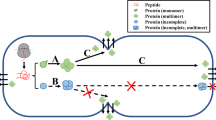
Abstract
Most bacterial proteins that are destined to leave the cytoplasm are exported across the cell membrane to their sites of function. These proteins are generally exported via the classical secretion pathway, in which the signal peptide plays a central role. However, some bacterial proteins have been found in the extracellular milieu without any apparent signal peptide. As none of the classical secretion systems is involved in their secretion, this occurrence is termed non-classical protein secretion. The mechanism or mechanisms responsible for non-classical secretion are contentious. This review compiles evidence from the debate over whether the release of the non-classically secreted proteins is the result of cell lysis and discusses how these proteins are exported to the exterior of the cell.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteins can play their proper role only when they are delivered to the appropriate destination. Following the nascence of protein synthesis in the cytosol, many bacterial proteins must be transported across a membrane to reach their sites of function. The protein export mechanism in prokaryotes has been extensively studied at the molecular level. Exported proteins are initially synthesized as precursors with an amino-terminal extension, the signal peptide. These preproteins are first targeted toward the translocation machinery in the cell membrane. The exported proteins are then transported through a proteinaceous channel in the membrane. Finally, the signal peptide is removed, leading to the release of the mature protein from the membrane. The fate of newly synthesized proteins is thought to be determined by the signal peptide, which distinguishes the exported proteins from the cytoplasmic ones and is needed to target the proteins toward the export pathway [45, 51, 55, 60, 61].
Although many of the proteins that secrete into the extracellular milieu have signal peptides, some cytoplasmic proteins with no known signals or secretion motifs can also be found in extracellular locations in bacteria. As none of the classical secretion systems appears to participate in their secretion, this type of secretion is referred to as non-classical protein secretion [7, 37]. With the development of proteomics and protein characterization technologies, the number of these proteins is steadily growing. Although their number varies by experimental condition and bacterial species, non-classically secreted proteins have been shown to exist across a wide range of bacteria, and some are common in specific bacteria [7, 54].
The presence of non-classically secreted proteins in the extracellular environment can be attributed simply to cell lysis. Although several reports support such attribution [38, 59], an increasing number of experiments cast doubt on this point. One of the most elegant and direct pieces of evidence in this regard is that provided by Boël et al. [10, 11], who showed that the export of enolase and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) can be inhibited by preventing the 2-phosphoglycerate-dependent automodification of enolase and inserting a hydrophobic tail in the GAPDH protein, respectively. Although the mechanism or mechanisms responsible for non-classical secretion remain unknown, considerable effort has been devoted to identifying the proteins that are secreted in this manner. Aguilera et al. reported that GAPDH is secreted via the LEE-encoded type III secretion system in enteropathogenic Escherichia coli [1], and the staphylococcal major autolysin is known to be involved in the secretion of non-classically secreted proteins in Staphylococcus aureus [42]. Ten non-classically secreted proteins have also been identified to involve SecA2-dependent secretion in Listeria monocytogenes [32]. However, in the latter study, it was not ascertained whether these proteins were directly or indirectly involved in the export of non-classically secreted proteins. More studies are required to confirm whether different transport mechanisms are used for the same protein in different bacterial species or for various non-classically secreted proteins.
Cell Lysis or Secretion?
The Evidence Supporting the Cell Lysis Hypothesis
As proteins identified in the extracytoplasmic milieu of bacteria are traditionally thought to be strictly cytosolic, and there is no known mechanism for their secretion, they have been assumed to result primarily from the cell lysis. Bacterial surface localization is the result of the reassociation of the released enzymes from the lysed cell. There is some research to support this view (Table 1).
First, most of these non-classically secreted proteins are abundant proteins of the cytoplasmic proteome and are secreted in the stationary phase [2, 34, 53]. Tullius et al. [57] provided evidence to show that the extracellular abundance of Mycobacterium tuberculosis glutamine synthetase and superoxide dismutase is due simply to their high level of expression and extracellular stability. Further, the amount of cytosolic proteins in the culture is very small in some species. For example, in one study, cytosolic proteins were found to account for less than 1 % of the total proteins in the culture supernatant of Bacillus cereus and Bacillus thuringiensis [19].
Second, the cell lysis hypothesis is also supported by evidence showing the levels of cytoplasmic proteins in the extracellular proteome to increase in some mutants of Bacillus subtilis [2]. The binding of these extracellular cytoplasmic proteins from the medium onto the bacteria surface has also been reported [3, 9, 26]. Oliveira et al. provided direct evidence to support the cell lysis hypothesis. They found an impaired GAPDH presence on the surface and in the supernatant of group B streptococcus to be associated with a lower level of bacterial lysis [38].
Research Supporting the Secretion Hypothesis
The foregoing evidence provides some support for the cell lysis hypothesis. However, following the development of proteomics and protein characterization technologies, an increasing number of studies have indicated that the release of cytoplasmic proteins is not simply mediated by cell lysis, at least in some special bacteria (Table 1). The following evidence provides support for the alternative secretion hypothesis.
