Abstract
As bacterial natural products have been proved to be the most important source of many therapeutic medicines, the need to discover novel natural products becomes extremely urgent. Despite the fact that the majority of bacterial species are yet to be cultured in a laboratory setting, and that most of the bacterial natural product biosynthetic genes are silent, “metagenomics technology” offers a solution to help clone natural product biosynthetic genes from environmental samples, and genetic engineering enables the silent biosynthetic genes to be activated. In this work, a type II polyketide biosynthetic gene cluster was identified from a soil metagenomic library and was activated by over-expression of a SARP regulator gene in the gene cluster in Streptomyces hosts. A new tetracenomycin type compound tetracenomycin Y was identified from the fermentation broth. This study shows that metagenomics and genetic engineering could be combined to provide access to new natural metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria are an important source of natural products and many currently used therapeutic agents are bacterial natural products or their derivatives. However, the vast majority of bacteria in environmental samples remain recalcitrant to culturing; as a result, uncultured bacteria have not yet been extensively explored for the production of novel natural products [1]. Although most of the bacteria cannot be cultured in current laboratory condition, the natural product biosynthetic genes on the genomes of the uncultured bacteria (environmental DNA, eDNA) could be extracted, ligated with a suitable vector, and functionally expressed in cultured hosts for new compound discovery [2]. Using metagenomics strategies, some novel natural products with diverse bioactivities have been identified, including aromatic polyketides, nonribosomal peptides, tryptophan dimers, polyenes, and topoisomerase inhibitors [3,4,5,6].
Aromatic polyketides synthesized by type II polyketide synthetases (PKSs), including some current clinically valuable antibiotics or chemotherapeutic agents, such as doxorubicin (antitumor), tetracycline (antibiotic), and mithramycin (immunosuppressant), are structure-diverse compounds with various bioactivities [7, 8]. Because of their remarkable bioactivities, aromatic polyketides became one of the main target compounds of metagenomics strategies. eDNA derived aromatic polyketides with distinguish antibacterial activity have been discovered, including turbomycins with broad-spectrum antibacterial effect [9], and fasamycins with high activity against methicillin resistant Staphylococcus aureus (MRSA) and vancomycin resistant enterococcus (VRE) [10]. Moreover, investigation on eDNA derived biosynthetic genes revealed that there may be many novel aromatic polyketides hidden in the environment and metagemomics could be an important approach to the aromatic polyketides discovery [11]. Stevenson et al. had reported the identification of an aureolic acid biosynthetic gene cluster (BGC) and its product (metathramycin) from metagenome library which possessed potent bioactivity against a human colon carcinoma cell line [12]. Environmental DNA library is a large bacterial genetic reservoir from which new natural product BGCs can be systematically recovered and studied. The BGSs for different types of natural products can be identified from the library using different corresponding probes. On the other hand, because the length of a type II PKS gene cluster is normally less than 40 kb, it is possible to capture the intact BGC in an individual cosmid clone.
Uncultured bacteria are considered as the biggest remaining pool for natural product discovery, and metagenomics approach has been considered as one of the promising strategies for discovering novel natural product from uncultured bacteria [13, 14]. Although most of BGCs are not expressed under standard laboratory condition, several methods for the activation of silent BGCs have been developed, including heterologous expression, promoter engineering, ribosome engineering, and engineering of transcriptional regulators [15]. An ideal heterologous host should display high natural propensity to support the expression of diverse BGCs [16]. As Streptomycetes have a very high GC content and highly complex regulatory networks that direct expression of BGCs, Streptomyces represent potential hosts for heterologous expression of BGCs derived from other Streptomycetes or related genera with high GC content [16]. Regulator genes are often located in microbial natural product biosynthetic gene clusters, and play critical roles in determining the onset and production level of each natural product [17]. The biosynthesis of nature products in Streptomyces is regulated by precise regulatory systems, in which transcription factors regulate the initiation level of transcription by binding to DNAs [15]. Streptomyces antibiotic regulatory protein (SARP) family regulators are positive pathway-specific transcriptional regulators which can bind to the promoter DNA of the biosynthetic gene in the BGC to activate the gene [18]. Chen et al. improved the fredericamycin A titer 5.6-fold relative to that of wild-type S. griseus by overexpression of a SARP family regulator gene fdmR1 [19]. Beck et al. have activated a silent BGC in Streptomyces sp. CA-256286 by overexpression of a set of SARP family transcriptional regulators [20]. A metagenomic library of DNA extracted from soil collected in Yunnan, China had previously been constructed [21]. Type II PKS gene clusters contain a conserved minimal PKS that is composed of two ketosynthase genes (KSα and KSβ) and an acyl carrier protein (ACP). Here, the conserved KSα gene in the type II PKS gene cluster was used as the probe to identify and recover cosmid clones from the soil eDNA library. The recovered clones were retrofitted with the genetic elements necessary for conjugation and integration into Streptomyces. Using different Streptomyces hosts S. coelicolor M1146 and S. venezuela, and over-expression of a SARP family regulator gene, we activated an eDNA derived type II polyketide biosynthetic gene cluster cosmidYN01. The cosmidYN01 gene cluster produced tetracenomycin type compounds and a new compound (TCM Y) was identified.
