Abstract
In this mini-review, we present a perspective on the recent findings relating Spo0M structure and function that will stimulate and guide further studies in the characterization of this interesting protein. Cell division and sporulation constitute two of the best studied processes in the model organism Bacillus subtilis; however, there are many missing pieces in the giant regulatory puzzle that governs the independent and shared networks between them. Spo0M is a little studied protein that has been related to both, cell division and sporulation, but its biochemical function and its direct interactions have not been yet defined. Structural analysis of Spo0M revealed the presence of an arrestin-like domain and an FP domain (a dimerization domain present in proteasome elements), motifs more commonly found in eukaryotic proteins. The aim of this perspective is to present open questions regarding the functional and structural features of Spo0M that make this protein a good candidate for the ancestor of arrestins in bacteria and an important element in developmental and differentiation processes of Bacillus subtilis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Spo0M background
Spo0M is a regulator of sporulation whose role has recently been extended to include cell division in Bacillus subtilis (Vega-Cabrera et al. 2017). Interestingly, this protein contains motifs that were only previously found in eukaryotes, and the analysis of the mutant phenotype, the possible interaction partners of the protein, and its structure, could shed light on the role of this multi-functional protein in bacteria and explain its presence in several genera of non-sporulating species.
Adaptive bacterial responses to environmental stress have led to the evolution of highly robust cell survival mechanisms. Sporulation constitutes a cell differentiation program that bacteria activate to contend with harsh conditions and promote the survival of its progeny. Sporulation in Bacillus subtilis is a strictly regulated process that begins with the phosphorylation of the master regulator Spo0A by sensor histidine kinases KinA-B in response to environmental and metabolic signals (Al-Hinai et al. 2015). The level of phosphorylation of Spo0A regulates a continuous transition between different pathways established to assure the survival of the bacteria, including biofilm formation, motility or sporulation, which is the ultimate fate (De Hoon et al. 2010; Grau et al. 2015).
After the decision to sporulate is taken, at stage zero, asymmetric cell division proceeds generating a small prespore and a large mother cell. From this point, two different genetic programs are established in each compartment that direct the pathway they will follow until the release of the mature spore (Piggot and Hilbert 2004). Many of the key regulators of the sporulation program have been studied in depth and the mechanisms governing activation of successive stages along the sporulation pathway are well characterized. Despite this, gaps in our knowledge remain, possibly due to lack of phenotype when proteins with redundant functions are mutated.
Our knowledge of Spo0M function represents one of these gaps. Almost 20 years ago, Spo0M was first described as a sporulation control protein after its role in this process of Bacillus subtilis was demonstrated in the absence and overexpression of the protein. Loss of function mutation of Spo0M resulted in lower expression of the master regulator Spo0A (Han et al. 1998), although how this effect is mediated is unknown. Intriguingly, homologs of Spo0M are present in several non-sporulating bacterial species (Vega-Cabrera et al. 2017), which hints at further roles for Spo0M beyond regulation of sporulation.
Recently, it was demonstrated that Spo0M participates in the cell division process of Bacillus subtilis, constituting the first report of its function in the vegetative stage of the bacteria. Spo0M null mutants, besides being affected in sporulation, are elongated and have membrane and nucleoid anomalies, when compared to wild-type cells. Using immunoprecipitation analysis, it was determined that Spo0M is in the same molecular complex as the septum proteins FtsZ and ZapA, and co-localization experiments demonstrated overlap in their intracellular spatial distributions (Vega-Cabrera et al. 2017). It remains to be shown whether these interactions are relevant to the cell division defects observed in the spo0M mutants and could yet provide the basis to explain the presence of Spo0M in non-sporulating bacteria.
Moreover, proteomic analysis revealed a sizable group of proteins that include proteases, chaperones and late sporulation proteins, that apparently interact with Spo0M (Vega-Cabrera et al. 2017). It remains to be determined which of those interactions are crucial for Spo0M function and in which mechanisms, yet such a diverse array of potential interaction partners, does hint at a complex and probably multi-functional role for Spo0M in bacterial cell biology. Insights from the structure of Spo0M may prove informative in this regard.
