Os03g0203700 was similar to Calcium-transporting ATPase 2, which has been found irrelevant to plant branching so far, and its expression showed no significant difference between two parents. Instead, Os03g0203200 (D88) encoded esterase D14/D88 homologous to the α/β-fold hydrolase superfamily protein, which has been reported affecting plant branching and significant difference in its expression level was detected here between 93-11 and PA64s. Only one substitution (T959 to G959) resulted in amino acid change from valine in 93-11 to glycine in PA64s (Fig. 3). Therefore, Os03g0203200 (D88) gene was suggested to be a candidate gene for qPPB3.
Comparison of genomic sequences and amino acid sequences of rice qPPB3 between two parents. a The qPPB3 gene is composed of three exons and two introns. Black and white boxes indicate coding and untranslated region (UTR), respectively, and lines represent introns. Arrows indicate the locations of the 93-11 mutation. b The nucleotide substitution for change of amino acid is marked with a red box. c Multiple deduced amino acid sequence alignment of protein coded by qPPB3 of Nipponbare, 93-11, and PA64s. A red box indicates the altered amino acid
Expression comparison of D88 and panicle structure related genes in initial inflorescence of parents
RNA was extracted from inflorescences with length less than 5 mm at the formation stage of panicle primary branch. At the initial formation stage of panicle, the expression level of D88 in PA64s was significantly lower than that in 93-11 (Fig. 4), which is consistent with the previous report that lower expression of D88 gene led to higher tillers but smaller panicle in the d88 mutant (Gao et al. 2009). With the same material, the expression of APO1 and IPA1 in 93-11 also significantly increased compared with PA64s, while the expression of GN1a and DST, two negative regulators of grain number, enhanced in PA64s significantly.
Subcellular localization of D88
To define the intracellular localization of D88 in rice cells, we introduced D88::GFP into rice by using Agrobacterium transformation. We found that D88::GFP fusion protein was localized to both nucleus and cytoplasm in rice root cells, weakly appeared in the plasma membrane (Fig. 5).
Discussion
Panicle structure is one of the most important factors for rice yield. To date, 54 QTLs for PPB and 33 QTLs for SPB have been reported distributed on 12 chromosomes (Supplemental Fig. 1). Here, two QTLs were mapped for SPB, qSPB1 and qSPB9. Two QTLs, sbr1.1 and qSNB1-1 for SPB on chromosome 1 were mapped to the same interval of qSPB1 (Li et al. 2006; Cui et al. 2002). The qSPB9 was also reported previously (Li et al. 2006), and qSN9 for SN was found in the region of qSPB9. In this study, two QTLs for PPB, qPPB3 and qPPB8 were located on chromosome 3 and chromosome 8, respectively. The region of qPPB8 was also reported to be related to the QTLs for PPB and SN in other studies (Lin et al. 1996; Xu et al. 2001). One QTL (QPbn3a) for PPB and another QTL (QSbn3a) for SPB were detected in the same genomic region of qPPB3 (Xu et al. 2001). The qPPB3 was further mapped to a region of 34.6-kb on a BAC clone AC146702, where four candidate genes (Os03g0203100, Os03g0203200, Os03g0203700, Os03g0203800) existed. The translational product of Os03g0203200 was D88/D14, an esterase that confirmed to control rice tillering, with d88/d14 mutants exhibits high tillers but dwarfism (Arite et al. 2009; Gao et al. 2009).
Panicle primary branches developed from primary branch meristems, which are produced by inflorescence meristem. During reproductive development, inflorescence branching is under genetic control affected by hormones that include brassinosteroid (BR), cytokinin (CK), auxin and strigolactone (SL) (Beveridge 2006; McSteen and Leyser 2005; Ongaro and Leyser 2008; Dun et al. 2009; Wang et al. 2013). Like strigolactones insensitive mutants, PA64s also exhibited more tillers and less panicle branches than 93-11. In the study, qPPB3 was fine mapped to a region covering the D88 gene, whose product believed to be an important receptor in strigolactone signaling (Jiang et al. 2013). Strigolactones or related compounds were reported to inhibit shoot branching in plants (Gomez-Roldan et al. 2008). At the initial development stage of panicle, the expression level of D88 was significantly higher in 93-11 than PA64s. And the PPB of 93-11 was larger than PA64s. Therefore, strigolactones or their derivants might be the receptible hormone by D88 in cytoplasm or nucleus to affect the PPB, though regulatory mechanism of branching in panicle is different from that in shoot. In contrast to SL, CK directly promote bud growth and cell proliferation. We tested the expression of two CK related genes, GN1a/OsCKX2 and DST. The expression level of GN1a/OsCKX2 was significantly lower in 93-11 with more SN, suggesting CK might be accumulated higher in 93-11 than PA64s. Expression level of DST was also higher in PA64s, which in accord with its direct regulation of GN1a/OsCKX2 to control CK level in the reproductive SAM and, as a result, affects panicle branching and grain number (Li et al. 2013).
