Abstract
Bacterial cell division is a highly controlled process regulated accurately by a diverse array of proteins spatially and temporally working together. Among these proteins, FtsZ is recognized as a cytoskeleton protein because it can assemble into a ring-like structure called Z-ring at midcell. Z-ring recruits downstream proteins, thus forming a multiprotein complex termed the divisome. When the Z-ring scaffold is established and the divisome matures, peptidoglycan (PG) biosynthesis and chromosome segregation are triggered. In this review, we focus on multiple interactions between FtsZ and its accessory proteins in bacterial cell cytokinesis, including FtsZ localization, Z-ring formation and stabilization, PG biosynthesis, and chromosome segregation. Understanding the interactions among these proteins may help discover superior targets on treating bacterial infectious diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Most bacteria divide and proliferate through binary fission by splitting into two new daughter cells with equal sizes. A large protein complex called divisome is substantially formed at the future division site. The core component of divisome is the conserved tubulin homologue FtsZ, which regulates bacterial cytokinesis mainly with the help of a series of accessory proteins appearing in a hierarchical manner.
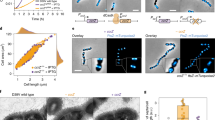
The Min and nucleoid occlusion (NO) system proteins inhibit FtsZ assembly in other places except midcell and localize this protein to the nascent division plane (Fig. 1) to ensure that two progeny cells are equal in size (Shih and Zheng 2013; Schumacher and Zeng 2016). FtsZ monomers then polymerize into single protofilament and form a ring-like structure (Z-ring) at midcell (Fig. 1). Zap proteins stabilize lateral interactions among FtsZ protofilaments and promote FtsZ assembly into an integrated Z-ring (Galli and Gerdes 2010; Durand-Heredia et al. 2011; Roach et al. 2016). This ring is also stabilized by anchor proteins FtsA and ZipA, which tether FtsZ to the inner face of the plasma membrane. Z-ring also acts as a dynamic scaffold to recruit downstream proteins with critical regulatory functions of cytokinesis, such as peptidoglycan (PG) remodeling and chromosome segregation (den Blaauwen et al. 2017).
Schematic model of Min system, nucleoid occlusion (NO) system, and Z-ring. Min system forms bipolar gradients that suppress FtsZ (red spheres) polymerization at both polar sites. As an important protein of NO system, SlmA (green) stretches along the non-terminus-containing (Ter) region of chromosome (black) to inhibit FtsZ assembly. Min and NO systems drive FtsZ monomers to assemble into Z-ring at midcell where Ter region (blue) is located
The function of FtsZ and its accessory proteins has been clarified, but their coordinated roles must be further explored. In this review, we summarize the potential properties of FtsZ and its accessory proteins and their multiple interactions to help discover possible targets for the development of novel antimicrobials.
FtsZ localization
FtsZ shares high structural similarity to tubulin because of their ancestral homology. The structure of FtsZ consists of an N-terminal domain (NTD) and C-terminal domain (CTD) linked by central core H7 helices. A globular structure exists within the NTD, which contains the nucleotide-binding site (Ortiz et al. 2016). Following the H7 helices is the catalytic T7 loop at the base of the CTD. The CTD can also be divided into an intrinsically disordered C-terminal linker and a C-terminal conserved peptide, which play vital roles in recognizing several FtsZ accessory proteins, such as FtsA, ZipA, SlmA, MinC and ZapD (Schumacher and Zeng 2016; Sundararajan and Goley 2017; Vega and Margolin 2019; Blasios et al. 2013; Huang et al. 2016). When the T7 loop of an FtsZ monomer inserts into the nucleotide-binding site of another FtsZ monomer, the GTPase-active site for GTP hydrolysis is formed at the junction of two FtsZ monomers (Matsui et al. 2014). Linear protofilaments are created when GTP binds to the globular domains of FtsZ monomers.
