Abstract
Saccharomyces cerevisiae adapts to oxidative, osmotic stress and nutrient deprivation through transcriptional changes, decreased proliferation, and entry into other developmental pathways such as pseudohyphal formation and sporulation. Inositol pyrophosphates are necessary for these cellular responses. Inositol pyrophosphates are molecules composed of the phosphorylated myo-inositol ring that carries one or more diphosphates. Mutations in the enzymes that metabolize these molecules lead to altered patterns of stress resistance, altered morphology, and defective sporulation. Mechanisms to alter the synthesis of inositol pyrophosphates have been recently described, including inhibition of enzyme activity by oxidation and by phosphorylation. Cells with increased levels of 5-diphosphoinositol pentakisphosphate have increased nuclear localization of Msn2 and Gln3. The altered localization of these factors is consistent with the partially induced environmental stress response and increased expression of genes under the control of Msn2/4 and Gln3. Other transcription factors may also exhibit increased nuclear localization based on increased expression of their target genes. These transcription factors are each regulated by TORC1, suggesting that TORC1 may be inhibited by inositol pyrophosphates. Inositol pyrophosphates affect stress responses in other fungi (Aspergillus nidulans, Ustilago maydis, Schizosaccharomyces pombe, and Cryptococcus neoformans), in human and mouse, and in plants, suggesting common mechanisms and possible novel drug development targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The yeast Saccharomyces cerevisiae responds to cellular stress through dramatic and global changes in gene expression, translation, and protein/enzyme activity. The environmental stress response (ESR) lies at the center of these changes and represents a common gene expression program that is triggered by numerous distinct external stresses (Gasch 2003, 2007; Gasch et al. 2000). These gene expression changes include ~ 900 genes that have similar responses to osmolytes, oxidants, thermal shock, nitrogen depletion, amino acid starvation, reducing conditions (DTT), and lead to adaption and/or progression into stationary phase (Gasch 2003, 2007; Gasch et al. 2000). In general, the ESR genes are categorized as repressed genes (rESR), such as ribosome biogenesis (RiBi) genes that are associated with proliferation, and induced genes (iESR), such as protein chaperones that provide adaptive responses to intracellular stress. Each of these stresses also generates a specific transcriptional response in addition to the ESR (Brewster and Gustin 2014; Morano et al. 2012). In response to nitrogen and carbon deprivation, cells can also enter into the developmental pathways of sporulation and pseudohyphal formation instead of progression into stationary phase (Gancedo 2001; Neiman 2011). Unexpectedly and as described below, the cellular responses to these stresses require inositol pyrophosphates.
Inositol pyrophosphates (PP-InsPs) are composed of a phosphorylated myo-inositol ring that has one or more β-phosphates (pyrophosphates; see Fig. 1). We refer readers to several excellent reviews to learn more about the enzymes that metabolize inositol polyphosphates, the cellular processes affected by them, and their multiple mechanisms of action (Eskes et al. 2017; Mutlu and Kumar 2019; Shears 2015, 2017; Thota and Bhandari 2015; Wickner et al. 2018; Wilson et al. 2013). Here, we focus on the functional roles for PP-InsPs in the cellular responses of S. cerevisiae to stress with a brief review of the key metabolic enzymes.
Schematic depicting the metabolism of inositol pyrophosphates. The soluble inositol phosphate pathway begins with cleavage of phosphatidylinositol (4,5) bisphosphate by phospholipase C (not shown) to generate inositol (1,4,5) trisphosphate (IP3). Arg82 (inositol phosphate multikinase) phosphorylates the 3 and 6 positions to generate inositol (1,3,4,5,6) pentakisphosphate (IP5). Ipk1 (inositol phosphate kinase) phosphorylates the 2 position to generate inositol (1,2,3,4,5,6) hexakisphosphate (IP6). The preferred pathway for synthesis of inositol pyrophosphates with heavy arrows (top): IP6 is pyrophosphorylated by Kcs1 (IP6K) on the 5 position to generate 5PP-InsP5 (aka, 5-IP7), and Vip1 (PP-IP5K) pyrophosphorylates 5-IP7 to generate 1,5PP-InsP4 (IP8). Siw14 specifically dephosphorylates the β-phosphate on the 5 position, and Vip1 and Ddp1 dephosphorylate the β-phosphate on the 1 position. Below is the less preferred pathway, in which Vip1 acts first on the 1 position of IP6 to generate 1PP-InsP5 (1-IP7) and Kcs1 pyrophosphorylates the 5 position on 1PP-InsP5 to generate 1,5PP-InsP4 (IP8)
The soluble inositol polyphosphate pathway begins with the cleavage of membrane-associated phosphoinositide, phosphatidylinositol 4,5-bisphosphate (PtdIns(4,5)P2) by phospholipase C (PLC1) to form inositol (1,4,5) trisphosphate (abbreviated as InsP3 and further shortened to IP3). IP3 is phosphorylated by inositol multikinase (ARG82) and inositol phosphate kinase (IPK1) to produce inositol (1,2,3,4,5,6) hexakisphosphate (IP6). The kinase Kcs1 (aka, IP6K) phosphorylates the C5 α-phosphate of IP6 to generate 5PP-InsP5 (shortened to 5-IP7) (Saiardi et al. 1999), and the phosphatase Siw14 reverses this reaction by hydrolyzing the C5 β-phosphate (Steidle et al. 2016). Vip1 has dual kinase and phosphatase domains and, therefore, both adds and removes the C1 β-phosphate on 5-IP7 to generate 1,5PP-InsP4 (IP8) or it produces 1PP-InsP5 (1-IP7) from InsP6 (Lee et al. 2007; Mulugu et al. 2007). The DDP1 gene encodes a nonspecific polyphosphate phosphatase with many substrates, including 1-IP7, IP8, polyphosphate, and diadenosine polyphosphates (Kilari et al. 2013; Safrany et al. 1999; Yang et al. 1999). As such, the vip1∆ mutant cannot synthesize 1-IP7 or IP8 and has increased levels of its substrate 5-IP7 (Onnebo and Saiardi 2009; Steidle et al. 2016). The kcs1∆ mutant produces neither 5-IP7 nor IP8 (Dubois et al. 2002). The kcs1∆ vip1∆ double mutant has undetectable levels of all inositol pyrophosphates (Mulugu et al. 2007). The siw14∆ mutant has increased levels of both 5-IP7 and IP8 (Steidle et al. 2016). Many studies use the mutants described above to vary the levels of PP-InsPs to examine isoform-specific phenotypes and to characterize mechanisms of action.
