Abstract
Myeloid-derived suppressor cells (MDSC) constitute the major cell population that regulates immune responses. They are known to accumulate in tumors, chronic inflammatory and autoimmune diseases. Previous data indicate that high mobility group box 1(HMGB1) facilitates MDSC differentiation from bone marrow, suppresses NK cells, CD4+ and CD8+ T cells and is involved in cancer development. However, it remains unclear what potential mechanisms of HMGB1 facilitate MDSC differentiation. In the present work, we clearly demonstrate that HMGB1 secreted by cancer cells is N-glycosylated at Asn37, which facilitates monocytic (M)-MDSC differentiation from bone marrow via the p38/NFκB/Erk1/2 pathway and also contributes to conversion of monocytes into MDSC-like cells; HMGB1 blockade by a monoclonal antibody against the HMGB1 B box obviously reduced the accumulation of M-MDSC in tumor-bearing mice, delaying tumor growth and development; additionally, MDSC expansion and HMGB1 up-regulation were also found in breast cancer patients. All these data indicate that HMGB1 might be a potential tumor immunotherapy target.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer development is dependent on intrinsic changes as well as inflammatory factors in the tumor microenvironment. The inflammatory milieu contributes to cancer progression. Increasing reports have demonstrated that tumor milieu is immunosuppressive; which abrogates the beneficial immune response, and nourishes tumors. Cumulative evidences indicate that the most “powerful players” in turning off the immune response are immature myeloid cells (IMCs), also termed non-polarized or “resting” myeloid-derived suppressor cells (MDSC) [1]. MDSC represent a heterogeneous population that consists of myeloid progenitors and differentiates into mature macrophages, dendritic cells (DCs), and neutrophils while losing their suppressive phenotype [1, 2]. In mice, MDSC are characterized by CD11b+Gr-1+ and can be subdivided into two different subsets: CD11b+Ly6G+Ly6Clow (granulocytic-MDSC, G-MDSC) and CD11b+Ly6G−Ly6Chigh (monocytic-MDSC, M-MDSC) [3]. In contrast, in cancer patients, MDSC are defined by expression of the common myeloid marker CD33 but lack expression of mature myeloid and lymphoid cell markers [4].
Polarized MDSC infiltrate and accumulate at inflammatory sites, and contribute to systemic and site-specific immunosuppression. It is not yet fully clear how MDSC expand and transform into harmful immunosuppressive MDSC in the periphery. However, it is well established that inflammatory factors such as, TNF-α, IFN-γ, IL-1β, IL-6, and TGF-β [5–8], play critical roles in inducing of MDSC polarization and blocking their differentiation. The published data also indicate that high mobility group box protein 1 (HMGB1), as a chaperone or an inducer for many proinflammatory molecules, can drive MDSC differentiation [9]. HMGB1, as a non-histone chromosomal-binding protein, is organized into two DNA-binding domains (A box and B box) and a negatively charged C-terminus. The A box and B box are similar in conformation. Structure–function analysis showed that the B box confers proinflammatory activity, whereas the A box acts as a specific antagonist by attenuating HMGB1 B box induced secretion of proinflammatory cytokines [10, 11]. Additionally, HMGB1 has a significant role in maintaining a stable nuclear structure, contributing to development of inflammation and cell differentiation [12, 13]. As a crucial hallmark of cancers [14], HMGB1 has been verified in many cancers, such as breast cancer [15], prostate cancer [16], hepatocellular carcinoma [17], gastric cancer [18], and lung cancer [19].
Previous data also indicate that HMGB1 facilitates MDSC differentiation from bone marrow and enhances MDSC immune suppression activity, such as suppressing CD4+ and CD8+ T cells as well as NK cells [9, 20]. However, some questions remain unclear for example: (1) the mechanisms by which HMGB1 facilitates MDSC differentiation from bone marrow; (2) the other potentiality of HMGB1 to induce MDSC expansion in cancer tissues; and (3) whether HMGB1 preferentially drives G-MDSC or M-MDSC differentiation. In the present work, we clearly demonstrated that: (1) N-glycosylated HMGB1 secreted by cancer cells facilitated M-MDSC differentiation from bone marrow via p38/NFκB/Erk1/2 pathway and also contributed to conversion of monocytes into MDSC-like cells; (2) HMGB1 blockade obviously reduced the accumulation of M-MDSC in tumor-bearing mice, delaying tumor growth and development; (3) MDSC expansion and HMGB1 up-regulation were also found in breast cancer patients.
Materials and methods
Mice and cell lines
BALB/c and nude BALB/c female mice (6 weeks old) were purchased from the Animal Center of Yangzhou University and maintained in the Animal Center of Jiangsu University in compliance with the Guide for the Care and Use of Laboratory Animals” [NIH,76 FR 91 (May 11, 2011)]. The experimental protocol was approved by Jiangsu University ethics committee. The human breast cancer cell line (MCF-7) was cultured in DMEM supplemented with 10% fetal bovine serum (FBS; Hyclone, Logan, UT, USA) and 0.01 mg/mL insulin at 37 °C in a humidified atmosphere of 5% CO2.