Materials and Methods
Bacterial Strains, Plasmids, Media and Culture Conditions
S. coelicolor M1146 and S. venezuela were used as hosts for gene heterologous expression. Escherichia coli EPI100 was used as host of metagenomic library. E. coli JTU007 (pUZ8002) were used as donor strain in E. coli/Streptomyces conjugation. E. coli strains were grown in LB medium. For sporulation, Streptomyce strains were grown in MS medium. For fermentation, Streptomyce strains harboring the YN01 type II polyketide BGC were grown at 28 °C in seed R5 medium for 4 days, and the 40 mL seed culture was inoculated into 400 mL of ISP4 fermentation medium containing 5% HP-20 resin for 7 days, or into 400 mL of R5 fermentation medium for 5 days. Plasmid pWEB-TNCTM was used for metagenomic library construction, plasmid pOJ436 for conjugation, and plasmid pUWL201PW-OriT for endogenous expression. All primers used are listed in supplement Table S1.
Construction and Screening of Metagenomic Cosmid Library
The soil sample was collected from Yunnan, China (25° 20′ N, 102° 30′ E) [21], and the high-quality eDNA was extracted following a reported protocol [22]. Briefly, soil was heated (70 °C) in lysis buffer [100 mM Tris–HCl, 100 mM EDTA, 1.5 M NaCl, 1% (wt/vol) cetyltrimenthyl ammonium bromide, 2% (wt/vol) SDS (pH 8.0)] for 2 h. After soil particulates were removed from the crude lysate by centrifugation, eDNA was precipitated from the supernatant with the addition of 0.7 vol isopropanol. Crude eDNA was collected by centrifugation, washed with 70% ethanol, and resuspended in TE [10 mM Tris HCl, 1 mM EDTA (pH 8.0)]. Then eDNA was gel purified (1% agarose) and the gel purified high-molecular-weight eDNA was blunt ended, ligated into SmaI site of pWEB-TNC (Epicenter), and packaged. Then the packaged products were transfected into E. coli EPI 100 to construct a metagenomic library.
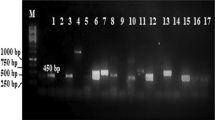
Degenerate primers (540-Fw/1100-Re) based on conserved regions of KSα genes were used to screen the metagenomic library by PCR (95 °C for 3 min, followed by 35 cycles of 94 °C for 45 s, 64 °C for 40 s, 72 °C for 75 s, and finally 72 °C for 10 min) [23]. Amplicons of the correct predicted size (560 bp) were gel purified, sequenced, and compared to deposited KSα genes from National Center for Biotechnology Information (NCBI) database. Unique KSα genes were used as probes to recover type II PKS containing clones by serial dilution method using the following touchdown protocol: denaturation (95 °C, 4 min), 10 touchdown cycles (95 °C, 40 s; 65 °C [−1 °C per cycle until 55 °C], 40 s; 72 °C, 1 min); 30 standard cycles (95 °C, 40 s; 55 °C, 40 s; 72 °C, 40 s) and a final extension step (72 °C, 10 min).
