Abstract
Over the last decade, advances in transcriptomics have revealed that the pervasive transcription of eukaryotic genomes produces plethora of long noncoding RNAs (lncRNAs), which are now recognized as major regulators of multiple cellular processes. Although they have been thought to lack any protein-coding potential, recent ribosome-profiling data indicate that lncRNAs can interact with the translation machinery, leading to the production of functional peptides in some cases. In this perspective, we have explored the idea that translation can be part of the fate of cytoplasmic lncRNAs, raising the possibility for them to work as bifunctional RNAs, endowed with dual coding and regulatory functions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction: the arrival on the scene of the non-coding RNAs
With protein-coding genes representing only 2 % of the human genome, the other 98 % have been considered for a long time as inactive material, regions of several mega-bases without any function, so-called ‘junk DNA’ (Taft et al. 2007). However, the overwhelming development of high-density micro-arrays and high-throughput sequencing technologies, as well as of bioinformatics analyses, has enabled to go deeper into transcriptomes, revealing that eukaryotic genomes are pervasively transcribed (Berretta and Morillon 2009). For instance, the ENCODE project revealed that up to 75 % of the human genome is transcribed in at least one cell line or condition (Djebali et al. 2012).
The pervasive transcription of eukaryotic genomes produces thousands of non-coding transcripts, that are commonly classified into small and long (l)ncRNAs, and that are now recognized as major regulators involved in multiple cellular processes, including cell differentiation and development, chromosome dosage compensation, imprinting, regulation of gene expression, cell cycle control and adaptation to environment changes (Rinn and Chang 2012; Sole et al. 2015; Wery et al. 2011). In addition, lncRNAs show tissue-specificity (Djebali et al. 2012), indicating that their expression is tightly regulated. Furthermore, dysregulation of ncRNAs has been associated to human diseases, such as cancer or neurodegenerative disorders (Taft et al. 2010).
In most eukaryotes, small and long regulatory ncRNAs coexist and even cooperate. However, the budding yeast Saccharomyces cerevisiae constitutes an exception to this paradigm as it has lost the RNA interference system and is, therefore, devoid of small-interfering (si)RNAs and micro (mi)RNAs (Drinnenberg et al. 2009). In this respect, S. cerevisiae is a unique model to specifically study the regulatory effects of lncRNAs, which in other organisms might be partially hidden by the effects of the small RNAs (such as siRNAs and miRNAs).
Cytoplasmic lncRNAs are targeted by a translation-dependent surveillance pathway
Over the last years, thousands of lncRNAs have been annotated in S. cerevisiae (Tisseur et al. 2011). Strikingly, most of them were found to be cryptic due to their rapid and extensive degradation in the nucleus or in the cytoplasm (Tisseur et al. 2011; Tudek et al. 2015). Notably, the nuclear and cytoplasmic RNA degradation machineries target distinct types of lncRNAs. For instance, the nuclear exosome-dependent 3′–5′ decay pathway degrades the so-called Cryptic Unstable Transcripts (CUTs), a class of lncRNAs that are mainly transcribed from divergent bidirectional promoters (Neil et al. 2009; Xu et al. 2009). On the other hand, the cytoplasmic Xrn1-dependent 5′–3′ RNA decay pathway is specialized into the degradation of another class of lncRNAs referred to as Xrn1-sensitive unstable transcripts (XUTs), most of which are antisense to protein-coding genes (Van Dijk et al. 2011).
XUTs are synthesized by RNA Polymerase II as capped and poly-adenylated transcripts, similar to mRNAs (Van Dijk et al. 2011; Wery et al. 2016). But what determines their instability has remained unclear until the recent finding that the majority of them are specifically targeted by the Nonsense-Mediated Decay (NMD) pathway (Malabat et al. 2015; Wery et al. 2016). NMD is a translation-dependent RNA decay pathway that targets mRNAs with aberrant translation termination, such as mRNAs with premature stop codon (Muhlrad and Parker 1994) and long 3′-UTR (Muhlrad and Parker 1999). The sensitivity of XUTs to NMD suggests that once in the cytoplasm, they associate to the translation machinery and undergo at least a pioneer round of translation. Supporting this hypothesis, ribosome profiling data revealed the presence of small open reading frames (smORFs) on yeast lncRNAs (Smith et al. 2014), including a large set of XUTs (Malabat et al. 2015; Wery et al. 2016), and some of them were shown to be translated into peptides in vivo (Smith et al. 2014). Strikingly, ribosomes on NMD-sensitive XUTs are restricted to a short 5′-proximal region, followed by a long ribosome-free 3′-UTR, which probably constitutes the NMD-activating signal (Wery et al. 2016).