A bacterial protein with eukaryotic features
Recently, the structure of Spo0M was published. Surprisingly, Sp0M was found to contain two domains with similarity to motifs which were only previously found in eukaryotic proteins: an N-terminal arrestin-like domain and a C-terminal region with homology to the FP dimerization domain of the human proteasome inhibitor PI31 subunit (Sonoda et al. 2015).
The arrestin-like domain and its relation to the cell division/sporulation phenotype
The arrestin protein is a ubiquitous family of eukaryotic proteins that are involved in endocytosis, receptor desensitization, signal transduction and regulation of gene expression among others activities (Lefkowitz 2013). This family of proteins is divided into two categories: α-arrestins or visual arrestins, which include proteins that contain arrestin-like domains, such as ARRDC1-4, TXNIP or Vsp26, and β-arrestin proteins, both sharing greater structural similarities than sequence identity (Aubry and Klein 2013). Until now, it was thought that the arrestin family of proteins is only present in eukaryotes. Arrestins have two domains, a C-terminal domain involved in dimerization and endocytosis-related protein interactions, and an N-terminal domain involved in receptor recognition. Arrestin proteins have been shown to interact with a wide range of binding partners, as befits their diversity of function (Gurevich and Gurevich 2013).
As previously mentioned, a null mutant in spo0M is impaired in sporulation and cell division, and has membrane and nucleoid abnormalities when compared with the wild-type strain (Vega-Cabrera et al. 2017). Cell division defects could be in part explained by the interaction of Spo0M with a molecular complex that includes the cell division proteins FtsZ and ZapA. However, the decondensed nucleoid and the wide distribution of cell lengths of the elongated mutant strain are more difficult to explain. Among the possible binding partners of Spo0M, its interaction with DivIVA is worth exploring in this respect.
DivIVA is a topological determinant that is active during both, vegetative growth and sporulation (Fig. 1). DivIVA recruits the MinCDJ system to the cell poles and the septum region preventing the formation of aberrant septa during the vegetative stage (Fig. 1a) (Eswaramoorthy et al. 2011). During sporulation, DivIVA recruits RacA and facilitates its interaction with oriC, promoting the formation of the axial filament (Fig. 1b) (Eswaramoorthy et al. 2014). In a recent report, it was demonstrated that DivIVA forms a molecular complex at the cell poles along with the MinD/MinJ, ComN, and Spo0J/Soj (homologs of ParA/ParB) proteins that is necessary for the proper segregation of the chromosome during vegetative growth and sporulation (Fig. 1c) (Kloosterman et al. 2016). Most probably, the element responsible for the chromosome segregation is the complex Spo0J/Soj which could interact with specific regions of the DNA near the oriC and the structural chromosome maintenance (SCM) system to ensure the proper distribution of the chromosome and regulate its condensation state (Touzain et al. 2011).
DivIVA protein complexes that regulate different stages of the B. subtilis life cycle. a During the vegetative stage, DivIVA localizes the MinCDJ system to the cell poles and the nascent septum to avoid formation of aberrant septa by inhibition of the assembly of the Z ring. b During sporulation, DivIVA recruits RacA to the cell poles and facilitates assembly of the axial filament. c During sporulation and vegetative cell division, DivIVA forms a cell pole protein complex that is necessary for the proper segregation of the chromosomes. N-terminal arrestin-like domain of Spo0M could participate in this protein complex through interaction with the PDZ domain of MinJ
The arrestin domain of Spo0M could mediate an interaction with the pole protein complex (Fig. 1c), as it was reported that β-arrestins associate with PDZ domain-containing proteins (Gallon et al. 2014). PDZ domain function has been linked to the interaction of receptor proteins in the plasma membrane with cytoskeletal elements (Lee and Zheng 2010) and the organization of signal transduction complexes (Jeleń et al. 2003). The PDZ domain is present in MinJ, but the biochemical function of this domain and the entire protein is still under study.
