Abstract
Tylosin polyketide synthase (Tyl PKS) was heterologously expressed in an engineered strain of Streptomyces venezuelae bearing a deletion of pikromycin PKS gene cluster using two compatible low-copy plasmids, each under the control of a pikAI promoter. The mutant strain produced 0.5 mg/l of the 16-membered ring macrolactone, tylactone, after a 4-day culture, which is a considerably reduced culture period to reach the maximum production level compared to other Streptomyces hosts. To improve the production level of tylactone, several precursors for ethylmalonyl-CoA were fed to the growing medium, leading to a 2.8-fold improvement (1.4 mg/ml); however, switching the pikAI promoter to an actI promoter had no observable effect. In addition, a small amount of desosamine-glycosylated tylactone was detected from the extract of the mutant strain, revealing that the native glycosyltransferase DesVII displayed relaxed substrate specificity in accepting the 16-membered ring macrolactone to produce the glycosylated tylactone. These results demonstrate a successful attempt for a heterologous expression of Tyl PKS in S. venezuelae and introduce S. venezuelae as a rapid heterologous expression system for the production of secondary metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyketides are useful natural products that include important therapeutic agents, such as antibacterial, immunosuppressive, and cholesterol-lowering compounds (O’Hagan 1991). The number of cloned and sequenced genes involved in polyketide biosynthesis has increased rapidly during the past decade, and increasing knowledge on genetic information has enabled the manipulation of polyketide biosynthetic pathways to yield novel compounds. However, many of the original polyketide producers are difficult to manipulate genetically, have poor growth characteristics, or yield poor titers. The transfer of biosynthetic genes from the original producers to a more amenable and robust heterologous host therefore becomes an attractive alternative to producing high levels of desired compounds or to providing a basis for subsequent combinatorial biosynthetic approaches (Pfeifer and Khosla 2001; Rude and Khosla 2004; Wenzel and Muller 2005). The most widely used heterologous hosts for polyketide production are Streptomyces coelicolor (McDaniel et al. 1995) and Streptomyces lividans (Ziermann and Betlach 1999). In these strains, a number of metabolic pathways have been successfully engineered to increase the titer of natural products themselves and to generate a variety of novel unnatural products. Saccharopolyspora erythraea (Gaisser et al. 2000; Martin et al. 2003) and Streptomyces fradiae (Rodriguez et al. 2004; Reeves et al. 2004) have been also engineered to create strains that could be used for the production of novel polyketides through heterologous expression and manipulation of polyketide biosynthetic pathways. However, these heterologous hosts have a slow growth rate and need relatively prolonged culture periods (6–9 days) to reach the maximum production level of metabolites in the fermentation process. In contrast, Streptomyces venezuelae has been recently developed as a heterologous host (Yoon et al. 2002; Jung et al. 2003; Hong et al. 2004) that requires a short culture period (3–4 days) for metabolite production compared to other Streptomyces species (Xue and Sherman 2001). It is also amenable to genetic manipulation and has high transformation efficiency. These characteristics make S. venezuelae an alternative system for a rapid heterologous production of polyketides from benchtop genetic manipulation to product fermentation (Xue and Sherman 2001).
Tylosin polyketide synthase (Tyl PKS) encoded by tylGI–V of S. fradiae is responsible for the biosynthesis of the macrolactone, tylactone, which is further modified by oxidation at C-20 and C-23 and glycosylation of mycaminose, mycinose, and mycarose to produce tylosin (Fig. 1a). Tyl PKS has a biochemical architecture similar to that of the pikromycin (Pik) PKS of S. venezuelae (Fig. 1b). Based on this similarity between Tyl PKS and Pik PKS, the hybrid Tyl–Pik modular PKS system in S. venezuelae has been developed in a previous study (Yoon et al. 2002). The expression of several hybrid PKS resulted in the production of macrolactones with predicted structural alterations and further elaborated structures by the action of post-PKS-modifying enzymes of S. venezuelae, DesVII glycosyltransferase, and PikC hydroxylase.