Min system
MinC and MinD synergistically suppress Z-ring assembly at both polar sites. Considering the roles of these proteins, MinD may function as trucks, whereas MinC serve as cargos. MinC is a critical effector protein in the Min system with two different functional domains. The NTDs of MinC can inhibit the assembly of FtsZ monomers into protofilaments, and the CTDs can weaken the association among FtsZ protofilaments (LaBreck et al. 2019; Park et al. 2018). MinD is tethered to the membrane when bound to ATP, forming a polar zone at one side of the two cell poles. MinC then attaches to ATP–MinD through its CTD, forming a MinCD complex that binds to the CTD of FtsZ (Conti et al. 2015). In this case, the concentration of MinCD is sufficiently high at one cell pole, eventually preventing a division septum from forming at the polar site. In this process, MinD may interfere with cell division by acting as a MinC propeller.
MinE is designated as a restriction factor against the MinCD complex, which guarantees that MinCD only functions at both cell poles. In Escherichia coli (E. coli), the MinD and MinE proteins self-organize into a pole-to-pole standing wave oscillator on the membrane to facilitate division initiation at midcell (Conti et al. 2015; Park et al. 2017). At the polar zone, MinC outcompetes MinE for binding with MinD because of its strong binding activity. Thus, MinC can maintain the inhibition of FtsZ assembly at the polar site. When the polar zone covers half of the cell, MinE assembles into an E-ring, which inhibits zone extension, and moves towards the polar site. During this movement, MinE activates the ATPase activity of MinD, which then separates from the membrane. When the polar zone is completely disassembled, the E-ring also dissociates (Loose et al. 2011). Subsequently, MinC, MinD and MinE appear at the opposite pole again and conduct the same procedure. Thus, the old zone disappears at one site, and a new zone develops at the other site. This periodicity of oscillatory behaviour averages approximately 20–50 s. In brief, the MinDE complex is suggested to serve as a reaction–diffusion device that drives oscillation, whereas MinC is like a passenger in this oscillation process (Mizuuchi and Vecchiarelli 2018). Given the pole-to-pole oscillation activity of these Min proteins, the MinC concentration is regulated to be high at polar sites but low at the midcell by the MinDE complex, which ensures the assembly of FtsZ at the midcell.
NO system
In E. coli, the NO system is mediated by the SlmA protein, an FtsZ antagonist and DNA-binding protein. SlmA binds to the specific SlmA-binding sequences (SBSs) dispersed on the chromosome, except for the terminus-containing (Ter) region stretched along the cell’s short axis (Fig. 1) (Schumacher and Zeng 2016; Tonthat et al. 2013). Given that the Ter region is the last chromosomal region to split, only FtsZ is allowed to polymerize at the midcell. The activated SlmA–SBS complex can bind to the CTD of FtsZ without affecting its GTPase activity. However, by stimulating FtsZ–GDP formation, SlmA–SBSs promote the disassembly of preassembled FtsZ polymers (Tonthat et al. 2013; Cabre et al. 2015; Cho et al. 2011). After SlmA binds with DNA, its conformational change and its antagonistic activity are enhanced by the SBSs. In Bacillus subtilis (B. subtilis) and Staphylococcus aureus, the Noc proteins act as SlmA and participate in the NO system (Pang et al. 2017).
Z-ring formation and stabilization
Early in cytokinesis, FtsZ polymers migrate to the midcell and then assemble into a highly dynamic Z-ring at the division site (Fig. 1). The Z-ring may contain a cluster of short and overlapping filaments that are roughly perpendicular to the long axis of the cell (Huecas et al. 2017). These single-stranded filaments can assemble laterally and form bundles or sheets with the help of other regulatory proteins, such as ZipA, ZapA and ZapC (Guan et al. 2018; Ruiz-Martinez et al. 2018; Bhattacharya et al. 2015). Before cell division is initiated, FtsZ filaments on the inner membrane are arranged loosely and are not completely aligned. The Z-ring also exhibits a loose helical structure rather than a closed ring. However, when division begins, FtsZ filaments condense, and the Z-ring becomes a tight structure (Viola et al. 2017; Wang and Wingreen 2013).