Sequencing and In Silico Analysis of YN01 Gene Cluster
Cosmid from the positive clone was isolated and sequenced by Invitrogen Company. After the quality control, sequence data were assembled, corrected and optimized, and the final cosmid sequence was obtained. Because type II polyketide BGCs are relatively conserved, the functions of related genes were deduced based on sequence similarity using Basic Local Alignment Search Tool (BLAST) and Rapid Annotation Subsystem Technology (RAST) in the database.
Retrofitting and Conjugation of eDNA Derived Cosmidyn01 Gene Cluster into Streptomyces Hosts
YN01 cosmid was digested with AanI and ligated with the 6.8 kb DraI fragment from pOJ436 which contains an origin of transfer, an apramycin resistant gene and the ФC31 phage integration system. The retrofitted cosmid in E. coli JTU007 (pUZ8002) was conjugated into S. coelicolor M1146 or S. venezuela to generate S. coelicolor/YN01 and S. venezuela/YN01, respectively [10]. Plasmid pOJ436 was also conjugated into S. coelicolor M1146 and S. venezuela to generate S. coelicolor/pOJ436 and S. venezuela/pOJ436, respectively, which would be used as negative controls in the fermentation.
Construction of SARP Regulator Construct
The SARP regulator gene in YN01 gene cluster was PCR-amplified using the primer pair SARP-Fw/SARP-Re. PCR was conducted using the following touchdown protocol: denaturation (95 °C, 4 min), 10 touchdown cycles (95 °C, 40 s; 68 °C (-1 °C per cycle), 40 s; 72 °C, 1 min; 30 standard cycles (95 °C, 40 s; 58 °C, 40 s; 72 °C, 40 s) and a final extension step (72 °C, 10 min). The PCR amplicon was double digested with NdeI and HindIII and inserted into the NdeI/HindIII site of pUWL201PW-OriT to generate pYN01/SARP. Then pYN01/SARP was conjugated into S. coelicolor YN01 and S. venezuela YN01 to generate S. coelicolor M1146/YN01/SARP and S. venezuela/YN01/SARP, respectively.
Isolation of Clone-Specific Compounds from Fermentation Broth
HP-20 resin in ISP4 fermentation medium was collected and the metabolites on the resin were dissolved by methanol. The metabolites in R5 fermentation medium were extracted twice with an equal volume of ethyl acetate. The extracts were then dried by rotary evaporator and dissolved in methanol. The methanol eluents were analyzed by HPLC (1 mL/min) using a linear gradient of 80:20/H2O:MeOH to 100% MeOH over 30 min. For compound purification, 48 mg crude extract from 48 L R5 fermentation medium was initially fractionated by silica gel flash chromatography using a CH2Cl2:MeOH step gradient, and then purified by semi-preparative HPLC (4 mL/min) using isocratic gradient at 65% methanol. Finally, 76 mg of compound 1, and 1.7 mg compound 2 were obtained.
Structure Determination by NMR and LC–MS
NMR study was performed at room temperature on BTUKER AVANCE III 600. For LC–MS detection, the extracts were dissolved in chromatographic grade methanol and centrifuged for 10 min. The resulting clear supernatant (10 μL) was used for LC–MS analysis. The study was performed on Agilent 1200-6410B liquid mass spectrometry instrument.
Nucleotide Sequence Accession Numbers
The sequence data of YN01 gene cluster were deposited in GeneBank under the accession no. MK158260-158,275.
Results
Construction of a Soil Metagenomic Cosmid Library and Recovery of YN01 Clone Harboring a type II PKS Gene Cluster from the Library
A metagenomic library was constructed using soil sample from Yunnan, China. The library contained more than 10 million cosmid clones with an average insertion size of approximately 38 kb. Eighteen clones harboring type II polyketide biosynthetic gene clusters were obtained after screening 500, 000 clones in the library.
