Abstract
Purpose
The purpose of this study was to evaluate the use of CT radiomics features and machine learning analysis to identify aggressive tumor features, including high nuclear grade (NG) and sarcomatoid (sarc) features, in large renal cell carcinomas (RCCs).
Methods
CT-based volumetric radiomics analysis was performed on non-contrast (NC) and portal venous (PV) phase multidetector computed tomography images of large (> 7 cm) untreated RCCs in 141 patients (46W/95M, mean age 60 years). Machine learning analysis was applied to the extracted radiomics data to evaluate for association with high NG (grade 3–4), with multichannel analysis for NG performed in a subset of patients (n = 80). A similar analysis was performed in a sarcomatoid rich cohort (n = 43, 31M/12F, mean age 63.7 years) using size-matched non-sarcomatoid controls (n = 49) for identification of sarcomatoid change.
Results
The XG Boost Model performed best on the tested data. After manual and machine feature extraction, models consisted of 3, 7, 5, 10 radiomics features for NC sarc, PV sarc, NC NG and PV NG, respectively. The area under the receiver operating characteristic curve (AUC) for these models was 0.59, 0.65, 0.69 and 0.58 respectively. The multichannel NG model extracted 6 radiomic features using the feature selection strategy and showed an AUC of 0.67.
Conclusions
Statistically significant but weak associations between aggressive tumor features (high nuclear grade, sarcomatoid features) in large RCC were identified using 3D radiomics and machine learning analysis
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As the volume of computed tomography (CT) performed for a variety of indications continues to increase, the incidence of renal cell carcinoma (RCC) has also continued to rise [1,2,3,4,5,6]. Spatial heterogeneity is a common feature of RCC, with multiple studies demonstrating variability within tumors with respect to pathologic features, genomics, and RNA/protein expression [7,8,9]. This heterogeneity gives rise to a spectrum of biologic and clinical behaviors, with an increasingly less aggressive management approach in more indolent disease and nephron sparing approaches in cases where intervention is warranted [10,11,12]. Pathologic markers of tumor aggressiveness such as higher nuclear grade (NG) or presence of sarcomatoid (sarc) features may only be present in a small portion of the tumor but may profoundly impact treatment decisions and prognosis [13,14,15]. These small areas can be challenging to identify on biopsy, and although radiomic features provide more global tumor assessment and have shown some promise in non-invasively capturing and characterizing tumor heterogeneity, some aggressive tumor features have remained elusive at imaging [9, 16,17,18,19,20,21,22,23,24,25,26,27,28,29]. If aggressive features could be reliably identified in advance of surgery, either through more targeted biopsies or non-invasive assessment, it could have immediate clinical impact on treatment decisions and prognostication. Recently, multiple groups have used machine learning analysis applied to radiomics features in an attempt to improve performance in identification of aggressive features such as high nuclear grade on imaging, with some success [30,31,32,33,34,35,36]. Identification of sarcomatoid features has remained challenging from CT imaging [24]. The purpose of this study is to evaluate the use of CT radiomics features and machine learning analysis to identify aggressive tumor features, including high nuclear grade and sarcomatoid features, in large RCCs. For nuclear grade, this would be an attempt to reproduce other groups’ results and for sarcomatoid features, to identify an as yet unidentified radiomics imaging signature.
Methods
This study was IRB approved and HIPAA compliant.
Patient selection and CT images
The CT images obtained between 2000 and 2013 of 141 patients (46 women and 95 men, mean age 60 years) with large (> 7 cm) RCCs were obtained from the surgical database of the Department of Urology and were retrospectively reviewed. All patients in the cohort had a CT scan performed before undergoing surgery or receiving any other treatment. Subsequent removal of the primary tumor and pathologic analysis that included histologic subtyping and nuclear grading were performed for all patients. CT texture analysis data from these patients were previously analyzed in [19], where single-slice analysis was used [19] and a multi-platform radiomics analysis was performed by Dreyfuss et al. in 2019, where both single-slice and volumetric platforms were applied [37]. We specifically targeted large RCCs to increase the likelihood that aggressive features would be present.
Analysis of both portal venous phase images (n = 124, 44 with portal venous phase only) and non-contrast images (n = 97; 17 had non-contrast images only, 80 had both non-contrast and portal venous phase images) were performed. 74 of 124 portal venous (59.7%) CT examinations were performed at institutions other than the study institution. All scans were performed using MDCT scanners and the imaging parameters were as follows: a tube potential of 100–140 kV (with 110 of 124 (89.4%) scans using a tube potential of 120 kV) and a matrix of 512 × 512 × 16. Most CT scans were performed using automated or variable tube current, and the slice thickness used for 122 of 125 scans was 2–5 mm. Although the non-contrast and portal venous analyses were performed separately, a multi-channel analysis of patients who had both datasets was additionally performed. This cohort of 141 patients with large RCCs was used in the assessment of imaging features of nuclear grade. Patients with nuclear grade of 3–4 were considered high grade, while grades 1–2 were considered low grade.