ZapA
ZapA consists of an N-terminal globular head domain and a C-terminal coiled-coil domain in B. subtilis, E. coli and Pseudomonas aeruginosa (Galli and Gerdes 2010; Nogueira et al. 2015). The amino acid residue K42 within the N-terminal globular domain of ZapA is required to interact directly with residues K51 and K66, which are close to the GTP-binding site of FtsZ (Roseboom et al. 2018). The tetramer of ZapA cross-links adjacent FtsZ protofilaments and promotes Z-ring stability by inhibiting GTPase activity (Ruiz-Martinez et al. 2018; Low et al. 2004). However, GTPase activity is not affected after ZapA directly binds to FtsZ in Caulobacter crescentus, indicating that the interaction site of ZapA and FtsZ varies across different bacterial species (Woldemeskel et al. 2017).
ZapB
ZapB exists as a dimer that contains a 100% coiled-coil domain. High-resolution 3D reconstruction images indicate that ZapB forms a highly ordered structure concentric to the Z-ring (Soderstrom and Daley 2017). ZapB indirectly binds to FtsZ via ZapA, the N terminus of which is responsible for its interaction with ZapA. The dimers of ZapB cross-link ZapA molecules between two FtsZ protofilaments, which further stabilize Z-ring assembly (Buss et al. 2017; Galli and Gerdes 2012). ZapB directly connects with the MatP protein, which binds to the Ter region of the chromosomes. MatP, ZapB and ZapA interact with one another in a sequential order and form a MatP–ZapB–ZapA structure, which anchors the Ter region of the chromosomes to the Z-ring (Mannik et al. 2016; Espeli et al. 2012). ZapB serves as a linker to stabilize the Z-ring and coordinate chromosome segregation between the chromosome and Z-ring within this structure.
ZapC
ZapC is a small cytoplasmic monomeric protein. It does not share any sequence identity to ZapA or ZapB but exhibits similar phenotypic and functional characteristics to other Zap proteins (Durand-Heredia et al. 2011; Schumacher et al. 2016). As an early divisome protein, ZapC co-localizes with FtsZ at the midcell in an FtsZ-dependent manner. Both the N- and C-domains of ZapC possess hydrophobic cavities that serve as FtsZ-binding sites. ZapC binds close to the GTPase globular core of FtsZ with high affinity through its hydrophobic pockets. ZapC can bind and stabilize FtsZ protofilaments by inhibiting GTPase activity of full-length FtsZ polymers. Superfluous ZapC leads to lethal filamentous morphology or cell division block (Bhattacharya et al. 2015; Ortiz et al. 2015; Hale et al. 2011).
ZapD
ZapD forms a symmetrical dimeric structure that consists of an α-helical domain and β-strand domain in E. coli. Residues R116, R221 and R225 in ZapD are critical for forming a positively charged binding pocket, which is required for bundling FtsZ protofilaments (Roach et al. 2016; Schumacher et al. 2017). As a molecular cross-linking reagent, ZapD binds to the CTD of FtsZ with a set of ZapD arginine residues. Similar to ZapC, FtsZ is also required for ZapD to localize at the midcell and inhibit GTPase activity. ZapD is also believed to participate in the condensation of loose FtsZ filaments (Roach et al. 2016; Huang et al. 2016; Choi et al. 2016). ZapD overexpression causes the separation of FtsZ away from the midcell, leading to elongated cells.
FtsA and ZipA
In many bacterial species, FtsZ filaments are tethered to the cytoplasmic membrane by interacting with the anchor proteins FtsA and ZipA, forming a functional cytokinetic ring. FtsA is an actin-related protein that can insert itself into the bacterial membrane through its C-terminal amphipathic helix. ZipA, a bitopic membrane protein restricted to Gammaproteobacteria, integrates into the membrane through its N-terminal transmembrane domain (Krupka et al. 2019; Pichoff and Lutkenhaus 2005). Cross-linking data show that FtsA and ZipA interact directly with each other through the exposed surface of the FtsA helix 7 (Vega and Margolin 2019). Both FtsA and ZipA bind to the conserved CTD of FtsZ.