Reagents
Phospho-antibodies (Abs) against Erk1/2 (p44/42), P38, NF-κB (p65), stat3 and corresponding total antibodies are obtained from Cell Signaling Technology (Danvers, MA, USA). U0126, SB203580, PDTC and niclosamide (inhibitors of Erk1/2, P38, NF-κB and Stat3, respectively) were purchased from univ-bio (Shanghai, China); ethyl pyruvate (EP), an inhibitor of HMGB1 secretion, was obtained from GenePharma (Shanghai, China); recombinant HMGB1 (rHMGB1) was obtained from HMGbiotech, with a purity of >95% and free from lipopolysaccharide (LPS) (Cambrex Limulus Amoebocyte Assay QCL-1000, <0.4 ng LPS per mg protein). Bio-HMGB1 was purified from MCF-7 cell conditioned medium by saturated ammonium sulfate precipitation and further purification by affinity chromatography. The isoform of HMGB1 was disulfide; endotoxin activity was detected by Limulus Amoebocyte Lysate (LAL) test, and the result was <1 EU/μg, which indicates that the protein is LPS free. IL-10 and HMGB1 detection kits were obtained from R&D Systems (Minneapolis, MN, USA) and Chondrex (Shanghai, China), respectively.
Tumor inoculations, tumor measurements, and treatment
Nude mice were inoculated subcutaneously in the flank with 1 × 107 MCF-7 breast cancer cells. For HMGB1 blockade on Day 1 (D1) or Day 14 (D14) after inoculation, 100 μg/mouse i.p. neutralizing monoclonal antibody (mAb) against HMGB1 B box or 100 μg/mouse i.p. IgG2 mAb was administered every other day according to our laboratory protocol until the mice were killed [21]. The neutralizing mAb against HMGB1 B box was produced by the hybridoma technique and purified by protein-A affinity chromatography from ascites of BALB/c mice. The valence was 106, and the isotype of mAb was IgG2a, κ chain. Tumors were measured in two perpendicular diameters every three days. Tumor volume = (a × b2)/2 with “a” as the long diameter and “b” as the short diameter. All tumors were weighed when the mice were killed.
Patients and samples
Peripheral blood was collected from 10 breast cancer patients (42 ± 3 years; 100% female) enrolled in the Affiliated Fourth People’s Hospital of Jiangsu University (Zhenjiang, China). All the patients were diagnosed with stage IV primary breast cancer without distant metastasis. Five healthy volunteers were also included in this study. All procedures performed in studies involving human participants were in accordance with the ethical standards of the Committee for Ethical Affairs of Jiangsu University (Zhenjiang, China) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Additional informed consent was obtained from all individual participants for whom identifying information is included in this article.
Western blotting
Total protein extracts were prepared using the Total Protein Extraction Kit (KeyGEN BioTECH, Shanghai, China). Equal amounts of protein were separated by 12% sodium dodecyl sulfate polyacrylamide electrophoresis (SDS-PAGE) before being transferred to polyvinylidene difluoride (PVDF) membranes. The membranes were blocked with 5% non-fat dried milk in tris-buffered saline (TBS) containing 0.1% Tween 20 and incubated with specific primary antibodies against HMGB1, p-Erk1/2, Erk1/2, p-p38, p38, p-p65, p-65, p-Stat3, Stat3 and GAPDH or β-actin overnight at 4 °C. After washing, HRP-labeled secondary antibodies were added for 1 h at 37 °C. Detection was performed by electrochemiluminescence (ECL) and relevant blots quantified by densitometry using the accompanying computerized image analysis program (Amercontrol Biosciences, San Francisco, CA, USA).
MDSC generation from bone marrow cells
Bone marrow was flushed aseptically from femurs with serum-free medium using a syringe, and red blood cells (RBCs) were lysed with red blood cell lysis buffer. Cells were cultured with 1 × 106 cells/mL in 6-well plates at 37 °C in a humidified atmosphere of 5% CO2 in medium consisting of DMEM supplemented with 10% FBS, 80 ng/mL IL-6, and 80 ng/mL GM-CSF. After 2 days of culture, all the cells were gated and stained by fluorochrome-conjugated CD11b and Ly6C mAbs, and the ratios (%double positive cells/gated cells) were determined by flow cytometry. For the MDSC frequency of tissues from cancer-bearing mice, all the cells were collected following digestion of the tissues by trypsin and lysis of red blood cells with red blood cell lysis buffer. The analysis of MDSC frequency was performed by flow cytometry. The details are as follows: according to the manufacturer’s protocol, single-cell suspensions were stained with relevant fluorochrome-conjugated mAbs against CD11c, F4/80, CD11b, Ly6C (Becton–Dickinson, Shanghai, China) and incubated for 30 min at 37 °C. Flow cytometry was performed using a FACSCalibur flow cytometer (Becton–Dickinson, Shanghai, China).
Identification of glycosylation sites by LC–MS/MS
HMGB1 was fractionated by 12% SDS-PAGE, and the protein bands were visualized by Coomassie blue staining. The lanes were excised for in-gel deglycosylation. HMGB1 gel pieces were soaked in 50 mM ammonium bicarbonate containing 100 U/mL peptide N-glycosidase (PNGase) overnight at 37 °C. Then, the gel pieces were dried with 100% acetonitrile and rehydrated with 10 ng/μL trypsin (sequencing grade, Promega, Madison, WI, USA) in 25 mM ammonium bicarbonate or 25 ng/μL chymotrypsin (Roche Molecular Biochemicals, Basel, Switzerland). Enzymatic digestion was performed overnight at 37 °C (trypsin) or 25 °C (chymotrypsin). The deglycosylated peptides were analyzed by an LC-20AB system (Shimadzu Corp., Kyoto, Japan) connected to an LTQ-Orbitrap XL mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) interfaced with an online nano-electrospray ion source (Michrom Bioresources, Auburn, CA, USA). All MS/MS data were searched against the human SWISS-PROT database using the Sequest algorithm incorporated into Bioworks software (Version 3.3.1, Thermo Fisher).