A second sarcomatoid rich dataset was created, using CT imaging obtained between 2001 and 2018, including 43 RCCs with sarcomatoid features (31 M, 12F, mean age 63.7 years) with 49 size-matched non-sarcomatoid RCCs from the nuclear grade cohort above to serve as controls (30 M, 19F, mean age 64.4 yrs) with extraction of radiomics features using the method described above. As with the nuclear grade analysis, both non-contrast (n = 28, n = 3 non-contrast only) and portal venous phase CT (n = 40, n = 15 pv only, n = 25 both pv and non-contrast) images in patients with sarcomatoid RCCs were evaluated. Size-matched non-sarcomatoid controls came from the large RCC nuclear grade dataset and had similar distribution of non-contrast and portal venous exams (pv n = 49, non-con = 36, both n = 31). For a subset of 25 patients in the sarcomatoid cohort, the percentage of sarcomatoid features present in the tumor was quantified by the surgical pathologist. Additional analysis of this subset of patients was performed where a threshold of 10% sarcomatoid features was applied, and included tumors were reanalyzed to see whether higher tumoral fraction of aggressive features improved performance of the model. This is further detailed under SMOTE analysis.
Radiomics platform
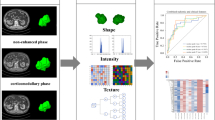
Radiomics features were extracted using Healthmyne Radiomic Precision Metrics (https://www.healthmyne.com/radiomic-precision-metrics/, Madison, WI, USA), a server-based platform that performs volumetric CT radiomic analysis. Healthmyne does not perform a filtration step and analyzes unfiltered data. This software extracts over 300 radiomics features, including first-order texture features (mean gray-level intensity, entropy, standard deviation) and second-order texture features derived using gray-level co-occurrence matrix (GLCM). Second-order metrics allow quantification of the spatial relationship between pixels [38]. It also extracts a variety of anatomic and morphologic tumor descriptors including tumor volume, surface area, sphericity, etc. Some features are locations in the image used to calculate distances (long axis, short axis, etc.). These do not extract meaningful image data, only reflect coordinates and were manually excluded from the analysis (the calculated distances from these coordinates reflecting tumor measurements were included).
Region of interest (ROI) selection
The process of ROI selection for the 3-dimensional platform (Healthmyne) is as follows. First, the CT scan of interest is opened in the platform. The index slice at the level at the largest overall transverse tumor diameter is identified. The tumor is traced at this level with care to maintain the outer margins of the ROI just within the boundaries of the tumor. Once the single-slice ROI has been traced, automatic segmentation is performed. During automatic segmentation, the entire volume of tumor as seen on cross-sectional CT imaging is automatically segmented by the platform. Following automatic segmentation, the user must manually refine the tumor boundaries in order to ensure non-tumor tissues are excluded from analysis. Once correct tumor margins have been verified, the radiomics metrics are extracted. All segmentations were created by a trained medical student under the direct supervision of a fellowship trained abdominal radiologist with 11 years of experience.
Data processing and cleaning
As discussed above in Radiomics Platform section, an initial pass was made through the data with manual exclusion of categories that were not extracting meaningful image data (coordinates, etc.). The data extracted from the CT scans were to the best of our knowledge and resources available, and had some missing data. To avoid data loss, we used different imputation methods to fit the data. As we were aware of no advantages to more complex methods, when imputation was needed for a method we chose a simple imputation scheme of replacing data for a component in a feature by the mean of values of other features, making use of the SimpleImputer package available in scikit-learn [39]. We made sure that no data leakage occurred in fitting of the data by performing imputation on the training data and transforming the test data before prediction. For machine learning methods with built in imputation schemes we used those schemes.
Data visualization
Before developing any model, we try to get an estimate of the data distribution and analyze if there is a clear and evident margin of classification. We visualized our data to understand the distribution over both the classes on a 2-dimensional plot using t-SNE (t-distributed Stochastic Neighbor Embedding). This is a non-linear technique used for dimensionality reduction of high dimensional data and is widely used for cancer detection applications. The data were imputed and normalized before transforming to 2-dimensional data. The t-SNE plots are shown in the Results section.
Machine learning analysis
The goal of our model was to evaluate for association between high nuclear grade (grade 3–4) and imaging features in our first cohort and between presence/amount of sarcomatoid features and imaging features in our second cohort. For the purpose of our study, we tested our model with gradient boosted trees (implemented in XGBoost (XGB), and Random Forest (RF) and Support Vector Machine (SVM) (implements in Scikit-learn [39]). The Scikit-learn implementations of RF and SVM do not allow for missing data while XGB has a built in imputation scheme. Therefore, we have used data imputation (see Sec. Data Processing and Cleaning) on training data for SVM and RF during model development. The performance of the models were evaluated on six metrics.