The conformation of the FtsA protein can affect the formation of the Z-ring and the process of bacterial division. Purified E. coli FtsA protein can be assembled into a mini-ring structure to inhibit FtsZ bundling and ring formation (Krupka and Margolin 2018; Krupka et al. 2017). The gain-of-function variant protein FtsA* cannot be normally polymerized, and a mini-ring structure cannot be formed on the lipid monolayer, thereby suppressing the inhibition of the FtsZ bundling process. The monomeric FtsA protein can promote FtsZ bundling and recruitment of downstream division proteins into the Z-ring. In normal bacteria, ZipA can prevent the FtsA protein from polymerizing to maintain monomeric forms or oligomers (Schoenemann et al. 2018; Pichoff et al. 2012). The E. coli FtsA protein can remodel the bacterial membrane structure and is essential for membrane division. Given that the C-terminal amphipathic helix of FtsA can be inserted into membrane phospholipids, FtsA protein conformation changes when FtsA binds to ATP; this change causes cell membrane deformation, which eventually leads to contraction of the middle membrane (Conti et al. 2018).
Cytokinesis after divisome assembly
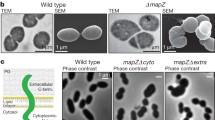
The Z-ring functions as a scaffold and recruits another set of conserved proteins in a hierarchical and roughly temporal order when Z-ring is assembled at the division site with the help of the aforementioned FtsZ localization and stabilization proteins (Fig. 2). This large protein complex is defined as a divisome or division machine, which is not a static structure. FtsZ filaments within the Z-ring bind with PG synthases, treadmill on the inner face of the cell membrane and promote PG synthesis (Bisson-Filho et al. 2017; Ramirez-Diaz et al. 2018). Besides, the ring indirectly binds to chromosomes and participates in its segregation.
Overview of the apparatus of divisome. Z-ring, which is composed of FtsZ protofilaments, is anchored to the membrane by FtsA and ZipA within the divisome. Peptidoglycan (PG) synthase (PBP1b–FtsW–FtsI subcomplex) is linked to the Z-ring by FtsQLB subcomplex, dominating PG synthesis. PG synthesis will not be initiated until FtsN appears and relieves the inhibition of PG synthase by FtsQLB. Transmembrane FtsEX is responsible for septal PG hydrolysis when the cell wall needs to split. FtsKc and MatP–ZapB–ZapA bind to the Ter region and participate in chromosome segregation. The Fts family proteins are represented by capital letters
PG biosynthesis
PG is one of the main components of the bacterial cell wall. During cell division, PG biosynthesis occurs at the midcell mainly under the control of the downstream proteins FtsW, FtsI, FtsQLB, FtsN and FtsEX (Soderstrom et al. 2019). FtsW is an integral membrane protein that belongs to the shape, elongation, division and sporulation (SEDS) family. FtsW facilitates the transport of cytoplasmic lipid II, the PG precursor, into the existing cell wall for PG synthesis. FtsW also acts as a PG polymerase, which polymerizes lipid II into PG (Taguchi 2019; Perez et al. 2019). FtsI is a penicillin-binding protein (PBP3) with monofunctional transpeptidase activity. In E. coli, FtsI is required for septal PG synthesis in coordination with PBP1b, which demonstrates both transglycosylase and transpeptidase activities. FtsW directly interacts with FtsI and PBP1b, forming a PBP1b–FtsW–FtsI synthase subcomplex within the divisome (Boes et al. 2019; Fraipont et al. 2011; Wissel and Weiss 2004). FtsQ, FtsL and FtsB are all bitopic membrane proteins that assemble into an FtsQLB subcomplex. Without enzymatic activity, FtsQLB appears to serve as a link between the Z-ring and PG synthase (Condon et al. 2018; Kureisaite-Ciziene et al. 2018; Villanelo et al. 2011). FtsQLB also inhibits the glycosyltransferase activity of PBP1b and the transpeptidase domain of PBP3 and eventually suppresses PG synthesis. FtsN is the last protein that arrives at the division site, which is believed to trigger cell constriction. When FtsN accumulation at the midcell relieves the inhibition of PBP1b and FtsI by FtsQLB, PG synthesis starts (Weiss 2015; Liu et al. 2015; Pichoff et al. 2018).