Notably, ribosome-profiling approaches also identified smORFs on transcripts annotated as non-coding in other Eukaryotes, including Drosophila (Aspden et al. 2014), zebrafish (Bazzini et al. 2014) and mouse (Ingolia et al. 2011), although there is debate on the extent to which ribosome-footprints detection reflects genuine on-going translation (Chew et al. 2013; Guttman et al. 2013).
The observation of ribosome binding to smORFs on transcripts annotated as lncRNAs challenges the initial assumption that these transcripts are really noncoding and raises the fundamental question of the function of the peptides produced upon translation of such smORFs. In this regard, recent works described lncRNAs producing smORFs peptides that control heart activity in Drosophila (Magny et al. 2013) and mammals (Nelson et al. 2016), or cell movement during embryogenesis in zebrafish (Pauli et al. 2014). In yeast, the evolutionary conservation of a subset of lncRNAs smORFs within yeast species indicates that the encoded peptides might have biological importance (Smith et al. 2014).
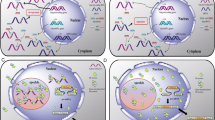
Thus, many transcripts initially thought to lack coding potential are likely to bear smORFs, that can be translated and give rise to functional peptides (Fig. 1). On the other hand, we speculate that a fraction of smORFs-bearing transcripts, reminiscent of the yeast NMD-sensitive XUTs, will also be targeted by the NMD in other eukaryotic cells (Fig. 1). In this respect, NMD inhibition in mouse embryonic stem cells has been shown to result in stabilization of a subset of annotated lncRNAs (Smith et al. 2014). Conceptually, one can also imagine that a cryptic transcript targeted to the NMD in one condition could escape the NMD and be stabilized in another condition (see below), possibly giving rise to a functional smORF peptide. Additional work will be required to define the comprehensive landscape of NMD-sensitive lncRNAs in different eukaryote models, but given the extent of ribosome association to lncRNAs, we anticipate the NMD to be recognized in the future as a major regulator of cytoplasmic lncRNAs.
Translation of lncRNAs smORFs can lead to RNA degradation or functional peptide production. Translation of smORF (red box) on a transcript annotated as lncRNA might target this transcript to the degradation via the NMD pathway, or alternatively give rise to the production of a small peptide, possibly functional (see examples in main text)
NMD as an additional layer in lncRNA-mediated regulation of gene expression
In yeast, antisense XUTs can regulate paired-sense gene expression, at the transcriptional level, through histone modifications (Berretta et al. 2008; Van Dijk et al. 2011) and constitute to date the only class of lncRNAs for which the associated gene-regulation is thought to depend on the lncRNA per se, rather than its transcription. Interestingly, NMD specifically and exclusively targets this class of regulatory lncRNAs (Wery et al. 2016) and might be considered in that way as a novel player in the lncRNA-dependent buffering of genome expression.
NMD not only acts as a surveillance pathway targeting aberrant mRNAs and cryptic lncRNAs to degradation, but it also directly regulates physiological mRNAs in yeast, Drosophila and human (Peccarelli and Kebaara 2014). NMD itself is tightly regulated, and its activity is modulated in response to multiple stresses, including hypoxia, amino-acid or nutrient deprivation (Karam et al. 2013; Lykke-Andersen and Jensen 2015). Interestingly, many stress-related mRNAs are targeted by the NMD under normal physiological conditions but are stabilized upon stress, due to NMD activity inhibition (Lykke-Andersen and Jensen 2015). Note that under stress conditions, global translation also decreases, selectively preserving translation of stress-related mRNAs to the detriment of “housekeeping” mRNAs (Yamasaki and Anderson 2008). Another consequence of such a stress-mediated reduction of translation is that transcripts (aberrant mRNAs or lncRNAs) that are normally targeted by the NMD will be stabilized, since transcripts evading translation escape NMD.