Sequence Analysis of YN01 Gene Cluster
To achieve the complete characterization of type II polyketide biosynthetic gene clusters, eighteen cosmids were sequenced. Sequence analysis showed that cosmidYN01 contained an intact type II polyketide biosynthetic gene cluster. Sequence analysis of the inserted DNA in YN01 cosmid revealed 21 complete ORFs. The corresponding deduced proteins were compared with other known proteins in the databases by BLAST (Table 1). YN01 gene cluster shared high homology with Tcm gene cluster from Amycolatopsis sp. A23 strain (Fig. 1). Although the eDNA deduced proteins shared high identities of 97%–99% with the counterpart Tcm proteins in Amycolatopsis sp. A23, difference between the two gene clusters could be identified. The location and the orientation of the SARP and O-methyltransferase (P) genes were different, as well as a GntR family regulator gene located in YN01 gene cluster instead of a AsfR family regulator gene in Tcm gene cluster. Moreover, the deduced protein of a gene (Q) downstream YN01 cluster shared 99% identity with a methyltransferase YktD that located in another gene cluster in Amycolatopsis sp. A23. The differences between cosmidYN01 BGC and Tcm BGC in Amycolatopsis sp. A23 imply that cosmidYN01 gene cluster might produce novel tetracenomycin analogs.
Chromosomal arrangement of the tetracenomycin biosynthesis gene clusters in Amycolatopsis sp. A23 and cosmidYN01. Arrows show the direction of transcription. Orthologous genes are shown with matching colors and letters. All the counterpart genes in the two gene cluster are located at the same positions except P and two regulator genes, and a methyltransferase gene Q is only located in YN01 gene cluster, which was predicted to relate with the O-methylation on C4 of 2. Functional annotations of the respective genes are provided in supplementary Table S2
Heterologous Expression of YN01 Type II Polyketide Biosynthetic Gene Cluster in Streptomyces Hosts
To uncover the products of YN01 gene cluster, cosmidYN01 was ligated with pOJ436 vector, and then the recombinant DNA was conjugated from E.coli JTU007 (pUZ8002) into S. coelicolor M1146 and S. venezuelae to obtain S. coelicolor M1146/YN01 and S. venezuelae/YN01, respectively. Then the Streptomyces strains harboring YN01 gene cluster were fermented in ISP4 and R5 medium and the fermentation broths were analyzed for clone-specific peaks related to the gene cluster by HPLC. No specific peak could be identified from S. venezuelae/YN01 cultured in both ISP4 and R5 media, while only a small special peak (1) could be identified from R5 fermentation broth of S. coelicolor M1146/YN01 (Fig. 2). Because the yield of 1 in S. coelicolorM1146/YN01 was low, we then over-expressed a SARP gene located in YN01 gene cluster to regulate the biosynthesis of 1 in Streptomyces hosts. The 912 bp SARP gene was cloned under control of an ermE* promoter in pUWL201-OriT to obtain pUWSA. Plasmid pUWSA was conjugated into S. coelicolor M1146/YN01 and S. venezuelae/YN01 to get S. coelicolor M1146/YN01/SARP and S. venezuelae/YN01/SARP, respectively. Then S. coelicolor M1146/SARP and S. venezuelae/YN01/SARP were fermented and the fermentation broths were HPLC-analyzed. The yield of 1 was improved distinctly. S. coelicolor M1146/YN01/SARP was fermented in 48 L of R5 medium and 1 from the fermentation broth was isolated and purified by chromatography and its structure was elucidated by NMR and MS. Apart from 1, another new peak could also be observed during purification of 1. This compound (2) was purified and its structure was also elucidated. Compound 1 was isolated as a yellow powder with the molecular formula C24H22O11 ([M + H]+, m/z 487.1247, calcd. for 487.1240) (supplement Fig. S1) as determined by HRESIMS. Analysis of 1H and 13C NMR data suggested that 1 was a known aromatic polyketide, tetracenomycin X (supplement Figs. S3~7 and Table 2). The molecular formula of 2 was determined as C24H22O11 ([M + H]+, m/z 487.1233, calcd. for 487.1240) (supplement Fig. S2), same formula as 1. Analysis of 1H,13C NMR and HMBC NMR data suggested that 2 was an isomer of TCM X (supplement Figs. S8~12 and Table 2). The main difference was attributed to ring A, in which the methoxy group at C-12a in 1 was transferred to C-4 in 2. The structures of compound 1 and compound 2 are shown in Fig. 2.