Inositol polyphosphates currently have three known mechanisms of action. They serve as structural elements enabling proteins to fold into biologically active molecules (Dick et al. 2018; Macbeth et al. 2005; Millard et al. 2013; Ouyang et al. 2016; Watson et al. 2012, 2016). They act as regulatory molecules to affect signaling pathways, most commonly by competing with membrane-associated phosphatidylinositides for protein binding (Gokhale et al. 2013; Luo et al. 2003; Nagata et al. 2010; Wild et al. 2016). Finally, inositol pyrophosphates can donate the β-phosphate group to phosphoserine in proteins—without requiring a kinase—to alter protein activity (Azevedo et al. 2009; Bhandari et al. 2007; Chanduri et al. 2016; Pulloor et al. 2014; Ravi et al. 2019; Szijgyarto et al. 2011; Thota et al. 2015).
Inositol pyrophosphates are required for cellular responses to stress
Several studies have shown that inositol pyrophosphates are required for cell survival in response to stress. In one of the earliest studies, the kcs1∆ mutant was discovered to be sensitive to osmotic and cell wall (caffeine) stresses (Dubois et al. 2002). The kcs1Δ mutant is also sensitive to heat stress (Auesukaree et al. 2009; Gibney et al. 2013; Horigome et al. 2009; Jarolim et al. 2013; Ruiz-Roig et al. 2010; Saiardi et al. 2000; Sinha et al. 2008). Perhaps unexpectedly, both the kcs1∆ and vip1∆ mutants are resistant to hydrogen peroxide (Onnebo and Saiardi 2009). Although cell survival phenotypes for the kcs1∆ vip1∆ double mutant have not been reported, this strain is unable to activate the ESR in response to oxidative and osmotic stresses as measured by global gene expression assays (Worley et al. 2013).
We recently reported that the siw14Δ mutant was resistant to osmotic, heat, and oxidative stresses and nutrient deprivation (Steidle et al. 2020), which was also found during high-throughput analyses (Brown et al. 2006; Garay et al. 2014; Jarolim et al. 2013). Unlike the kcs1∆ vip1∆ double mutant, the siw14∆ mutant can generate the ESR; in fact, the siw14∆ mutant has an amplified transcriptional response to stress compared to the wild-type (Steidle et al. 2020). These results in the siw14∆, kcs1∆, and vip1∆ mutants indicate that formation of 5-IP7 (or possibly IP8) is important for the adaptive stress responses, although the different stress responses may utilize the PP-InsPs uniquely.
Cells also require inositol pyrophosphates to undergo the developmental changes of sporulation and pseudohyphal development in response to carbon and nitrogen starvation. The kcs1Δ mutant is defective for both sporulation (Enyenihi and Saunders 2003; Kloimwieder and Winston 2011) and pseudohyphal formation (Norman et al. 2018), and the vip1Δ strain is unable to form pseudohyphae (Pöhlmann et al. 2014). Increased levels of 5-IP7 due to overexpression of KCS1 or deletion of siw14Δ result in hyperfilamentation (Norman et al. 2018). Thus, the phenotypes of the siw14Δ mutant are largely the reverse of those in the kcs1Δ mutant, suggesting that synthesis of 5-IP7 is important for the adaptive stress responses as well as the developmental and morphological changes in response to nutrient deprivation.
Mechanisms to change inositol pyrophosphate levels in response to stress
Given the requirement for inositol pyrophosphates in the cellular responses to stress, the question was raised whether the levels of PP-InsPs are altered upon stress. Two studies showed opposite results when cells were challenged with hydrogen peroxide. In one, hydrogen peroxide treatment led to a decrease in IP7 levels to ~ 20% of the wild-type in vivo, and activity of recombinant Kcs1 was inhibited by oxidation in vitro, consistent with the lower levels of IP7 (Onnebo and Saiardi 2009). In the other, hydrogen peroxide treatment led to a twofold increase in IP7 levels and a fivefold increase in IP8 levels in vivo, and activity of recombinant Siw14 was reversibly inhibited by oxidation in vitro, consistent with the higher levels of IP7 (Steidle et al. 2020). The differences in the findings from the two studies have not been resolved; however, the differences could be due to strain backgrounds or experimental conditions. Both reports suggested that direct oxidation of the enzymes could occur within cells to alter inositol pyrophosphate levels.
Phosphorylation of enzymes is another possible mechanism to alter inositol pyrophosphates levels. IP7 levels increase under phosphate- and nitrogen-limiting conditions (Lee et al. 2007; Norman et al. 2018). Pseudohyphal growth depends upon several signaling pathways and their kinases, including the mitogen-activated protein kinase (MAPK) Kss1, the AMP-activated kinase Snf1, and the Ras/Protein Kinase A (PKA) pathway kinase Tpk1/2 (Celenza and Carlson 1984; Cook et al. 1996; Cullen and Sprague 2012; Erdman and Snyder 2001; Pan and Heitman 1999; Robertson and Fink 1998; Vyas et al. 2003). Both Kcs1 and Vip1 (but not Siw14) are phosphorylated under pseudohyphal-inducing conditions by the Kss1 and Snf1 kinases (Shively et al. 2015). Importantly, the ratio of 5-IP7 to 1-IP7 increased under pseudohyphal-inducing conditions (Norman et al. 2018), suggesting that phosphorylation of Kcs1 increases its catalytic activity and/or that phosphorylation of Vip1 either decreases its kinase activity or increases its phosphatase activity. These results suggest that phosphorylation could be a cellular mechanism to regulate inositol pyrophosphate levels; however, a direct demonstration of altered enzymatic activity is lacking. Further research will be necessary to determine if there are additional mechanisms to modulate inositol pyrophosphate levels.