Statistical analysis
All statistical analyses were performed using Graph Pad Prism (Graphpad Software Inc., La Jolla, CA, USA). Data are expressed as the mean ± standard deviation (SD). Student’s t test or one-way ANOVA with Bonferroni correction was used to determine statistical significance. For all tests, p < 0.05 was considered statistically significant.
Results
Bio-HMGB1 characterized by N-glycosylation preferentially drove M-MDSC differentiation from bone marrow progenitor cells
First, the level of bio-HMGB1 was analyzed. As shown in Fig. 1a, MCF-7 cells secreted high levels of bio-HMGB1 in the culture supernatant; however, after addition of EP, bio-HMGB1 secretion was obviously inhibited. Furthermore, an N-glycosylation site at Asn37 of bio-HMGB1 has been also identified. Similar N-glycosylation was identified in serum of cancer-bearing mice and patients (Fig. 1b). Additionally, two N-glycosylation sites: Asn37 and Asn 115 were also found on HMGB1 secreted by monocytes/macrophages in experimental autoimmune myocarditis (data not shown).
Bio-HMGB1 from MCF-7 cells was characterized by N-glycosylation and drove M-MDSC differentiation from bone marrow progenitor cells. a HMGB1 expression in cell lysates and cell culture supernatant. MCF-7 cells were cultured in medium for 24 h, then the supernatants were harvested, and cells were lysed. Protein levels were assessed by western blotting. β-actin was used as a loading control. Representative blots are shown. b HMGB1 expression and identification of glycosylation sites. Symbols used are as follows: blue box N-acetylhexosamine (HexNAc); green circle, mannose (Hexose); yellow circle galactose (Hexose); pep peptide. c HMGB1 drove MDSC differentiation from bone marrow cells. Bone marrow cells were harvested from the femurs of healthy BALB/c mice cultured for 2 d with 250 ng/mL rHMGB1, GM-CSF + IL-6, bio-HMGB1, S-E or S-A; then the cells were stained for Ly6C and CD11b to assess the frequency of MDSC by flow cytometry. d Bio-HMGB1 preferentially drove M-MDSC differentiation from bone marrow progenitor cells. Bone marrow cells were harvested from the femurs of healthy BALB/c mice cultured for 2 d with 250 ng/mL rHMGB1, bio-HMGB1 or S-E; then, the cells were stained for Ly6G, Ly6C and CD11b to assess the percentage of MDSC by flow cytometry. e Agr-1 and iNOS expression in cell lysis. Bone marrow cells were treated with 250 ng/mL rHMGB1 or bio-HMGB1 for 48 h; and then cells were harvested by lysis. The levels of proteins were assessed by western blotting. β-actin was used as a loading control. Representative blots are shown. C control, I-G IL-6 + GM-CSF, H rHMGB1, B-H bio-HMGB1, S-E MCF-7 culture supernatant with EP, S-A MCF-7 culture supernatant with anti-HMGB1 B box mAb
To determine whether HMGB1 contributes to MDSC expansion, bone marrow progenitor cells were exposed to bio-HMGB1, IL-6 + GM-CSF, or recombinant HMGB1. As negative controls MCF-7 cell supernatant with EP (S-E) and MCF-7 cell supernatant with anti-HMGB1 B box mAb (S-A) were used. As shown in Fig. 1c, rHMGB1, bio-HMGB1 and IL-6 + GM-CSF all obviously increased the proportions of MDSC (15.49 ± 1.28, 18.51 ± 1.07 and 16.27 ± 1.15%, respectively) from bone marrow progenitor cells compared with control (5.31 ± 1.39%; p < 0.05). There were no obvious differences among the rHMGB1, IL-6 + GM-CSF and bio-HMGB1 groups. In contrast, in the S-E and S-A groups, the proportions of MDSC (8.12 ± 1.77%, 7.95 ± 1.43%, respectively) were significantly decreased compared with the bio-HMGB1 group.
Furthermore, bio-HMGB1 preferentially drove M-MDSC (10.47 ± 1.23%) differentiation from bone marrow progenitor cells compared with the rHMGB1 group (5.27 ± 0.86; p < 0.05); although the proportion of G-MDSC was also increased in the bio-HMGB1 and rHMGB1 groups (7.50 ± 0.89, 6.40 ± 0.66%, respectively); there was no significant difference between the groups. Remarkably, in the S-E group, only the proportion of M-MDSC was significantly decreased (4.37 ± 0.67%, p < 0.05). The frequency of G-MDSC was also decreased; however, the difference was not statistically significant (Fig. 1d). Since bio-HMGB1 drove M-MDSC differentiation, arginase-1 (Arg-1) and inducible nitric oxide synthase (iNOS) were detected in the bio-HMGB1 treated group. As shown in Fig. 1e, Arg-1 and iNOS expression were up-regulated in the bio-HMGB1 group compared with the rHMGB1 and control groups. Taken together, these results clearly demonstrated that bio-HMGB1 was characterized by N-glycosylation at Asn37 from cancer cells and preferentially drove M-MDSC expansion.