The APO1/SCM2, GN1a/OsCKX2 and IPA1/OsSPL14 were cloned QTLs associated with panicle structure. The line contained APO1 ORF and the proximal promoter region controlled only the number of PPB but not the number of grains per panicle. However, the line included only the distal promoter region that could increase the grain number and harvest index (Terao et al. 2010). The present study also confirmed that the higher PPB in 93-11 exhibited higher expression level of APO1. IPA1 was considered important not only for plant architecture but also for panicle structure. Higher expression of IPA1/OsSPL14 at the reproductive stage promoting panicle branching is consistent with our results (Miura et al. 2010; Jiao et al. 2010).
Dense panicle structure has been a major target for improvement of rice grain yield because of its relationship with grain number. The utilization of QTLs panicle structure, such as dep1, ipa1, and apo1 has increased the crop productivity. Therefore, pyramiding of these elite QTLs, including qPPB3 mapped here, will certainly benefit high-yield breeding by marker-assisted selection (MAS) for rice in the future.
Abbreviations
- APO1 :
-
Aberrant panicle organization 1
- BAC:
-
Bacterial artificial chromosome
- BR:
-
Brassinosteroid
- CK:
-
Cytokinin
- CSSL:
-
Chromosome segment substitution line
- DEP1 :
-
Dense erect panicle 1
- DST :
-
Drought and salt tolerance
- GFP:
-
Green fluorescence protein
- GN1a :
-
Grain number 1 a
- IPA1 :
-
Ideal plant architecture 1
- LAX1, LAX2 :
-
Lax panicle 1, lax panicle 2
- LYP9:
-
Liang-you-pei-jiu
- MAS:
-
Marker-assisted selection
- Moc1 :
-
Mono-culm 1
- ORF:
-
Open reading frame
- OsCKX2 :
-
Cytokinin oxidase 2
- OsSPL14 :
-
Squamosa promoter binding protein-like 14
- PPB:
-
Primary panicle branch number
- qPCR:
-
Quantitative PCR
- QTL:
-
Quantitative trait locus
- RIL:
-
Recombinant inbred line
- SL:
-
Strigolactone
- SN:
-
Spikelet number per panicle
- SP1 :
-
Small panicle 1
- SPB:
-
Secondary panicle branch number
References
Arite T, Umehara M, Ishikawa S, Hanada A, Maekawa M, Yamaguchi S, Kyozuka J (2009) d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol 50(8):1416–1424
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Science 309(5735):741–745
Beveridge CA (2006) Axillary bud outgrowth: sending a message. Curr Opin Plant Biol 9(1):35–40
Cui KH, Peng SB, Xing YZ, Yu SB, Xu CG (2002) Genetic analysis of the panicle traits related to yield sink size of rice. Yi Chuan Xue Bao 29(2):144–152
Dun EA, Brewer PB, Beveridge CA (2009) Strigolactones: discovery of the elusive shoot branching hormone. Trends Plant Sci 14(7):364–372
Gao Z, Qian Q, Liu X, Yan M, Feng Q, Dong G, Liu J, Han B (2009) Dwarf 88, a novel putative esterase gene affecting architecture of rice plant. Plant Mol Biol 71(3):265–276
Gao ZY, Zhao SC, He WM, Guo LB, Peng YL, Wang JJ, Guo XS, Zhang XM, Rao YC, Zhang C, Dong GJ, Zheng FY, Lu CX, Hu J, Zhou Q, Liu HJ, Wu HY, Xu J, Ni PX, Zeng DL, Liu DH, Tian P, Gong LH, Ye C, Zhang GH, Wang J, Tian FK, Xue DW, Liao Y, Zhu L, Chen MS, Li JY, Cheng SH, Zhang GY, Qian Q (2013) Dissecting yield-associated loci in super hybrid rice by resequencing recombinant inbred lines and improving parental genome sequences. Proc Natl Acad Sci USA 110(35):14492–14497
Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pages V, Dun EA, Pillot JP, Letisse F, Matusova R, Danoun S, Portais JC, Bouwmeester H, Becard G, Beveridge CA, Rameau C, Rochange SF (2008) Strigolactone inhibition of shoot branching. Nature 455(7210):189–194
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6(2):271–282
Huang X, Qian Q, Liu Z, Sun H, He S, Luo D, Xia G, Chu C, Li J, Fu X (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41(4):494–497
