Abstract
The novel Enterobacter strains TS1L and TS3, isolated from textile wastewater, showed a good ability to decolourise Basic Red 9 (BR9). The effects of various physicochemical parameters on decolourisation efficiency were evaluated using both single and mixed culture of Enterobacter sp. The optimal conditions for the decolourising activity of strains TS1L, TS3 and a mixed culture were as follows: textile wastewater as sole substrate without glucose addition, pH 7.0, 150 rpm, 35 °C and 12 h of incubation. The highest decolourisation rate was observed at 81.15% for the single culture of strain TS1L. Moreover, TS1L not only reduced BR9 in wastewater, but also improved the quality of the water under optimal conditions. The treated wastewater met the criteria of the Water Quality Standard (Thailand). Based on gas chromatography-mass spectrometry, TS1L completely degraded BR9 and converted it into organic compounds. To our knowledge, this is the first report of Enterobacter with the ability to decolourise BR9 dye.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
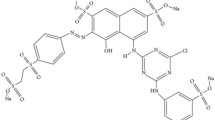
Synthetic dyes, organic compounds, have been widely used in various industries since 1856 [1, 2]. The use of synthetic dyes and pigments increases constantly and is estimated to reach a value of 63 billion USD in 2022, with a 29% of growth rate [3]. There are several types of synthetic dyes, including Basic Red 9 (BR9) Monohydrochloride (also known as benzene-mine 4-(4aminophenyl)(4-imino-2,5-cyclohexadian-1-ylidene)-methyl monohydro- chloride), which is a popular dye used in the textile and medical industry. It is an inflammable triphenylmethane dye used as a colouring agent [4], characterised as a solid, red colour with a green metallic sheen, slightly soluble in water and ether. However, it shows good solubility in ethylene glycol methyl ether, ethanol, benzyl alcohol and methanol [5]. As BR9 is classified as carcinogenic, its use is of particular concern, especially regarding treatment and final disposal [6].
The affectation of hydric systems is one of the main problems generated as a consequence of population growth and industrialisation. Every day a significant volume of domestic effluent to develop and optimise systems that, in addition to mitigating the adverse effects caused by its discharge, meet the provisions established in the corresponding environmental regulations. There are several methods for the treatment of BR9, including physical, chemical and biological methods [1, 7, 8]. Physical (adsorption, precipitation, coagulation, filtration, electrolysis, photodegradation) and chemical (oxidation) methods are most frequently used because of their high decolourisation efficiency. However, the disadvantages include high operation costs, limited versatility, interference with other wastewater compounds, formation of dangerous intermediates and high energy requirement [9].
Due to its overall environmental impact, the residual dye in the wastewater from the synthetic dye manufacturing and textile industries is a global concern. The discharge contains a high content of pigments and other additives, possessing complex structures. As per the requirement for dyed clothing, dyestuff in the effluent is less susceptible to acids, bases, and oxygen. Thus, conventional physical and chemical methods are not always efficient in degrading the dyes. Some microorganisms growing in an area affected with textile effluent have the capability to utilise the dyes as a source of carbon or nitrogen or both. As a very clean, inexpensive, and sufficient alternative, bioremediation of textile wastewater using these microorganisms has gained major popularity [10, 11]. Therefore, biological treatment options are currently receiving increased interest as they are highly effective, can completely convert synthetic dye into CO2 and water, are environmentally friendly and have low operation costs [12, 13]. In Thailand, BR9 is mostly used for dyeing woven bulrush mats. After dye processing, BR9-containing wastewater is normally disposed off in open ponds and treated via physical methods. However, this conventional approach requires long treatment times and results in the release of residues into natural water bodies, with the potential accumulation of toxic substances. Biological treatment of BR9 has been reported using Trametes hirsuta and immobilised laccase from T. hirsuta. Using immobilised laccase, up to 80% of toxicity was removed from synthetic dye [14]. However, there are few studies on the degradation of BR9 by biological methods using Enterobacter species. It has been reported that Enterobacter species can be decolorisation/biodegradation of BR9 and metabolise related/other structures of triphenylmethane (such as Basic Green 1, Acid Blue 93, Basic Violet 3 and Basic Green 4) using oxidoreductive enzymes [15].
We determined the optimisation of various parameters (glucose amount, pH, temperature and agitation speed) to achieve maximum BR9 degradation using single and mixed cultures of a novel isolate of Enterobacter. The various intermediates formed were analysed during the degradation of BR9 using gas chromatography-mass spectrometry (GC–MS) and ultraviolet–visible (UV–Vis) techniques. Textile wastewater before and after treatment with Enterobacter was characterised, and the parameters were compared with the guidelines of the Thailand Water Quality Standard.