Organization of Tyl PKS genes and pikromycin biosynthetic gene cluster. a Organization of Tyl PKS genes and structures of tylactone and tylosin. b Organization of pikromycin biosynthetic gene cluster and structures of narbonolide and pikromycin. Each arrow represents an open reading frame (ORF). The direction of transcription and the relative sizes of ORFs are indicated
In this study, we describe the expression of the entire Tyl PKS from two compatible plasmids in a Pik-PKS-deleted strain of S. venezuelae in order to probe the applicability of S. venezuelae as a heterologous host for the expression of a whole PKS gene cluster. The resulting strain produced a 16-membered ring macrolactone, tylactone, in a considerably reduced culture period (4 days) compared to those of other Streptomyces heterologous hosts. Interestingly, tylactone was further modified probably by the action of flexible glycosyltransferase DesVII, leading to its desosamine-glycosylated derivative. Furthermore, the production level of tylactone was enhanced considerably through the feeding of several precursors of ethylmalonyl-CoA, which is expected to be the limiting substrate of Tyl PKS in S. venezuelae. This work demonstrates the unique capacity of S. venezuelae as a new system for the rapid expression of heterologous PKS and the creation of hybrid polyketides.
Materials and methods
Bacterial strains, culture conditions,and genetic manipulation
S. venezuelae ATCC15439 was used for the construction of mutant strains. The transformants of S. venezuelae were selected on R2YE agar plates (Kieser et al. 2000) by overlaying with an appropriate antibiotic or a combination of antibiotics (Yoon et al. 2002). SGGP liquid medium (Kieser et al. 2000) was used for the propagation of S. venezuelae. S. fradiae ATCC19609 grown in tryptic soy broth medium (Kieser et al. 2000) was used for the preparation of genomic DNA. Genetic manipulation was performed in Escherichia coli DH10B using standard protocols (Sambrook et al. 2001). Transformation of DNA into protoplasts of S. venezuelae followed standard protocols (Kieser et al. 2000). Litmus28 (New England Biolabs) and pGEM-3Zf(+) (Promega) were used for subcloning, and Southern blot hybridization was performed according to the protocol recommended by a digoxigenin labeling and detection kit (Roche).
Construction of pikA deletion mutant
A gene replacement plasmid (pYJ230) derived from pKC1139 (Bierman et al. 1992), which contains an apramycin-resistant marker, was constructed for the replacement of pikAI–IV genes by hygromycin resistance gene (Elhai and Wolk 1988). Plasmid pYJ230 was constructed by cloning the hygromycin resistance gene flanked by two 1-kb fragments homologous to the upstream and downstream regions of pikA. The following primers with engineered restriction sites (underlined) were used to generate the two 1-kb fragments by polymerase chain reaction (PCR): a HindIII–XbaI fragment containing the upstream region of pikAI(forward,5′-AAAAAGCTTGGGGATGTGGCGCCGGAGGATCTC-3′;reverse,5′-TTTTCTAGACATCCGGCTCCGTCTCCGGAAGCC-3′) and a KpnI–EcoRI fragment containing pikAV and part of desVIII (forward, 5′-AAAGGTACCGAGGGGGCGGGCAAGTGACCGACAGAC-3′; reverse, 5′-AAAGAATTCGGACAGCGGCCGGAGCCGCTCCAG-3′). The plasmid pYJ230 was introduced into a previously constructed pikAIII–IV deletion mutant of S. venezuelae YJ004 (Yoon et al. 2002), for plasmid-mediated homologous recombination, as described previously (Yoon et al. 2002). Several double crossover mutants were identified on the basis of their phenotype of hygromycin resistance and apramycin sensitivity, and their genotype by Southern blot hybridization. The resulting strain was designated DHS2001.