Additional support for this hypothesis comes from (1) the location pattern of Spo0M being very similar to that of DivIVA/MinCDJ (Strahl and Hamoen 2012; Vega-Cabrera et al. 2017); (2) MinJ interacts with several elements of the divisome (Bramkamp and van Baarle 2009); (3) minJ mutants are filamentous and form septa, albeit of aberrant function or irregularly located, and cells produce approximately 30% fewer spores than the wild-type strain (Meeske et al. 2016) these latter traits are similar to those reported for Spo0M mutants.
The interaction of Spo0M with a PDZ domain could also explain the poor resistance of the spores generated by the spo0M mutant strain. Resistance to heat, desiccation, radiation, and chemical hazards is a well-known characteristic of wild-type B. subtilis spores. In the spo0M null mutant, this diminished resistance was attributed to a possible involvement of Spo0M in advanced stages of spore maturation (Vega-Cabrera et al. 2017), and not just at stage zero, as previously reported (Han et al. 1998). In addition, proteins involved in cortex and coat formation, and germination, were identified as possible interactors with Spo0M (Vega-Cabrera et al. 2017).
Proteins that form the coat and cortex (the resistant structures of the spores) and elements necessary for germination are synthesized from the mother cell-specific factors SigmaE and SigmaK (Driks and Eichenberger 2016). Specifically, SigmaK is produced as a pro-sigma factor that requires processing to attain activity by the serine proteases SpoIVB and CtpB, of which both, incidentally, contain PDZ domains (Mastny et al. 2013).
The arrestin–PDZ domain interaction is not restricted to β-arrestins, but it is also shared by the Vps26 α-arrestin, an element of the heterotrimeric complex of the retromer that mediates recycling of proteins from endosomes to the trans Golgi network and the plasma membrane. The groove region between the two arrestin domains of Vps26 interacts with the PDZ domain of sorting nexin 27 (SNX27); another component of the retromer. This conservative interaction between different members of the arrestin family and PDZ domain-containing proteins could be shared by the N-terminal domain of Spo0M and would support its function as a true arrestin.
Finally, another conserved activity of the arrestin family of proteins in Spo0M could be the regulation of metabolism. α and β-Arrestins are related to different metabolic cues (Xiao et al. 2010). Of special interest is the involvement of TXNIP in glucose metabolism. This α-arrestin is highly overexpressed in diabetes, which makes it a good target for strategies to control this disease (Patwari et al. 2009). In a similar fashion, it has been reported that Spo0M might interact with the carbon metabolism regulator CcpA (Wünsche et al. 2012), but the physiological implications of this interaction and the participation of the N-terminal arrestin-like domain of Spo0M should be studied.
The FP domain and the stress response
Sonoda et al. (2015) have proposed that the homology of the FP domain of the PI31 protein in Spo0M could be indicative of a common evolutionary origin and could allow Spo0M to form dimers (Sonoda et al. 2015). FP domains have been identified in the PI31 and the F-box 7 proteins, both proteasome elements, and it is known that they facilitate their homo- and heterodimerization (Shang et al. 2014).
In eukaryotes, the proteasome system contributes to the quality control mechanism of protein synthesis; it constitutes a compartmentalized complex of proteases that degrade misfolded proteins (often tagged with a ubiquitin mark). The proteasome is composed of the core particle (CP) 20S that contains the active cleavage site, and the regulatory particle (RP) 19S that removes the ubiquitin tag, unfolds the proteins, and directs them to the 20S CP (Livneh et al. 2016). Bacteria lack a homolog of the proteasome with the exception of some members of the Actinomycetales and Nitrospirales orders that have structurally similar proteasomes to eukaryotes, but the assembly and activation paths of the subunits are different and the ubiquitination mark is substituted for an analogous pupylation signal (from Prokaryotic Ubiquitin-like Protein) (Jastrab and Darwin 2016).
Notwithstanding the absence of a proteasome, most bacteria contain quality control systems that eliminate misfolded proteins and contribute to the regulation of specific protein abundance, such as the heat-shock response or complexes of AAA + ATP-dependent proteases.