FtsE (a cytoplasmic ATPase) and FtsX (an integral membrane protein) form a widely conserved ATP-binding cassette transporter-like complex, which functions as transmembrane regulator for septal PG hydrolysis. When PG synthesis is complete, the newly synthesized cell wall must be equally split into two daughter cells by hydrolytic enzymes called amidases (Arends et al. 2009; Yang et al. 2011). In E. coli, the amidases AmiA, AmiB and AmiC are responsible for cleaving bonds between stem peptides and glycan strands at the septum. To hydrolyse PG efficiently, these amidases must be activated by EnvC (for AmiA and AmiC) and by NlpD (for AmiC), both of which contain the critical LytM domains for cell separation. EnvC is recruited to the division site by FtsEX (Ercoli et al. 2015; Rued et al. 2019; Dubey and Priyadarshini 2018).
Chromosome segregation
DNA replication and chromosome segregation are crucial for bacterial reproduction. Several proteins are associated with this process, such as the aforementioned SlmA and MatP. During cytokinesis, FtsK is another key protein responsible for chromosome segregation. FtsK is a membrane-bound DNA translocase that belongs to the FtsK/SpoIIIE/Tra family. The multifunctional FtsK coordinates chromosome unlinking, segregation and cytokinesis. The structure of FtsK is mainly divided into three domains, namely the N-terminal domain (FtsKN), linker domain (FtsKL) and CTD (FtsKC) (Grainge 2010; Berezuk et al. 2018). FtsKN is a membrane anchor domain with four transmembrane helices and is involved in septum constriction. FtsKL is poorly conserved with various length sequences among species and is required for the recruitment of FtsI. FtsKC is an ATP-dependent DNA translocase and facilitates the disassembly of chromosome dimers in coordination with the XerCD/dif resolvase system (Sherratt et al. 2010; Misra et al. 2018). The dif sites are located in the Ter region of the chromosome, which is close to the midcell. Upon stimulation by FtsK, the tyrosine recombinases XerC and XerD resolve the dimeric chromosomes into monomers at the dif sites. The chromosome monomers can then be segregated into daughter cells (Kennedy et al. 2008; Galli et al. 2017).
Conclusions and perspectives
Cytokinesis is important to living organisms because any aberrant fission leads to abnormal cell morphology or even cell death. Bacterial cell division is a sophisticated process, in which the emergence of proteins is strictly ordered, and the amount of proteins is precisely regulated. FtsZ plays a major role during cytokinesis by recruiting other accessory proteins to immobilize and stabilize the Z-ring and then accurately triggering and regulating bacterial cytokinesis.
The position of Z-ring needs to be limited precisely at the future division site by the Min system and NO system. The Min system proteins, namely MinC, MinD and MinE, and the NO system protein SlmA are responsible for directing FtsZ localization to midcell. FtsZ cannot function alone during bacterial cytokinesis. A diverse array of accessory proteins appears in a hierarchical manner and binds to FtsZ. Zap proteins, namely ZapA, ZapB, ZapC and ZapD, stabilize lateral interactions among FtsZ protofilaments and facilitate the Z-ring formation. The Z-ring, tethered on the inner membrane by FtsA and ZipA, acts as a dynamic scaffold for recruiting downstream proteins with regulatory functions of cytokinesis, forming a multiprotein machine termed the divisome. When the divisome matures, it dominates septum PG synthesis, chromosome segregation, and triggers the constrictive force for cell division.
This review focuses on the potential properties of FtsZ and its accessory proteins, as well as their multiple interactions. Investigating this problem may help reveal bacterial binary fission in depth and develop novel cytokinesis inhibitors in the future.