Expression of Tyl PKS under the controlof pikAI promoter
Tyl PKS genes were individually cloned into two compatible vectors, pDHS702 (Xue and Sherman 2000) and pDHS618 (Zhao et al. 1999), which are E.coli–Streptomyces shuttle vectors containing a pikAI promoter (Xue and Sherman 2001), SCP2* origin, and different antibiotic resistance genes. The tylGI–III genes were cloned into pDHS702 as a MunI–NsiI fragment by combining an EcoRI–SphI partial fragment of tylGI–III isolated from pSET506 (a gift from Eli Lilly Co.; GenBank accession number SFU78289), a MunI–EcoRI fragment of the 5′ region, and an SphI–NsiI fragment of the 3′ region of tylGI–III genes amplified by PCR to generate pBB155 (Fig. 1a). Both ends of tylGI–III were obtained by PCR using the following primers: a MunI–EcoRI fragment containing the 5′ end of tylGI (forward, 5′-AAACAATTGACACAGCTCGGTGACACGGCAGCC-3′; reverse, 5′-AAAGAATTCCTCGACACCGGCCGCGCCCGGCAG-3′) and an SphI–NsiI fragment containing the 3′ end of tylGIII (forward, 5′- AAAGCATGCAACGGCGCCTGAACCGGAGCGGCA-3′; reverse, 5'-AAAATGCATGTGTTTTCACCTGTGCTGTCGCTG-3′) (restriction sites are underlined). The plasmid pDHS3003 (Fig. 2b) expressing tylGIV–V under the control of the pikAI promoter was constructed as described previously (Yoon et al. 2002). Transformation of the DHS2001 strain by the plasmids pBB155 and pDHS3003 generated the mutant strain YJ005.
Expression of Tyl PKS expression under the controlof actI promoter
An actI promoter and a pathway-specific activator gene (actII–ORF4) (McDaniel et al. 1993) were also used for the expression of Tyl PKS. A 1.43-kb PacI–KpnI fragment containing an actI/actII–ORF4 promoter–activator pair was amplified from pHGF7505 (Yu et al. 2001) by PCR and ligated to a 313-bp KpnI–EcoRI fragment corresponding to the 5′ end of tylGI amplified by PCR from the genomic DNA of S. fradiae. The resulting PacI–EcoRI fragment was ligated to the same sites of pBB155 to generate pYJ153. The following primers with engineered restriction sites (underlined) were used: a PacI–KpnI fragment (forward, 5′-AAGCTTTTAATTAAAGTTTAAACAGCTCGGATT-3′; reverse, 5′-ATCCTTGGTACCTCGATGCGTTCGTCCGGTGGT-3′) and a KpnI–EcoRI fragment (forward, 5′-GCCGCAGGTACCGTTGGCAGGCTCACCGGCACG-3′; reverse, 5′-TTCCCAGAATTCCTCGACACCGGCCGCGCCCGG-3). The EcoRI–XbaI fragment containing tylGIV–V genes from plasmid pDHS3003 was transferred to the same sites of a pRM5 derivative (McDaniel et al. 1993), which contains the actI/actII–ORF4 promoter–activator pair and apramycin resistance gene. The resulting plasmid was designated pYJ154.
Production and analysis of heterologous polyketides
All strains were cultured on R2YE solid medium (Kieser et al. 2000) at 30°C for 4 days with appropriate antibiotic selection. Agar medium production gives more reproducible results than liquid fermentation, although liquid fermentation typically supports the same level of antibiotic production. Agar-grown culture was diced and extracted with 2 vol of methanol. The extract was washed with water, extracted again with 1 vol of chloroform, and concentrated with methanol. The extract was analyzed for polyketide production on a reverse-phase C18 column (4.6×250 mm Watchers 120 ODS-BP, 5 μm; Daiso Watchers) with a binary mobile phase (solvent A: acetonitrile/methanol=4:1 with 5 mM ammonium acetate; solvent B: 5 mM ammonium acetate in H2O; A: 20–75%, 30 min; 75%, 10 min; 75–20%, 10 min) at a flow rate of 1 ml/min (Rodriguez et al. 2004) by the combination of liquid chromatography/mass spectrometry (LC/MS) and mass spectrometry/mass spectrometry (MS/MS) using a Finnigan LCQ-DecaXP with an electrospray source. A purified tylactone standard (kindly provided by Professor Eric Cundliffe at University of Leicester) was used to generate a calibration curve for titer determination. The production level of tylactone was averaged from five separate cultivations and extractions.