Material and Methods
Sample Collection
Textile wastewater was collected from the Ban Phraek Weaving Group (Phatthalung, Thailand). Samples were kept on ice and transferred to the laboratory. We determined the parameters pH, total dissolved solids (TDS), settleable solids, total suspended solids (TSS), oil and grease, sulphide, total Kjeldahl nitrogen (TKN), chemical oxygen demand (COD) and colour.
Textile Wastewater Characterisation
Textile wastewater was characterised using the standard method. The pH was measured using a pH meter. The parameters TDS, settleable solid, TSS, TKN and oil and grease were determined following American Public Health Association (APHA) [16, 17]. Sulphide colouring reagent and hydrochloric acid were used to determine the sulphide content. The sulphide concentration was measured with a spectrophotometer at a wavelength 667 nm [18]. Wastewater colour was also determined using a spectrophotometric method (OD 200–800 nm) [19]. The COD was evaluated using a Spectroquant® COD Cell Test (Germany) following the manufacturer´s recommendations. Textile wastewater was added to standard and miniaturised COD ampules and diluted if necessary. Subsequently, the samples were incubated at 150 °C for 2 h in a dry incubator and kept at room temperature. The COD concentration was measured using a UV–visible spectrophotometer at a wavelength of 600 nm. A standard or semi-micro cuvette was used to maintain a 1 cm path length of the sample [20].
Bacterial Isolates and Inoculum Preparation
Two bacterial isolates of the Enterobacter sp. strains TS1L and TS3 (GenBank numbers MN508471 and MN508473, respectively) were used throughout this study. These strains have been isolated from textile wastewater from the Ban Phraek Weaving Group (Phatthalung, Thailand) by Rakkan et al. [21]. The isolates were kept in polyhydroxyalkanoate (PHA) producing agar plate containing 2 g/L (NH4)2SO4, 13.3 g/L KH2PO4, 1.2 g/L MgSO4.7H2O, 1.7 g/L citric acid, 10 mL/L trace element and 15 g/L agar at pH 7 [22]. The inoculum was prepared in 250 mL Erlenmeyer flasks containing 50 mL of PHA producing medium; 10% of each isolate was added to the medium and incubated at 35 °C, 150 rpm for 24 h.
Effects of Cultivation Parameters on Decolourisation Efficiency
The effects of cultivation parameters on decolourisation efficiency were determined in Erlenmeyer flasks containing 50 mL of textile wastewater medium (TWM) containing 100% textile wastewater as sole substrate without any supplements. We added 10% of inoculum containing 1 × 106 cells/mL or an OD600 nm of 0.6. The effect of glucose addition (0–12 g/L) was determined while the other parameters were fixed. Subsequently, pH (6–8), temperature (20–45 °C) and agitation speed (100–300 rpm) were evaluated. The optimal condition was also studied for both single and mixed Enterobacter sp. (TS1L and TS3) cultures. For the mixed culture, 5% of each strain was mixed and inoculated into TWM. Samples were collected every 12 h to measure decolourisation efficiency.
Decolourisation Measurement
The decolourisation efficiency and the concentration of BR9 were measured in a UV–Vis spectrophotometer at a wavelength of 540 nm. The samples were collected and centrifuged at 10,000 × g for 10 min prior to measurements [23]. Decolourisation efficiency (expressed as % of decolourisation) was calculated as follows:
To determine the BR9 concentration, at first, a calibration cure (R2 = 0.996) was drawn based on the absorbance levels corresponding to samples of known concentration. Then, the following expression was used to calculate the dye concentration:
Total Cell Mass Analysis
Total cell mass was determined by weighing the cell dry mass (CDM). Briefly, 10 mL samples were collected and centrifuged at 13,000×g at 4 °C for 15 min. The pellet was resuspended in 10 mL of distilled water and centrifuged again for washing. Washed cells were dried at 105 °C for 24 h in a hot air oven and then cooled down in a desiccator [24]. The samples were weighted, and the total cell mass was reported in g/L.
Determination of Decolourisation and Biodegradation Metabolites
A UV–Vis spectrophotometer and GC–MS were used for analysis of products generated during decolourisation and biodegradation by Enterobacter sp. Samples were taken at different time points and centrifuged at 10,000 × g for 10 min. A TWM without bacteria was used as control. An aliquot of 5 mL was analysed in a Perkin–Elmer Lambda 4 B Accessory Interface UV–Vis Spectrophotometer at 540 nm [23, 25].