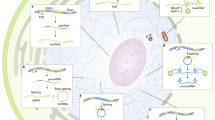
On this basis, we propose a model where a stress that results in translation inhibition and/or NMD inhibition will lead to stabilization of regulatory antisense NMD-sensitive lncRNAs (such as yeast XUTs), which could in turn repress the transcription of their paired-sense genes (Fig. 2). This would prevent the synthesis of mRNAs that could probably not be translated, avoiding the cell to waste an energy that could be crucial to survive the stress. Alternatively but not exclusively, the regulatory antisense lncRNAs could also regulate the paired-sense mRNAs at the post-transcriptional level, potentially through the formation of double-stranded (ds) RNA structures. In yeast, sense mRNAs and antisense XUTs have been shown to form dsRNA in vivo, and this protects XUTs from NMD (Wery et al. 2016). Reciprocally, formation of dsRNA with a stabilized antisense lncRNA might affect sense mRNA stability, as suggested by a recent study of mRNA isoforms half-lives (Geisberg et al. 2014). Besides RNA stability, regulatory antisense lncRNAs might also interfere with mRNA splicing, localization, or translation. Future work will be needed to decipher these potential lncRNA-mediated regulatory mechanisms and determine whether they can be integrated within larger stress-activated signalling networks (Ho and Gasch 2015).
NMD buffers genome expression by controlling levels of regulatory antisense lncRNAs. NMD contributes to genome expression buffering by restricting the levels of a class of cytoplasmic antisense regulatory lncRNAs (red) that can attenuate the transcription of their paired-sense protein-coding genes (blue). In this model, any stress leading to translation and/or NMD inhibition would result in the stabilization of the lncRNA and transcriptional attenuation of the paired-sense gene
Concluding remarks
Over the years, lncRNAs have been recognized as major regulators of multiple cellular processes. However, the initial assumption that they are devoid of coding potential is now challenged. Conceptually, coding a peptide/protein in specific circumstances and functioning as a regulatory RNA molecule in others are not exclusive possibilities for a transcript, whatever it has been primarily annotated as a lncRNA or mRNA. Examples of bifunctional RNAs with dual coding and regulatory functions have been reported (Ulveling et al. 2011). In this respect, in yeast, convergent mRNAs can regulate each other at the RNA level providing additional intriguing cases where mRNAs can switch their initial coding function into regulatory RNAs (Sinturel et al. 2015). But how many among the thousands of lncRNAs annotated in the different eukaryotic models correspond to such bifunctional RNAs remains to be determined. The classical distinction between coding and non-coding RNAs might, therefore, become less strict in the future, if the possibility to switch between regulatory and coding functions in response to specific stimuli appears to be a common feature of “lncRNAs”.
References
Aspden JL, Eyre-Walker YC, Phillips RJ, Amin U, Mumtaz MA, Brocard M, Couso JP (2014) Extensive translation of small open reading frames revealed by poly-ribo-seq. Elife 3:e03528
Bazzini AA et al (2014) Identification of small ORFs in vertebrates using ribosome footprinting and evolutionary conservation. EMBO J 33:981–993
Berretta J, Morillon A (2009) Pervasive transcription constitutes a new level of eukaryotic genome regulation. EMBO Rep 10:973–982
Berretta J, Pinskaya M, Morillon A (2008) A cryptic unstable transcript mediates transcriptional trans-silencing of the Ty1 retrotransposon in S. cerevisiae. Genes Dev 22:615–626
Chew GL, Pauli A, Rinn JL, Regev A, Schier AF, Valen E (2013) Ribosome profiling reveals resemblance between long non-coding RNAs and 5′ leaders of coding RNAs. Development 140:2828–2834
Djebali S et al (2012) Landscape of transcription in human cells. Nature 489:101–108
Drinnenberg IA, Weinberg DE, Xie KT, Mower JP, Wolfe KH, Fink GR, Bartel DP (2009) RNAi in budding yeast. Science 326:544–550
Geisberg JV, Moqtaderi Z, Fan X, Ozsolak F, Struhl K (2014) Global analysis of mRNA isoform half-lives reveals stabilizing and destabilizing elements in yeast. Cell 156:812–824