Results
Generation of Pik PKS deletion mutant of S. venezuelae DHS2001
A strain for the expression of heterologous PKS (DHS2001) was constructed by deleting the native Pik PKS from the chromosome of S. venezuelae and by replacing with the hygromycin resistance gene through a plasmid-mediated homologous recombination. A pikAV gene encoding a type II thioesterase desosamine (des) biosynthetic gene cluster, a pikC gene encoding a cytochrome P450 hydroxylase, and a pikD gene encoding a pathway-specific activator (Xue et al. 1998a,b; Wilson et al. 2001) downstream of Pik PKS (Fig. 1b) remain intact in DHS2001. The genomic DNA of the resulting mutant strain was analyzed by Southern blot hybridization to confirm double crossover (data not shown). The culture extract of strain DHS2001 was analyzed, and no antibiotic production was observed. Feeding the pikromycin aglycone, narbonolide, to the growing medium of DHS2001 restored the production of pikromycin, which is synthesized through the attachment of desosamine to narbonolide at C-5 oxygen by DesVII glycosyltransferase and by the addition of a hydroxyl group at C-12 position by PikC hydroxylase (Fig. 1b). This result indicates that the remaining des cluster, the monooxygenase pikC, and the pathway-specific activator pikD genes are functional in DHS2001.
Heterologous expression of the entire Tyl PKS in S. venezuelae DHS2001
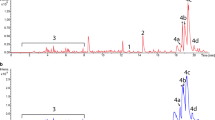
A previous study revealed that two SCP2*-derived plasmids carrying different antibiotic markers were compatible and stable during growth (Xue et al. 1999). For the expression of the entire Tyl PKS, two plasmids containing SCP2* replication origin and two different resistance genes were used. The expression plasmids pBB155 (carrying tylGI–III; Fig. 2a) and pDHS3003 (carrying tylGIV–V; Fig. 2b) were constructed and transformed into the pikA deletion mutant DHS2001. Transformants resistant to apramycin and thiostrepton were isolated and designated YJ005. The organic extract of YJ005 was analyzed by LC/MS and MS/MS for identification of tylactone. A major peak was observed at a retention time of 30.7 min with m/z=395 and 377, consistent with the calculated molecular weight for tylactone and its dehydrated form, respectively, from LC/MS analysis (Fig. 3a). MS/MS fragmentation of this compound with m/z=395 produced a pattern identical to the characteristic fragmentation pattern of a tylactone standard (data not shown), thus confirming the presence of tylactone in the growing medium of YJ005. In addition, a peak with m/z=552 was observed at a retention time of 43.0 min from LC/MS analysis (Fig. 3a), corresponding to desosaminyl tylactone, which is known to be a biologically active compound against the majority of Gram-positive bacteria (Furumai et al. 1977). The compound with m/z=552 fragmented into peaks at m/z=534, 395, 377, and 158, corresponding to the dehydrated form of desosaminyl tylactone, tylactone, dehydrated form of tylactone, and oxonium ion fragment derived from desosamine (Butler et al. 2002), respectively (Fig. 3b). This fragmentation pattern is consistent with the previous report in which 5-O-desosaminyl tylactone was synthesized by feeding tylactone to a mutant of Sa. erythraea that is deficient in eryA-encoding erythromycin PKS and both eryCIII and eryBV glycosyltransferases but containing the tylosin tylMII glycosyltransferase gene (Gaisser et al. 2000). The yield of tylactone in S. venezuelae YJ005 was approximately 0.5 mg/l, and the percentage conversion of tylactone into desosaminyl tylactone was approximately 8.3% based on the peak intensity from LC/MS analysis. The maximum level of tylactone production was reached after a 4-day culture (Fig. 4).
LC/MS and MS/MS analyses of compounds from the extract of YJ005. a A major peak was observed at a retention time of 30.7 min with m/z=395 and 377, consistent with the calculated molecular weight for tylactone and its dehydrated form, respectively. A minor peak was detected at 43.0 min with m/z=552, which corresponds to desosaminyl tylactone. b The compound with m/z=552 fragmented into peaks at m/z=158, 377, 395, and 534, corresponding to characteristic desosamine moiety ion (oxonium ion), a dehydrated form of tylactone, tylactone, and a dehydrated form of desosaminyl tylactone, respectively
Feeding several precursors for ethylmalonyl-CoAto the culture medium of YJ005
Besides malonyl-CoA and methylmalonyl-CoA, the biosynthesis of tylactone requires ethylmalonyl-CoA as an extender unit, which is incorporated into the tylosin polyketide backbone by an ethylmalonyl-specific acyltransferase in module 5 of Tyl PKS. However, Pik PKS does not require ethylmalonyl-CoA as an extender unit for the biosynthesis of pikromycin. Successful production of tylactone in S. venezuelae suggests that this strain certainly contains the biosynthetic pathway for ethylmalonyl-CoA. However, it is uncertain whether S. venezuelae contains a sufficient pool of intracellular ethylmalonyl-CoA to support the efficient biosynthesis of tylactone. To address this question, S. venezuelae YJ005 was grown on R2YE medium containing precursors for ethylmalonyl-CoA, such as butanol, butyrate, and diethylmalonate (Stassi et al. 1998). Consequently, supplementation with the precursors for ethylmalonyl-CoA into the growing medium resulted in enhancement of tylactone productivity. When 50 mM butanol, 50 mM butyrate, and 10 mM diethylmalonate were added to the growing medium during inoculation, the production of tylactone increased by about 1.4-fold (0.7 mg/l), 1.6-fold (0.8 mg/l), and 2.8-fold (1.4 mg/l), respectively.