Bio-HMGB1 drove M-MDSC differentiation from bone marrow progenitor cells via p38-, NF-κB- and Erk1/2-dependent pathway
Next, the potential mechanisms by which bio-HMGB1 could drive M-MDSC differentiation were investigated. Published data indicate that p38, stat3, Erk1/2 and NF-κB activation are all involved in different stages of MDSC development [7, 22–24]. Therefore, p38, stat3, Erk1/2 and NF-κB activation were detected. As shown in Fig. 2a–d, bio-HMGB1 treatment resulted in a transient increase in phosphorylation of stat3, p38, p65 and Erk1/2 within bone marrow progenitor cells, peaking at 15, 30, 60 and 90 min, respectively. To confirm that stat3, p38, p65 and Erk1/2 are involved in bio-HMGB1 induction of M-MDSC differentiation, U0126, an Erk1/2 inhibitor, SB203580, a p38 inhibitor, PDTC, an NF-κB inhibitor and niclosamide, a stat3 inhibitor were employed to pre-treat cells before bio-HMGB1 or rHMGB1 treatment. As shown in Fig. 2e, the frequency of M-MDSC was 5.29 ± 0.35, 4.27 ± 0.55 and 4.73 ± 0.97% in the presence of U0126, SB203580 or PDTC, respectively. The frequency of M-MDSC was obviously decreased compared with the bio-HMGB1 group (p < 0.05). In contrast, niclosamide was not effectively able to inhibit bio-HMGB1 induction of M-MDSC differentiation (9.52 ± 0.73%). Furthermore, there was no obvious difference among the groups treated with HMGB1 with/without inhibitors. Taken together, these results clearly demonstrated that bio-HMGB1 drove M-MDSC differentiation from bone marrow progenitor cells via p38, NF-κB and Erk1/2 dependent pathway.
Bio-HMGB1 drove M-MDSC differentiation from bone marrow progenitor cells via p38, NF-κB and Erk1/2 dependent pathways. a–d Erk1/2, p38, stat3 and p65 were activated during HMGB1 stimulation of M-MDSC differentiation from bone marrow cells. Bone marrow cells were treated with bio-HMGB1. At the points indicated, cells were harvested and phosphorylated Erk1/2, p38, stat3 and p65 levels were assessed by western blotting. Representative blots are shown above and densitometric analyses below. e bio-HMGB1 drove M-MDSC differentiation via p38, NF-κB and Erk1/2. Bone marrow cells were pre-cultured with PDTC (NF-κB), niclosamide (Stat3), SB203580 (p38) and U0126 (Erk1/2) for 1 h before treatment with bio-HMGB1 and rHMGB1. The percentage of M-MDSC was assessed by flow cytometry
Bio-HMGB1 facilitated conversion of monocytes into TGF-β- and IL-10-producing MDSC-like cells
Previous evidence has shown that PGE2 contributes to the conversion of monocytes into MDSC-like cells [25, 26]. To determine whether HMGB1 can influence monocyte conversion, monocytes were isolated from the spleen of BALB/c mice and treated with bio-HMGB1, S-E and S-A. The results demonstrated that changes in the proportion of MDSC-like cells were statistical significance in the bio-HMGB1 group compared with the rHMGB1, S-E and S-A groups (p < 0.05) (Fig. 3a). Furthermore, changes in the proportion of F4/80 cells were also detected. The results showed that there were no differences in the number of F4/80 cells between the bio-HMGB1 (4.20 ± 0.80 folds), S-E group (4.87 ± 0.50 folds) and S-A group (4.12 ± 0.56 folds), whereas, in the rHMGB1 treatment group, the population of F4/80 was obviously increased (7.87 ± 0.50 folds) compared with the bio-HMGB1, S-E and S-A groups, p < 0.05, respectively (Fig. 3b). The TGF-β and IL-10 levels in supernatant, as shown in Fig. 3c, d, were analyzed among the rHMGB1, bio-HMGB1, S-E and S-A groups. TGF-β and IL-10 level were obviously increased in the bio-HMGB1 group compared with the rHMGB1, S-E and S-A groups, p < 0.05. Taken together, these results clearly demonstrated that bio-HMGB1 from cancer cells facilitated conversion of monocytes into TGF-β and IL-10 producing MDSC-like cells, whereas rHMGB1 preferentially drove the monocytes to differentiate into F4/80 macrophages.
Bio-HMGB1 facilitated conversion of monocytes into MDSC-like cells. Monocytes from the spleen were treated with 250 ng/mL rHMGB1 or bio-HMGB1 and then stained for Ly6C and CD11b or F4/80 to assess the percentage of MDSC or F4/80 cells by flow cytometry. a Changes in the population of MDSC versus fresh monocytes. b F4/80 cell changes versus fresh monocytes; c, d TGF-β and IL-10 levels in culture supernatant. The treated cells were harvested and cultured in normal medium for 24 h, and then the culture supernatant was collected for detection of TGF-β and IL-10. Data are mean ± SD from three independent experiments. C control, H rHMGB1, B-H bio-HMGB1, S-E MCF-7 culture supernatant with EP, S-A MCF-7 culture supernatant with anti-HMGB1 B box mAb
Bio-HMGB1 blockade reduced the frequency of M-MDSC in MCF-7-bearing mice and delayed tumor growth
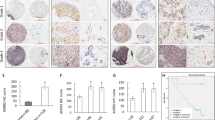
In vivo, anti-HMGB1 B box mAbs were employed to block HMGB1 on the first day (D1) or on the fourteenth day (D14) after MCF-7 inoculation. Following HMGB1 blockade, the results showed that the tumor volume was obviously reduced compared with IgG2 mAb groups and the weights were also significantly decreased compared with IgG2 mAb groups (p < 0.05; Fig. 4a, c).