First, in several studies, the apparent molecular weights and isoelectric points of the cytoplasmic proteins located in the extracellular milieu and identified in 2-D gels were not significantly different from their calculated values, and no modification was observed in the vast majority of these proteins other than the removal of the N-terminal methionine by the determined N-terminal sequence [31, 42, 47, 52]. Cytoplasmic proteins were not identified in the culture (although some were abundant in the cytoplasmic proteome), and neither were the proteolytic fragments of the corresponding proteins detected therein [42].
Second, several research groups have used a number of cytoplasmic marker macromolecules to assess cell integrity in various bacterial species. In E. coli and Staphylococcus agalactiae, for example, the protein preparations were examined for the presence of DNA contamination [23, 63]. Dreisbach et al. [17] employed TrxA as the cytoplasmic marker protein to identify the reliable surfacome of S. aureus and to check for cell integrity. Other researchers measured the lactate dehydrogenase activity in the supernatant of the cell suspension to confirm cytosolic protein leakage from the cells in Lactobacillus plantarum and Lactococcus lactis [6, 27]. The strictly cytoplasmic enzyme aminopeptidase C was used as an indicator of cytoplasmic contamination in L. monocytogenes [52]. In all of these examinations, no or very few cytoplasmic markers were found in the extracellular milieu of the cells, and thus the presence of cytosolic proteins in the culture supernatants is unlikely to be the result of cell lysis. The constant cell density and viability counts in the stationary phase provided further evidence to suggest that the release of these proteins is not the result of cell lysis. In addition to monitoring cell integrity, some researchers have taken special precautions during experiments to prevent the contamination of cytosolic proteins, by cultivating the cultures in a fermenter, growing the cells in a medium supplemented with glucose, and/or harvesting the cultures prior to entry into the stationary phase. In each case, non-classically secreted proteins were also found in the cultures. Another report discussed the comprehensive coverage of the extracellular proteins of Corynebacterium pseudotuberculosis identified with a high degree of confidence by a newly combined approach [39]. Intriguingly, 19 of the 70 proteins in the exoproteome of C. pseudotuberculosis strain 1002 were primarily regarded as cytoplasmic, but only 2 of 67 in the exoproteome of the strain C231. The authors found that 13 of the 19 proteins in the exoproteome of the strain 1002 could be secreted by non-classical mechanisms [39], which strongly suggests that the secretion of these proteins is not due to cell lysis.
Third, a number of studies have used 2-D gel electrophoresis to compare the profiles and abundances of extracellular and intracellular proteins separately to determine whether the extracellular proteins are the result of cell lysis. In Staphylococcus pneumoniae, the distribution of the major protein spots in the cytoplasmic extract was found to be significantly different from that of the cell wall extract when an equivalent amount of protein from the total cytoplasmic protein extract and cell wall extract was separated via 2-D gel electrophoresis [33]. Choi et al. [14] found a very low degree of correlation between the cytosolic and secreted fraction proteins of S. pneumoniae using the DAnTE program, and found the amount of cytosolic protein in the exoproteome to not differ significantly at each growth stage. Walz et al. [62] found that two-thirds of all protein spot positions and relative abundances between the cytosol and exoproteome in Bacillus anthracis are distinctively different. In another study, the six separation profiles of Lactobacillus rhamnosus surface-associated proteins extracted using six different methods were similar to one another but markedly different from the total protein extracts analyzed by gel electrophoresis [50]. It is notable that, in contrast to B. subtilis, some of the non-classically secreted proteins did not increase in the mutants. In S. aureus, the amounts of the main non-classically secreted proteins (Eno, Gap, EF-Tu, EF-G, DnaK, GroEL, Tkt, and PdhD) were not influenced by a mutation in agr or sigB [65]. Only four of the 16 most common non-classically secreted proteins (Tkt, Ppi, Tig, and EF-Tu) were more abundant in the exoproteome of the Mycobacterium smegmatis Δlgt mutant than in the exoproteome of the wild-type. Five of these proteins (Gap, DnaK, DnaK, GpmA, and Adh) were more abundant in the exoproteome of the parental strains [56]. Hence, increased amounts of these non-classically secreted proteins in the mutants should not contribute to the mutants’ susceptibility to cell lysis. Cell lysis may not be the major contributor to protein accumulation in the exoproteome. Unlike the case B. cereus and B. thuringiensis, the amount of cytosolic protein in the culture was very large in B. subtilis and B. anthracis, with the total spot volume of cytosolic proteins in the culture supernatant more than 10 % of the total spot volume [2, 19]. When recombinant M. tuberculosis GS and SOD were expressed in M. smegmatis, 95 % GS and 66 % SOD were exported into the culture [20, 21]. When carboxylesterase Est55 from Geobacillus stearothermophilus, which lacks a classical signal peptide, was expressed in B. subtilis, more Est55 was found in the medium compared to the intracellular content during the late stationary phase [64]. Using a subcellular fractionation approach followed by quantitative Western blot analyses, Vanet and Labigne found the supernatant protein profiles to be very different from those of the cell pellets and a typical cytoplasmic protein. A beta-galactosidase homolog was found to be exclusively associated with the pellets of the whole cell extracts, and no traces were found in the supernatant [58]. These studies strongly suggest that the release of cytoplasmic proteins is not simply mediated by cell lysis.