Inositol pyrophosphates affect the ESR through both Rpd3L and Msn2/4
Two studies sought to determine the mechanism(s) by which inositol pyrophosphates influence the stress response, given the phenotypes associated with these molecules (Steidle et al. 2020; Worley et al. 2013). The general stress response in yeast depends on two distinct regulators, Msn2/4 and Rpd3L. Msn2 and Msn4 (abbreviated Msn2/4) are two partially redundant transcription factors that bind to stress response elements to promote the expression of their target genes (Görner et al. 1998; Sadeh et al. 2011). Rpd3L is a histone deacetylase (HDAC) that activates stress response genes and represses RiBi genes in response to stress (Alejandro-Osorio et al. 2009; Ruiz-Roig et al. 2010).
As noted above, one mechanism of action for inositol phosphates is structural. Mammalian Class I HDACs require IP4 to assemble into fully active histone deacetylase complexes (Millard et al. 2013; Watson et al. 2012, 2016). However, the authors note that it is possible that InsP4 regulates the enzymatic activity of the Class I HDACs because the binding pocket is accessible to soluble InsP4 and the apparent Kd for binding inositol phosphates is consistent with catalytic activity being affected by small changes in intracellular concentrations (Millard et al. 2013). The yeast Rpd3 protein has a putative inositol phosphate binding pocket based on sequence conservation with the mammalian enzymes (Worley et al. 2013). An important difference between yeast and mammalian HDACs is that full activity of the yeast Rpd3L complex requires inositol pyrophosphates (Worley et al. 2013). Substitution mutations in the residues proposed to bind inositol phosphate in Rpd3 (rpd3ibs) led to a pattern of gene expression similar to that of strains that either lack the HDAC (rpd3Δ) or are unable to synthesize inositol pyrophosphates (kcs1Δ vip1Δ) (Worley et al. 2013). In support of this, the Rpd3L target genes were not deacetylated upon osmotic stress in the kcs1∆ vip1∆ mutant. These findings demonstrated that inositol pyrophosphates are required for Rpd3L deacetylase activity (Worley et al. 2013). However, current results from the experiments in yeast cannot determine whether the PP-InsPs are exchangeable and regulate the HDAC activity or serve solely a structural role (Steidle et al. 2020; Worley et al. 2013).
Under normal growth conditions, Msn2 is phosphorylated and sequestered in the cytoplasm with low frequency shuttling into the nucleus; however, upon stress Msn2 is dephosphorylated and shuttles into the nucleus at higher frequency (Cai et al. 2008; Gonze et al. 2008; Jacquet et al. 2003; Petrenko et al. 2013). The siw14∆ mutant shows increased nuclear localization of Msn2-GFP and increased expression of Msn2 target genes under non-stress conditions (Steidle et al. 2020). These results indicated that high levels of PP-InsPs likely enhance activation of the iESR genes through Msn2/4. Further experiments are necessary to determine how inositol pyrophosphates influence Msn2/4 activity, considering that Msn2/4 are regulated by multiple pathways, including TORC1, PKA, Snf1, and Psr1/2-Whi2 (Beck and Hall 1999; Boy-Marcotte et al. 1998; Görner et al. 1998; Kaida et al. 2002; Lee et al. 2013; Smith et al. 1998).
Do inositol pyrophosphates affect signaling through TORC1?
The Target of Rapamycin (TOR) kinase is a key regulator that integrates signals of nutrient abundance and cellular stress to regulate proliferation and stress responses (reviewed in Deprez et al. 2018; Zaman et al. 2008). The TORC1 complex regulates proliferation by affecting ribosome synthesis, gene expression, and translation initiation (Fig. 2). During favorable conditions, TORC1 phosphorylates the S6 kinase Sch9 to activate it (Urban et al. 2007); in turn, Sch9 phosphorylates Dot6 and Tod6 to prevent repression of Ribi genes (Huber et al. 2011). Under stress conditions, TORC1 inhibits Sch9 activity (Urban et al. 2007), thus allowing Dot6/Tod6 to repress RiBi genes. Under favorable conditions, TORC1 also phosphorylates Tap42 to inhibit the activity of the associated Pph21/22 phosphatases (Di Como and Arndt 1996; Wang and Jiang 2003). Upon TORC1 inhibition, the Tap42 complex dissociates allowing for activation of transcription factors, such as Gln3/Gat1 and Msn2/4, and expression of the target genes (Beck and Hall 1999).
Simplified depiction of the TORC1 regulated genes. (Left) Under favorable conditions, TORC1 is active to phosphorylate Tap42. Tap42 interacts with the PP2A phosphatases to inhibit their activity, leading to lower activity of the downstream transcription factors. Active TORC1 phosphorylates the Sch9 kinase to active it; the repressors Dot6/Tod6 are inactivated by Sch9 phosphorylation. (Right) Under unfavorable conditions, TORC1 is inactive, Tap42 releases the PP2A phosphatases that dephosphorylate the downstream targets to activate the transcription factors, and inactive Sch9 does not inhibit the repressors Dot6/Tod6, leading to RiBi gene repression. High levels of inositol pyrophosphates in the siw14Δ led to increased expression of target genes in the Tap42 branch and no difference in the Sch9 branch (Steidle et al. 2020), consistent with partial activation of the Tap42 branch. Lack of inositol pyrophosphates in the kcs1Δ vip1Δ double mutant led to no change in the phosphorylation of Dot6/Tod6 (Worley et al. 2013) under favorable growth conditions
In our analysis of microarray data obtained for an siw14Δ strain that overproduces 5-IP7 and IP8 (Steidle et al. 2020), we noted that approximately 75% of genes misregulated in the absence of stress in the siw14Δ mutant are regulated by transcription factors that are downstream of TORC1. In that report, we focused on the role of Msn2/4 because of the connection with stress-resistance phenotypes and stress-induced transcriptional changes (Steidle et al. 2020). However, target genes for the transcription factors Gln3/Gat1, Rap1, Yap1, Gcn4, Gis1, and Sfp1 also showed increased expression that was not seen in wild-type cells treated with osmotic or oxidative stresses (Fig. 3, Table 1). Increased nuclear localization of Gln3 in the siw14Δ mutant was previously reported (Hirasaki et al. 2010), and our data showing altered expression of target genes is consistent with this. Gln3 is inhibited by TORC1 (Beck and Hall 1999) through the Npr1 kinase (Tate et al. 2006), and the siw14Δ and npr1Δ mutants have opposing effects on Gln3 localization that correspond to increased or decreased activation of Gln3 target genes, respectively (Hirasaki et al. 2007). Multiple studies have found that the siw14Δ mutant is sensitive to caffeine (Alic et al. 2003; Brown et al. 2006; Care et al. 2004; Hirasaki et al. 2007; Romá-Mateo et al. 2011; Sakumoto et al. 2002). Caffeine imposes a cell wall stress through the inhibition of the cell wall integrity pathway as it intersects with TORC1 (Yan et al. 2012). Therefore, the high levels of PP-InsPs present in the siw14Δ strain could partially inhibit TORC1, leading to increased nuclear localization of downstream transcription factors (e.g., Msn2, Gln3) and increased transcription of some of their target genes.