Tyl PKS expression under the control of actI promoter
The actI promoter and its cognate activator gene actII–ORF4 from the actinorhodin biosynthetic gene cluster of S. coelicolor (McDaniel et al. 1993) have been widely used to express several large PKS genes in genetically engineered “clean” host strains of S. coelicolor and S. lividans (Pfeifer and Khosla 2001). Erythromycin PKS was expressed under the control of the actI promoter after integration into the chromosome by homologous recombination in a mutant strain of Sa. erythraea that is deficient in native PKS, and the resulting strain produced a high quantity of erythromycin when compared with the wild-type Sa. erythraea (Rowe et al. 1998). In contrast, use of this promoter–activator in a Sa. erythraea overproducer resulted in a >20-fold decrease in erythromycin titers compared to the strain using a native eryAI promoter. (Rodriguez et al. 2003). To investigate the effect of promoters on the productivity of heterologous polyketide in S. venezuelae, the actI promoter and the actII–ORF4 activator gene were tested to express Tyl PKS in the S. venezuelae mutant strain. The plasmids containing Tyl PKS under the control of the actI promoter were constructed and introduced into the pikA deletion mutant DHS2001 to generate YJ048. The organic extract of YJ048 was analyzed by LC/MS and MS/MS. The calculated molecular weights corresponding to tylactone and desosaminyl tylactone were detected. The amount of tylactone produced from YJ048 was similar to levels produced from the YJ005 strain, demonstrating that substitution of the pikAI promoter with the actI promoter did not affect the production level of heterologous polyketides in this host–vector system.
Discussion
S. venezuelae is a suitable heterologous host for the expression of polyketide biosynthetic genes due to its a fast growth, which allows large quantities of cell mass and metabolites to be harvested relatively quickly compared to other Streptomyces species (Xue and Sherman 2001). Development of the genetically engineered strain of S. venezuelae DHS2001 and rapid production of tylactone in the mutant strain validated the unique advantage of S. venezuelae as an attractive host for the production of heterologous polyketides. In addition, biosynthetic pathways for the three most common precursors found in modular polyketide biosynthesis (malonyl-CoA, methylmalonyl-CoA, and ethylmalonyl-CoA) seem to be present in S. venezuelae as proven by the production of tylactone from the YJ005 strain, which requires all of the three common precursors. The presence of a range of precursors for the biosynthesis of polyketides in S. venezuelae would facilitate the heterologous expression of a diverse class of PKS.
The flexibility of DesVII glycosyltransferase in accepting new sugar donors as substrate has been demonstrated from previous reports (Zhao et al. 1998a,b, 2001; Yamase et al. 2000; Borisova et al. 1999; Hong et al. 2004). DesVII has also been shown to be flexible toward hybrid aglycones, to which the sugar is attached (Yoon et al. 2002). The unique property of DesVII has allowed researchers to explore the possibility of generating novel macrolide derivatives attached with new sugars. The current study takes this a step further by showing that DesVII, which normally accepts both 12- and 14-membered ring macrolactones, is capable of accepting a 16-membered ring, tylactone, as an aglycone substrate. The successful generation of desosaminyl tylactone has further expanded the usefulness of glycosyltransferases as a tool for the generation of new hybrid polyketide structures. In another study, a pikromycin producer (Streptomyces sp. no. 4,900, in which the activity of the KS domain of PKS was inhibited by cerulenin) also successfully converted tylactone (200 mg) into 5-O-desosaminyl tylactone (30 mg) with a 15% conversion yield (Omura et al. 1980), which is approximately 2-fold higher than that of S. venezuelae (DesVII). The glycosyltransferase of another pikromycin producer Streptomyces sp. no. 4,900 seems to have a higher efficiency in converting tylactone to desosaminyl tylactone than DesVII, and its catalytic efficiency toward unnatural substrates would be worth investigating further.