Degradation metabolites were also determined using an Agilent 7890B Series with an Agilent 5977 mass selective detector. The mass spectrometer was operated in the electron impact mode with an electron current of 70 eV. For this, 1 mL aliquots were injected with an auto sampler (AUC20i) in split mode via a GC inlet (injector temperature 250 ͦ C). A HP-5MS capillary column (30 m × 0.25 mm ID, 0.25 µm film thickness) was connected directly to the ion source of the mass spectrometer. The oven temperature was kept isothermal for 1 min at 50 °C and then increased to 270 °C at a rate of 10 °C/min. The injector, MS source and MS quad temperatures were 250, 230 and 150 °C, respectively. The GC–MS system was operated in full scan (m/z 50–500). The biodegradation products were identified by comparison of retention time and fragmentation patterns with mass spectra in the NIST spectral library [23, 26].
Statistical Analysis
All experiments were run in triplicate. A completely randomised design was used throughout this study. Data were subjected to analysis of variance (ANOVA), and mean comparison was carried out using Duncan’s multiple range test [27]. All analyses were performed using the statistical package for social science, SPSS (SPSS 24 for windows, SPSS Inc., Chicago, IL, USA).
Results and Discussion
Characterisation of Textile Wastewater Prior to Degradation
Textile wastewater from the Ban Phraek Weaving Group (Phatthalung, Thailand) was collected after the dyeing and weaving of bulrush mat products. The characteristics of the wastewater are shown in Table 1. Prior to degradation, the characteristics of the wastewater were as follows: pH 6.92, a dark red colour, high TDS 407 mg/L, settleable 3.0 mg/L, TSS 20 mg/L, oil and grease 2 mg/L, sulphide 0.02 mg/L, TKN 40 mg/L and COD 5,600 mg/L. In addition, the initial concentration of BR9 in textile wastewater was 80.94 mg/L. The optimum pH for bacterial growth is 6.5–7.5 [28]. Because of its optimal pH and high carbon and nitrogen contents, the textile wastewater from the Ban Phraek Weaving Group was suitable for bacterial growth and was therefore used as sole substrate for Enterobacter sp. without prior pre-treatment.
Effects of Cultivation Parameters on Decolourisation Efficiency
In our previous research, we isolated five novel strains of Enterobacter sp. (TW1L, TS1P, TS1L, TS3 and TW1P) from textile wastewater containing BR9. All strains were capable of growing in synthetic medium containing BR9, with the ability to decolourise the wastewater. Among them, TS1L and TS3 showed the highest dye decolourisation efficiencies of 79.15 and 63.43%, respectively [20] and were therefore used in the current study. Cultivation parameters affected the decolourisation and degradation of BR9, including glucose addition, pH, temperature, aeration and agitation to achieve maximum BR9 degradation using single and mixed cultures of a novel isolate of Enterobacter.
Effect of Glucose Addition
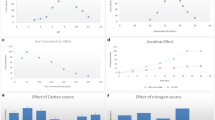
The effect of glucose was evaluated using 100% TWM with glucose addition of 0–12 g/L. The other cultivation parameters were kept constant at 150 rpm and 35 °C for 48 h. A sample was collected every 12 h for determination of cell mass and decolourisation efficiency. The maximum textile dye decolourisation, with a decolorisation of 81.15% (corresponding to final BR9 concentration at 15.26 mg/L), was observed under TWM without glucose addition (Fig. 1a). When the glucose concentration was increased to 2–12 g/L, the decolourisation efficiency decreased. This suggests that the bacteria prefer to degrade glucose because of its less complex structure compared to dye molecules [29,30,31]. This result may contrast with the degradation of triphenylmethane dyes (Basic violet 3 and Malachite green) and Amido black 10B, which require the addition of glucose for biological degradation [26, 32]. However, only low doses of glucose were added, while the decolourisation of Amido black 10B by P. chrysosporium decreased when the glucose concentration was increased to more than 0.5% [33].