Ikeda K, Ito M, Nagasawa N, Kyozuka J, Nagato Y (2007) Rice ABERRANT PANICLE ORGANIZATION 1, encoding an F-box protein, regulates meristem fate. Plant J 51(6):1030–1040
Jiang L, Liu X, Xiong G, Liu H, Chen F, Wang L, Meng X, Liu G, Yu H, Yuan Y, Yi W, Zhao L, Ma H, He Y, Wu Z, Melcher K, Qian Q, Xu HE, Wang Y, Li J (2013) DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 504(7480):401–405
Jiao Y, Wang Y, Xue D, Wang J, Yan M, Liu G, Dong G, Zeng D, Lu Z, Zhu X, Qian Q, Li J (2010) Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat Genet 42(6):541–544
Komatsu K, Maekawa M, Ujiie S, Satake Y, Furutani I, Okamoto H, Shimamoto K, Kyozuka J (2003) LAX and SPA: major regulators of shoot branching in rice. Proc Natl Acad Sci USA 100(20):11765–11770
Li X, Qian Q, Fu Z, Wang Y, Xiong G, Zeng D, Wang X, Liu X, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li J (2003) Control of tillering in rice. Nature 422(6932):618–621
Li C, Zhou A, Sang T (2006) Genetic analysis of rice domestication syndrome with the wild annual species Oryza nivara. New Phytol 170(1):185–193
Li S, Qian Q, Fu Z, Zeng D, Meng X, Kyozuka J, Maekawa M, Zhu X, Zhang J, Li J, Wang Y (2009) Short panicle1 encodes a putative PTR family transporter and determines rice panicle size. Plant J 58(4):592–605
Li S, Zhao B, Yuan D, Duan M, Qian Q, Tang L, Wang B, Liu X, Zhang J, Wang J, Sun J, Liu Z, Feng YQ, Yuan L, Li C (2013) Rice zinc finger protein DST enhances grain production through controlling Gn1a/OsCKX2 expression. Proc Natl Acad Sci USA 110(8):3167–3172
Lin HX, Qian HR, Zhuang JY, Lu J, Min SK, Xiong ZM, Huang N, Zheng KL (1996) RFLP mapping of QTLs for yield and related characters in rice (Oryza sativa L.). Theor Appl Genet 92(8):920–927
Mccouch SR, Chen X, Panaud O, Temnykh S, Xu Y, Cho YG, Huang N, Ishii T, Blair M (1997) Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Mol Biol 35(1–2):89–99
Mcsteen P, Leyser O (2005) Shoot branching. Annu Rev Plant Biol 56:353–374
Miura K, Ikeda M, Matsubara A, Song XJ, Ito M, Asano K, Matsuoka M, Kitano H, Ashikari M (2010) OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat Genet 42(6):545–549
Ongaro V, Leyser O (2008) Hormonal control of shoot branching. J Exp Bot 59(1):67–74
Tabuchi H, Zhang Y, Hattori S, Omae M, Shimizu-Sato S, Oikawa T, Qian Q, Nishimura M, Kitano H, Xie H, Fang X, Yoshida H, Kyozuka J, Chen F, Sato Y (2011) LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 23(9):3276–3287
Terao T, Nagata K, Morino K, Hirose T (2010) A gene controlling the number of primary rachis branches also controls the vascular bundle formation and hence is responsible to increase the harvest index and grain yield in rice. Theor Appl Genet 120(5):875–893
Vandesompele J, De Paepe A, Speleman F (2002) Elimination of primer-dimer artifacts and genomic coamplification using a two-step SYBR green I real-time RT-PCR. Anal Biochem 303(1):95–98
Wang Y, Sun S, Zhu W, Jia K, Yang H, Wang X (2013) Strigolactone/MAX2-induced degradation of brassinosteroid transcriptional effector BES1 regulates shoot branching. Dev Cell 27(6):681–688
Xu JL, Xue QZ, Luo LJ, Li ZK (2001) QTL dissection of panicle number per plant and spikelet number per panicle in rice (Oryza sativa L.). Yi Chuan Xue Bao 28(8):752–759
Acknowledgments
This work was supported by Grants from the National Natural Science Foundation of China (Grant Nos. 31071485 and 31221004).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Kang Chong.
Y. Peng and Z. Gao contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Peng, Y., Gao, Z., Zhang, B. et al. Fine mapping and candidate gene analysis of a major QTL for panicle structure in rice. Plant Cell Rep 33, 1843–1850 (2014). https://doi.org/10.1007/s00299-014-1661-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1661-0