a HPLC analysis of the extracts of R5 fermented broths of S. coelicolor M1146 and S. venezuelae derived strains (UV at 254 nm). S. venezuelae/YN01, S. venezuelae harbors YN01 gene cluster. S. venezuelae/YN01/SARP, S. venezuelae/YN01 harbors a SARP overexpressed plasmid. S. venezuelae/pOJ436, S. venezuelae harbors pOJ436. S. coelicolor M1146/YN01, S. coelicolor M1146 harbors YN01 gene cluster. S. coelicolor M1146/YN01/SARP, S. coelicolor M1146/YN01 harbors a SARP overexpressed plasmid. S. coelicolor M1146/pOJ436, S. venezuelae harbors pOJ436. Metabolite 1 is a clone-specific compound with the characteristic UV absorbance spectrum of aromatic polyketide. b Chemical structure of 1. c Chemical structure of 2. The arrows showed the HMBC
Proposed Biosynthetic Pathway of 1 and 2
The biosynthetic pathway of 1 and 2 was proposed based on the known biosynthetic genes and pathway of tetracenomycins (Fig. 3) [24]. At first, polyketide chain was synthesized by ORF12 (TcmK), ORF13 (TcmL), and ORF14 (TcmM) using acetyl-CoA and malonyl-CoA as the chain starter and extender units, respectively; next, ORF15 (TcmN), ORF11 (TcmJ)and ORF10 (TcmI) acted as aromatase and cyclases to catalyze the formation of the aromatic rings, followed by ORF9 (TcmH) catalyzing the C-5 carbonylation; then, hydroxylation and methylation occurred, and final O-methylation at C-12a and C-4 formed 1 and 2, respectively.
Discussion
The tetracenomycins are a family of aromatic polyketides with antibacterial activity against gram-positive microorganisms and antitumor activity against a variety of mammalian cancer cell lines [25]. The tetracenomycins inhibiting peptide translation via binding to the large ribosomal subunit in cancer cells [26]. In this study, we obtained a cosmidYN01 harboring a type II polyketide BGC. The in silico analysis revealed that YN01 gene cluster shared high similarity with tcm gene cluster form Amycolatopsis sp. A23, except that it has one more methyltransferase gene. Activation expression of cosmidYN01 gene cluster in Streptomyces hosts resulted in a new tetracenomycin type compound 2. Three O-methyltransferase genes, tcmN, tcmO and tcmP, are located in Tcm biosynthetic gene cluster, while four O-methyltransferase genes, ORF2, ORF16, ORF17 and ORF18 are located in cosmidYN01 gene cluster (Fig. 1). Among the three O-methyltransferase in 1 and 2 biosynthesis, ORF2, ORF16, and ORF17 share identities of 99%, 98% and 97% with TcmP, TcmN, and TcmO inAmycolatopsis sp. A23, respectively. ORF18 shares identity of 99% with a methyltransferase and thus proposed to catalyze the C-4 O-methylation of TCM C to produce 2. To reveal the detailed biosynthesis of 2, more research work needs to be performed next in the future.
With the development of sequencing technologies, more and more DNA sequences from both cultured microbes and uncultured samples are reported. Analysis of these sequence data revealed many potential natural product BGCs, however, most of them are silent under standard laboratory growth conditions [27,28,29]. Heterologous expression is an efficient and established approach to unlock the gene silence [15]. Selection of suitable heterologous hosts to express natural product biosynthetic genes is a promising approach for discovering novel microbial natural products. There are a number of potential Actinomycete hosts identified from molecular engineering, combinatorial biosynthesis, and gene cluster expression studies. S. coelicolor M1146 is commonly used as a host strain for engineering of secondary metabolite production. Shi et al. heterologously expressed the chuangxinmycin BGC from A. tsinanensis using S. coelicolor M1146 host [30], while Nguyen et al. expressed the elloramycin BGC in the heterologous host S. coelicolor M1146 to facilitate the downstream production of tetracenomycin analogs [31]. S. venezuelae has been recently developed as a heterologous host [32, 33] that requires a short culture period (3 ~ 4 days) for metabolite production compared to other Streptomyces [34]. It is also amenable to genetic manipulation and has high transformation efficiency. These characteristics make S. venezuelae an alternative system for a rapid heterologous production of polyketides from genetic manipulation to product fermentation [34, 35].