Effect of Initial pH Value on Decolourisation
The pH is another important factor impacting decolourisation. Figure 1b shows that Enterobacter sp. TS1L grown on TWM had the highest decolourisation (81.59%) at pH 8, although there was no statistical difference to the decolourisation efficiency obtained at pH 7 (81.15%). Corresponding with the BR9 concentration was also decrease from 80.94 mg/L to 14.82 and 15.26 mg/L when strain TS1L grown at pH 7 and 8, respectively. In addition, the pH was measured every 6 h during experiment. The pH was slightly decreased to 6.9 when initial pH was 7. Similar trend was also observed with other pH. We therefore suggest that strain TS1L is an efficient decolouriser of TWM at neutral and mildly alkaline pH levels (pH 7–8). However, its decolourisation efficiency at pH 6 was poor, reaching only 32.78%. This can be explained by the positive charge at acidic pH levels, preventing the binding of the positively charged BR9 molecule, while at pH 7–8, the bacterial surface was negatively charged, resulting in an increased electrostatic force between the bacterial cell and the positively charged BR9 molecule [34]. Based on previous studies, the optimum pH for biological decolourisation ranges between 7 and 8 [26, 35,36,37]. Malachite green could be effectively be decolourised at pH 7–8 using Penicillium ochrochloron and the Enterobacter asburiae strain XJUHX-4TM [35, 37]. In another study, the degradation rate of acid violet 19 by P. aeruginosa BCH reached 98% within 30 min at pH 7 [35].
Effect of Incubation Temperature
The decolourisation efficiency is high at temperatures optimal for bacterial growth. Higher cultivation temperatures may increase respiration and substrate use of bacteria, which may increase the decolourisation efficiency. Therefore, in our experiment, the initial incubation temperature of 20–35 °C was adjusted. The highest decolourisation of 81.15% was achieved at 35 °C. However, decolourisation decreased (70–75%) when the cultivation temperature increased (Fig. 1c). High concentration of BR9 (20.40–24.28 mg/L) was observed under high cultivation temperature. High temperature may inactivate the enzyme responsible for decolourisation, cause loss of cell viability and affect cell structures such as the cell membrane, leading to a lower decolourisation efficiency [38,39,40]. According to a previous study, optimal degradation efficiency can be achieved at 30 to 35 °C. The use of a bacterial consortium for Congo Red degradation also showed the highest decolourisation rate (98%) at a temperature of 37 °C [41, 42]. Based on our results, the temperature of 35 °C is the optimal cultivation temperature for Enterobacter sp.
Effect of Agitation Speed
Biodegradation and decolourisation of the dyes by aerobic bacteria require oxygen for cell growth and maintenance of cell viability. Therefore, agitation is necessary to provide the oxygen and to increase the oxygen gas–liquid mass transfer [41, 42]. Hence, the decolourisation efficiency of TWM by strain TS1L was determined at different agitation speeds (100–300 rpm). Figure 1d shows that the decolourisation rate increased with an increase in agitation speed from 100 to 150 rpm. The maximum decolourisation efficiency was 81.15% at an agitation speed of 150 rpm. Similarly, Fu and Viraraghavan, [43] and Parshetti et al. [44] reported that a lower agitation rate between 100 and 150 rpm was more beneficial than static conditions in achieving better decolourisation results with fungi. However, an increase in agitation speed to 200 rpm drastically affected the decolourisation activity of the strain TS1L, which was significantly decreased at 200, 250 and 300 rpm. Based on this, the optimum agitation speed for Enterobacter sp. is 150 rpm.
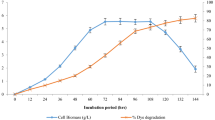
Moreover, Fig. 1a–d indicated that BR9 removal was mainly occurred by biodegradation using Enterbacter culture. The decolourisation efficiency increased significantly from 0% at 0 h to > 70% after 12 h of incubation. However, the decolourisation of textile wastewater may be caused by biosorption by biomass of Enterobacter. Rani et al. [45] and Harry-asobara and Kame [46] stated that biosorption of dyes essentially occurs through adsorption by physical forces, entrapment in inner spaces of Enterobacter mycelia, precipitation, complexation, ion exchange owing to surface ionisation and hydrogen bond formation. The crystal violet was removed by biosorption on the mycelia of Enterobacter sp. TN3W-14 [46]. However, it is not unusual to demonstrate only enzyme-mediated degradation or both biosorption and enzyme degradation in the decolorisation of textile dye.
Effect of Single and Mixed Enterobacter sp.