References
Arends SJ, Kustusch RJ, Weiss DS (2009) ATP-binding site lesions in FtsE impair cell division. J Bacteriol 191(12):3772–3784
Berezuk AM et al (2018) Outer membrane lipoprotein RlpA is a novel periplasmic interaction partner of the cell division protein FtsK in Escherichia coli. Sci Rep 8(1):12933
Bhattacharya A et al (2015) ZapC promotes assembly and stability of FtsZ filaments by binding at a different site on FtsZ than ZipA. Int J Biol Macromol 81:435–442
Bisson-Filho AW et al (2017) Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division. Science 355(6326):739–743
Blasios V et al (2013) Genetic and biochemical characterization of the MinC-FtsZ interaction in Bacillus subtilis. PLoS One 8(4):e60690
Boes A et al (2019) Regulation of the peptidoglycan polymerase activity of PBP1b by antagonist actions of the core divisome proteins FtsBLQ and FtsN. MBio 10(1):e01912-18
Buss JA et al (2017) ZapA and ZapB form an FtsZ-independent structure at midcell. Mol Microbiol 104(4):652–663
Cabre EJ et al (2015) The nucleoid occlusion SlmA protein accelerates the disassembly of the FtsZ protein polymers without affecting their GTPase activity. PLoS One 10(5):e0126434
Cho H et al (2011) Nucleoid occlusion factor SlmA is a DNA-activated FtsZ polymerization antagonist. Proc Natl Acad Sci USA 108(9):3773–3778
Choi H et al (2016) Structural and biochemical studies reveal a putative FtsZ recognition site on the Z-ring stabilizer ZapD. Mol Cells 39(11):814–820
Condon S et al (2018) The FtsLB subcomplex of the bacterial divisome is a tetramer with an uninterrupted FtsL helix linking the transmembrane and periplasmic regions. J Biol Chem 293(5):1623–1641
Conti J, Viola MG, Camberg JL (2015) The bacterial cell division regulators MinD and MinC form polymers in the presence of nucleotide. FEBS Lett 589(2):201–206
Conti J, Viola MG, Camberg JL (2018) FtsA reshapes membrane architecture and remodels the Z-ring in Escherichia coli. Mol Microbiol 107(4):558–576
den Blaauwen T, Hamoen LW, Levin PA (2017) The divisome at 25: the road ahead. Curr Opin Microbiol 36:85–94
Dubey A, Priyadarshini R (2018) Amidase activity is essential for medial localization of AmiC in Caulobacter crescentus. Curr Genet 64(3):661–675
Durand-Heredia JM et al (2011) Identification and characterization of ZapC, a stabilizer of the FtsZ ring in Escherichia coli. J Bacteriol 193(6):1405–1413
Ercoli G et al (2015) LytM proteins play a crucial role in cell separation, outer membrane composition, and pathogenesis in nontypeable Haemophilus influenzae. MBio 6(2):e02575
Espeli O et al (2012) A MatP-divisome interaction coordinates chromosome segregation with cell division in E. coli. EMBO J 31(14):3198–3211
Fraipont C et al (2011) The integral membrane FtsW protein and peptidoglycan synthase PBP3 form a subcomplex in Escherichia coli. Microbiology 157(Pt 1):251–259
Galli E, Gerdes K (2010) Spatial resolution of two bacterial cell division proteins: ZapA recruits ZapB to the inner face of the Z-ring. Mol Microbiol 76(6):1514–1526
Galli E, Gerdes K (2012) FtsZ-ZapA-ZapB interactome of Escherichia coli. J Bacteriol 194(2):292–302
Galli E et al (2017) Fast growth conditions uncouple the final stages of chromosome segregation and cell division in Escherichia coli. PLoS Genet 13(3):e1006702
Grainge I (2010) FtsK–a bacterial cell division checkpoint? Mol Microbiol 78(5):1055–1057
Guan F et al (2018) Lateral interactions between protofilaments of the bacterial tubulin homolog FtsZ are essential for cell division. Elife 7:35578
Hale CA et al (2011) Identification of Escherichia coli ZapC (YcbW) as a component of the division apparatus that binds and bundles FtsZ polymers. J Bacteriol 193(6):1393–1404
Huang KH et al (2016) Characterization of the FtsZ C-Terminal variable (CTV) region in Z-ring assembly and interaction with the Z-ring stabilizer ZapD in E. coli cytokinesis. PLoS One 11(4):e0153337