Although the glycosylation site of tylactone was not determined due to this low conversion yield, it is likely that desosamine was introduced to the oxygen at position C-5 of tylactone, according to the intrinsic catalytic specificity of DesVII. Further analysis, such as by nuclear magnetic resonance spectroscopy, is required to confirm the glycosylation site of tylactone. No hydroxylated forms of tylactone and desosaminyl tylactone were detected, although the monooxygenase PikC of S. venezuelae was known to have considerably relaxed substrate specificity (Yoon et al. 2002; Lee et al. 2005) toward alternative substrates, indicating that tylactone and desosaminyl tylactone may be poor substrates for PikC.
The level of production of tylactone in S. venezuelae is relatively low and requires further improvement. In this study, two simple strategies were employed to enhance the production level of tylactone. First, the limiting precursor was added to the growing medium. Ethylmalonyl-CoA, which is required for the biosynthesis of tylactone, was expected to be limiting in the S. venezuelae system since pikromycin production in wild type does not require ethylmalonyl-CoA as a precursor. Supplementation with several precursors for ethylmalonyl-CoA, such as butanol, butyrate, or diethylmalonate, into the growing medium led to improvement in tylactone production, suggesting that ethylmalonyl-CoA may be a limiting factor for the biosynthesis of tylactone in S. venezuelae. Second, Tyl PKS was expressed under the control of the actI promoter. Similar levels of tylactone production were observed when Tyl PKS was expressed under the control of either an actI promoter or a pikAI promoter. Switching promoters had no observable effect, although expression levels of Tyl PKS may have been different in using the actI promoter or the pikAI promoter. Taken together, the major factor limiting the production level of tylactone seems to be deficiency in the intracellular pool of ethylmalonyl-CoA rather than the expression level of PKS in this system. Previous work revealed that Tyl PKS genes have an independent expression pattern in which tylGI and tylGIII–V are under differential regulation in S. fradiae (Stratigopoulos and Cundliffe 2002). According to this observation, tylGIII expression in pBB155 and pYJ153 could be expressed independently under the control of its intrinsic promoter in S. venezuelae, which would be another explanation for the fact that switching the pikA promoter by actI/actII–ORF4 had no observable effect on tylactone production.
The newly developed S. venezuelae system described in this study has both promising and limiting factors as a heterologous host. The limiting factor is the relatively low production yield of heterologous polyketides compared to other well-developed Streptomyces host. Features of S. venezuelae, including rapid growth rate and quick production of metabolites, are favorable characteristics for heterologous polyketide production that cannot be easily endowed by genetic engineering. Reducing the time for bacteria culture has important implications for the production of metabolites through the fermentation process; thus, this unique advantage of S. venezuelae as a fast-growing bacteria (doubling time, approximately 1 h) is an attractive feature in developing a heterologous host. Further works on enhancing the production level of heterologous polyketides through metabolite engineering strategies will facilitate the development of S. venezuelae as a competent heterologous host.
