Abstract
Purpose
To predict the Fuhrman grade of clear cell renal cell carcinoma (ccRCC) with a machine learning classifier based on single- or three-phase computed tomography (CT) images.
Materials and methods
Patients with pathologically proven ccRCC from February 1, 2009 to September 31, 2018 who were not treated were retrospectively collected for machine learning-based analysis. The texture features were extracted and ranked from precontrast phase (PCP), corticomedullary phase (CMP), nephrographic phase (NP) and three-phase CT images, and open-source gradient boosting from the decision tree library of CatBoost was used to establish a machine learning classifier to differentiate low- from high-grade ccRCC. The performances of machine learning classifiers based on features from single- and three-phase CT images were compared with each other.
Results
A total of 231 patients with 232 pathologically proven ccRCC lesions were retrospectively collected. 35, 36, 41, and 22 Features were extracted and ranked from PCP, CMP, NP, and three-phase CT images, respectively. The machine learning model based on three-phase CT images [area under the ROC curve (AUC) = 0.87] achieved the best diagnostic performance for differentiating low- from high-grade ccRCC, followed by single-phase NP (AUC = 0.84), CMP (AUC = 0.80), and PCP images (AUC = 0.82).
Conclusion
Machine learning classifiers can be promising noninvasive techniques to differentiate low- and high-Fuhrman nuclear grade ccRCC, and classifiers based on three-phase CT images are superior to those based on features from each single phase.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Renal cell carcinoma (RCC) is the tenth most common cancer and the most common kidney tumor in adults and accounts for 2–3% of all malignant tumors [1, 2]. Clear cell RCC (ccRCC) is the predominant subtype of RCC, and different grades of ccRCC have diverse biological behaviors and variable prognoses, which lead to different management strategies in clinical practice [3]. Minimally invasive techniques are feasible management considerations for low-grade ccRCC, but radical operations are more acceptable for high-grade ccRCC [4, 5]. Therefore, there is increasing interest in accurately differentiating low- and high-grade ccRCC in recent years.
Biopsy is the gold standard for evaluating the grade of ccRCC before surgery. However, patients who undergo biopsy are at risk of complications, such as hemorrhage and infection. Several noninvasive techniques have been used for the preoperative assessment of the grade of ccRCC [6,7,8,9]. Although tumor size, enhancement pattern on computed tomography (CT), attenuation on unenhanced CT, wash-in index on magnetic resonance imaging (MRI), apparent diffusion coefficient (ADC) value, and multiple kurtosis metrics based on functional MRI may be valuable in grading ccRCC before surgery, there is significant overlap in tumor size and imaging features between low- and high-grade ccRCC. In addition, the accuracy and reproducibility of these metrics still need to be improved.
Machine learning is a branch of artificial intelligence and is considered a promising technique to analyze medical images because it enables the identification of the best image feature combinations for making medical decisions [10,11,12]. A few previous studies have shown that CT textures or machine learning classifiers based on single- or multiphase CT images are valuable for distinguishing different subtypes of RCC [13,14,15]. However, to our knowledge, CT is widely used to stage RCC preoperatively, but no study has differentiated high-grade ccRCC from low-grade ccRCC using machine learning based on three-phase CT images with a large population.
Therefore, the aim of this study is to investigate an efficient machine learning classifier based on three-phased CT images to predict high-grade ccRCC.
Materials and methods
Patients
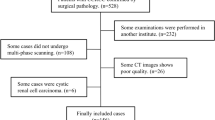
We retrospectively collected patients who underwent surgical resection for a renal mass from February 1, 2009 to September 31, 2018. The exclusion criteria were as follows: (1) tumor with serious hemorrhage or necrosis, (2) images with severe motion artifacts, and (3) lack of three-phase CT images. This retrospective study was approved by the research ethics board of our institution, which waived the requirement for informed consent. After retrieval of data from the institutional pathology database, the pathological diagnoses were reconfirmed by one pathologist with 10 years of genitourinary pathology experience. The Fuhrman grading system was adopted in the pathological analysis [16].