Huecas S et al (2017) Self-organization of FtsZ polymers in solution reveals spacer role of the disordered C-terminal tail. Biophys J 113(8):1831–1844
Kennedy SP, Chevalier F, Barre FX (2008) Delayed activation of Xer recombination at dif by FtsK during septum assembly in Escherichia coli. Mol Microbiol 68(4):1018–1028
Krupka M, Margolin W (2018) Unite to divide: oligomerization of tubulin and actin homologs regulates initiation of bacterial cell division. F1000Res 7:235
Krupka M et al (2017) Escherichia coli FtsA forms lipid-bound minirings that antagonize lateral interactions between FtsZ protofilaments. Nat Commun 8:15957
Krupka M et al (2019) Escherichia coli ZipA organizes FtsZ polymers into dynamic ring-like protofilament structures. MBio 9(3):e01008-18
Kureisaite-Ciziene D et al (2018) Structural analysis of the interaction between the bacterial cell division proteins FtsQ and FtsB. MBio 9(5):e01346-18
LaBreck CJ et al (2019) MinC N- and C-domain interactions modulate FtsZ assembly, division site selection, and MinD-dependent oscillation in Escherichia coli. J Bacteriol 201(4):e00374-18
Liu B et al (2015) Roles for both FtsA and the FtsBLQ subcomplex in FtsN-stimulated cell constriction in Escherichia coli. Mol Microbiol 95(6):945–970
Loose M et al (2011) Min protein patterns emerge from rapid rebinding and membrane interaction of MinE. Nat Struct Mol Biol 18(5):577–583
Low HH, Moncrieffe MC, Lowe J (2004) The crystal structure of ZapA and its modulation of FtsZ polymerisation. J Mol Biol 341(3):839–852
Mannik J et al (2016) The role of MatP, ZapA and ZapB in chromosomal organization and dynamics in Escherichia coli. Nucleic Acids Res 44(3):1216–1226
Matsui T et al (2014) Structural change in FtsZ Induced by intermolecular interactions between bound GTP and the T7 loop. J Biol Chem 289(6):3501–3509
Misra HS et al (2018) Interdependence of bacterial cell division and genome segregation and its potential in drug development. Microbiol Res 208:12–24
Mizuuchi K, Vecchiarelli AG (2018) Mechanistic insights of the Min oscillator via cell-free reconstitution and imaging. Phys Biol 15(3):031001
Nogueira ML et al (2015) Backbone and side chain NMR assignments of Geobacillus stearothermophilus ZapA allow identification of residues that mediate the interaction of ZapA with FtsZ. Biomol NMR Assign 9(2):387–391
Ortiz C et al (2015) Crystal structure of the Z-ring associated cell division protein ZapC from Escherichia coli. FEBS Lett 589(24 Pt B):3822–3828
Ortiz C et al (2016) The keepers of the ring: regulators of FtsZ assembly. FEMS Microbiol Rev 40(1):57–67
Pang T et al (2017) The nucleoid occlusion factor Noc controls DNA replication initiation in Staphylococcus aureus. PLoS Genet 13(7):e1006908
Park KT et al (2017) MinE conformational dynamics regulate membrane binding, MinD interaction, and Min oscillation. Proc Natl Acad Sci USA 114(29):7497–7504
Park KT et al (2018) MinC and FtsZ mutant analysis provides insight into MinC/MinD-mediated Z ring disassembly. J Biol Chem 293(16):5834–5846
Perez AJ et al (2019) Movement dynamics of divisome proteins and PBP2x:FtsW in cells of Streptococcus pneumoniae. Proc Natl Acad Sci USA 116(8):3211–3220
Pichoff S, Lutkenhaus J (2005) Tethering the Z ring to the membrane through a conserved membrane targeting sequence in FtsA. Mol Microbiol 55(6):1722–1734
Pichoff S et al (2012) FtsA mutants impaired for self-interaction bypass ZipA suggesting a model in which FtsA’s self-interaction competes with its ability to recruit downstream division proteins. Mol Microbiol 83(1):151–167
Pichoff S, Du S, Lutkenhaus J (2018) Disruption of divisome assembly rescued by FtsN-FtsA interaction in Escherichia coli. Proc Natl Acad Sci USA 115(29):E6855–E6862
Ramirez-Diaz DA et al (2018) Treadmilling analysis reveals new insights into dynamic FtsZ ring architecture. PLoS Biol 16(5):e2004845
Roach EJ et al (2016) Structure and mutational analyses of Escherichia coli ZapD reveal charged residues involved in FtsZ filament bundling. J Bacteriol 198(11):1683–1693