-
1.
Accuracy: This metric is the fraction of correct predictions made by the model.
$${\text{Accuracy}}= \frac{{\text{Number}}\; {\text{of}}\; {\text{correct}}\; {\text{predictions}} }{{\text{Total}}\; {\text{number}}\; {\text{of}}\; {\text{predictions}}}$$ -
2.
Precision: Also known as positive predictive value, this metric gives the fraction of true positive predictions among total positive predictions.
$${\text{Precision}}= \frac{{\text{True}}\; {\text{positive}}\; {\text{predictions}}}{{\text{True}}\; {\text{positive}}\; {\text{predictions}}+{\text{false}}\; {\text{positive}}\; {\text{predictions}}}$$ -
3.
Recall: Also known as sensitivity, gives the fraction of true positive predictions among actual positive elements.
$${\text{Recall}}= \frac{{\text{True}}\; {\text{positive}}\; {\text{predictions}}}{{\text{True}}\; {\text{positive}}\; {\text{predictions}}+{\text{false}}\; {\text{negative}}\; {\text{predictions}}}$$ -
4.
f1_score: This metric is a measure of test’s accuracy and is defined as the harmonic mean of precision and recall.
$$f1=2\cdot\frac{{\text{precision}}\cdot{\text{recall}}}{{\text{precision}}+{\text{recall}}}$$ -
5.
AUC: Receiver operating characteristic (ROC) curve is a graph of true positive rate with false positive rate. This is a measure of a classifier’s performance plotted for different classification thresholds of the classifier. The metric (AUC) is the area under the ROC curved provides an aggregate measure of the classifier’s performance
-
6.
Geometric mean: To counter a 2-class imbalance in the dataset, another metric to determine the classification accuracy is the geometric mean score which is just the geometric mean of the true positive rate and true negative rate
We calculate the mean and standard error for these metrics using a set of values determined from 20 iterations of fivefold cross-validated scores, where each score is determined for each fold, leading to 100 samples for each statistic being used to find mean and standard deviation.
Further, we performed permutation testing to determine the statistical significance of the model. We tested the model for f1 score averaged over 20 iterations of fivefold cross-validation and then ran 100 random permutations of the target data pairings to input features to estimate the p-value score of the model. We observed the mean of the final model score (i.e., the f1 score for every run), and mean and standard deviation of p-values (which is a measure of fraction of how many random permutation runs performed better than the original model) over a set of 5 permutation test runs (where each run included 100 permutations, as noted just above).
Feature ranking and selection strategy
The datasets had a high feature-to-sample ratio so dimensional reduction and feature selection were an important step to filter out unwanted features. We explored two approaches, including our own custom feature ranking algorithm and the algorithm used in XGBoost to assign a normalized importance score to each of the features.
The evaluation of features was done in two phases. In the first phase, or ‘feature ranking phase’, we performed fivefold cross-validation on the dataset for 20 times each time with a different train-test split. In essence, the model ran for 100 independent iterations and assigned an importance score each time. The feature list was sorted based on the cumulative importance score after 100 runs. In the second phase, or ‘feature selection phase’, a fivefold CV score averaged over 20 runs was observed over the entire dataset with just the highest ranked feature from phase 1. This was repeated with the 2 highest ranked features and the average fivefold CV score was observed. The average fivefold CV score vs number of features was plotted and the list of features for which the best CV score was obtained gives us the optimal set of features which we used for model optimization later. To avoid data leakage, during model assessment this feature ranking and selection was performed only for data subsets using nested cross-validation, as described below.
Nested cross-validation
As we are dealing with small datasets, a commonly known problem of data leakage often arises and can heavily impact or bias the result. To avoid this, we have evaluated the model performance using a nested cross-validation approach. We use a fivefold loop which forms our ‘outer loop’. The training data which forms the ‘inner loop’, in every fold, goes through the feature selection strategy described above to give the optimal feature list to be used for model optimization. We observe the scoring metrics on the test data using this optimal feature list. Each fold produces its own feature list and scoring metrics and we average the scoring metrics over the 5 folds.
Synthetic minority oversampling technique (SMOTE) analysis
For the sarcomatoid rich dataset, the samples included had a non-zero percentage of sarcomatoid features. However, a concern was that even if sarcomatoid features have an imaging signature it might be overwhelmed by the background features of the tumor if only a small percentage of sarcomatoid features are present. To address this concern and give the modeling the best chance of success, we used a filter on the percentage of sarcomatoid features present, taking only values with \(\ge 10\%\) (this was the median percentage in our cohort, n = 25). As many of the samples had a sarcomatoid percentage less than 10%, these samples were filtered out, causing an imbalance in the samples of each class. We used SMOTE on the minority class and performed naive classification using XGBoost. To apply SMOTE, it was essential to use imputation (using the methods from Sec. Data Processing and Cleaning) before performing classification.