© The authors E.A. Steidle, V.A. Morrissette, K. Fujimaki, L. Chong, A.C. Resnick, A.P. Capaldi, and R.J. Rolfes
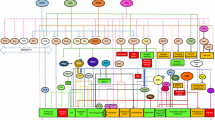
Heat map showing differential gene expression. Heat maps comparing differentially expressed genes from DNA microarray analysis. Differentially expressed genes with lower expression (blue) or higher expression (yellow) on a Log2 scale. There are 444 genes misregulated in the siw14∆ mutant (top bar), aligned with the corresponding genes in the wild-type strain stressed with either 1 mM H2O2 (middle) or 1.3 M KCl (bottom). Genes with increased expression are regulated by Msn2/4 (left portion in the siw14Δ/WT comparison) or by multiple transcription factors—Gln3/Gat1, Rap1, Yap1, Gcn4, Gis1, and Sfp1—that are regulated by TORC1 (under the label “TFs, Table 1”). There are other genes regulated by TORC1 but the factors are not annotated. Finally, some differentially expressed genes are affected by PP-InsPs and are not regulated by TORC1. The gene expression data for this figure was originally published in the Journal of Biological Chemistry. Steidle et al. (2020).
Capaldi and colleagues examined the consequences of the absence of inositol pyrophosphates on the TOR pathway (Worley et al. 2013). They focused on the phosphorylation of the repressors Dot6 and Tod6, as well as expression of RiBi genes in the kcs1Δ vip1Δ double mutant (see Fig. 2). They found that phosphorylation of Dot6/Tod6 in the kcs1Δ vip1Δ double mutant was the same as in the wild-type strain (Worley et al. 2013), indicating that signaling through the Sch9 branch was unaffected by the loss of inositol pyrophosphates. This finding is not unexpected if the signal for stress is increased PP-InsPs, and this interpretation is consistent with the wild-type expression of RiBi genes in the siw14Δ mutant (Steidle et al. 2020). Together, the results from both reports (Steidle et al. 2020; Worley et al. 2013) suggest that inositol pyrophosphates do not affect the Sch9 branch, but instead may affect the Tap42 branch of TORC1 signaling (Fig. 2). The mechanism(s) by which inositol pyrophosphates affect the TOR pathway is not yet known but could be through direct binding of PP-InsPs or pyrophosphorylation. Further research is necessary to determine the mechanism(s) by which inositol pyrophosphates affect TORC1 signaling.
Beyond Saccharomyces
Inositol pyrophosphates regulate and affect cellular responses to stress in not only Saccharomyces but they also play similarly important roles in other eukaryotes. In fungi, the focus has been on developmental changes. Strains of the fungal pathogens Aspergillus nidulans and Ustilago maydis with mutations in the VIP1 ortholog (ASP1) displayed abnormal cell morphologies, defects in polarized cell growth, and defects in filamentation (Pöhlmann et al. 2014). In Schizosaccharomyces pombe, mutations in asp1 (vip1) blocked the dimorphic switch (Pöhlmann and Fleig 2010). In the pathogenic yeast Cryptococcus neoformans, kcs1Δ mutants were avirulent and displayed lower melanization, decreased laccase activity, and increased capsule formation (Lev et al. 2015; Li et al. 2016a; Liao et al. 2018). These results indicate that disruptions in the synthesis of the different types of inositol pyrophosphates has dramatic effects on morphology and virulence. Determining the mechanisms of action will lead to increased understanding of these important cellular processes and provide potentially new therapeutic targets (Kang 2018; Li et al. 2016b; Saiardi et al. 2018).
Inositol pyrophosphates also affect the stress responses in multicellular organisms. The human antiviral immune response requires the production of 1-InsP7 to activate the transcription factor IRF3 and increase expression of Type 1 interferon (Pulloor et al. 2014). Murine skeletal muscle cells express the KCS1 orthologue IP6K3 and gene expression increases in response to nutrient deprivation (fasting) (Moritoh et al. 2016). Murine IP6K3 is also implicated in controlling glucose and insulin levels (Moritoh et al. 2016). Mammalian cells respond to osmotic and thermal stresses by increasing the synthesis of InsP8 (Choi et al. 2005; Pesesse et al. 2004). In plants, inositol pyrophosphates play an important role in polarized growth (Heilmann 2016), jasmonate perception, and responses to herbivorous insects (Laha et al. 2015). Thus, the role(s) for inositol pyrophosphates in stress responses are found broadly across eukaryotes, although there is not a single common response.