Guttman M, Russell P, Ingolia NT, Weissman JS, Lander ES (2013) Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154:240–251
Ho YH, Gasch AP (2015) Exploiting the yeast stress-activated signaling network to inform on stress biology and disease signaling. Curr Genet 61:503–511
Ingolia NT, Lareau LF, Weissman JS (2011) Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147:789–802
Karam R, Wengrod J, Gardner LB, Wilkinson MF (2013) Regulation of nonsense-mediated mRNA decay: implications for physiology and disease. Biochim Biophys Acta 1829:624–633
Lykke-Andersen S, Jensen TH (2015) Nonsense-mediated mRNA decay: an intricate machinery that shapes transcriptomes. Nat Rev Mol Cell Biol 16:665–677
Magny EG, Pueyo JI, Pearl FM, Cespedes MA, Niven JE, Bishop SA, Couso JP (2013) Conserved regulation of cardiac calcium uptake by peptides encoded in small open reading frames. Science 341:1116–1120
Malabat C, Feuerbach F, Ma L, Saveanu C, Jacquier A (2015) Quality control of transcription start site selection by nonsense-mediated-mRNA decay. Elife 4:e06722
Muhlrad D, Parker R (1994) Premature translational termination triggers mRNA decapping. Nature 370:578–581
Muhlrad D, Parker R (1999) Aberrant mRNAs with extended 3′ UTRs are substrates for rapid degradation by mRNA surveillance. RNA 5:1299–1307
Neil H, Malabat C, d’Aubenton-Carafa Y, Xu Z, Steinmetz LM, Jacquier A (2009) Widespread bidirectional promoters are the major source of cryptic transcripts in yeast. Nature 457:1038–1042
Nelson BR et al (2016) A peptide encoded by a transcript annotated as long noncoding RNA enhances SERCA activity in muscle. Science 351:271–275
Pauli A et al (2014) Toddler: an embryonic signal that promotes cell movement via Apelin receptors. Science 343:1248636
Peccarelli M, Kebaara BW (2014) Regulation of natural mRNAs by the nonsense-mediated mRNA decay pathway. Eukaryot Cell 13:1126–1135
Rinn JL, Chang HY (2012) Genome regulation by long noncoding RNAs. Annu Rev Biochem 81:145–166
Sinturel F, Navickas A, Wery M, Descrimes M, Morillon A, Torchet C, Benard L (2015) Cytoplasmic control of sense–antisense mRNA pairs. Cell Rep 12:1853–1864
Smith JE et al (2014) Translation of small open reading frames within unannotated RNA transcripts in Saccharomyces cerevisiae. Cell Rep 7:1858–1866
Sole C, Nadal-Ribelles M, de Nadal E, Posas F (2015) A novel role for lncRNAs in cell cycle control during stress adaptation. Curr Genet 61:299–308
Taft RJ, Pheasant M, Mattick JS (2007) The relationship between non-protein-coding DNA and eukaryotic complexity. BioEssays 29:288–299
Taft RJ, Pang KC, Mercer TR, Dinger M, Mattick JS (2010) Non-coding RNAs: regulators of disease. J Pathol 220:126–139
Tisseur M, Kwapisz M, Morillon A (2011) Pervasive transcription—lessons from yeast. Biochimie 93:1889–1896
Tudek A, Candelli T, Libri D (2015) Non-coding transcription by RNA polymerase II in yeast: hasard or necessite? Biochimie 117:28–36
Ulveling D, Francastel C, Hube F (2011) When one is better than two: rNA with dual functions. Biochimie 93:633–644
Van Dijk EL et al (2011) XUTs are a class of Xrn1-sensitive antisense regulatory non coding RNA in yeast. Nature 475:114–117
Wery M, Kwapisz M, Morillon A (2011) Noncoding RNAs in gene regulation. Wiley Interdiscip Rev Syst Biol Med 3:728–738
Wery M, Descrimes M, Vogt N, Dallongeville AS, Gautheret D, Morillon A (2016) Nonsense-mediated decay restricts LncRNA levels in yeast unless blocked by double-stranded RNA structure. Mol Cell 61:379–392
Xu Z et al (2009) Bidirectional promoters generate pervasive transcription in yeast. Nature 457:1033–1037
Yamasaki S, Anderson P (2008) Reprogramming mRNA translation during stress. Curr Opin Cell Biol 20:222–226
Acknowledgments
We thank all the members of our lab for discussions and critical reading of the manuscript. A. Morillon’s lab is supported by the ANR “DNA-Life” and ERC “DARK” consolidator grants.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
de Andres-Pablo, A., Morillon, A. & Wery, M. LncRNAs, lost in translation or licence to regulate?. Curr Genet 63, 29–33 (2017). https://doi.org/10.1007/s00294-016-0615-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0615-1