Bio-HMGB1 blockade decreased M-MDSC accumulation and delayed tumor growth in MCF-7-bearing mice. a, c Tumor volumes and weights. b, d Frequency of M-MDSC in tumor tissue and spleen. The frequency of M-MDSC in tumor tissue and spleen was measured by flow cytometry when mice were killed on day 25 or day 41. Each group comprised five mice. H-B HMGB1 blockade by anti-HMGB1 B box mAbs
Furthermore, after HMGB1 blockade on D1 or D14, M-MDSC accumulation in spleen or tumor tissues was significantly decreased comparing with IgG2 mAb groups, p < 0.05 (Fig. 4b, d). Additionally, the proportion of G-MDSC was also analyzed; however, there was no significant difference (data not shown). Taken together, these results demonstrated that bio-HMGB1 blockade decreased M-MDSC accumulation and delayed tumor growth in MCF-7-bearing mice.
Bio-HMGB1 was up-regulated and the proportion of MDSC increased in breast cancer patients
The bio-HMGB1 level in patients is shown in Supplementary Figure 1a. Levels of MDSC were also analyzed in peripheral blood of breast cancer patients. The results showed that the proportion of MDSC was 15.61 ± 4.23%, which was significantly increased compared to volunteers (2.30 ± 0.67%) (Supplementary Figure 1b). Taken together, these results clearly showed that MDSC also accumulated in breast cancer patients.
Discussion
Increasing evidence supports that MDSC accumulation occurs in cancer patients. [1, 27, 28]. HMGB1 could contribute to MDSC differentiation, enhance their suppressive activity and be involved in cancer development [28, 29]. However, the potential mechanisms by which HMGB1 induces MDSC differentiation remain unclear. HMGB1 is mainly localized in the nucleus in almost all cell types and can be rapidly shuttled to other sites, such as the cytoplasm and mitochondria. Additionally, HMGB1 can also be actively or passively secreted into the extracellular space following cytokine stimulation, addition of LPS or cell death. Furthermore, HMGB1 contributes to the development of several inflammatory disorders such as inflammatory diseases, autoimmune diseases and cancers [30–32]. In the present work, we demonstrated that bio-HMGB1 from cancer cells was N-glycosylated at Asn37. Bio-HMGB1 preferentially drove M-MDSC differentiation via the p38, NF-κB and Erk1/2 pathways (Figs. 1, 2). Bio-HMGB1 could also facilitate conversion of spleen monocytes into TGF-β- and IL-10-producing MDSC-like cells. Furthermore, in vivo, bio-HMGB1 blockade by anti-HMGB1 B box mAbs decreased M-MDSC accumulation and delayed tumor growth (Fig. 4). However, rHMGB1 did not have a similar function to bio-HMGB1. Although rHMGB1 also induced bone marrow progenitor cell differentiation into MDSC, there was no difference between G-MDSC and M-MDSC; furthermore, rHMGB1 contributed to monocyte differentiation into M1 macrophages [33] (Fig. 3). All the above data indicated that HMGB1 from cancer cells and rHMGB1 plays different roles; which may be associated with post-translational modifications of HMGB1.
HMGB1 contains three cysteine residues (Cys23, Cys45 and Cys45); two of these cysteine residues (Cys23 and Cys45) can form a disulfide bond, and all three are sensitive to oxidation status in the environment. Therefore, the three major HMGB1 isoforms have been termed ‘disulfide HMGB1’, ‘thiol HMGB1’ and ‘oxidized HMGB1’ [32, 34]. The thiol isoform is a chemokine-like molecule, located in the nucleus and passively released by necrotic cells; the oxidized HMGB1 isoform is released by apoptotic cell and currently considered to be noninflammatory, while the disulfide HMGB1 is actively secreted by cells following external stress and is the main isoform found in the extracellular space and in serum. In the present work, we also identified that HMGB1 from cancer cells is the disulfide form. As a proinflammatory cytokine, this isoform could activate macrophages/monocytes and other cells and could be involved in inflammatory diseases, autoimmune diseases and cancer development.
Since there was no novel isoform existence; whether HMGB1 had undergone other modifications apart from phosphorylation, acetylization and ubiquitylation [11, 35], which are mainly associated with its' secretion or shuttle between nucleus and cytoplasm. In the present study, we found a novel post-translational modification of HMGB1: N-glycosylated at Asn37 in the serum of cancer-bearing mice and patients and in the culture supernatant of cancer cells (Fig. 1). Furthermore, we also found that HMGB1 secreted by monocytes/macrophages under external stress had two N-glycosylation sites: Asn37 and Asn 115. N-glycosylated HMGB1 (Asn37/115) contributed to macrophage reprogramming as well as cardiac fibroblast activation and collagen expression (data not shown). Data described here indicate that N-glycosylated HMGB1 produced by cancer cells functions as an inducer of MDSC expansion. Of course, further work will be needed to confirm the present function of HMGB1 associated with N-glycosylated at Asn37 in future; for example, whether N-glycosylation HMGB1 can bind advanced glycation end-products (RAGE) [36], or exogenous toll like receptor 2/4/9 (TLR2/4/9) [37, 38] and CD24/Siglec-10 [39], and whether other potential receptors exist.