Fourth, the secretion of several specific non-classically secreted proteins has been induced in special experimental conditions, similar to eukaryocytes, where non-classical protein export is tightly regulated and usually induced by specific stimuli, such as various forms of cell stress [43]. It has been suggested that the aim of non-classical proteins released via the non-classical secretion pathway is to protect the organism in question from their undesirable effects. The secretion of KatA is H2O2 inducible in B. subtilis, and is essential to protect B. subtilis cells from oxidative assault [34]. Similarly, in Legionella pneumophila, the levels of periplasmic KatA have been shown to increase steadily in the presence of H2O2, and the inactivation of katA reduced the stationary phase survival of 100–10,000-fold [5]. A greater amount of the DnaK protein was released in the presence of bile salts in Bifidobacterium animalis. The secretion of DnaK induced by bile salts is thought to facilitate the colonization of the human host B. animalis in the gut bile environment [13]. In M. tuberculosis culture filtrates, FbaA and Ald were found in increased amounts under a low oxygen concentration, and hypoxic conditions are generally believed to be the environment that the pathogen M. tuberculosis localizes in the central part of the granuloma [48]. It is thus reasonable to suppose that these proteins may be involved in bacterial virulence. In the presence of a low level of glucose, L. plantarum exhibits an elevated level of cell wall-associated GAPDH, and further experiments have shown the levels of such GAPDH, cell membrane permeability, and carbon source availability to be interdependent parameters [49]. Similarly, under iron starvation conditions, an increase in GAPDH release was seen in Streptococcus pyogenes and Streptococcus gordonii [18, 36]. Such induction may be related to bacterial virulence, but further investigations are required to confirm that speculation.
Fifth, the most straightforward way of proving that cell lysis does not explain the presence of certain cytoplasmic proteins in the extracellular space of bacteria is to prevent their release. Research has confirmed that the 2-phosphoglycerate-dependent automodification of enolase and a hydrophobic alpha-helical domain within enolase are necessary for its export from the cytoplasm [10, 64]. In one study, C-terminal-deleted flagellin was confined to the intracellular domain in B. subtilis [29], and in another the S. pyogenes GAPDH was not exported to the cell surface when a hydrophobic tail was inserted at the C-terminal end of GAPDH [11]. In E. coli, GAPDH secretion is abolished in mutants that are defective in type III ATPase EscN [1]. Yang et al. [64] provided comprehensive evidence to show that the release of several cytoplasmic proteins into a growth medium is not the result of gross cell lysis in B. subtilis. The other evidence supporting the secretion hypothesis is related to the mechanisms of these proteins secretion (see the forthcoming discussion).
The various functions that the non-classically secreted proteins exercise in the cell exterior provide further evidence to suggest that the release of cytoplasmic proteins cannot simply be attributed to cell lysis. The cytoplasmic proteins present in the cell exterior are not extravagant, but they perform multiple functions. Since these proteins have multiple biologically unrelated autonomous functions and often localize to separate cellular compartments, they are also called moonlighting proteins [7, 8, 24]. The major moonlighting functions identified in pathogens and probiotics are adhesion to the host epithelia and host components, such as extracellular matrices and plasminogen, and modulation of host immune responses [4, 22, 41]. These proteins are thought to be involved in bacterial virulence or bacterial benefit.
Collectively, the evidence presented in this section indicates that cytoplasmic proteins that lack typical signal peptides and are present in the extracellular milieu are not simply the result of cell lysis and that their appearance is probably a general occurrence in bacteria.
The Potential Export Pathway
SecA2-Dependent Secretion Pathway in Gram-Positive Bacteria
To date, the protein secretion systems characterized in monoderm bacteria include the secretion (Sec), twin-arginine translocation (Tat), Flagella export apparatus (FEA), fimbrilin-protein exporter (FPE), hole forming (Holin), ATP-binding cassette (ABC) transporters, and WXG100 secretion system (Wss) pathways [15]. Owing to the lack of detectable secretion signals, these proteins should not be transported via the well-characterized Sec and Tat pathways. Only specific proteins are secreted through such pathways as FEA, FPE, and ABC. Hence, these proteins cannot be the pathways’ substrates. The Wss also known as the ESAT6-secretion system, secretes small antigenic proteins that reportedly share a WXG motif as identified by PSI-BLAST [40]. Pasztor et al. [42] proved that the presence or absence of prophages has little influence on the excretion of cytoplasmic proteins in S. aureus . Further, it has been predicted that none of the non-classically secreted proteins is exported via holins (that can form holes in the membrane) in L. monocytogenes [16, 44], and thus that holins cannot explain the release of these proteins.