References
Bierman M, Logan R, O’Brien K, Seno ET, Rao RN, Schoner BE (1992) Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces spp. Gene 116:43–49
Borisova SA, Zhao L, Sherman DH, Liu HW (1999) Biosynthesis of desosamine: construction of a new macrolide carrying a genetically designed sugar moiety. Org Lett 1:133–136
Butler AR, Bate N, Kiehl DE, Kirst HA, Cundliffe E (2002) Genetic engineering of aminodeoxyhexose biosynthesis in Streptomyces fradiae. Nat Biotechnol 20:713–716
Elhai J, Wolk CP (1988) A versatile class of positive-selection vectors based on the nonviability of palindrome-containing plasmids that allows cloning into long polylinkers. Gene 68:119–138
Furumai T, Maezawa I, Matsuzawa N, Yano S, Yamaguchi T, Takeda K, Okuda T (1977) Macrolide antibiotics M-4365 produced by micromonospora I. Taxonomy, production, isolation, characterization and properties. J Antibiot 30:443–449
Gaisser S, Reather J, Wirtz G, Kellenberger L, Staunton J, Leadlay P (2000) A defined system for hybrid macrolide biosynthesis in Saccharopolyspora erythraea. Mol Microbiol 36:391–401
Hong JSJ, Park SH, Choi CY, Sohng JK, Yoon YJ (2004) New olivosyl derivatives of methymycin/pikromycin from an engineered strain of Streptomyces venezuelae. FEMS Microbiol Lett 238:291–399
Jung WS, Kim ES, Kang HY, Choi CY, Sherman DH, Yoon YJ (2003) Site-directed mutagenesis on putative macrolactone ring size determinant in the hybrid pikromycin–tylosin polyketide synthase. J Microbiol Biotechnol 13:823–827
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hoopwood DA (2000) Practical Streptomyces genetics. John Innes Centre, Norwich
Lee SK, Basnet DB, Hong JSJ, Jung WS, Choi CY, Lee HC, Sohng JK, Ryu KG, Kim DJ, Ahn JS, Kim BS, Oh HC, Sherman DH, Yoon YJ (2005) Structural diversification of macrolactones by substrate-flexible cytochrome P450 monooxygenase. Adv Synth Catal 347:1369–1378
Martin CJ, Timoney MC, Sheridan RM, Kendrew SG, Wilkinson B, Staunton J, Leadlay PF (2003) Heterologous expression in Saccharopolyspora erythraea of a pentaketide synthase derived from the spinosyn polyketide synthase. Org Biomol Chem 1:4144–4147
McDaniel R, Ebert-Khosla S, Hopwood DA, Khosla C (1993) Engineered biosynthesis of novel polyketides. Science 262:1546–1550
McDaniel R, Ebert-Khosla S, Hopwood DA, Khosla C (1995) Rational design of aromatic polyketide natural products by recombinant assembly of enzymatic subunits. Nature 375:549–554
O’Hagan D (1991) The polyketide metabolites. Ellis Horwood, Chichester
Omura S, Ikeda H, Matsubara H, Sadakane N (1980) Hybrid biosynthesis and absolute configuration of macrolide antibiotic M-4365 G1. J Antibiot 33:1570–1572
Pfeifer BA, Khosla C (2001) Biosynthesis of polyketides in heterologous hosts. Microbiol Mol Biol Rev 65:106–118
Reeves CD, Ward SL, Revill WP, Suzuki H, Marcus M, Petrakovsky OV, Marquez S, Fu H, Dong SD, Katz L (2004) Production of hybrid 16-membered macrolides by expressing combinations of polyketide synthase genes in engineered Streptomyces fradiae hosts. Chem Biol 11:1465–1472
Rodriguez E, Hu Z, Ou S, Volchegursky Y, Hutchinson CR, McDaniel R (2003) Rapid engineering of polyketide overproduction by gene transfer to industrially optimized strains. J Ind Microbiol Biotechnol 30:480–488
Rodriguez E, Ward S, Fu H, Revil WP, McDaniel R, Katz L (2004) Engineered biosynthesis of 16-membered macrolides that require methoxymalonyl-ACP precursors in Streptomyces fradiae. Appl Microbiol Biotechnol 66:85–91
Rowe CJ, Cortes J, Gaisser S, Staunton J, Leadlay PF (1998) Construction of new vectors for high level expression in actinomycetes. Gene 216:215–223
Rude MA, Khosla C (2004) Engineered biosynthesis of polyketides in heterologous hosts. Chem Eng Sci 59:4693–4701
Sambrook J, Fritsch EF, Maniatis T (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Stassi DL, Kakavas SJ, Reynolds KA, Gunawardana G, Swanson S, Zeidner D, Jackson M, Liu H, Buko A, Katz L (1998) Ethyl-substituted erythromycin derivatives produced by directed metabolic engineering. Proc Natl Acad Sci U S A 95:7305–7309