The heat-shock response is a ubiquitous homeostasis maintenance system that helps the cells eliminate misfolded proteins. In B. subtilis, the chaperones DnaK, GroESL, and the SigmaB regulon are activated in the heat-shock response. Both chaperones have been reported as possible interactors of Spo0M (Vega-Cabrera et al. 2017) and both generate a filamentous phenotype when mutated in E. coli due to its possible interaction with the cell division protein FtsZ (Sugimoto et al. 2008). On the other hand, the Sigma B regulon is kept inactive in physiological conditions by the anti-sigma factor RbsW (Benson and Haldenwang 1993), a kinase also reported to be an interactor of Spo0M. In heat-shock and other physical stresses like acid, salt, and ethanol exposure, RsbW is sequestered by the anti-anti-sigma factor RsbV that releases the active SigmaB factor that goes on to regulate the transcription of approximately 200 genes that contend with general conditions of stress (Hecker et al. 2007). Given that Spo0M has been reported to be a member of the SigmaW stress response regulon (Zweers et al. 2012), it is possible that its FP domain contributes to this function through regulation of the “proteasome-like” response in bacteria. Sustained stress can lead bacteria to sporulate, and participation in the stress response may explain the role of Spo0M at the beginning of this differentiation process.
AAA + ATP-dependent proteases recognize their substrate through the ATPase domain; they unfold the protein and transfer it to the catalytic domain. Some complexes contain additional adaptor proteins that are modified in response to environmental or developmental signals that direct the proteolysis of particular substrates (Battesti and Gottesman 2013). An example of this type of complex is the ClpCP system that is a possible interactor of Spo0M (Vega-Cabrera et al. 2017) and is involved in competence, motility, and sporulation (Msadek et al. 1998).
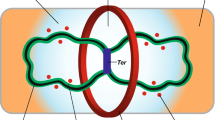
A notable example of a protein belonging to both, the heat-shock response and the specific protease system, is the ATP-dependent metalloprotease FtsH. It is known that FtsH degrades Spo0M in vitro and it is proposed that this activity maintains the optimal levels of Spo0M abundance in vivo (Thi Nguyen and Schumann 2012). This protease is induced in heat shock and is involved in sporulation, cell division, and biofilm formation (Yepes et al. 2012); moreover, it has been suggested that FtsH could degrade EzrA (Mielich-Süss et al. 2013) the main negative regulator of the polymerization of FtsZ in vivo (Land et al. 2014). It could be that Spo0M activity is regulated by a feedback loop with FtsH through its FP domain (Fig. 2), which might in turn explain part of the spo0M mutant phenotype, such as the sporulation defect or the filamentation resulting from increased levels of EzrA. Although twofold overexpression of EzrA in wild-type cells has no apparent effect (Haeusser et al. 2004), raising EzrA levels beyond this level has to have an impact on cell regulation, which we speculate could be the reason behind the partial phenotype observed in the spo0M mutant, where just approximately 30% of the cells are elongated (Vega-Cabrera et al. 2017). Furthermore, EzrA is linked to cell morphology through regulation of PBP1 (Fig. 2) (Claessen et al. 2008) and the serine/threonine protein kinase PrkC (Pompeo et al. 2015), both also reported to be possible interactors of Spo0M (Vega-Cabrera et al. 2017). The participation of Spo0M in a complex that regulates EzrA could also explain the cell membrane abnormalities present in the spo0M mutant.