Roseboom W et al (2018) Mapping the contact sites of the escherichia coli division-initiating proteins FtsZ and ZapA by BAMG cross-linking and site-directed mutagenesis. Int J Mol Sci 19(10):2928
Rued BE et al (2019) Structure of the large extracellular loop of FtsX and Its Interaction with the essential peptidoglycan hydrolase PcsB in Streptococcus pneumoniae. MBio 10(1):e02622-18
Ruiz-Martinez A et al (2018) Efficient models of polymerization applied to FtsZ ring assembly in Escherichia coli. Proc Natl Acad Sci USA 115(19):4933–4938
Schoenemann KM et al (2018) Gain-of-function variants of FtsA form diverse oligomeric structures on lipids and enhance FtsZ protofilament bundling. Mol Microbiol 109(5):676–693
Schumacher MA, Zeng W (2016) Structures of the nucleoid occlusion protein SlmA bound to DNA and the C-terminal domain of the cytoskeletal protein FtsZ. Proc Natl Acad Sci USA 113(18):4988–4993
Schumacher MA et al (2016) Structural and functional analyses reveal insights into the molecular properties of the Escherichia coli Z ring stabilizing protein. ZapC. J Biol Chem 291(5):2485–2498
Schumacher MA et al (2017) Structure of the Z ring-associated protein, ZapD, bound to the C-terminal domain of the tubulin-like protein, FtsZ, suggests mechanism of Z ring stabilization through FtsZ cross-linking. J Biol Chem 292(9):3740–3750
Sherratt DJ et al (2010) The Escherichia coli DNA translocase FtsK. Biochem Soc Trans 38(2):395–398
Shih YL, Zheng M (2013) Spatial control of the cell division site by the Min system in Escherichia coli. Environ Microbiol 15(12):3229–3239
Soderstrom B, Daley DO (2017) The bacterial divisome: more than a ring? Curr Genet 63(2):161–164
Soderstrom B, Chan H, Daley DO (2019) Super-resolution images of peptidoglycan remodelling enzymes at the division site of Escherichia coli. Curr Genet 65(1):99–101
Sundararajan K, Goley ED (2017) The intrinsically disordered C-terminal linker of FtsZ regulates protofilament dynamics and superstructure in vitro. J Biol Chem 292(50):20509–20527
Taguchi A et al (2019) FtsW is a peptidoglycan polymerase that is functional only in complex with its cognate penicillin-binding protein. Nat Microbiol 4:587–594
Tonthat NK et al (2013) SlmA forms a higher-order structure on DNA that inhibits cytokinetic Z-ring formation over the nucleoid. Proc Natl Acad Sci USA 110(26):10586–10591
Vega DE, Margolin W (2019) Direct interaction between the Two Z ring membrane anchors FtsA and ZipA. J Bacteriol 201(4):e00579-18
Villanelo F et al (2011) A model for the Escherichia coli FtsB/FtsL/FtsQ cell division complex. BMC Struct Biol 11:28
Viola MG et al (2017) Proteolysis-dependent remodeling of the tubulin homolog FtsZ at the division septum in Escherichia coli. PLoS One 12(1):e0170505
Wang S, Wingreen NS (2013) Cell shape can mediate the spatial organization of the bacterial cytoskeleton. Biophys J 104(3):541–552
Weiss DS (2015) Last but not least: new insights into how FtsN triggers constriction during Escherichia coli cell division. Mol Microbiol 95(6):903–909
Wissel MC, Weiss DS (2004) Genetic analysis of the cell division protein FtsI (PBP3): amino acid substitutions that impair septal localization of FtsI and recruitment of FtsN. J Bacteriol 186(2):490–502
Woldemeskel SA et al (2017) A conserved coiled-coil protein pair focuses the cytokinetic Z-ring in Caulobacter crescentus. Mol Microbiol 105(5):721–740
Yang DC et al (2011) An ATP-binding cassette transporter-like complex governs cell-wall hydrolysis at the bacterial cytokinetic ring. Proc Natl Acad Sci USA 108(45):E1052–E1060
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Nos. 81673477, 81471997 and 81001460).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, M., Fang, C., Ma, B. et al. Regulation of cytokinesis: FtsZ and its accessory proteins. Curr Genet 66, 43–49 (2020). https://doi.org/10.1007/s00294-019-01005-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-01005-6