It should be noted that some Gram-positive bacteria have an accessory SecA protein known as SecA2. SecA2 plays an important role in the export of certain proteins with or without signal peptides, and is believed to contribute to pathogenesis [46]. Using comparative secretomic analysis of wild-type and SecA2 mutant M. tuberculosis on 2-D-PAGE, Braunstein et al. [12] identified SodA and KatG as SecA2-dependent secreted proteins, whereas the non-classically secreted proteins RplL and CspA were more abundant in culture filtrates in SecA2-mutant M. tuberculosis. In another experiment, 17 SecA2-dependent secreted and surface proteins of L. monocytogenes were identified using a proteomics approach. Of these proteins, seven contained signal peptides. The 10 SecA2-dependent surface proteins that lacked signal peptides had well-described cytosolic functions [32]. Oliveira et al. reported that GAPDH was slightly more abundant at the surface and in the culture supernatant of the SecA2 mutant compared with the WT strain [38] (Table 2).
However, these investigations failed to determine whether SecA2 was directly or indirectly involved in the export of these cytoplasmic proteins. It is likely that their excretion was not regulated by the SecA2-dependent pathway, but it may have been related to the pathway’s authentic substrates. It is possible that p60 and MurA, which are the substrates of the SecA2-dependent pathway with signal peptides, induced altruistic autolysis, which contributed to the SecA2-dependent release of cytosolic bacterial proteins in L. monocytogenes [32]; since the authors proved that p60 autolysin is not required for the release of other SecA2-dependent proteins, that release may be related to the MurA that is required for cell separation, just like the major autolysin (Atl) of S. aureus, which reportedly plays an important role in the excretion of cytoplasmic proteins [42] (see forthcoming details). The ability of SecA2 to coinstantaneously transport proteins with and without signal peptides would be surprising. Hence, further investigation is required to determine whether SecA2 can export these non-classically secreted proteins directly.
Major Autolysin-Related Pathway in Gram-Positive Bacteria
While Pasztor et al. [42] demonstrated that the major autolysin (Atl) plays a crucial role in the excretion of non-classically secreted proteins, the most abundant proteins in the cytoplasm were not found in the exoproteome of S. aureus (Table 2), which implies the existence of a selection mechanism in cytoplasmic protein excretion. The enhanced expression of other autolysins in Atl mutation cannot compensate for the defect in the excretion of cytoplasmic proteins in S. aureus [42]. The p60 autolysin is not required for the release of these proteins in L. monocytogenes [32], and the lytC lytD double-mutant B. subtilis lacking two major autolysins has no effect on their secretion [64]. This evidence suggests that the excretion of cytoplasmic proteins is related only to special autolysins. It should also be mentioned that Atl is targeted to the cell septa region for the next cell division site. In another study, the non-classically secreted proteins: enolase, GAPDH, GS, and glucose-6-phosphate isomerase showed localized binding to the cell division septa and poles of Lactobacillus crispatus [26]. In B. subtilis, enolase exhibited a diffuse profile in the exponential phase but localized very strongly to one pole in the stationary phase when secreted abundantly [25]. The cellular localization of enolase was controlled by the BY-kinase PtkA [25], but PtkA’s effect on the release of non-classically secreted proteins was not examined. Taken together, this evidence suggests that the specific localization of autolysins may be an essential factor in determining cytoplasmic protein excretion and that these proteins are preferentially released at the cell division septa or poles during septum formation. This assumption is worthy of additional research, particularly to define the relationship between the autolysins that localize at different subcellular locations of bacteria and cytoplasmic protein excretion and to determine the effect of the localization of non-classically secreted proteins in the cell on their secretion. It is possible that the cytoskeleton or components of the cell wall synthesis machinery are involved, but much more work is required to identify the possible secretion mechanisms.
Other Potential Pathway in Gram-Positive Bacteria
Saad et al. [49] demonstrated that the concentration of cell wall GAPDH is closely related to membrane permeability in L. plantarum (Table 2). The authors found that free GAPDH was not observed in the culture supernatant at any time during growth and, further, because the provoked cell lysis was not concomitant with any re-association, that cell lysis could not be the reason for the presence of GAPDH on the cell surface [49]. Using flow cytometry measurement and the double labeling of L. plantarum with anti-GAPDH antibodies and propidium iodide, they established a close relationship between plasma membrane integrity and cw-GAPDH concentration [49]. A limited number of proteins were found on the cell wall of L. plantarum, which suggests the existence of a selection mechanism in the efflux of cytoplasmic proteins that we know nothing about.
In another study, the large conductance mechanosensitive channel protein MscL of B. subtilis was found to prevent the specific release of cytoplasmic proteins during hypo-osmotic shock. MscL did not affect such secretion under normal growth conditions. However, under hypo-osmotic shock conditions, specific and normal cytoplasmic proteins were selectively released by MscL mutant cells and the presence of MscL prevented the specific release of these proteins. The authors further confirmed that the specific protein release could not be attributed to cell death or lysis [28]. These results show that there is an unidentified pathway for their selective release.