Schematic representation of the proposed participation of Spo0M in a regulatory feedback loop with FtsH. FtsH is a metalloprotease that degrades Spo0M in vitro and possibly maintains its physiological levels in vivo. FtsH is involved in sporulation, cell division, and biofilm formation and it has been reported to degrade EzrA, the main negative regulator of FtsZ polymerization that is also involved in cell morphology maintenance through regulation of PBP1. The C-terminal domain of Spo0M, with homology to an FP domain, could mediate a regulatory feedback loop with FtsH, which would explain some of the alterations observed in the spo0M null mutant, for example sporulation, elongation, and cell membrane defects
It has been recently proposed that proteins which share an evolutionary origin and conserved structural motifs could preserve functional interaction through these structural elements (Derouiche et al. 2016). Spo0M could represent an example of this phenomenon, preserving the capacity of its arrestin and FP domain to interact with potential partners that have so far been observed only in eukaryotic organisms. The functionality of the FP domain should be experimentally tested and differentiated from the function of the arrestin-like domain. Full domain deletions or loss of function mutations would help to corroborate or eliminate the proposals presented.
Conclusions and future remarks
Assigning a specific function to a newly discovered protein is an arduous task that requires analysis of its structure, genomic context, genomic origins, homologies, mutant phenotype, structure, and interactors. Even when all these features have been analyzed, it is still often impossible to entirely define all such functions that a particular protein may possess. In this mini-review we have presented our perspective of the relation between the structure of Spo0M and its role in multiple processes during the life cycle of B. subtilis. The functional characterization of Spo0M is likely to be complicated, as it contains two domains with not-known homologs in bacteria and a partial, but pleiotropic phenotype. We propose that analyses of the possible interactors in the context of the homologies of the identified Spo0M domains constitute the best approach to determine the functions of Spo0M. We speculate that Spo0M will share functional as well as structural homology to eukaryotic arrestins and could be considered the bacterial ancestor of this family of proteins with important regulatory functions.
Without doubt, much work remains to be performed before we may definitively assign a function or functions to Spo0M. Once achieved, the broad distribution of Spo0M among bacteria genera may enable targeting of specific physiological and developmental programs, like cell division, in pathogenic bacteria that carry a Spo0M copy such as Vibrio cholerae, to mention just one example.
References
Al-Hinai MA, Jones SW, Papoutsakis ET (2015) The Clostridium sporulation programs: diversity and preservation of endospore differentiation. Microbiol Mol Biol Rev 79:19–37. doi:10.1128/MMBR.00025-14
Aubry L, Klein G (2013) True arrestins and arrestin-fold proteins: a structure-based appraisal. In: Khalil RA (ed) Progress in molecular biology and translational science, vol 118. Elsevier Inc, London, UK, pp 21–56
Battesti A, Gottesman S (2013) Roles of adaptor proteins in regulation of bacterial proteolysis. Curr Opin Microbiol 16:140–147. doi:10.1038/jid.2014.371
Benson AK, Haldenwang WG (1993) Bacillus subtilis SigmaB is regulated by a binding protein (RsbW) that blocks its association with core RNA polymerase. Proc Natl Acad Sci USA 90:2330–2334
Bramkamp M, van Baarle S (2009) Division site selection in rod-shaped bacteria. Curr Opin Microbiol 12:683–688. doi:10.1016/j.mib.2009.10.002