Abbreviations
- PtdIns(4,5)P2 :
-
Phosphatidylinositol 4,5-bisphosphate
- IP3 and InsP3 :
-
Inositol (1,4,5) trisphosphate
- IP4 and InsP4 :
-
Inositol (1,3,4,5) tetraphosphate
- IP5 and InsP5 :
-
Inositol (1,3,4,5,6) pentaphosphate
- IP6 and InsP6 :
-
Inositol (1,2,3,4,5,6) hexakisphosphate
- 5-IP7 and 5PP-InsP5 :
-
5-Diphosphoinositol pentakisphosphate
- 1-IP7 and 1PP-InsP5 :
-
1-Diphosphoinositol pentakisphosphate
- IP8 and 1,5PP-InsP4 :
-
1,5-Diphosphoinositol tetrakisphosphate
- IP7 :
-
Either of the two isoforms (5-IP7 or 1-IP7)
- PP-InsPs:
-
General term to refer to all of inositol pyrophosphates together, IP7 and IP8
- HDAC:
-
Histone deacetylase complex
- ESR:
-
Environmental stress response
- iESR:
-
Induced ESR
- rESR:
-
Repressed ESR
- RiBi:
-
Ribosome biogenesis
References
Alejandro-Osorio AL, Huebert DJ, Porcaro DT, Sonntag ME, Nillasithanukroh S, Will JL, Gasch AP (2009) The histone deacetylatse Rpd3p is required for transient changes in genomic expression in response to stress. Genome Biol 10:R57. https://doi.org/10.1186/gb-2009-10-5-r57
Alic N, Higgins VJ, Pichova A, Breitenbach M, Dawes IW (2003) Lipid hydroperoxides activate the mitogen-activated protein kinase Mpk1p in Saccharomyces cerevisiae. J Biol Chem 278:41849–41855. https://doi.org/10.1074/jbc.M307760200
Auesukaree C, Damnernsawad A, Kruatrachue M, Pokethitiyook P, Boonchird C, Kaneko Y, Harashima S (2009) Genome-wide identification of genes involved in tolerance to various enrionmental stresses in Saccharomyces cerevisiae. J Appl Genet 50:301–310. https://doi.org/10.1007/BF03195688
Azevedo C, Burton A, Ruiz-Mateos E, Marsh M, Saiardi A (2009) Inositol pyrophosphate mediated pyrophosphorylation of AP3B1 regulates HIV-1 Gag release. Proc Natl Acad Sci USA 106:21161–21166. https://doi.org/10.1073/pnas.0909176106
Beck T, Hall MN (1999) The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature 402:689–692. https://doi.org/10.1038/45287
Bhandari R et al (2007) Protein pyrophosphorylation by inositol pyrophosphates is a post-translational event. Proc Natl Acad Sci USA 104:15305–15320. https://doi.org/10.1073/pnas.0707338104
Boy-Marcotte E, Perrot M, Bussereau F, Boucherie H, Jacquet M (1998) Msn2p and Msn4p control a large number of genes induced at the diauxic transition which are repressed by cyclic AMP in Saccharomyces cerevisiae. J Bacteriol 180:1044–1052
Brewster JL, Gustin MC (2014) Hog 1: 20 years of discovery and impact. Sci Signal 7:re7. https://doi.org/10.1126/scisignal.2005458
Brown JA et al (2006) Global analysis of gene function in yeast by quantitative phenotypic profiling. Mol Syst Biol 2(2006):0001. https://doi.org/10.1038/msb4100043
Cai L, Dalal CK, Elowitz MB (2008) Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455:485–490. https://doi.org/10.1038/nature07292
Care A, Vousden K, Binley K, Radcliffe P, Trevethick J, Mannazzu I, Sudbery P (2004) A synthetic lethal screen identifies a role for the cortical actin patch/endocytosis complex in the response to nutrient deprivation in Saccharomyces cerevisiae. Genetics 166:707–719. https://doi.org/10.1534/genetics.166.2.707
Celenza J, Carlson M (1984) Cloning and genetic mapping of SNF1, a gene required for expression of glucose-repressible genes in Saccharomyces cerevisiae. Mol Cell Biol 4:49–53
Chanduri M, Rai A, Malla AB, Wu M, Fiedler D, Mallik R, Bhandari R (2016) Inositol hexakisphosphate kinase 1 (IP6K1) activity is required for cytoplasmic dynein-driven transport. Biochem J 473:3031–3047. https://doi.org/10.1042/BCJ20160610
Choi K, Mollapour E, Shears SB (2005) Signal transduction during environmental stress: InsP8 operates within highly restricted contexts. Cell Signal 17:1533–1541. https://doi.org/10.1016/j.cellsig.2005.03.021
Cook JG, Bardwell L, Kron SJ, Thorner J (1996) Two novel targets of the MAP kinase Kss1 are negative regulators of invasive growth in the yeast Saccharomyces cerevisiae. Genes Dev 10:2831–2848
Cullen PJ, Sprague GFJ (2012) The regulation of filamentous growth in yeast. Genetics 190:23–49
Deprez M-A, Eskes E, Winderickx J, Wilms T (2018) The TORC1-Sch9 pathway as a crucial mediator of chronological lifespan in the yeast Saccharomcyes cerevisiae. FEMS Yeast Res 18:foy048. https://doi.org/10.1093/femsyr/foy048
Di Como CJ, Arndt KT (1996) Nutrients, via the Tor proteins, stimulate the associate of Tap42 with type 2A phosphatases. Genes Dev 10:1904–1916. https://doi.org/10.1101/gad.10.15.1904
Dick RA et al (2018) Inositol phosphates are assembly co-factors for HIV-1. Nature 560:509–512. https://doi.org/10.1038/s41586-018-0396-4
Dubois E, Scherens B, Vierendeels F, Ho MM, Messenguy F, Shears SB (2002) In Saccharomyces cerevisiae, the inositol polyphosphate kinase activity of Kcs1p is required for resistance to salt stress, cell wall integrity, and vacuolar morphogenesis. J Biol Chem 277:23755–23763. https://doi.org/10.1074/jbc.M202206200
Enyenihi AH, Saunders WS (2003) Large-scale functional genomic analysis of sporulaiton and meiosis in Saccharomyces cerevisie. Genetics 163:47–54
Erdman S, Snyder M (2001) A filamentous growth response mediated by the yeast mating pathway. Genetics 159:919–928
Eskes E, Deprez M-A, Wilms T, Winderickx J (2017) pH homeostasis in yeast; the phosphate perspective. Curr Genet 64:155–161. https://doi.org/10.1007/s00294-017-0743-2
Gancedo JM (2001) Control of pseudohyphae formation in Saccharomyces cerevisiae. FEMS Microbiol Rev 25:107–123. https://doi.org/10.1111/j.1574-6976.2001.tb00573.x