Conclusions
Bio-HMGB1 secreted by cancer cells drove M-MDSC differentiation from bone marrow progenitor cells via the p38, NF-κB and Erk1/2 pathway and facilitated conversion of monocytes into MDSC-like cells; in vivo bio-HMGB1 blockade decreased M-MDSC accumulation and delayed tumor growth (Fig. 5). Furthermore, HMGB1 up-regulation and MDSC expansion were also found in breast cancer patients; which indicates that HMGB1 is a potential therapeutic target in cancer.
Bio-HMGB1 contributed to breast cancer development by inducing M-MDSC differentiation from bone marrow progenitor cells and facilitated conversion of monocytes into MDSC-like cells. HMGB1 secreted by cancer cells was characterized by N-glycosylation (Asn37) and contributed to M-MDSC expansion. The expanded M-MDSC contributed to breast cancer progression by secretion of cytokines (e.g. IL-10, TGF-β) and inhibition of T cells and NK cells
Change history
24 January 2020
The Editor-in-Chief has retracted this article.
24 January 2020
The Editor-in-Chief has retracted this article.
Abbreviations
- Arg-1:
-
Arginase-1
- bio-HMGB1:
-
HMGB1 purify from the MCF-7 cell culturing (FBS free) supernatant
- ECL:
-
Electrochemiluminescence
- EP:
-
Ethyl pyruvate
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- G-MDSC:
-
Granulocytic-MDSC
- HMGB1:
-
High mobility group box 1
- IMCs:
-
Immature myeloid cells
- iNOS:
-
Inducible nitric oxide synthase
- LAL:
-
Limulus amebocyte lysate
- mAb:
-
Monoclonal antibody
- MCF-7:
-
Human breast cancer cell line
- M-MDSC:
-
Monocytic-MDSC
- PNGase:
-
Peptide N-glycosidase
- PVDF:
-
Polyvinylidene difluoride
- RAGE:
-
Advanced glycation end-products
- rHMGB1:
-
Recombinant HMGB1
- S-A:
-
MCF-7 cell culturing supernatant anti-HMGB1 B box mAb
- SD:
-
Standard deviation
- S–E:
-
MCF-7 cell culturing supernatant with EP
References
Meirow Y, Kanterman J, Baniyash M (2015) Paving the road to tumor development and spreading: myeloid-derived suppressor cells are ruling the fate. Front Immunol 6:523. doi:10.3389/fimmu.2015.00523
Gabrilovich DI, Nagaraj S (2009) Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol 9(3):162–174. doi:10.1038/nri2506
Movahedi K, Guilliams M, Van den Bossche J, Van den Bergh R, Gysemans C, Beschin A, De Baetselier P, Van Ginderachter JA (2008) Identification of discrete tumor-induced myeloid-derived suppressor cell subpopulations with distinct T cell-suppressive activity. Blood 111(8):4233–4244. doi:10.1182/blood-2007-07-099226
Almand B, Clark JI, Nikitina E, van Beynen J, English NR, Knight SC, Carbone DP, Gabrilovich DI (2001) Increased production of immature myeloid cells in cancer patients: a mechanism of immunosuppression in cancer. J Immunol 166(1):678–689
Sade-Feldman M, Kanterman J, Ish-Shalom E, Elnekave M, Horwitz E, Baniyash M (2013) Tumor necrosis factor-alpha blocks differentiation and enhances suppressive activity of immature myeloid cells during chronic inflammation. Immunity 38(3):541–554. doi:10.1016/j.immuni.2013.02.007
Zhan X, Fang Y, Hu S, Wu Y, Yang K, Liao C, Zhang Y, Huang X, Wu M (2015) IFN-gamma differentially regulates subsets of Gr-1(+) CD11b(+) myeloid cells in chronic inflammation. Mol Immunol 66(2):451–462. doi:10.1016/j.molimm.2015.05.011
Draghiciu O, Lubbers J, Nijman HW, Daemen T (2015) Myeloid derived suppressor cells—an overview of combat strategies to increase immunotherapy efficacy. Oncoimmunology 4(1):e954829. doi:10.4161/21624011.2014.954829
Shvedova AA, Kisin ER, Yanamala N, Tkach AV, Gutkin DW, Star A, Shurin GV, Kagan VE, Shurin MR (2015) MDSC and TGFbeta are required for facilitation of tumor growth in the lungs of mice exposed to carbon nanotubes. Cancer Res 75(8):1615–1623. doi:10.1158/0008-5472.CAN-14-2376
Sims GP, Rowe DC, Rietdijk ST, Herbst R, Coyle AJ (2010) HMGB1 and RAGE in inflammation and cancer. Annu Rev Immunol 28:367–388. doi:10.1146/annurev.immunol.021908.132603
Bianchi ME, Manfredi A (2004) Chromatin and cell death. Biochim Biophys Acta 1677(1–3):181–186. doi:10.1016/j.bbaexp.2003.10.017
Dumitriu IE, Baruah P, Manfredi AA, Bianchi ME, Rovere-Querini P (2005) HMGB1: guiding immunity from within. Trends Immunol 26(7):381–387. doi:10.1016/j.it.2005.04.009