Claessen D, Emmins R, Hamoen LW et al (2008) Control of the cell elongation-division cycle by shuttling of PBP1 protein in Bacillus subtilis. Mol Microbiol 68:1029–1046. doi:10.1111/j.1365-2958.2008.06210.x
De Hoon MJL, Eichenberger P, Vitkup D (2010) Hierarchical evolution of the bacterial sporulation network. Curr Biol. doi:10.1016/j.cub.2010.06.031
Derouiche A, Shi L, Kalantari A, Mijakovic I (2016) Evolution and tinkering: what do a protein kinase, a transcriptional regulator and chromosome segregation/cell division proteins have in common? Curr Genet 62:67–70. doi:10.1007/s00294-015-0513-y
Driks A, Eichenberger P (2016) The spore coat. Microbiol Spectr 4(2). doi:10.1128/microbiolspec.TBS-0023-2016
Eswaramoorthy P, Erb ML, Gregory JA et al (2011) Cellular architecture mediates DivIVA ultrastructure and regulates min activity in Bacillus subtilis. MBio. doi:10.1128/mBio.00257-11
Eswaramoorthy P, Winter PW, Wawrzusin P et al (2014) Asymmetric division and differential gene expression during a bacterial developmental program requires DivIVA. PLoS Genet. doi:10.1371/journal.pgen.1004526
Gallon M, Clairfeuille T, Steinberg F et al (2014) A unique PDZ domain and arrestin-like fold interaction reveals mechanistic details of endocytic recycling by SNX27-retromer. Proc Natl Acad Sci USA 111:E3604–E3613. doi:10.1073/pnas.1410552111
Grau RR, De Oña P, Kunert M et al (2015) A duo of potassium-responsive histidine kinases govern the multicellular destiny of Bacillus subtilis. MBio. doi:10.1128/mBio.00581-15
Gurevich VV, Gurevich EV (2013) Structural determinants of arrestin function. In: Khalil RA (ed) Progress in molecular biology and translational science, vol 118. Elsevier Inc, London, UK, pp 57–92
Haeusser DP, Schwartz RL, Smith AM et al (2004) EzrA prevents aberrant cell division by modulating assembly of the cytoskeletal protein FtsZ. Mol Microbiol 52:801–814. doi:10.1111/j.1365-2958.2004.04016.x
Han W-D, Kawamoto S, Hosoya Y et al (1998) A novel sporulation-control gene (spo0M) of Bacillus subtilis with a σH-regulated promoter. Gene 217:31–40. doi:10.1016/S0378-1119(98)00378-3
Hecker M, Pané-Farré J, Uwe V (2007) SigB-dependent general stress response in Bacillus subtilis and related gram-positive bacteria. Annu Rev Microbiol 61:215–236. doi:10.1146/annurev.micro.61.080706.093445
Jastrab JB, Darwin KH (2016) Bacterial proteasomes. Annu Rev Microbiol 69:109–127. doi:10.1038/nbt.3121.ChIP-nexus
Jeleń F, Oleksy A, Śmietana K (2003) PDZ domains—common players in the cell signaling. Acta Biochim Pol 50(4):985–1017
Kloosterman TG, Lenarcic R, Willis CR et al (2016) Complex polar machinery required for proper chromosome segregation in vegetative and sporulating cells of Bacillus subtilis. Mol Microbiol 101:333–350. doi:10.1111/mmi.13393
Land AD, Luo Q, Levin PA (2014) Functional domain analysis of the cell division inhibitor EzrA. PLoS One 9:e102616. doi:10.1371/journal.pone.0102616
Lee H-J, Zheng JJ (2010) PDZ domains and their binding partners: structure, specificity, and modification. Cell Commun Signal 8:8. doi:10.1186/1478-811X-8-8
Lefkowitz RJ (2013) Arrestins come of age: a personal historical perspective. In: Khalil RA (ed) Progress in molecular biology and translational science, vol 118. Elsevier Inc, London, UK, pp 3–18
Livneh I, Cohen-kaplan V, Cohen-rosenzweig C et al (2016) The life cycle of the 26S proteasome: from birth, through regulation and function, and onto its death. Cell Res 26:869–885. doi:10.1038/cr.2016.86
Mastny M, Heuck A, Kurzbauer R et al (2013) XCtpB assembles a gated protease tunnel regulating cell–cell signaling during spore formation in Bacillus subtilis. Cell. doi:10.1016/j.cell.2013.09.050