Potential Export Pathway in Gram-Negative Bacteria
Protein secretion has been more thoroughly investigated in Gram-negative bacteria than Gram-positive bacteria. To date, six major protein secretion systems, T1SS to T6SS, have been discovered and named [15]. However, only a few Gram-negative species have been reported to release non-classically secreted proteins, and such secretion depends on the growth conditions. Few studies have focused on the mechanism directing that secretion. In eukaryocytes, certain proteins that lack signal peptides, such as IL1β, can apparently be incorporated into autophagosomes and then released in exosomes [43]. The transport of such cytoplasmic proteins, such as GroEL, EF-G, and GlnA to the supernatants via membrane vesicles (MVs) has also been reported, but the proteomic profiling of the native outer MVs derived from E. coli has shown significant differences and little specificity in comparison with the extracellular proteomes of E. coli analyzed by 2D-PAGE [30, 35, 63]. It is difficult to assess the degree to which MVs contribute to the release of non-classically secreted proteins. In their most recent study, Aguilera et al. elegantly demonstrated the secretion of GAPDH through T3SS by enteropathogenic E. coli (EPEC) grown in Dulbecco’s modified eagle’s medium (DMEM) and proved that GAPDH secretion is abolished in mutants that are defective in the type III ATPase EscN and restored by escN gene complementation [1]. However, when grown in Luria broth (LB), GAPDH is secreted to the supernatant of escN mutant strain CVD452, indicating that a secretion system other than T3SS is responsible for GAPDH secretion when EPEC is grown in LB. Further studies have shown that gut microbiota E. coli strains such as EcoR12, EcoR26, and Nissle 1917 that do not contain the T3SS-encoding genes do not secrete GAPDH when grown in DMEM. Only probiotic strain Nissle 1917 in the gut microbiota E. coli strains tested was found to secrete GAPDH when grown in LB cultures. These results prove that there are at least two alternative pathways for GAPDH secretion in E. coli. One is pathogen-specific T3SS in cells grown in DMEM, and the other is the widespread, but unidentified, pathway in pathogenic and non-pathogenic E. coli grown in LB [1] (Table 2).
The export mechanisms of non-classically secreted proteins appear to be even more complex, as the aforementioned evidence indicates the existence of various kinds of distinct non-classical export routes. Further research is needed to confirm whether different bacterial species use different transport mechanisms or whether different such mechanisms are used for different non-classically secreted proteins. Since these non-classically secreted proteins play some important roles within the cytoplasm, their secretion must be strictly controlled no matter which pathway they follow.
Conclusions and Future Directions
This overview suggests that although there are different views on the release of non-classically secreted proteins, such release is unlikely to occur simply because of cell lysis. These proteins could be secreted via pathways that have yet to be comprehensively or systematically investigated. There are several key questions surrounding non-classically secreted proteins that require further investigation. Is SecA2 directly involved in the export of these cytoplasmic proteins? Are all of the autolysins involved in the excretion of these proteins? Is the cellular localization of such proteins in cytoplasm related to their secretion? We hope that this review arouses more research interest in tackling these and related questions. A greater understanding of non-classical protein secretion will help to improve bacterial efficiency.
References
Aguilera L, Ferreira E, Giménez R, Fernández FJ, Taulés M, Aguilar J, Vega MC, Badia J, Baldomà L (2012) Secretion of the housekeeping protein glyceraldehyde-3-phosphate dehydrogenase by the LEE-encoded type III secretion system in enteropathogenic Escherichia coli. Int J Biochem Cell Biol 44(6):955–962
Antelmann H, Van Dijl JM, Bron S, Hecker M (2006) Proteomic survey through secretome of Bacillus subtilis. Methods Biochem Anal 49:179–208
Antikainen J, Kuparinen V, Lähteenmäki K, Korhonen TK (2007) pH dependent association of enolase and glyceraldehyde-3-phosphate dehydrogenase of Lactobacillus crispatus with the cell wall and lipoteichoic acids. J Bacteriol 189:4539–4543