Here, we used S. coelicolor M1146 and S. venezuelae as heterologous hosts. The production level of 1 in S. coelicolor M1146 was extremely low, while the YN01 gene cluster seemed silent in S. venezuelae. The biosynthesis of natural products was strictly regulated in microbes and many regulation factors involved in natural product biosynthesis were identified. SARP belongs to pathway-specific activators that have been found in natural product BGCs [17] and was harnessed for increasing the production of natural products [36], even in an eDNA derived BGC to obtain two novel MRSA active compounds tetarimycin A and B [11]. Thus, a SARP regulator gene located in cosmidYN01 gene cluster was over-expressed in Streptomyces strains harboring cosmidYN01 gene cluster to activate the gene cluster. Finally, high yield of 1 was achieved in SARP over-expressed strains. Meanwhile, a new natural tetracenomycin derivative 2 was identified. This study provides proof-of-concept that genetic manipulation of regulation factors in potential biosynthetic pathways could lead to discovering more novel compounds.
Conclusions
In summary, an eDNA derived YN01 type IIpolyketide gene cluster was cloned from a metagenomic library. Over-expression of a SARP gene in Streptomyces hosts harboring YN01 gene cluster activated the gene cluster to produce a new tetracenomycin derivative, TCM Y. This study provided another example in eDNA derived biosynthetic gene activation and confirmed that gene manipulation was an effective way in activation of eDNA derived natural product biosynthetic genes.
References
Jung WS, Kim ES, Kang HY, Choi CY, Sherman DH, Yoon YJ (2003) Site-directed mutagenesis on putative macrolactone ringsize determinant in the hybrid pikromycin–tylosin polyketidesynthase. J Microbiol Biotechnol 13:823–827
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (1998) Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol 5(10):R245-249. https://doi.org/10.1016/s1074-5521(98)90108-9
Katz M, Hover BM, Brady SF (2016) Culture-independent discovery of natural products from soil metagenomes. J Ind Microbiol Biotechnol 43(2–3):129–141. https://doi.org/10.1007/s10295-015-1706-6
Owen JG, Charlop-Powers Z, Smith AG, Ternei MA, Calle PY, Reddy BV, Montiel D, Brady SF (2015) Multiplexed metagenome mining using short DNA sequence tags facilitates targeted discovery of epoxyketone proteasome inhibitors. Proc Natl Acad Sci USA 112(14):4221–4226. https://doi.org/10.1073/pnas.1501124112
Wang Z, Forelli N, Hernandez Y, Ternei M, Brady SF (2022) Lapcin, a potent dual topoisomerase I/II inhibitor discovered by soil metagenome guided total chemical synthesis. Nat Commun 13(1):842. https://doi.org/10.1038/s41467-022-28292-x
Li L, MacIntyre LW, Ali T, Russo R, Koirala B, Hernandez Y, Brady SF (2021) Biosynthetic interrogation of soil metagenomes reveals metamarin, an uncommon cyclomarin congener with activity against Mycobacterium tuberculosis. J Nat Prod 84(4):1056–1066. https://doi.org/10.1021/acs.jnatprod.0c01104
Weissman KJ (2009) Chapter 1 introduction to polyketide biosynthesis. Complex enzymes in microbial natural product biosynthesis, part B: polyketides, aminocoumarins and carbohydrates. Academic Press, Cambridge, pp 3–16
Zhang Z, Pan HX, Tang GL (2017) New insights into bacterial type II polyketide biosynthesis. F1000Research 6:172. https://doi.org/10.12688/f1000research.10466.1