Results
Patient cohorts
Two patient cohorts were evaluated. One was a group of 141 patients (46 women and 95 men, mean age 60 years) with large RCC (mean size 10 ± 3 cm, median 9 cm) who underwent non-contrast and/or portal venous phase CT used for identification of high nuclear grade (NG). This group contained mostly clear cell RCC (n = 118, 84%), with fewer non-clear cell (papillary n = 14, chromophobe n = 9). There was a slight majority of high grade tumors, (n = 75 nuclear grade 3, 4) with 63 low grade (nuclear grade 1, 2) and 3 tumors not graded (Fig. 1).
Renal Cell carcinoma of increasing nuclear grade in 4 different patients from the large RCC nuclear grade cohort. 51-year-old male with 7 cm homogeneous renal mass found on PV phase contrast enhanced CT (arrow, a), found to be nuclear grade 1 clear cell renal cell carcinoma (ccRCC) at tumor resection. He remains alive and disease free > 93 months later. A heterogeneous 10 cm tumor on CECT in a 76-year-old female (arrow, b) was nuclear grade 2 ccRCC, while the 12 cm tumor in a 79-year-old female (arrow, c) showed nuclear grade 3. An 8 cm left renal mass invading the left psoas on CECT in a 67-year-old female (d) was ccRCC, NG 4, and patient died of her disease approximately 17 months after surgery
The second was a group of 43 patients with RCCs with sarcomatoid features (31 M, 12F, mean age 63.7 yrs) who underwent non-contrast and/or portal venous phase CT with 49 size-matched non-sarcomatoid RCCs from the nuclear grade cohort above to serve as controls (30 M, 19F, mean age 64.4 yrs). Mean size of the sarcomatoid tumors was 9.8 ± 3 cm, median 10 cm; mean size of controls was 8.7 ± 2 cm, median 9 cm. Sarcomatoid tumors were predominantly clear cell (n = 35, 81%). A group of 25 tumors in the sarcomatoid cohort had an estimate of the percentage of tumor with sarcomatoid features. In this subcohort, the median was 10% sarcomatoid features, mean 21% ± 26%, range 1–90% (Fig. 2).
Renal cell carcinoma with increasing percentage of sarcomatoid features in 3 different patients from the sarcomatoid cohort. An 8 cm infiltrative renal mass on portal venous phase contrast enhanced CT (arrow, a) in a 64-year-old male was clear cell renal cell carcinoma with 10% sarcomatoid features. A different heterogeneous 8 cm mass with large intratumoral vessels found at CECT (arrow, b) in a 76-year-old male was ccRCC with 40% sarcomatoid features. A 65-year-old male presented with a 7 cm left renal mass at CECT (arrow, c), found to have ccRCC with 90% sarcomatoid features. This patient died of his disease within 1 year of diagnosis
Classification
We observed that XGB was the best performing classifier on each of the datasets when compared with RF and SVM. Summary of the classification results with XGB is detailed in Table 1. Non-contrast and portal venous phase CT datasets from sarcomatoid patients with size-matched controls were classified to distinguish the presence of sarcomatoid features whereas non-contrast and portal venous phase CT images in patients with large RCCs were classified to distinguish the presence of high (grade 3–4) nuclear grade compared to low (grade 1–2). In the portal venous phase large RCC dataset (PV_NG), the model achieved 58% accuracy for identification of high nuclear grade, with 69% achieved for the non-contrast CT dataset (Noncon_NG). In the portal venous phase sarcomatoid data set (PV_Sarc), accuracy of 66% was achieved for identifying sarcomatoid features compared to size-matched controls. For the non-contrast sarcomatoid dataset (Noncon_Sarc), accuracy of 60% was obtained. We also tested using multichannel analysis on a cohort with patients from both non-contrast CT and portal venous phase CT datasets and attained an accuracy of 67%.
Feature selection
Each dataset had a different number of features and samples available for feature selection. Among 318 texture features for Noncon_Sarc dataset, 463 texture features for PV_Sarc dataset, 317 texture features for Noncon_NG dataset and 49 features for PV_NG dataset. Those that were not clinically relevant (did not describe imaging data) were manually excluded leaving 80, 85, 82 and 49 features, respectively. Our feature selection strategy then selected 3, 7, 5 and 10 radiomics features, respectively (Table 2), that were sufficient to provide a comparable accuracy to when all features were considered together. Our feature selection strategy extracted 6 radiomic features on the multichannel cohort.
Nested cross-validation
Results for our fivefold nested CV approach are tabulated in Table 3. We found these average scores are comparable to classification results by fivefold CV with XGBoost. PV_Sarc dataset was able to achieve 67% accuracy while the Noncon_Sarc dataset accuracy was fairly low at 48%. Noncon_NG and PV_NG gave similar accuracy of 56% and 60%, respectively, while multichannel cohort gave 56% accuracy.