As mentioned previously, the single strain TS1L shows the highest decolourisation efficiency of 81.15% in TWM without glucose addition, pH 7, 150 rpm and 35 °C. However, the use of mixed bacterial cultures may increase the decolourisation efficiency [47,48,49]. We therefore evaluated the cultivation of single and mixed cultures of the newly isolated Enterobacter under optimal conditions. The highest decolourisation efficiency of each culture was similar conditions. However, the results indicate that the single culture of Enterobacter TS1L showed the highest decolourisation efficiency (81.15%), followed by the single culture of strain TS3 (79.45%) (Table 2). The lowest decolourisation efficiency (78.43%) was observed for the mixed culture of TS1L and TS3. Similar with the final concentration of BR9 at 15.26, 16.63–17.46 mg/L was observed using single and mixed culture, respectively. This is the first report of the high decolourisation capacity of a single strain of Enterobacter. The addition of mixed culture of Enterobacter to the TWM may inhibit the reciprocal bacterial activity, resulting in decreased dye degradation. Moreover, pure strains are often specific to a certain type of dye, and therefore, the cultivation of a single strain in a medium containing only one dye should be more efficient than the use of a mixed culture [50].
Characterisation of Textile Wastewater After Degradation
The parameters of the wastewater after biological degradation using Enterobacter sp. are shown in Table 1. The initial pH of 6.92 was lowered to 4.55 after degradation. This result is in agreement with the findings of the GC–MS analysis, indicating acidic compounds after degradation (Fig. 2). Moreover, the final BR9 concentration was decreased from 80.94 mg/L to 15.26 mg/L. BR9 degradation occurs in 2 mechanisms. The first mechanism of BR9 was changed to oxime-, methoxy-phenyl by oxidoreductase. Afterward, oxime-, methoxy-phenyl changed to pyruvate which can be converted into acetyl-CoA by dearomatisation and amino acid. The acetyl-CoA undergoes the tricarboxylic acid cycle (TCA cycle) to produce NADH2 and FADH2 (substrates of electron transport chain). The second mechanism, BR9 degradation, oxidative deamination and carboxylation lead to produce phthalic acid, di(2-propylpentyl) ester, which can be transformed into different fatty acids and aldehydes (carbonic acid, allyl nonyl ester, tetradecanoic acid, hexadecanoic acid methyl ester, hexadecanoic acid, bis(2-ethylhexyl) ester, 9-decanoic acid, squalene, n-hexadecanoic acid, octadecanoic acid and hexadecanoic acid ethyl ester). The phthalate, fatty acids and aldehydes can directly/indirectly enter into fatty acid oxidation reactions (beta-oxidation) to produce acetyl-CoA, NADH2 and FADH2 [10].
Total dissolved solids (mg/L), settleable solids (mg/L) and total suspended solids (mg/L) after degradation were 329, < 0.1 and 13, respectively. Notably, all parameters related to water quality were significantly decreased. The COD was decreased from 5,600 to 1,970 mg/L, indicating that Enterobacter sp. not only degraded BR9, but also decomposed solid and organic substrates in the wastewater. The quality of the wastewater after treatment was determined using the guidelines of the Quality Standard (Thailand). Regarding the parameters such as pH, TDS, settleable solids, TSS, oil and grease as well as sulphide, these guidelines were met (Table 1). The colour of the wastewater changed from red to colourless. According to our results, the quality of the textile wastewater was improved dramatically after biological degradation using Enterobacter sp. However, high value of nitrogen was detected after treatment which is over than the water quality standard and could, therefore, cause eutrophication if discharged untreated. Therefore, the study on nitrogen reduction using physical, chemical or biological methods will be studied before discharge the textile wastewater to environment.
The GC–MS indicated that BR9 was completely degraded into organic compounds within 12 h of cultivation. Based on the results of our GC–MS analysis of intermediate products, the degradation pathway of BR9 by strain TS1L is proposed in Fig. 2. Specifically, BR9 degradation occurs via the stepwise oxidation, ring fission and stepwise polymerisation. The sequence of products formed during degradation is as follows: carbonic acid, allyl nonyl ester, tetradecanoic acid, hexadecanoic acid methyl ester, hexadecanoic acid, bis(2-ethylhexyl) ester, 9-decanoic acid, squalene, n-hexadecanoic acid, octadecanoic acid, hexadecanoic acid ethyl ester, 2,3-butanediol, urea, 2-butanol, 2-propanone, 1-hydroxy and acetic acid methoxy. This is the first report on the BR9 degradation pathway using Enterobacter sp. However, when using the blue green algae Hydrocoleum oligotrichum and Oscillatoria limnetica for BR9 degradation, only two compounds were yielded, namely aminobenzoic acid and N-Dimethyl-p-phenylenedi-amine [10]. The decrease in BR9 was also confirmed via UV–visible spectral analysis. The UV–visible spectral scan (200–800 nm) of textile wastewater after Enterobacter degradation at different time intervals showed decolourisation and a decrease in dye concentration from the batch culture. The BR9 displayed one peak at a maximum wavelength (λmax) of 540 nm in the UV–Vis spectra. The intensity of this peak decreased significantly after the addition of the Enterobacter sp. strain TS1L due to decolourisation. Normally, bacterial decolourisation follows two pathways, namely adsorption or biodegradation [11, 51, 52]. Dye adsorption can be detected when investigating the bacterial cell, which will deepen its colour, while during degradation, the cell remains colourless [52]. In our study, Enterobacter sp. TS1L could decolourise BR9 via both pathways. After decolourisation, the TS1L cells were stained with BR9, indicating that BR9 was adsorbed on the cell surface. In addition, dye degradation was clearly identified by the changes in the UV-spectrum (Fig. 3).