Garay E, Campos SE, González de la Cruz J, Gaspar AP, Jinich A, Deluna A (2014) High-resolution profiling of stationary-phase survival reveals yeast longevity factors and their genetic interactions. PLoS Genet 10:e100468. https://doi.org/10.1371/journal.pgen.1004168
Gasch AP (2003) The environmental stress response: a common yeast response to diverse environmental stresses. In: Hohmann SMWH (ed) Yeast stress responses: topics in current genetics, vol 1. Springer, Berlin, Heidelberg, pp 11–70
Gasch AP (2007) Comparative genomics of the environmental stress response in ascomycete fungi. Yeast 24:961–976
Gasch AP et al (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11:4241–4257
Gibney PA, Lu C, Caudy AA, Hess DC, Botstein D (2013) Yeast metabolic and signaling genes are required for heat-shock survival and have little overlap with the heat-induced genes. Proc Natl Acad Sci USA 110:E4393–E4402. https://doi.org/10.1073/pnas.1318100110
Gokhale NA, Zaremba A, Janoshazi AK, Weaver JD, Shears SB (2013) PPIP5K1 modulates ligand competition between diphosphoinositol polyphosphates and PtdIns(3,4,5)P3 for polyphosphoinositide-binding domains. Biochem J 453:413–426. https://doi.org/10.1042/BJ20121528
Gonze D, Jacquet M, Goldbeter A (2008) Stochastic modelling of nucleocytoplasmic oscillations of the transcription factor Msn2 in yeast. J R Soc Interface 5:S95–S109. https://doi.org/10.1098/rsif.2008.0141.focus
Görner W et al (1998) Nuclear localization of the C2H2 zinc finger protein Msn2p is regulated by stress and protien kinase A activity. Genes Dev 12:586–597
Heilmann I (2016) Phosphoinositide signaling in plant development. Development 143:2044–2055. https://doi.org/10.1242/dev.136432
Hirasaki M, Kaneko Y, Harashima S (2007) Protein phosphatase Siw14 controls intracellular localization of Gln3 in cooperation with Npr1 kinase in Saccharomyces cerevisiae. Gene 409:34–43
Hirasaki M et al (2010) Deciphering cellular functions of protein phosphatases by comparison of gene expression profiles in Saccharomyces cerevisiae. J Biosci Bioeng 109:433–441
Horigome C, Ikeda R, Okada T, Takenami K, Mizuta K (2009) Genetic interaction between ribosome biogenesis and inositol polyphophate metabolism in Saccharomyces cerevisiae. Biosci Biotech Biochem 73:443–446. https://doi.org/10.1271/bbb.80599
Huber A et al (2011) Sch9 regulates ribosome biogenesis via Stb3, Dot6 and Tod6 and the histone deacetylase complex RPD3L. EMBO J 30:3052–3064. https://doi.org/10.1038/emboj.2011.221
Jacquet M, Renault G, Lallet S, De Mey J, Goldbeter A (2003) Oscillatory nucleocytoplasmic shuttling of the general stress response transcriptional activators Msn2 and Msn4 in Saccharomyces cerevisiae. J Cell Biol 161:497–505. https://doi.org/10.1083/jcb.200303030
Jarolim S et al (2013) Saccharomyces cerevisiae genes involved in survival of heat shock. G3 3:2321–2333. https://doi.org/10.1534/g3.113.007971
Kaida D, Yashiroda H, Toh-e A, Kikuchi Y (2002) Yeast Whi2 and Psr1-phosphatase form a complex and regulate STRE-mediated gene expression. Genes Cells 7:543–552
Kang HA (2018) Emerging roles of inositol pyrophosphates as key modulators of fungal pathogenicity. Virulence 9:563–565. https://doi.org/10.1080/21505594.2017.1421832
Kilari RS, Weaver JD, Shears SB, Safrany ST (2013) Understanding inositol pyrophosphate metabolism and function: kinetic characterization of the DIPPs. FEBS Lett 587:3464–3470. https://doi.org/10.1016/j.febslet.2013.08.035
Kloimwieder A, Winston F (2011) A screen for germination mutants in Saccharomyces cerevisiae. G3 1:143–149. https://doi.org/10.1534/g3.111.000323
Laha D et al (2015) VIH2 regulates the synthesis of inositol pyrophosphate InsP8 and jasmonate-dependent defenses in Arabidopsis. Plant Cell 27:1082–1097. https://doi.org/10.1105/tpc.114.135160
Lee P, Kim MS, Paik S-M, Choi S-H, Cho B-R, Hahn J-S (2013) Rim15-dependent activation of Hsf1 and Msn2/4 transcription factors by direct phosphorylation in Saccharomyces cerevisiae. FEBS Lett 587:3648–3655. https://doi.org/10.1016/j.febslet.2013.10.004
Lee YS, Mulugu S, York JD, O'Shea EK (2007) Regulation of a cyclin-CDK-CDK inhibitor complex by inositol pyrophosphates. Science 316:109–112. https://doi.org/10.1126/science.1139080
Lev S et al (2015) Fungal inositol pyrophosphate IP7 is crucial for metabolic adaptation to the host environment and pathogenicity. MBio 6:e00531–e1515. https://doi.org/10.1128/mBio.00531-15
Li C, Lev S, Saiardi A, Desmarini D, Sorrell TC, Djordjevic JT (2016a) Identification of a major IP5 kinase in Cryptococcus neoformans confirms that PP-IP5/IP7, not IP6, is essential for virulence. Sci Rep 6:23927. https://doi.org/10.1038/srep23927
Li C, Lev S, Saiardi A, Desmarini D, Sorrell TC, Djordjevic JT (2016b) Inositol polyphosphate kinaes, fungal virulence and drug discovery. J Fungi 2:E24. https://doi.org/10.3390/jof2030024
Liao G et al (2018) Role of the inositol pyrophsohate multikinase Kcs1 in Cryptococcus inositol metabolism. Fungal Genet Biol 113:42–51. https://doi.org/10.1016/j.fgb.2018.01.006
Luo HR et al (2003) Inositol pyrophosphates mediate chemotaxis in Dictyostelium via pleckstrin homology domain-PtdIns(3,4,5)P3 interactions. Cell 114:559–572
Macbeth MR, Schubert HL, Vandemark AP, Lingam AT, Hill CP, Bass BL (2005) Inositol hexakisphosphate is bound in the ADAR2 core and required for RNA editing. Science 309:1534–1539. https://doi.org/10.1126/science.1113150