Permutation tests
The statistical significance of our model predictive ability was assessed by performing permutation tests averaged over 5 runs. We used f1 score as the metric being assessed. The results are shown in Table 4. A low p-value score pertains to high significance of the model. Our results show that the mean p-value for each of the dataset is less than 0.10, demonstrating that the predictions are better than random with high probability.
t-SNE plots
The t-SNE plots (t-distributed Stochastic Neighbor Embedding) for each dataset can be seen in Fig. 3. As we observe each of the datasets, both classes are spread across evenly and there is no clear division between them. This suggests that the input features are not strongly correlated to the aggressiveness of the RCC, consistent with the results of machine learning fitting.
t-SNE plots. The top panel demonstrates plots for the nuclear grade dataset, with blue dots representing high grade tumors (nuclear grade 3–4) and red dots low grade on non-con, non-con + pv and pv datasets, showing a fairly even spread without clear delineating threshold. Similar results are seen in the lower panel for sarcomatoid features (blue sarcomatoid features present, red absent) on non-con, portal venous and thresholded (10% sarcomatoid features) data
SMOTE analysis
For the subgroup of patients in the sarcomatoid database with percentage of sarcomatoid features included (n = 25), the median was 10%. We further filtered those data samples keeping those with \(\ge 10\%\) sarcomatoid features to explore if tumors with more sarcomatoid features present might be better classified. However, we did not observe any improvement in the classification results using XGBoost classifier when comparing the full subgroup of patients in the sarcomatoid database with percentage of sarcomatoid features included and those thresholded with \(\ge 10\%\) sarcomatoid features. The results with SMOTE are shown in Table 5.
Discussion
Renal cell carcinoma is a heterogeneous tumor that can contain multiple different nuclear grades or genetic features in a single tumor [7]. Even if only a small portion of the tumor is grade 4, the overall nuclear grade assigned to the tumor will be 4 and that grade will drive treatment decisions and patient prognosis. Similarly, even if only a small portion of the tumor contains sarcomatoid features, if these features are identified prospectively, the identification can profoundly impact patient management and these patients are often not surgical candidates [14, 15]. However, sometimes these small areas may be missed at biopsy due to sampling error, and this uncertainty about the reliability of biopsy and the presence of aggressive tumor features may make prospective informed decision making about treatment challenging [8, 9, 40].
Global tumor assessment on imaging provides a non-invasive means for capturing tumor characteristics and radiomics features have shown promising associations with histopathologic features in RCC. Given the large number of radiomics features produced by many software packages, use of machine learning analysis can aid in feature extraction and robust analysis of feature and model performance. Several groups have recently looked specifically at the ability of radiomics data evaluated with machine learning to identify nuclear grade with some promising results [30, 32,33,34,35, 41, 42]. For example, Bektas et al. looked at a cohort of 54 clear cell RCCs (ccRCCs), roughly half high grade tumors. They used different machine learning classifiers of 279 2D texture features extracted from portal venous phase CT. In their series, the overall accuracy, sensitivity, specificity (for detecting high grade ccRCC) and overall AUC for the best model were 85.1%, 91.3%, 80.6% and 86%, respectively [30]. He et al. looked at 227 ccRCCs, extracted 14 conventional imaging features manually and 556 texture features using a software application, applied machine learning analysis, and found that the predictive models for high grade vs low grade tumors had accuracies ranging from approximately 90–94% [31].
Identification of sarcomatoid features has been challenging on CT imaging to date. Schieda et al. looked at a cohort of 20 sarcomatoid RCCs matched to 25 ccRCCs and manually extracted a variety of imaging features including tumor size, subjective tumor heterogeneity, tumor margin, presence of tumoral calcification and intra and peritumoral vascularity among other features. In addition, they extracted a variety of texture features. The best performing model combined textural features and subjective features demonstrated an AUC of 0.81 in identification of sarcomatoid features [24]. Meng et al. recently looked at a cohort of 29 sarcomatoid RCCs using both subjective and radiomics features and found widely variable model performance with AUCs ranging from 0.77–0.97 [43]. Our cohort of 43 sarcomatoid RCCs is one of the larger series to date.
However, even using a similar approach in both our cohort of 141 large RCCs and 43 size-matched sarcomatoid RCCs, we were unable to reproduce these results. We used an extensive feature selection process, applied multiple different machine learning models, tested with fivefold nested cross-validation, performed multichannel analysis of both non-contrast and pv phase post contrast data, used thresholded analysis of sarcomatoid features where quantitative data were available, and performed follow up permutation testing. There are several possible explanations for this performance. There is a growing body of literature that a variety of imaging parameters unrelated to biologic heterogeneity may impact selected radiomic features [44,45,46,47,48,49]. In addition, there is variability in the features extracted and even the values produced for the same types of feature depending on the software platform used [37]. There have been calls for standardization to make such automatically generated features a more viable clinical tool [50]. We also note that we used a 3D segmentation tool that incorporated the imaging features of the entire large tumor. It is possible that if only small areas of high nuclear grade or sarcomatoid features were present they may have been obscured by the dominant imaging features of the rest of the tumor. If other studies were more selective about where in the tumor the ROI was placed, or if 2D segmentation was used, this may have been less of a factor. However, even using a threshold of 10% sarcomatoid features to select tumors with a higher percentage of sarcomatoid change, our model performance did not improve. In a prior analysis using a similar dataset to this study and single-slice analysis where only a small portion of the tumor was evaluated, only weak associations with tumor grade and no association with sarcomatoid features were identified.