Conclusions
In this study, single and mixed cultures of Enterobacter sp. TS1L and TS3 demonstrated potential for textile wastewater degradation. The strain TS1L showed high decolourisation efficiency, with a decolourisation efficiency of 81.15% under the following conditions: 100% of textile wastewater without glucose addition, pH 7.0, 150 rpm, 35 °C. Moreover, the BR9 concentration in textile wastewater was significantly decreased from 80.94 mg/L to 15.26 mg/L. Based on these results Enterobacter sp. TS1L is able to decolourise textile wastewater within a short time (12 h). In addition, the products formed during biodegradation were analysed by GC–MS, and the results indicate that the strain TS1L completely degraded and converted wastewater compounds into organic compounds under optimal conditions. The treated wastewater met the requirements of the Water Quality Standard (Thailand). Enterobacter sp. TS1L can both decolourise and degrade BR9, making it a suitable bacterial strain for the physical and chemical processing of textile wastewater. Based on the successful laboratory results, the future work should then be scale up and evaluate in textile industrial. In addition to the study on genomics and proteomics, there is a possibility to increase the ability of bacterial of textile wastewater. Moreover, the reduction of nitrogen content after microbial treatment should be also evaluated in more detail. With all the positive research and developments, biological treatment is hope to be outstanding method in the dye and toxic chemical elimination in textile wastewater.
References
Ali H (2010) Biodegradation of synthetic dyes—a review. Water Air Soil Poll 213:251–273
Rauf MA, Ashraf SS (2012) Survey of recent trends in biochemically assisted degradation of dyes. Chem Eng J 209:520–530
Wood L (2019) Synthetic dyes and pigments. E.S.T. https://www.marketscreener.com/BASF-SE-6443227/news/Synthetic-Dyes-Pigments-Market-2019-Global-Report-By-Type-And-By-Key-Players-28949353/. Accessed 21 August 2020
Zargar B, Parham H, Hatamie A (2009) Modified iron oxide nanoparticles as solid phase extractor for spectrophotometeric determination and separation of basic fuchsin. Talanta 77:1328–1331. https://doi.org/10.1016/j.talanta.2008.09.011
Duman O, Tunc S, Polat TG (2015) Adsorptive removal of triarylmethane dye (Basic Red 9) from aqueous solution by sepiolite as effective and low-cost adsorbent. Microporous Mesoporous Mater 210:176–184
Scaringelli FP, Saltzman BE, Frey SA (1967) Spectrophotometric determination of atmospheric sulfur dioxide. Anal Chem 39:1709–1726. https://doi.org/10.1021/ac50157a031
Crini G (2006) Non-conventional low-cost adsorbents for dye removal: a review. Bioresour Technol 97:1061–1085
Verma AK, Dash RR, Bhunia P (2012) A review on chemical coagulation/flocculation technologies for removal of color from textile wastewaters. J Environ Manage 93:154–168
Kaushik P, Malik A (2009) Fungal dye decolourization: recent advances and future potential. Environ Int 35:127–141
Ajar M, Skakeel S, Rehman A (2020) Microbial use for azo dye degradation-a strategy. Int Microbiol 23:149–159
Jamee R, Siddique M (2019) Biodegradation of synthetic dyes of textile effluent by microorganisms: an environmentally and economically sustainable approach. Eur J Microbiol Immunol 9:114–118
dos Santos AB, Cervantes FJ, van Lier JB (2007) Review paper on current technologies for decolorization of textile wastewaters: perspectives for anaerobic biotechnology. Bioresour Technol 98:2369–2385
Sarayu K, Sandhya S (2012) Current technologies for biological treatment of textile wastewater a review. Appl Biochem Biotechnol 167:645–661
Abadulla E, Tzanov T, Costa S, Robra KH, Paulo AC, Gübitz GM (2000) Decolorization and detoxification of textile dyes with a laccase from Trametes hirsute. Appl Environ Microbiol 66:3357–3362. https://doi.org/10.1128/AEM.66.8.3357-3362.2000