Lotze MT, Tracey KJ (2005) High-mobility group box 1 protein (HMGB1): nuclear weapon in the immune arsenal. Nat Rev Immunol 5(4):331–342. doi:10.1038/nri1594
Muller S, Scaffidi P, Degryse B, Bonaldi T, Ronfani L, Agresti A, Beltrame M, Bianchi ME (2001) New EMBO members’ review: the double life of HMGB1 chromatin protein: architectural factor and extracellular signal. EMBO J 20(16):4337–4340. doi:10.1093/emboj/20.16.4337
Guo ZS, Liu Z, Bartlett DL, Tang D, Lotze MT (2013) Life after death: targeting high mobility group box 1 in emergent cancer therapies. Am J Cancer Res 3(1):1–20
Ladoire S, Enot D, Senovilla L, Ghiringhelli F, Poirier-Colame V, Chaba K, Semeraro M, Chaix M, Penault-Llorca F, Arnould L, Poillot ML, Arveux P, Delaloge S, Andre F, Zitvogel L, Kroemer G (2016) The presence of LC3B puncta and HMGB1 expression in malignant cells correlate with the immune infiltrate in breast cancer. Autophagy 12(5):864–875. doi:10.1080/15548627.2016.1154244
Ishiguro H, Nakaigawa N, Miyoshi Y, Fujinami K, Kubota Y, Uemura H (2005) Receptor for advanced glycation end products (RAGE) and its ligand, amphotericin are overexpressed and associated with prostate cancer development. Prostate 64(1):92–100. doi:10.1002/pros.20219
Jiang W, Wang Z, Li X, Fan X, Duan Y (2012) High-mobility group box 1 is associated with clinicopathologic features in patients with hepatocellular carcinoma. Pathol Oncol Res 18(2):293–298. doi:10.1007/s12253-011-9442-3
Akaike H, Kono K, Sugai H, Takahashi A, Mimura K, Kawaguchi Y, Fujii H (2007) Expression of high mobility group box chromosomal protein-1 (HMGB-1) in gastric cancer. Anticancer Res 27(1A):449–457
Liu PL, Tsai JR, Hwang JJ, Chou SH, Cheng YJ, Lin FY, Chen YL, Hung CY, Chen WC, Chen YH, Chong IW (2010) High-mobility group box 1-mediated matrix metalloproteinase-9 expression in non-small cell lung cancer contributes to tumor cell invasiveness. Am J Respir Cell Mol Biol 43(5):530–538. doi:10.1165/rcmb.2009-0269OC
Parker KH, Sinha P, Horn LA, Clements VK, Yang H, Li J, Tracey KJ, Ostrand-Rosenberg S (2014) HMGB1 enhances immune suppression by facilitating the differentiation and suppressive activity of myeloid-derived suppressor cells. Cancer Res 74(20):5723–5733. doi:10.1158/0008-5472.CAN-13-2347
Su Z, Sun C, Zhou C, Liu Y, Zhu H, Sandoghchian S, Zheng D, Peng T, Zhang Y, Jiao Z, Wang S, Xu H (2011) HMGB1 blockade attenuates experimental autoimmune myocarditis and suppresses Th17-cell expansion. Eur J Immunol 41(12):3586–3595. doi:10.1002/eji.201141879
Mace TA, Ameen Z, Collins A, Wojcik S, Mair M, Young GS, Fuchs JR, Eubank TD, Frankel WL, Bekaii-Saab T, Bloomston M, Lesinski GB (2013) Pancreatic cancer-associated stellate cells promote differentiation of myeloid-derived suppressor cells in a STAT3-dependent manner. Cancer Res 73(10):3007–3018. doi:10.1158/0008-5472.CAN-12-4601
Waight JD, Netherby C, Hensen ML, Miller A, Hu Q, Liu S, Bogner PN, Farren MR, Lee KP, Liu K, Abrams SI (2013) Myeloid-derived suppressor cell development is regulated by a STAT/IRF-8 axis. J Clin Investig 123(10):4464–4478. doi:10.1172/JCI68189
Tian J, Ma J, Ma K, Guo H, Baidoo SE, Zhang Y, Yan J, Lu L, Xu H, Wang S (2013) β-Glucan enhances antitumor immune responses by regulating differentiation and function of monocytic myeloid-derived suppressor cells. Eur J Immunol 43(5):1220–1230. doi:10.1002/eji.201242841
Mao Y, Sarhan D, Steven A, Seliger B, Kiessling R, Lundqvist A (2014) Inhibition of tumor-derived prostaglandin-e2 blocks the induction of myeloid-derived suppressor cells and recovers natural killer cell activity. Clin Cancer Res 20(15):4096–4106. doi:10.1158/1078-0432.CCR-14-0635
Obermajer N, Muthuswamy R, Lesnock J, Edwards RP, Kalinski P (2011) Positive feedback between PGE2 and COX2 redirects the differentiation of human dendritic cells toward stable myeloid-derived suppressor cells. Blood 118(20):5498–5505. doi:10.1182/blood-2011-07-365825
Albeituni SH, Ding C, Yan J (2013) Hampering immune suppressors: therapeutic targeting of myeloid-derived suppressor cells in cancer. Cancer J 19(6):490–501. doi:10.1097/PPO.0000000000000006
Baniyash M, Sade-Feldman M, Kanterman J (2014) Chronic inflammation and cancer: suppressing the suppressors. Cancer Immunol Immunother 63(1):11–20. doi:10.1007/s00262-013-1468-9