Millard CJ et al (2013) Class I HDACs share a common mechanism of regulation by inositol phosphates. Mol Cell 51:57–67. https://doi.org/10.1016/j.molcel.2013.05.020
Morano KA, Grant CM, Moye-Rowley WS (2012) The response to heat shock and oxidative stress in Saccharomyces cerevisiae. Genetics 190:1157–1195. https://doi.org/10.1534/genetics.111.128033
Moritoh Y, Oka M, Yasuhara Y, Hozumi H, Iwachidow K, Fuse H, Tozawa R (2016) Inositol hexakisphosphate kinase 3 regulates metabolism and lifespan in mice. Sci Rep 6:32072. https://doi.org/10.1038/srep32072
Mulugu S et al (2007) A conserved family of enzymes that phosphorylate inositol hexakisphosphate. Science 316:106–109
Mutlu N, Kumar A (2019) Messengers for morphogenesis: inositol polyphosphate signaling and yeast pseudohyphal growth. Curr Genet 65:119–125. https://doi.org/10.1007/s00294-018-0874-0
Nagata E et al (2010) Inositol hexakisphosphate kinases promote autophagy. Int J Biochem Cell Biol 42:2065–2071. https://doi.org/10.1016/j.biocel.2010.09.013
Neiman AM (2011) Sporulation in the budding yeast Saccharomcyes cerevisiae. Genetics 189:737–765. https://doi.org/10.1534/genetics.111.127126
Norman KL, Shively CA, De La Rocha AJ, Mutlu N, Basu S, Cullen PJ, Kumar A (2018) Inositol polyphosphates regulate and predict yeast pseudohyphal growth phenotypes. PLoS Genet 14:e1007493. https://doi.org/10.1371/journal.pgen.1007493
Onnebo SM, Saiardi A (2009) Inositol pyrophosphates modulate hydrogen peroxide signalling. Biochem J 423:109–118
Ouyang A, Zheng G, Tomchick DR, Luo X, Yu H (2016) Structural basis and IP6 requirement for Pds5-dependent cohesin dynamics. Mol Cell 62:248–259. https://doi.org/10.1016/j.molcel.2016.02.033
Pan X, Heitman J (1999) Cyclic AMP-dependent protein kinase regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Mol Cell Biol 19:4874–4887
Pesesse X, Choi K, Zhang T, Shears SB (2004) Signaling by higher inositol polyphosphates. Synthesis of bisdiphophoinositol tetrakisphosphate ("InsP8") is selectively activated by hyperosmotic stress. J Biol Chem 279:43378–43381. https://doi.org/10.1074/jbc.C400286200
Petrenko N, Chereji RV, McClean MN, Morozov AV, Broach JR (2013) Noise and interlocking signaling pahtways promote distinct transcription factor dynamics in response to different stresses. Mol Biol Cell 24:2045–2057. https://doi.org/10.1091/mbc.E12-12-0870
Pöhlmann J, Fleig U (2010) Asp1, a conserved 1/3 inositol polyphosphate kinase, regulate the dimorphic switch in Schizosaccharomyces pombe. Mol Cell Biol 30:4535–4547
Pöhlmann J et al (2014) The Vip1 inositol polyphosphate kinase family regulates polarized growth and modulates the microtubule cytoskeleton in fungi. PLoS Genet 10:e1004586. https://doi.org/10.1371/journal.pgen.1004586
Pulloor NK et al (2014) Human genome-wide RNAi screen identifies an essential role for inositol pyrophosphates in Type-I interferon response. PLoS Pathog 10:e1003981. https://doi.org/10.1371/journal.ppat.1003981
Ravi C, Gowsalya R, Nachiappan V (2019) Impaired GCR1 transcription resulted in defective inositol levels, vacuolar structure and autophagy in Saccharomyces cerevisiae. Curr Genet 65:995–1014. https://doi.org/10.1007/s00294-019-00954-2
Robertson LS, Fink GR (1998) The three yeast A kinases have specific signaling functions in pseudohyphal growth. Proc Natl Acad Sci USA 95:13783–13787
Romá-Mateo C et al (2011) Phylogenetic and genetic linkage between novel atypical dual-specificity phosphatases from non-metazoan organisms. Mol Genet Genom 285:341–354. https://doi.org/10.1007/s00438-011-0611-6
Ruiz-Roig C, Viéitez C, Posas F, de Nadal E (2010) The Rpd3L HDAC complex is essential for the heat stress response in yeast. Mol Microbiol 76:1049–1062. https://doi.org/10.1111/j.1365-2958.2010.07167.x
Sadeh A, Movshovich N, Volokh M, Ghever L, Aharoni A (2011) Fine tuning of the Msn2/4-mediated yeast stress response as revealed by systematic deletion of Msn2/4 partners. Mol Biol Cell 22:3127–3138. https://doi.org/10.1091/mbc.E10-12-1007
Safrany ST, Ingram SW, Cartwright JL, Falck JR, McLennan AG, Barnes LD, Shears SB (1999) The diadenosine hexaphosphate hydrolases from Schizosaccharomyces pombe and Saccharomyces cerevisiae are homologues of the human diphosphoinositol polyphosphate phosphohydrolase. Overlapping substrate specificities in a MutT-type protein. J Biol Chem 274:21735–21740
Saiardi A, Azevedo C, Desfougères Y, Portela-Torres P, Wilson MSC (2018) Microbial inositol polyphophate metabolic pathway as drug development target. Adv Biol Regul 67:74–83. https://doi.org/10.1016/j.jbior.2017.09.007
Saiardi A, Caffrey JJ, Snyder SH, Shears SB (2000) The inositol hexakisphosphate kinase family. Catalytic flexibility and function in yeast vacuole biogenesis. J Biol Chem 275:24686–24692. https://doi.org/10.1074/jbc.M002750200
Saiardi A, Erdjument-Bromage H, Snowman AM, Tempst P, Snyder SH (1999) Synthesis of diphophoinositol pentakisphosphate by a newly identified family of higher inositol pholyphosphate kinases. Curr Biol 9:1323–1326
Sakumoto N, Matsuoka I, Mukai Y, Ogawa N, Kaneko Y, Harashima S (2002) A series of double disruptants for protein phosphatase genes in Saccharomyces cerevisiae and their phenotypic analysis. Yeast 19:587–599
Shears SB (2015) Inositol pyrophosphates: why so many phosphates? Adv Biol Regul 57:203–216. https://doi.org/10.1016/j.jbior.2014.09.015
Shears SB (2017) Intimate connections: inositol pyrophosphates at the interface of metabolic regulation and cell signaling. J Cell Physiol 233:1897–1912. https://doi.org/10.1002/jcp.26017