Jasińska A, Paraszkiewicz K, Slaba M, Dłgoński J (2015) Microbial decolorization of triphenylmethane dyes. Singh SN (ed.), Microbial degradation of synthetic dyes in wastewaters, Environ Sci Eng. https://doi.org/10.1007/978-3-319-10942-8_8
APHA (1992) Standard Methods for the examination of water and wastewater, 18th edn. EEF, Washington, DC
APHA (1999) Standard methods for the examination of water and wastewater, 20th edn. EEF, Washington, DC
Sugahara S, Suzuki M, Kamiya H, Yamamuro M, Semura H, Senga Y, Egawa M, Seike Y (2016) Colorimetric determination of sulfide in microsamples. Anal Sci 32:1129–1131
Anouzla A, Abrouki Y, Souabi S, Safi M, Rhbal H (2009) Colour and COD removal of disperse dye solution by a novel coagulant: application of statistical design for the optimization and regression analysis. J Hazard Mater 166:1302–1306
Lapara TM, Alleman JE, Pope PG (2000) Miniaturized closed reflux, colorimetric method for the determination of chemical oxygen demand. Waste Manage 20:295–298
Rakkan T, Chana N, Chirapongsatonkul N, Utaynapun K, Sangkharak K (2020) Screening and identification of Basic Red 9-degrading bacteria from textile wastewater and their ability to produce medium- and long chain lengt-polyhydroxyalkanoate. (Submitted manuscript).
Sangkharak K, Prasertsan P (2012) The production of polyhydroxyalkanoate by Bacillus licheniformis using sequential mutagenesis and optimization. Biotechnol Bioprocess Eng 18:272–279
Martins AO, Canalli VM, Azevedo CMN, Pires M (2006) Degradation of pararosaniline (C.I. Basic Red 9 monohydrochloride) dye by ozonation and sonolysis. Dyes Pigm 68:227–234
Sangkharak K, Prasertsan P (2008) Nutrient optimization for production of polyhydroxybutyrate from halotolerant photosynthetic bacteria cultivated under aerobic-dark condition. Electron J Biotechnol 11:173–183
Ayed L, Chaieb K, Cheref A, Bakhrouf A (2010) Biodegradation and decolorization of triphenylmethane dyes by Staphylococcus epidermidis. Desalin 260:137–146
Deivasigamani Ch, Das N (2011) Biodegradation of Basic violet 3 by Candida krusei isolated from textile wastewater. Biodegradation 22:1169–1180
Steel RGD, Torrie JH (1980) Principles and procedures of statistics. McGraw-Hill, New York
Horikoshi K (1999) Alkaliphiles: some applications of their products for biotechnology. Microbiol Mol Biol Rev 63(4):735–750. https://doi.org/10.1128/MMBR.63.4.735-750.1999
Kadpan IK, Kargi F, Mullan GM, Marchant R (2000) Effect of environmental conditions on biological decolorization of textile dyestuff by C. versicolor. Enzyme Microb Technol 26:381–387
Ramsay JA, Goode C (2004) Decoloration of a carpet dye effluent using Trametes versicolor. Biotechnol Lett 26:197–201
Vahabzadeh F, Mehranian M, Saatari AR (2004) Color removal ability of Phanerochaete chrysosporium in relation to lignin peroxidase and manganese peroxidase produced in molasses wastewater. World J Microbiol Biotechnol 20:859–864
Jadhav JP, Govindwar SP (2006) Biotransformation of malachite green by Saccharomyces cerevisiae MTCC 463. Yeast 23:315–323
Senthilkumar S, Perumalsamy M, Prabhu HJ (2014) Decolourization potential of white-rot fungus Phanerochaete chrysosporium on synthetic dye bath effluent containing Amido black 10B. JSCS 18:845–853
Chen CY, Kuo JT, Cheng CY, Huang YT, Ho IH, Chung YC (2009) Biological decolorization of dye solution containing malachite green by Pandoraea pulmonicola YC32 using a batch and continuous system. J Hazard Mater 172:1439–1445
Shedbalkar U, Jadhav J (2011) Detoxification of malachite green and textile industrial effluent by Penicillium ochrochloron. Biotechnol Bioprocess Eng 16:196–204
Jadhav SB, Yedurkar SM, Phugare SS, Jadhav JP (2012) Biodegradation studies on acid violet 19, a triphenylmethane dye, by Pseudomonas aeruginosa BCH. Clean Soil Air Water 40:551–558
Mukherjee T, Das M (2014) Degradation of Malachite Green by Enterobacter asburiae strain XJUHX-4TM. Clean Soil Air Water 42:849–856
Shah MP (2013) Microbial degradation of textile dye (Remazol Black B) by Bacillus spp. ETL-2012. Appl Environ Microbiol 1:6–11