Antikainen J, Kuparinen V, Lähteenmäki K, Korhonen TK (2007) Enolases from Gram-positive bacterial pathogens and commensal lactobacilli share functional similarity in virulence-associated traits. FEMS Immunol Med Microbiol 51:526–534
Bandyopadhyay P, Steinman HM (2000) Catalase-peroxidases of Legionella pneumophila: cloning of the katA gene and studies of KatA function. J Bacteriol 182:6679–6686
Beck HC, Madsen SM, Glenting J, Petersen J, Israelsen H, Nørrelykke MR, Antonsson M, Hansen AM (2009) Proteomic analysis of cell surface-associated proteins from probiotic Lactobacillus plantarum. FEMS Microbiol Lett 297(1):61–66
Bendtsen JD, Kiemer L, Fausbøll A, Brunak S (2005) Non-classical protein secretion in bacteria. BMC Microbiol 5:58
Bendtsen JD, Wooldridge KG (2009) Non-classical secretion. In: Wooldridge K (ed) Bacterial secreted proteins: secretory mechanisms and role in pathogenesis. Caister Academic Press, Norfolk, UK, pp 193–223
Bergmann S, Rohde M, Chhatwal GS, Hammerschmidt S (2001) Alpha enolase of Streptococcus pneumoniae is a plasmin(ogen)-binding protein displayed on the bacterial cell surface. Mol Microbiol 40:1273–1287
Boël G, Pichereau V, Mijakovic I, Mazé A, Poncet S, Gillet S, Giard JC, Hartke A, Auffray Y, Deutscher J (2004) Is 2-phosphoglycerate-dependent automodification of bacterial enolases implicated in their export? J Mol Biol 337(2):485–496
Boël G, Jin H, Pancholi V (2005) Inhibition of cell surface export of group A streptococcal anchorless surface dehydrogenase affects bacterial adherence and antiphagocytic properties. Infect Immun 73:6237–6624
Braunstein M, Espinosa BJ, Chan J, Belisle JT, Jacobs WR Jr (2003) SecA2 functions in the secretion of superoxide dismutase A and in the virulence of Mycobacterium tuberculosis. Mol Microbiol 48:453–464
Candela M, Centanni M, Fiori J, Biagi E, Turroni S, Orrico C, Bergmann S, Hammerschmidt S, Brigidi P (2010) DnaK from Bifidobacterium animalis subsp. lactis is a surface-exposed human plasminogen receptor upregulated in response to bile salts. Microbiology 156:1609–1618
Choi CW, Lee YG, Kwon SO, Kim HY, Lee JC, Chung YH, Yun CY, Kim SI (2012) Analysis of Streptococcus pneumoniae secreted antigens by immuno-proteomic approach. Diagn Microbiol Infect Dis 72(4):318–327
Desvaux M, Hébraud M, Talon R, Henderson IR (2009) Secretion and subcellular localizations of bacterial proteins: a semantic awareness issue. Trends Microbiol 17(4):139–145
Desvaux M, Dumas E, Chafsey I, Chambon C, Hébraud M (2010) Comprehensive appraisal of the extracellular proteins from a monoderm bacterium: theoretical and empirical exoproteomes of Listeria monocytogenes EGD-e by secretomics. J Proteome Res 9(10):5076–5092
Dreisbach A, Hempel K, Buist G, Hecker M, Becher D, van Dijl JM (2010) Profiling the surfacome of Staphylococcus aureus. Proteomics 10:3082–3096
Eichenbaum Z, Green BD, Scott JR (1996) Iron starvation causes release from the group A streptococcus of the ADP-ribosylating protein called plasmin receptor or surface glyceraldehyde-3-phosphate-dehydrogenase. Infect Immun 64:1956–1960
Gohar M, Gilois N, Graveline R, Garreau C, Sanchis V, Lereclus D (2005) A comparative study of Bacillus cereus, Bacillus thuringiensis and Bacillus anthracis extracellular proteomes. Proteomics 5:3696–3711
Harth G, Horwitz MA (1997) Expression and efficient export of enzymatically active Mycobacterium tuberculosis glutamine synthetase in Mycobacterium smegmatis and evidence that the information for export is contained within the protein. J Biol Chem 272:22728–22735
Harth G, Horwitz MA (1999) Export of recombinant Mycobacterium tuberculosis superoxide dismutase is dependent upon both information in the protein and mycobacterial export machinery. A model for studying export of leaderless proteins by pathogenic mycobacteria. J Biol Chem 274:4281–4292
Henderson B, Martin A (2011) Bacterial virulence in the moonlight: multitasking bacterial moonlighting proteins are virulence determinants in infectious disease. Infect Immun 79:3476–3491
Hughes MJ, Moore JC, Lane JD, Wilson R, Pribul PK, Younes ZN, Dobson RJ, Everest P, Reason AJ, Redfern JM, Greer FM, Paxton T, Panico M, Morris HR, Feldman RG, Santangelo JD (2002) Identification of major outer surface proteins of Streptococcus agalactiae. Infect Immun 70(3):1254–1259
Jeffery CJ (2009) Moonlighting proteins-an update. Mol BioSyst 5:345–350
Jers C, Pedersen MM, Paspaliari DK, Schütz W, Johnsson C, Soufi B, Macek B, Jensen PR, Mijakovic I (2010) Bacillus subtilis BY-kinase PtkA controls enzyme activity and localization of its protein substrates. Mol Microbiol 77(2):287–299