An additional factor that could play a role is machine learning methodology. In particular, unless great care is taken, there is risk for data leakage, and the impact of even small amounts of data leakage can be significant, depending on the sample size and machine learning analysis applied. Therefore, very robust and rigorous methodology must be used. We ensured once we split the data, none of the strategies including imputation, normalization, feature selection and feature ranking were aware of any data point from the test data during fitting the model. Only once the model was ready, the test data were transformed as the training data before observing the prediction results. We note that by allowing some data leakage we can significantly improve our results. Specifically, if we fit to the whole dataset, obtain a ranked list of features, and then optimize the CV score using that list (this follows the approach in Sec. Feature Ranking and Selection Strategy but with the whole dataset) we obtain a 5–7% improvement in the results (see supplemental section Table 5). This improvement shows the importance of avoiding data leakage in the analysis.
The features that performed well in our model included things that make intuitive sense and are similar to those extracted in other series, including things like density, uniformity and GLCM features such as entropy as well as size metrics. It is possible that a study using more precise radiologic pathologic correlation to look directly at the imaging features of portions of the tumors known to have aggressive features may help better delineate the imaging signature of these areas or improve model performance. This is an area of investigation that warrants further study.
There are limitations to this study. This is a relatively small dataset for this type of analysis, but it is comparable to those used in other studies, with this sarcomatoid dataset one of the largest analyzed to date. There is some heterogeneity to the CT data, but the imaging parameters used to obtain the images were within a reasonable range, and data normalization was used. Both non-contrast and portal venous phase images were separately analyzed and multi-channel analysis was performed where the data were available. Portal venous phase CT was selected due to wide applicability, but other phases of contrast including corticomedullary or delayed phase images commonly used in renal imaging were not evaluated. Quantification of sarcomatoid features was only available in a subset of patients for this study. Finally, pathologic features were used as a surrogate for clinical outcomes, but this is not the only determinant of outcomes and more detailed analysis and modeling using clinical endpoints such as survival is ongoing.
Conclusion
Despite use of a robust radiomics platform and highly effective machine learning models, performance of models for identifying aggressive tumor features in RCC (high nuclear grade, sarcomatoid features) were quite poor. Our group was unable to reproduce results seen by other groups in the literature, possibly due to variability in CT data, radiomics platforms and machine learning analyses approaches, which limits the ability to widely apply these models in clinical practice until further standardization is performed. Further study using more precise radiologic pathologic correlation may be useful in better delineating the imaging signature of these aggressive tumor features.
References
Moreno CC, Hemingway J, Johnson AC, Hughes DR, Mittal PK, Duszak R, Jr. Changing Abdominal Imaging Utilization Patterns: Perspectives From Medicare Beneficiaries Over Two Decades. J Am Coll Radiol 2016; 13:894-903
Chow WH, Devesa SS, Warren JL, Fraumeni JF, Jr. Rising incidence of renal cell cancer in the United States. JAMA 1999; 281:1628-1631
Cho E, Adami HO, Lindblad P. Epidemiology of renal cell cancer. Hematol Oncol Clin North Am 2011; 25:651-665
Gandaglia G, Ravi P, Abdollah F, et al. Contemporary incidence and mortality rates of kidney cancer in the United States. Can Urol Assoc J 2014; 8:247-252
Nguyen MM, Gill IS, Ellison LM. The evolving presentation of renal carcinoma in the United States: trends from the Surveillance, Epidemiology, and End Results program. J Urol 2006; 176:2397–2400; discussion 2400
Smith-Bindman R, Kwan ML, Marlow EC, et al. Trends in Use of Medical Imaging in US Health Care Systems and in Ontario, Canada, 2000-2016. JAMA 2019; 322:843-856
Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 2012; 366:883-892
Ball MW, Bezerra SM, Gorin MA, et al. Grade Heterogeneity in Small Renal Masses: Potential Implications for Renal Mass Biopsy. J Urol 2014;
Halverson SJ, Kunju LP, Bhalla R, et al. Accuracy of determining small renal mass management with risk stratified biopsies: confirmation by final pathology. J Urol 2013; 189:441-446
Silverman SG, Israel GM, Herts BR, Richie JP. Management of the incidental renal mass. Radiology 2008; 249:16-31
Volpe A, Cadeddu JA, Cestari A, et al. Contemporary management of small renal masses. Eur Urol 2011; 60:501-515
Volpe A, Finelli A, Gill IS, et al. Rationale for percutaneous biopsy and histologic characterisation of renal tumours. Eur Urol 2012; 62:491-504
Kapur P, Pena-Llopis S, Christie A, et al. Effects on survival of BAP1 and PBRM1 mutations in sporadic clear-cell renal-cell carcinoma: a retrospective analysis with independent validation. Lancet Oncol 2013; 14:159-167
Shuch B, Bratslavsky G, Linehan WM, Srinivasan R. Sarcomatoid renal cell carcinoma: a comprehensive review of the biology and current treatment strategies. Oncologist 2012; 17:46-54
Shuch B, Bratslavsky G, Shih J, et al. Impact of pathological tumour characteristics in patients with sarcomatoid renal cell carcinoma. BJU Int 2012; 109:1600-1606
Karlo CA, Di Paolo PL, Chaim J, et al. Radiogenomics of clear cell renal cell carcinoma: associations between CT imaging features and mutations. Radiology 2014; 270:464-471
Lebret T, Poulain JE, Molinie V, et al. Percutaneous core biopsy for renal masses: indications, accuracy and results. J Urol 2007; 178:1184–1188; discussion 1188
Leng S, Takahashi N, Gomez Cardona D, et al. Subjective and objective heterogeneity scores for differentiating small renal masses using contrast-enhanced CT. Abdom Radiol (NY) 2016;
Lubner MG, Stabo N, Abel EJ, Del Rio AM, Pickhardt PJ. CT Textural Analysis of Large Primary Renal Cell Carcinomas: Pretreatment Tumor Heterogeneity Correlates With Histologic Findings and Clinical Outcomes. AJR Am J Roentgenol 2016; 207:96-105
Millet I, Curros F, Serre I, Taourel P, Thuret R. Can renal biopsy accurately predict histological subtype and Fuhrman grade of renal cell carcinoma? J Urol 2012; 188:1690-1694
Pierorazio PM, Hyams ES, Tsai S, et al. Multiphasic enhancement patterns of small renal masses (</=4 cm) on preoperative computed tomography: utility for distinguishing subtypes of renal cell carcinoma, angiomyolipoma, and oncocytoma. Urology 2013; 81:1265-1271
Raman SP, Chen Y, Schroeder JL, Huang P, Fishman EK. CT Texture Analysis of Renal Masses: Pilot Study Using Random Forest Classification for Prediction of Pathology. Acad Radiol 2014; 21:1587-1596
Raman SP, Johnson PT, Allaf ME, Netto G, Fishman EK. Chromophobe renal cell carcinoma: multiphase MDCT enhancement patterns and morphologic features. AJR Am J Roentgenol 2013; 201:1268-1276
Schieda N, Thornhill RE, Al-Subhi M, et al. Diagnosis of Sarcomatoid Renal Cell Carcinoma With CT: Evaluation by Qualitative Imaging Features and Texture Analysis. AJR Am J Roentgenol 2015; 204:1013-1023
Scrima AT, Lubner MG, Abel EJ, et al. Texture analysis of small renal cell carcinomas at MDCT for predicting relevant histologic and protein biomarkers. Abdom Radiol (NY) 2018;
Takeuchi M, Kawai T, Suzuki T, et al. MRI for differentiation of renal cell carcinoma with sarcomatoid component from other renal tumor types. Abdom Imaging 2014;
Wang W, Ding J, Li Y, et al. Magnetic Resonance Imaging and Computed Tomography Characteristics of Renal Cell Carcinoma Associated with Xp11.2 Translocation/TFE3 Gene Fusion. PLoS One 2014; 9:e99990
Young JR, Margolis D, Sauk S, Pantuck AJ, Sayre J, Raman SS. Clear cell renal cell carcinoma: discrimination from other renal cell carcinoma subtypes and oncocytoma at multiphasic multidetector CT. Radiology 2013; 267:444-453
Abel EJ, Culp SH, Matin SF, et al. Percutaneous biopsy of primary tumor in metastatic renal cell carcinoma to predict high risk pathological features: comparison with nephrectomy assessment. J Urol 2010; 184:1877-1881
Bektas CT, Kocak B, Yardimci AH, et al. Clear Cell Renal Cell Carcinoma: Machine Learning-Based Quantitative Computed Tomography Texture Analysis for Prediction of Fuhrman Nuclear Grade. Eur Radiol 2019; 29:1153-1163