Meeske AJ, Rodrigues CDA, Brady J et al (2016) High-throughput genetic screens identify a large and diverse collection of new sporulation genes in Bacillus subtilis. PLoS Biol. doi:10.1371/journal.pbio.1002341
Mielich-Süss B, Schneider J, Lopez D (2013) Overproduction of flotillin influences cell differentiation and shape in Bacillus subtilis. MBio 4:e00719-13. doi:10.1128/mBio.00719-13
Msadek T, Dartois V, Kunst F et al (1998) ClpP of Bacillus subtilis is required for competence development, motility, degradative enzyme synthesis, growth at high temperature and sporulation. Mol Microbiol 27:899–914. doi:10.1046/j.1365-2958.1998.00735.x
Patwari P, Chutkow WA, Cummings K et al (2009) Thioredoxin-independent regulation of metabolism by the α-arrestin proteins. J Biol Chem 284:24996–25003. doi:10.1074/jbc.M109.018093
Piggot PJ, Hilbert DW (2004) Sporulation of Bacillus subtilis. Curr Opin Microbiol 7:579–586. doi:10.1016/j.mib.2004.10.001
Pompeo F, Foulquier E, Serrano B et al (2015) Phosphorylation of the cell division protein GpsB regulates PrkC kinase activity through a negative feedback loop in Bacillus subtilis. Mol Microbiol 97:1–12. doi:10.1111/mmi.13015
Shang J, Wang G, Yang Y et al (2014) Structure of the FP domain of Fbxo7 reveals a novel mode of protein–protein interaction. Acta Crystallogr Sect D Biol Crystallogr 70:155–164. doi:10.1107/S1399004713025820
Sonoda Y, Mizutani K, Mikami B (2015) Structure of Spo0M, a sporulation-control protein from Bacillus subtilis. Acta Crystallogr Sect F Struct Biol Commun 71:1488–1497. doi:10.1107/S2053230X15020919
Strahl H, Hamoen LW (2012) Finding the corners in a cell. Curr Opin Microbiol 15:731–736. doi:10.1016/j.mib.2012.10.006
Sugimoto S, Saruwatari K, Higashi C, Sonomoto K (2008) The proper ratio of GrpE to DnaK is important for protein quality control by the DnaK–DnaJ–GrpE chaperone system and for cell division. Microbiology 154:1876–1885. doi:10.1099/mic.0.2008/017376-0
Thi Nguyen HB, Schumann W (2012) The sporulation control gene spo0M of Bacillus subtilis is a target of the FtsH metalloprotease. Res Microbiol 163:114–118. doi:10.1016/j.resmic.2011.10.011
Touzain F, Petit M-A, Schbath S, El Karoui M (2011) DNA motifs that sculpt the bacterial chromosome. Nat Rev Microbiol 9:15–26. doi:10.1038/nrmicro2477
Vega-Cabrera LA, Guerrero A, Rodríguez-Mejía JL et al (2017) Analysis of Spo0M function in Bacillus subtilis. PLoS One 12:e0172737. doi:10.1371/journal.pone.0172737
Wünsche A, Hammer E, Bartholomae M et al (2012) CcpA forms complexes with CodY and RpoA in Bacillus subtilis. FEBS J 279:2201–2214. doi:10.1111/j.1742-4658.2012.08604.x
Xiao K, Sun J, Kim J et al (2010) Global phosphorylation analysis of beta-arrestin-mediated signaling downstream of a seven transmembrane receptor (7TMR). Proc Natl Acad Sci USA 107:15299–15304. doi:10.1073/pnas.1008461107
Yepes A, Schneider J, Mielich B et al (2012) The biofilm formation defect of a Bacillus subtilis flotillin-defective mutant involves the protease FtsH. Mol Microbiol 86:457–471. doi:10.1111/j.1365-2958.2012.08205.x
Zweers JC, Nicolas P, Wiegert T et al (2012) Definition of the σW regulon of Bacillus subtilis in the absence of stress. PLoS One. doi:10.1371/journal.pone.0048471
Acknowledgements
We thank Dr. Enrique Merino and Dr. Rosa María Gutiérrez-Ríos for enriching discussions regarding the genetic context, homology, and physiology of Spo0M. We thank Shirley Ainsworth for bibliographical assistance. This work was supported partially by CONACyT 176381 and DGAPA IN 204016. Luz Adriana Vega-Cabrera was supported by a CONACyT and DGAPA scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Vega-Cabrera, L.A., Wood, C.D. & Pardo-López, L. Spo0M: structure and function beyond regulation of sporulation. Curr Genet 64, 17–23 (2018). https://doi.org/10.1007/s00294-017-0718-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0718-3