Kainulainen V, Loimaranta V, Pekkala A, Edelman S, Antikainen J, Kylväjä R, Laaksonen M, Laakkonen L, Finne J, Korhonen TK (2012) Glutamine synthetase and glucose-6-phosphate isomerase are adhesive moonlighting proteins of Lactobacillus crispatus released by epithelial cathelicidin LL-37. J Bacteriol 194(10):2509–2519
Katakura Y, Sano R, Hashimoto T, Ninomiya K, Shioya S (2010) Lactic acid bacteria display on the cell surface cytosolic proteins that recognize yeast mannan. Appl Microbiol Biotechnol 86(1):319–326
Kouwen TR, Antelmann H, van der Ploeg R, Denham EL, Hecker M, van Dijl JM (2009) MscL of Bacillus subtilis prevents selective release of cytoplasmic proteins in a hypotonic environment. Proteomics 9(4):1033–1043
LaVallie ER, Stahl ML (1989) Cloning of the flagellin gene from Bacillus subtilis and complementation studies of an in vitro-derived deletion mutation. J Bacteriol 171(6):3085–3094
Lee EY, Bang JY, Park GW, Choi DS, Kang JS, Kim HJ, Park KS, Lee JO, Kim YK, Kwon KH, Kim KP, Gho YS (2007) Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli. Proteomics 7:3143–3153
Lei B, Mackie S, Lukomski S, Musser JM (2000) Identification and immunogenicity of group A streptococcus culture supernatant proteins. Infect Immun 68:6807–6818
Lenz LL, Mohammadi S, Geissler A, Portnoy DA (2003) SecA2-dependent secretion of autolytic enzymes promotes Listeria monocytogenes pathogenesis. Proc Natl Acad Sci USA 100:12432–12437
Ling E, Feldman G, Portnoi M, Dagan R, Overweg K, Mulholland F, Chalifa-Caspi V, Wells J, Mizrachi-Nebenzahl Y (2004) Glycolytic enzymes associated with the cell surface of Streptococcus pneumoniae are antigenic in humans and elicit protective immune responses in the mouse. Clin Exp Immunol 138(2):290–298
Naclerio G, Baccigalupi L, Caruso C, De Felice M, Ricca E (1995) Bacillus subtilis vegetative catalase is an extracellular enzyme. Appl Environ Microbiol 61(12):4471–4473
Nandakumar MP, Cheung A, Marten MR (2006) Proteomic analysis of extracellular proteins from Escherichia coli W3110. J Proteome Res 5:1155–1161
Nelson D, Goldstein JM, Boatright K, Harty DW, Cook SL, Hickman PJ, Potempa J, Travis J, Mayo JA (2001) pH-regulated secretion of a glyceraldehyde-3-phosphate dehydrogenase from Streptococcus gordonii FSS2: purification, characterization, and cloning of the gene encoding this enzyme. J Dent Res 80:371–377
Nickel W (2003) The mystery of nonclassical protein secretion. A current view on cargo proteins and potential export routes. Eur J Biochem 270(10):2109–2119
Oliveira L, Madureira P, Andrade EB, Bouaboud A, Morello E, Ferreira P, Poyart C, Trieu-Cuot P, Dramsi S (2012) Group B streptococcus GAPDH is released upon cell lysis, associates with bacterial surface, and induces apoptosis in murine macrophages. PLoS ONE 7(1):e29963
Pacheco LG, Slade SE, Seyffert N, Santos AR, Castro TL, Silva WM, Santos AV, Santos SG, Farias LM, Carvalho MA, Pimenta AM, Meyer R, Silva A, Scrivens JH, Oliveira SC, Miyoshi A, Dowson CG, Azevedo V (2011) A combined approach for comparative exoproteome analysis of Corynebacterium pseudotuberculosis. BMC Microbiol 11(1):12
Pallen MJ (2002) The ESTA-6/WXG100 superfamily—and a new gram-positive secretion system? Trends Microbiol 10(5):209–212
Pancholi V, Chhatwal GS (2003) Housekeeping enzymes as virulence factors for pathogens. Int J Med Microbiol 293:391–401
Pasztor L, Ziebandt AK, Nega M, Schlag M, Haase S, Franz-Wachtel M, Madlung J, Nordheim A, Heinrichs DE, Götz F (2010) Staphylococcal major autolysin (Atl) is involved in excretion of cytoplasmic proteins. J Biol Chem 285:36794–36803
Prudovsky I, Kumar TK, Sterling S, Neivandt D (2013) Protein-phospholipid interactions in nonclassical protein secretion: problem and methods of study. Int J Mol Sci 14(2):3734–3772
Renier S, Micheau P, Talon R, Hébraud M, Desvaux M (2012) Subcellular localization of extracytoplasmic proteins in monoderm bacteria: rational secretomics-based strategy for genomic and proteomic analyses. PLoS One 7(8):e42982
Riezman H (1997) The ins and outs of protein translocation. Science 278:1728–1729
Rigel NW, Braunstein M (2008) A new twist on an old pathway-accessory Sec systems. Mol Microbiol 69(2):291–302
Rosenkrands I, Weldingh K, Jacobsen S, Hansen CV, Florio W, Gianetri I, Andersen P (2000) Mapping and identification of Mycobacterium tuberculosis proteins by two-dimensional gel electrophoresis, microsequencing and immunodetection. Electrophoresis 21(5):935–948