He X, Wei Y, Zhang H, et al. Grading of Clear Cell Renal Cell Carcinomas by Using Machine Learning Based on Artificial Neural Networks and Radiomic Signatures Extracted From Multidetector Computed Tomography Images. Acad Radiol 2020; 27:157-168
Sun X, Liu L, Xu K, et al. Prediction of ISUP grading of clear cell renal cell carcinoma using support vector machine model based on CT images. Medicine (Baltimore) 2019; 98:e15022
Shu J, Wen D, Xi Y, et al. Clear cell renal cell carcinoma: Machine learning-based computed tomography radiomics analysis for the prediction of WHO/ISUP grade. Eur J Radiol 2019; 121:108738
Haji-Momenian S, Lin Z, Patel B, et al. Texture analysis and machine learning algorithms accurately predict histologic grade in small (< 4 cm) clear cell renal cell carcinomas: a pilot study. Abdom Radiol (NY) 2020; 45:789-798
Lin F, Cui EM, Lei Y, Luo LP. CT-based machine learning model to predict the Fuhrman nuclear grade of clear cell renal cell carcinoma. Abdom Radiol (NY) 2019; 44:2528-2534
Kocak B, Durmaz ES, Ates E, Kaya OK, Kilickesmez O. Unenhanced CT Texture Analysis of Clear Cell Renal Cell Carcinomas: A Machine Learning-Based Study for Predicting Histopathologic Nuclear Grade. AJR Am J Roentgenol 2019:W1-W8
Dreyfuss LD LM, Nystrom J, Stabo N, Pickhart PJ, Abel EJ. Texture Analysis of Large Renal Cell Carcinoma: Comparing the Performance of Texture Analysis Platforms in Predicting Histologic Findings and Clinical Outcomes. AJR Am J Roentgenol 2021; in press
Lubner MG, Smith AD, Sandrasegaran K, Sahani DV, Pickhardt PJ. CT Texture Analysis: Definitions, Applications, Biologic Correlates, and Challenges. Radiographics 2017; 37:1483-1503
Pedregosa F, Varoquaux G, Gramfort A, et al. Scikit-learn: Machine Learning in Python. J Mach Learn Res 2011; 12:2825-2830
Abel EJ, Carrasco A, Culp SH, et al. Limitations of preoperative biopsy in patients with metastatic renal cell carcinoma: comparison to surgical pathology in 405 cases. BJU Int 2012; 110:1742-1746
Yan L, Chai N, Bao Y, Ge Y, Cheng Q. Enhanced Computed Tomography-Based Radiomics Signature Combined With Clinical Features in Evaluating Nuclear Grading of Renal Clear Cell Carcinoma. J Comput Assist Tomogr 2020; 44:730-736
Han D, Yu Y, Yu N, et al. Prediction models for clear cell renal cell carcinoma ISUP/WHO grade: comparison between CT radiomics and conventional contrast-enhanced CT. Br J Radiol 2020; 93:20200131
Meng X, Shu J, Xia Y, Yang R. A CT-Based Radiomics Approach for the Differential Diagnosis of Sarcomatoid and Clear Cell Renal Cell Carcinoma. Biomed Res Int 2020; 2020:7103647
Lu L, Ehmke RC, Schwartz LH, Zhao B. Assessing Agreement between Radiomic Features Computed for Multiple CT Imaging Settings. PLoS One 2016; 11:e0166550
Mackin D, Fave X, Zhang L, et al. Measuring Computed Tomography Scanner Variability of Radiomics Features. Invest Radiol 2015; 50:757-765
Shafiq-Ul-Hassan M, Zhang GG, Latifi K, et al. Intrinsic dependencies of CT radiomic features on voxel size and number of gray levels. Med Phys 2017;
Solomon J, Mileto A, Nelson RC, Roy Choudhury K, Samei E. Quantitative Features of Liver Lesions, Lung Nodules, and Renal Stones at Multi-Detector Row CT Examinations: Dependency on Radiation Dose and Reconstruction Algorithm. Radiology 2016; 279:185-194
van Timmeren JE, Leijenaar RTH, van Elmpt W, et al. Test-Retest Data for Radiomics Feature Stability Analysis: Generalizable or Study-Specific? Tomography 2016; 2:361-365
Meyer M, Ronald J, Vernuccio F, et al. Reproducibility of CT Radiomic Features within the Same Patient: Influence of Radiation Dose and CT Reconstruction Settings. Radiology 2019; 293:583-591
Zwanenburg A, Vallieres M, Abdalah MA, et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-based Phenotyping. Radiology 2020; 295:328-338
Funding
This work was supported by internal funding from the ML4MI program and Shapiro research program at University of Wisconsin.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosures
SG, DM, VJ, LD, MS, AD, EJA have no relevant disclosures. GL: Prior Grant funding Philips, Ethicon.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gurbani, S., Morgan, D., Jog, V. et al. Evaluation of radiomics and machine learning in identification of aggressive tumor features in renal cell carcinoma (RCC). Abdom Radiol 46, 4278–4288 (2021). https://doi.org/10.1007/s00261-021-03083-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-021-03083-y