Gillespie DE, Brady SF, Bettermann AD, Cianciotto NP, Liles MR, Rondon MR, Clardy J, Goodman RM, Handelsman J (2002) Isolation of antibiotics turbomycin a and B from a metagenomic library of soil microbial DNA. Appl Environ Microbiol 68(9):4301–4306. https://doi.org/10.1128/AEM.68.9.4301-4306.2002
Feng Z, Kallifidas D, Brady SF (2011) Functional analysis of environmental DNA-derived type II polyketide synthases reveals structurally diverse secondary metabolites. Proc Natl Acad Sci USA 108(31):12629–12634. https://doi.org/10.1073/pnas.1103921108
Kallifidas D, Kang HS, Brady SF (2012) Tetarimycin A, an MRSA-active antibiotic identified through induced expression of environmental DNA gene clusters. J Am Chem Soc 134(48):19552–19555. https://doi.org/10.1021/ja3093828
Stevenson LJ, Bracegirdle J, Liu L, Sharrock AV, Ackerley DF, Keyzers RA, Owen JG (2021) Metathramycin, a new bioactive aureolic acid discovered by heterologous expression of a metagenome derived biosynthetic pathway. RSC Chem Biol 2(2):556–567. https://doi.org/10.1039/d0cb00228c
Brady SF, Simmons L, Kim JH, Schmidt EW (2009) Metagenomic approaches to natural products from free-living and symbiotic organisms. Nat Prod Rep 26(11):1488–1503. https://doi.org/10.1039/b817078a
Lewis K, Epstein S, D’Onofrio A, Ling LL (2010) Uncultured microorganisms as a source of secondary metabolites. J Antibiot (Tokyo) 63(8):468–476. https://doi.org/10.1038/ja.2010.87
Liu Z, Zhao Y, Huang C, Luo Y (2021) Recent advances in silent gene cluster activation in Streptomyces. Front Bioeng Biotechnol 9:632230. https://doi.org/10.3389/fbioe.2021.632230
Baltz RH (2010) Streptomyces and Saccharopolyspora hosts for heterologous expression of secondary metabolite gene clusters. J Ind Microbiol Biotechnol 37(8):759–772. https://doi.org/10.1007/s10295-010-0730-9
Liu G, Chater KF, Chandra G, Niu G, Tan H (2013) Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol Rev 77(1):112–143. https://doi.org/10.1128/MMBR.00054-12
Bibb MJ (2005) Regulation of secondary metabolism in Streptomycetes. Curr Opin Microbiol 8(2):208–215. https://doi.org/10.1016/j.mib.2005.02.016
Chen Y, Wendt-Pienkowski E, Shen B (2008) Identification and utility of FdmR1 as a Streptomyces antibiotic regulatory protein activator for fredericamycin production in Streptomyces griseus ATCC 49344 and heterologous hosts. J Bacteriol 190(16):5587–5596. https://doi.org/10.1128/JB.00592-08
Beck C, Gren T, Ortiz-Lopez FJ, Jorgensen TS, Carretero-Molina D, Martin Serrano J, Tormo JR, Oves-Costales D, Kontou EE, Mohite OS, Mingyar E, Stegmann E, Genilloud O, Weber T (2021) Activation and identification of a Griseusin cluster in Streptomyces sp. CA-256286 by employing transcriptional regulators and multi-omics methods. Molecules. https://doi.org/10.3390/molecules26216580
Wang S, Gao X, Gao Y, Li Y, Cao M, Xi Z, Zhao L, Feng Z (2017) Tetracycline resistance genes identified from distinct soil environments in China by functional metagenomics. Front Microbiol 8:1406. https://doi.org/10.3389/fmicb.2017.01406
Brady SF (2007) Construction of soil environmental DNA cosmid libraries and screening for clones that produce biologically active small molecules. Nat Protoc 2(5):1297–1305. https://doi.org/10.1038/nprot.2007.195
Wawrik B, Kerkhof L, Zylstra GJ, Kukor JJ (2005) Identification of unique type II polyketide synthase genes in soil. Appl Environ Microbiol 71(5):2232–2238. https://doi.org/10.1128/AEM.71.5.2232-2238.2005
Hutchinson CR (1997) Biosynthetic studies of daunorubicin and tetracenomycin C. Chem Rev 97:2525–2535
Qiao X, Gan M, Wang C, Liu B, Shang Y, Li Y, Chen S (2019) Tetracenomycin X exerts antitumour activity in lung cancer cells through the downregulation of cyclin D1. Mar Drugs. https://doi.org/10.3390/md17010063