Stratigopoulos G, Cundliffe E (2002) Expression analysis of the tylosin–biosynthetic gene cluster: pivotal regulatory role of the tylQ product. Chem Biol 9:71–78
Wenzel SC, Muller R (2005) Recent developments towards the heterologous expression of complex bacterial natural product biosynthetic pathways. Curr Opin Biotechnol 16:1–13
Wilson DJ, Xue Y, Reynolds KA, Sherman DH (2001) Characterization and analysis of the PikD regulatory factor in the pikromycin biosynthetic pathway of Streptomyces venezuelae. J Bacteriol 183:3468–3475
Xue Y, Sherman DH (2000) Alternative modular polyketide synthase expression controls macrolactone structure. Nature 403:571–575
Xue Y, Sherman DH (2001) Biosynthesis and combinatorial biosynthesis of pikromycin-related macrolides in Streptomyces venezuelae. Metab Eng 3:15–26
Xue Y, Zhao L, Liu HW, Sherman DH (1998a) A gene cluster for macrolide antibiotic biosynthesis in Streptomyces venezuelae: architecture of metabolic diversity. Proc Natl Acad Sci U S A 95:12111–12116
Xue Y, Wilson D, Zhao L, Liu H, Sherman DH (1998b) Hydroxylation of macrolactone YC-17 and narbomycin in mediated by the pikC-encoded cytochrome P450 in Streptomyces venezuelae. Chem Biol 5:661–667
Xue Q, Ashley G, Hutchinson CR, Santi DV (1999) A multiplasmid approach to preparing large libraries of polyketides. Proc Natl Acad Sci U S A 96:11740–11745
Yamase H, Zhao L, Liu HW (2000) Engineering a hybrid sugar biosynthetic pathway: production of l-rhamnose and its implication on dihydrostreptose biosynthesis. J Am Chem Soc 122:12397–12398
Yoon YJ, Beck JB, Kim BS, Kang HY, Reynolds KA, Sherman DH (2002) Generation of multiple bioactive macrolides by hybrid modular polyketide synthases in Streptomyces venezuelae. Chem Biol 9:203–214
Yu TW, Muller R, Muller M, Zhang X, Draeger G, Kim CG, Leistner E, Floss HG (2001) Mutational analysis and reconstituted expression of the biosynthetic genes involved in the formation of 3-amino-5-hydroxybenzoic acid, starter unit of rifamycin biosynthesis in Amycolatopsis mediterranei S699. J Biol Chem 276:12546–12555
Zhao L, Sherman DH, Liu HW (1998a) Biosynthesis of desosamine: construction of a new methymycin/neomethymycin analogue by deletion of a desosamine biosynthetic gene. J Am Chem Soc 120:10256–10257
Zhao L, Que NLS, Xue Y, Sherman DH, Liu HW (1998b) Mechanistic studies of desosamine biosynthesis: C-4 deoxygenation precedes C-3 transamination. J Am Chem Soc 120:12159–12160
Zhao L, Ahlert J, Xue YQ, Thorson JS, Sherman DH, Liu HW (1999) Engineering a methymycin/pikromycin–calicheamicin hybrid: construction of two new macrolides carrying a designed sugar moiety. J Am Chem Soc 121:9881–9882
Zhao L, Borisova SA, Yeung SM, Liu HW (2001) Study of C-4 deoxygenation in the biosynthesis of desosamine: evidence implicating a novel mechanism. J Am Chem Soc 123:7909–7910
Ziermann R, Betlach MC (1999) Recombinant polyketide synthesis in Streptomyces: engineering of improved host strain. Biotechniques 26:106–110
Acknowledgements
We thank Dr. Brian J. Beck for providing pBB155 and Dr. Y.S. Hong for help in constructing S. venezuelae DHS2001. We are also grateful to Professor Eric Cundliffe (University of Leicester) for kindly providing a tylactone standard. This work was supported by a grant (20050410-034-682-149-00-00) from the BioGreen 21 Program, Rural Development Administration, the 21C Frontier Microbial Genomics and Application Center Program, Ministry of Science and Technology (grant MG05-0303-4-0), Republic of Korea, and the SRC program of the Korea Science and Engineering Foundation (KOSEF) through the Center for Intelligent Nano-Bio Materials at Ewha Woman’s University (grant R11-2005-008-00000-0).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jung, W.S., Lee, S.K., Hong, J.S.J. et al. Heterologous expression of tylosin polyketide synthase and production of a hybrid bioactive macrolide in Streptomyces venezuelae . Appl Microbiol Biotechnol 72, 763–769 (2006). https://doi.org/10.1007/s00253-006-0318-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-006-0318-5