Panswad T, Luangdilok W (2000) Decolorization of reactive dyes with different molecular structures under different environmental conditions. Water Res 34:4177–4184
Cetin D, Donmez G (2006) Decolorization of reactive dyes by mixed cultures isolated from textile effluent under anaerobic conditions. Enzyme Microb Technol 38:926–930
Lalnunhlimi S, Krishnaswamy V (2016) Decolorization of azo dyes (Direct Blue 151 and Direct Red 31) by moderately alkaliphilic bacterial consortium. Braz J Microbiol 47:39–46
Swamy J, Ramsay JA (1999) The evaluation of white rot fungi in the decoloration of textile dyes. Enzyme Microb Technol 24:130–137
Fu Y, Viraraghavan T (2001) Fungal decolorization of dye wastewaters: a review. Bioresour Technol 79:251–262
Parshetti GK, Kalme SD, Gomare SS, Govindwar SP (2007) Biodegradation of Reactive blue-25 by Aspergillus ochraceus NCIM-1146. Bioresour Technol 98:3638–3642. https://doi.org/10.1016/j.biortech.2006.11.017
Rani B, Kumar V, Singh J, Bisht S, Teotia P, Sharma S, Ritu K (2014) Bioremediation of dyes by fungi isolated from contaminated dye effluent sites for bio-usability. Braz J Microbiol 45:1055–1630
Harry-asobara JL, Kamei I (2019) Characteristics of white-rot fungus Phlebia brevispora TMIC33929 and its growth-promoting bacterium Enterobacter sp. TN3W-14 in the decolorization of dye-contaminated water. Appl Biochem Biotechnol 189:1183–1194
Khehra MS, Saini HS, Sharma DK, Chadha BS, Chimni SS (2005) Decolorization of various azo dyes by bacterial consortium. Dyes Pigm 67:55–61. https://doi.org/10.1016/j.dyepig.2004.10.008
Nascimento C, Magalhaes DP, Brandao M, Santos AB, Chame M, Baptista D, Nishikawa M, da Silva M (2011) Degradation and detoxification of three textile azo dyes by mixed fungal cultures from semi-arid region of brazilian norteast. Braz Arch Biol Technol 54(3):621–628
Mikeskova H, Novotny C, Svobodova K (2012) Interspecific interactions in mixed microbial cultures in a biodegradation perspective. Appl Microbiol Biotechnol 95:861–870
Niam P, Banat IM, Singh D, Marchant R (1996) Microbial process for the decolorization of textile effluent containing azo, diazo and reactive dyes. Process Biochem 31:435–442
Banat IM, Nigam P, Singh D, Marchant R (1996) Microbial decolorization of textile-dye-containing effluents: a review. Bioresour Technol 58(3):217–227
Azmi W, Sani RK, Banerjee UC (1998) Biodegradation of triphenylmethane dyes. Enzyme Microb Technol 22(3):185–191
Ministry of Natural Resources and Environment (2005) Notification of Ministry of Natural Resources and Environment (MONRE): building effluents standards, date November 7, B.E. 2005, published in the Royal Government Gazette, vol. 122 Part 125D.
Acknowledgements
The authors would like to acknowledge the support of Thailand Science Research and Innovation (TSRI) through the Royal Golden Jubilee Ph.D. (RGJ#PHD) Program through grant number PHD/00073/2559 for RGJ#PHD. Acknowledgement is also made to the Department of Chemistry, Faculty of Science, Thaksin University, Phatthalung Campus, Thailand. We would like to thank Associated Professor Dr. Ken’ichiro MATSUMOTO and Assistant Prof. Dr. Netnapa Chana for a support and comments.
Author information
Authors and Affiliations
Contributions
TR carried out the field sampling, the experiments, data analysis and the writing of original manuscript. KS took part in design this work and reviewed data analysis and editing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical Approval
This manuscript is an original work and never has been published elsewhere in any form or language.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rakkan, T., Sangkharak, K. Enhanced Decolourisation and Biodegradation of Textile Wastewater Using Single and Mixed Cultures of a Newly Isolated Enterobacter Strain. Curr Microbiol 77, 4085–4094 (2020). https://doi.org/10.1007/s00284-020-02246-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02246-2