Diener KR, Al-Dasooqi N, Lousberg EL, Hayball JD (2013) The multifunctional alarmin HMGB1 with roles in the pathophysiology of sepsis and cancer. Immunol Cell Biol 91(7):443–450. doi:10.1038/icb.2013.25
Magna M, Pisetsky DS (2014) The role of HMGB1 in the pathogenesis of inflammatory and autoimmune diseases. Mol Med 20:138–146. doi:10.2119/molmed.2013.00164
Kang R, Zhang Q, Zeh HJ 3rd, Lotze MT, Tang D (2013) HMGB1 in cancer: good, bad, or both? Clin Cancer Res 19(15):4046–4057. doi:10.1158/1078-0432.CCR-13-0495
Harris HE, Andersson U, Pisetsky DS (2012) HMGB1: a multifunctional alarmin driving autoimmune and inflammatory disease. Nat Rev Rheumatol 8(4):195–202. doi:10.1038/nrrheum.2011.222
Su Z, Zhang P, Yu Y, Lu H, Liu Y, Ni P, Su X, Wang D, Wang J, Shen H, Xu W, Xu H (2016) HMGB1 facilitated macrophage reprogramming towards a proinflammatory M1-like phenotype in experimental autoimmune myocarditis development. Sci Rep 6:21884. doi:10.1038/srep21884
Antoine DJ, Harris HE, Andersson U, Tracey KJ, Bianchi ME (2014) A systematic nomenclature for the redox states of high mobility group box (HMGB) proteins. Mol Med 20:135–137. doi:10.2119/molmed.2014.00022
Janko C, Filipovic M, Munoz LE, Schorn C, Schett G, Ivanovic-Burmazovic I, Herrmann M (2014) Redox modulation of HMGB1-related signaling. Antioxid Redox Signal 20(7):1075–1085. doi:10.1089/ars.2013.5179
Hori O, Brett J, Slattery T, Cao R, Zhang J, Chen JX, Nagashima M, Lundh ER, Vijay S, Nitecki D et al (1995) The receptor for advanced glycation end products (RAGE) is a cellular binding site for amphoterin. Mediation of neurite outgrowth and co-expression of rage and amphoterin in the developing nervous system. J Biol Chem 270(43):25752–25761
Ivanov S, Dragoi AM, Wang X, Dallacosta C, Louten J, Musco G, Sitia G, Yap GS, Wan Y, Biron CA, Bianchi ME, Wang H, Chu WM (2007) A novel role for HMGB1 in TLR9-mediated inflammatory responses to CpG-DNA. Blood 110(6):1970–1981. doi:10.1182/blood-2006-09-044776
Park JS, Svetkauskaite D, He Q, Kim JY, Strassheim D, Ishizaka A, Abraham E (2004) Involvement of toll-like receptors 2 and 4 in cellular activation by high mobility group box 1 protein. J Biol Chem 279(9):7370–7377. doi:10.1074/jbc.M306793200
Chen GY, Tang J, Zheng P, Liu Y (2009) CD24 and Siglec-10 selectively repress tissue damage-induced immune responses. Science 323(5922):1722–1725. doi:10.1126/science.1168988
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 81370084, 81502663), the Social Development Foundation of Jiangsu Province (Grant No. BE2015668). The six talent peaks project in Jiangsu Province (2013-WSN-002, 2015-WSN-005); the maternal and child project in Jiangsu Province (F201511). We thank Huaxi Xu and the International Science Editing Company for editing and proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
The Editor-in-Chief has retracted this article [1] because a number of the images in Figures 4a and 4c, appear to have been duplicated then either rotated, cropped or stretched to make them appear to be different tumors. The Editor-in-Chief no longer has confidence in the results and conclusions presented in this article. Jia Wang agrees with this retraction. Seidu A. Richard does not agree with this retraction. Zhaoliang Su, Ping Ni, Peng She, Yueqin Liu, Wenlin Xu and Haitao Zhu have not responded to correspondence about this retraction.
[1] Zhaoliang Su, Ping Ni, Peng She, Yueqin Liu, Seidu A. Richard,Wenlin Xu, Haitao Zhu, Jia Wang. Bio-HMGB1 from breast cancer contributes to M-MDSC differentiation from bone marrow progenitor cells and facilitates conversion of monocytes into MDSC-like cells. Cancer Immunol Immunother 2017 66(3): 391-401, doi: 10.1007/s00262-016-1942-2.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Su, Z., Ni, P., She, P. et al. RETRACTED ARTICLE: Bio-HMGB1 from breast cancer contributes to M-MDSC differentiation from bone marrow progenitor cells and facilitates conversion of monocytes into MDSC-like cells. Cancer Immunol Immunother 66, 391–401 (2017). https://doi.org/10.1007/s00262-016-1942-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-016-1942-2