Rosenkrands I, Slayden RA, Crawford J, Aagaard C, Barry CE 3rd, Andersen P (2002) Hypoxic response of Mycobacterium tuberculosis studied by metabolic labeling and proteome analysis of cellular and extracellular proteins. J Bacteriol 184(13):3485–3491
Saad N, Urdaci M, Vignoles C, Chaignepain S, Tallon R, Schmitter JM, Bressollier P (2009) Lactobacillus plantarum 299v surface-bound GAPDH: a new insight into enzyme cell walls location. J Microbiol Biotechnol 19:1635–1643
Sánchez B, Bressollier P, Chaignepain S, Schmitter JM, Urdaci MC (2009) Identification of surface-associated proteins in the probiotic bacterium Lactobacillus rhamnosus GG. Int Dairy J 19(2):85–88
Schatz G, Dobberstein B (1996) Common principles of protein translocation across membranes. Science 271:1519–1526
Schaumburg J, Diekmann O, Hagendorff P, Bergmann S, Rohde M, Hammerschmidt S, Jänsch L, Wehland J, Kärst U (2004) The cell wall subproteome of Listeria monocytogenes. Proteomics 4(10):2991–3006
Scott JR, Barnett TC (2006) Surface proteins of gram-positive bacteria and how they get there. Annu Rev Microbiol 60:397–423
Sibbald M, van Dij JML (2009) Secretome mapping in Gram-positive pathogens. In: Wooldridge K (ed) Bacterial secreted proteins: secretory mechanisms and role in pathogenesis. Caister Academic Press, Norfolk, pp 193–223
Tjalsma H, Antelmann H, Jongbloed JD, Braun PG, Darmon E, Dorenbos R, Dubois JY, Westers H, Zanen G, Quax WJ, Kuipers OP, Bron S, Hecker M, van Dijl JM (2004) Proteomics of protein secretion by Bacillus subtilis: separating the “secrets” of the secretome. Microbiol Mol Biol Rev 68(2):207–233
Tschumi A, Grau T, Albrecht D, Rezwan M, Antelmann H, Sander P (2012) Functional analyses of mycobacterial lipoprotein diacylglyceryl transferase and comparative secretome analysisof a mycobacterial lgt mutant. J Bacteriol 194(15):3938–3949
Tullius MV, Harth G, Horwitz MA (2001) High extracellular levels of Mycobacterium tuberculosis glutamine synthetase and superoxide dismutase in actively growing cultures are due to high expression and extracellular stability rather than to a protein-specific export mechanism. Infect Immun 69(10):6348–6363
Vanet A, Labigne A (1998) Evidence for specific secretion rather than autolysis in the release of some Helicobacter pylori proteins. Infect Immun 66(3):1023–1027
Vitikainen M, Lappalainen I, Seppala R, Antelmann H, Boer H, Taira S, Savilahti H, Hecker M, Vihinen M, Sarvas M, Kontinen VP (2004) Structure function analysis of PrsA reveals roles for the parvulin-like and flanking N- and C-terminal domains in protein folding and secretion in Bacillus subtilis. J Biol Chem 279:19302–19314
von Heijne G (1990) Protein targeting signals. Curr Oppin Cell Biol 2:604–608
von Heijne G (1998) Life and death of a signal peptide. Nature 396:111–113
Walz A, Mujer CV, Connolly JP, Alefantis T, Chafin R, Dake C, Whittington J, Kumar SP, Khan AS, DelVecchio VG (2007) Bacillus anthracis secretome time course under host-simulated conditions and identification of immunogenic proteins. Proteome Sci 5:11
Xia XX, Han MJ, Lee SY, Yoo JS (2008) Comparison of the extracellular proteomes of Escherichia coli B and K-12 strains during high cell density cultivation. Proteomics 8(10):2089–2103
Yang CK, Ewis HE, Zhang X, Lu CD, Hu HJ, Pan Y, Abdelal AT, Tai PC (2011) Nonclassical protein secretion by Bacillus subtilis in the stationary phase is not due to cell lysis. J Bacteriol 193:5607–5615
Ziebandt AK, Becher D, Ohlsen K, Hacker J, Hecker M, Engelmann S (2004) The influence of agr and sigma B in growth phase dependent regulation of virulence factors in Staphylococcus aureus. Proteomics 4(10):3034–3047
Acknowledgments
This work was supported by the National Science Fund for Distinguished Young Scholars (31125021), the National High Technology Research and Development Program of China (2011AA100905), the National Natural Science Foundation of China (No. 31171636, No. 31000752), the Key program of National Natural Science Foundation of China (No. 20836003), the National Basic Research Program of China 973 Program (2012CB720802), the 111 project B07029, and the Fundamental Research Funds for the Central Universities (No. JUSRP51320B).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, G., Chen, H., Xia, Y. et al. How are the Non-classically Secreted Bacterial Proteins Released into the Extracellular Milieu?. Curr Microbiol 67, 688–695 (2013). https://doi.org/10.1007/s00284-013-0422-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-013-0422-6