Shively CA et al (2015) Large-scale analysis of kinase signaling in yeast pseudohyphal development identifies regulation of ribonucleoprotein granules. PLoS Genet 11:e1005564. https://doi.org/10.1371/journal.pgen.1005564
Sinha H et al (2008) Sequential elimination of major-effect contributors identifies additional quantitative trait loci conditioning high-temperature growth in yeast. Genetics 180:1661–1670. https://doi.org/10.1534/genetics.108.092932
Smith A, Ward MP, Garrett S (1998) Yeast PKA represses Msn2p/Msn4p-dependent gene expression to regulate growth, stress response and glycogen accumulation. EMBO J 17:3556–3564. https://doi.org/10.1093/emboj/17.13.3556
Steidle EA, Chong LS, Wu M, Crooke E, Fiedler D, Resnick AC, Rolfes RJ (2016) A novel inositol pyrophosphate phosphatase in Saccharomyces cerevisiae: Siw14 protein selectively cleaves the b-phosphate from 5-diphosphoinositol pentakisphosphate (5PP-IP5). J Biol Chem 291:6772–6783. https://doi.org/10.1074/jbc.M116.714907
Steidle EA, Morrissette VA, Fujimaki K, Chong L, Resnick AC, Capaldi AP, Rolfes RJ (2020) The InsP7 phosphatase Siw14 regulates inositol pyrophosphate levels to control localization of the general stress response transcription factor Msn2. J Biol Chem 295:2043–2056. https://doi.org/10.1074/jbc.RA119.012148
Szijgyarto Z, Garedew A, Azevedo C, Saiardi A (2011) Influence of inositol pyrophosphates on cellular energy dynamics. Science 334:802–805. https://doi.org/10.1126/science.1211908
Tate JJ, Rai R, Cooper TG (2006) Ammonia-specific regulation of Gln3 localization in Saccharomyces cerevisiae by protein kinase Npr1. J Biol Chem 281:28460–28469. https://doi.org/10.1074/jbc.M604171200
Thota SG, Bhandari R (2015) The emerging roles of inositol pyrophoshates in eukaryotic cell physiology. J Biosci 40:593–605. https://doi.org/10.1007/s12038-015-9549-x
Thota SG, Unnikannan CP, Thampatty SR, Manorama R, Bhandari R (2015) Inositol pyrophosphates regulate RNA polymerase I-mediated rRNA transcription in Saccharomyces cerevisiae. Biochem J 466:105–114. https://doi.org/10.1042/BJ20140798
Urban J et al (2007) Sch9 is a major target of TORC1 in Saccharomyces cerevisiae. Mol Cell 26:663–674. https://doi.org/10.1016/j.molcel.2007.04.020
Vyas VK, Kuchin S, Berkey CD, Carlson M (2003) Snf1 kinases with different beta-subunit isoforms play distinct roles in regulating haploid invasive growth. Mol Cell Biol 23:1342–1348
Wang H, Jiang Y (2003) The Tap42-protein phosphatase type 2A catalytic subunit complex is required for cell cycle-dependent distribution of actin in yeast. Mol Cell Biol 23:3116–3125. https://doi.org/10.1128/MCB.23.9.3116-3125.2003
Watson PJ, Fairall L, Santos GM, Schwabe JW (2012) Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature 481:335–340. https://doi.org/10.1038/nature10728
Watson PJ et al (2016) Insights into the activation mechanism of Class I HDAC complexes by inositol phosphates. Nat Commun 7:11262. https://doi.org/10.1038/ncomms11262
Wickner RB, Edskes HK, Bezsonov EE, Son M, Ducatez M (2018) Prion propagation and inositol polyphosphates. Curr Genet 64:571–574. https://doi.org/10.1007/s00294-017-0788-2
Wild R et al (2016) Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352:986–990. https://doi.org/10.1126/science.aad9858
Wilson MS, Livermore TM, Saiardi A (2013) Inositol pyrophosphates: between signaling and metabolism. Biochem J 452:369–379. https://doi.org/10.1042/BJ20130118
Worley J, Luo X, Capaldi AP (2013) Inositol pyrophosphates regulate cell growth and the environmental stress response by activating the HDAC Rpd3L. Cell Rep 3:1476–1482. https://doi.org/10.1016/j.celrep.2013.03.043
Yan G, Lai Y, Jiang Y (2012) The TOR complex 1 is a direct target of Rho1 GTPase. Mol Cell 45:743–753. https://doi.org/10.1016/j.molcel.2012.01.028
Yang X, Safrany ST, Shears SB (1999) Site-directed mutagenesis of diphosphoinositol polyphosphate phosphohydrolase, a dual specificity NUDT enzyme that attacks diadenosine polyphosphates and diphosphoinositol polyphosphates. J Biol Chem 274:35434–35440. https://doi.org/10.1074/jbc.274.50.35434
Zaman S, Lippman SI, Zhao X, Broach JR (2008) How Saccharomyces responds to nutrients. Annu Rev Genet 42:27–81
Acknowledgements
We thank Alan Hinnebusch, Andrew Capaldi, and Elizabeth Steidle for comments on the manuscript. This work was supported by funds from the Georgetown University College of Arts and Sciences.
Funding
This work was supported by Georgetown University College of Arts & Sciences for publication costs.
Author information
Authors and Affiliations
Contributions
Analysis of DNA microarray results and comparison with transcription factor profiles were performed by VAM; figures were prepared by VAM; literature search was conducted by VAM and RJR; drafting and editing the text was conducted by VAM and RJR; project development was overseen by RJR; and, Current Genetics editor MK approached RJR to write this mini-review.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Availability of data and material
Microarray data have been previously published and are accessible through GEO Series number GSE135546.
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Morrissette, V.A., Rolfes, R.J. The intersection between stress responses and inositol pyrophosphates in Saccharomyces cerevisiae. Curr Genet 66, 901–910 (2020). https://doi.org/10.1007/s00294-020-01078-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01078-8