Osterman IA, Wieland M, Maviza TP, Lashkevich KA, Lukianov DA, Komarova ES, Zakalyukina YV, Buschauer R, Shiriaev DI, Leyn SA, Zlamal JE, Biryukov MV, Skvortsov DA, Tashlitsky VN, Polshakov VI, Cheng J, Polikanov YS, Bogdanov AA, Osterman AL, Dmitriev SE, Beckmann R, Dontsova OA, Wilson DN, Sergiev PV (2020) Tetracenomycin X inhibits translation by binding within the ribosomal exit tunnel. Nat Chem Biol 16(10):1071–1077. https://doi.org/10.1038/s41589-020-0578-x
Walsh CT, Fischbach MA (2010) Natural products version 2.0: connecting genes to molecules. J Am Chem Soc 132(8):2469–2493. https://doi.org/10.1021/ja909118a
Rutledge PJ, Challis GL (2015) Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol 13(8):509–523. https://doi.org/10.1038/nrmicro3496
Mao D, Okada BK, Wu Y, Xu F, Seyedsayamdost MR (2018) Recent advances in activating silent biosynthetic gene clusters in bacteria. Curr Opin Microbiol 45:156–163. https://doi.org/10.1016/j.mib.2018.05.001
Jiang ZB, Ren WC, Shi YY, Li XX, Lei X, Fan JH, Zhang C, Gu RJ, Wang LF, Xie YY, Hong B (2018) Structure-based manual screening and automatic networking for systematically exploring sansanmycin analogues using high performance liquid chromatography tandem mass spectroscopy. J Pharm Biomed Anal 158:94–105. https://doi.org/10.1016/j.jpba.2018.05.024
Nguyen JT, Riebschleger KK, Brown KV, Gorgijevska NM, Nybo SE (2021) A BioBricks toolbox for metabolic engineering of the tetracenomycin pathway. Biotechnol J. https://doi.org/10.1002/biot.202100371
Yoon YJ, Beck BJ, Kim BS, Kang HY, Reynolds KA, Sherman DH (2002) Generation of multiple bioactive macrolides by hybrid modular polyketide synthases in Streptomyces venezuelae. Chem Biol 9(2):203–214. https://doi.org/10.1016/s1074-5521(02)00095-9
Hong JS, Park SH, Choi CY, Sohng JK, Yoon YJ (2004) New olivosyl derivatives of methymycin/pikromycin from an engineered strain of Streptomyces venezuelae. FEMS Microbiol Lett 238(2):391–399. https://doi.org/10.1016/j.femsle.2004.08.002
Xue Y, Sherman DH (2001) Biosynthesis and combinatorial biosynthesis of pikromycin-related macrolides in Streptomyces venezuelae. Metab Eng 3(1):15–26. https://doi.org/10.1006/mben.2000.0167
Jung WS, Lee SK, Hong JS, Park SR, Jeong SJ, Han AR, Sohng JK, Kim BG, Choi CY, Sherman DH, Yoon YJ (2006) Heterologous expression of tylosin polyketide synthase and production of a hybrid bioactive macrolide in Streptomyces venezuelae. Appl Microbiol Biotechnol 72(4):763–769. https://doi.org/10.1007/s00253-006-0318-5
Martin JF, Liras P (2010) Engineering of regulatory cascades and networks controlling antibiotic biosynthesis in Streptomyces. Curr Opin Microbiol 13(3):263–273. https://doi.org/10.1016/j.mib.2010.02.008
Funding
This study was supported by the National Natural Science Foundation of China under Grant Number 31770049.
Author information
Authors and Affiliations
Contributions
ZF designed the experiments; LN, YG and QJ carried out the experiments; MC performed NMR measurements; SW, YL, and ZF analyzed the experimental results; ZF and LN wrote the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nie, L., Cao, M., Ji, Q. et al. Regulated Expression of an Environmental DNA-Derived Type II Polyketide Gene Cluster in Streptomyces Hosts Identified a New Tetracenomycin Derivative TCM Y. Curr Microbiol 79, 336 (2022). https://doi.org/10.1007/s00284-022-03039-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03039-5