CT technique
All patients underwent an abdominal CT scan using a multidetector CT scanner (SOMATOM Force, Siemens Healthcare, Forchheim, Germany; SOMATOM Sensation 16, Siemens Healthcare, Forchheim, Germany; TOSHIBA Aquilion 64, Toshiba Medical Systems, Tokyo, Japan). All CT scans were performed with the same parameters and reconstruction used in daily clinical practice (slice thickness = 1.0 mm or 3.0 mm, matrix = 512 × 512, pixel size = 0.625 × 0.625 mm2). All subjects also underwent a three-phase CT scan including a precontrast phase (PCP), corticomedullary phase (CMP, 30-s delay after contrast injection), and nephrographic phase (NP, 90-s delay after contrast injection). Seventy to one hundred milliliters of contrast material (Iopamidol, Bracco, Italy; Iohexol, Yangtze River, China) was intravenously administered with a power injector at a rate of 3 ml/s.
Tumor segmentation
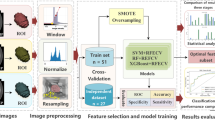
CT images were retrieved from the picture archiving and communication system. ITK-SNAP [17] (version 3.6.0, www.itksnap.org) was used for spatial matching and segmentation of tumors. A defined polygonal region of interest (ROI) was delineated on the center slice for low-grade ccRCC, but the slices for high-grade ccRCC were oversampled by selecting multiple slices at intervals of 15 mm (starting 10 mm from the apex and ending 10 mm from the bottom of the mass) (Fig. 1). To avoid a partial volume effect from the paratumoral renal parenchyma and perinephric fat, the ROI was carefully delineated and maintained at an approximate distance of 3 mm from the tumor margin. Two radiologists with more than 10 years of experience in abdominal imaging who were blinded to clinical and pathological information drew the ROIs without any divergence.
Texture analysis and machine learning
Texture analysis and machine learning were conducted using Python (version 3.6.5, www.python.org). The radiomic features extracted included the following (18): first-order features, shape features, gray-level cooccurrence matrix (GLCM) features, gray-level run-length matrix (GLRLM) features, gray-level size-zone matrix (GLSZM) features, gray-level dependence matrix (GLDM) features, and gray-tone difference matrix (NGTDM) features. All image calculations were performed for the PCP, CMP, and NP images separately. Features were named according to PyRadiomics [18] and the Imaging Biomarker Standardization Initiative (IBSI) [19], and a prefix (“pcp_,” “cmp_,” or “np_”) was added for the different scan phases.
CatBoost [20, 21], which is a state-of-the-art open-source gradient boosting decision tree library, was used to establish a machine learning model. Data related to patients were trained and tested using 5-fold cross-validation.
Statistical analysis
Performance results such as the true positive rate (TPR), specificity (SPC), positive predictive value (PPV), negative predictive value (NPV), accuracy (ACC), and area under the receiver operating characteristic (ROC) curve (AUC) were calculated for each phase and three-phase CT images. Additionally, feature importance scores and feature interaction scores were computed. ROC curve analysis was performed using Python (version 3.6.5, package scikit-learn).
Results
Demographics
Ultimately, 231 patients with 232 pathologically proven ccRCC lesions (low-grade ccRCC: 103 grade I lesions and 86 grade II lesions; high-grade ccRCC: 38 grade III lesions and 5 grade IV lesions) (one patient had two lesions in the left kidney) were included in the machine learning cohort. The mean ages of the low- and high-grade groups were 54.95 ± 11.94 years old and 53.07 ± 12.59 years old, respectively. There was no significant difference between these two groups in terms of patient characteristics.
Texture features ranking
In total, 35, 36, 41, and 22 features were extracted and ranked from PCP, CMP, NP, and three-phase CT images, respectively. The rankings of the texture features based on images of each phase and three-phase CT images are shown in Fig. 2.
Performance of the machine learning model
The TPR, SPC, PPV, NPV, ACC, and AUC for 5-fold cross-validation are shown in Table 1. The machine learning model based on three-phase CT images achieved the best diagnostic performance, followed by the single-phase NP, PCP, and CMP models. The ROC curves of the models based on images of each phase and three-phase CT images for differentiating low- from high-grade ccRCC are shown in Fig. 3.
Contribution of the combined features
The top five feature interaction rankings in the machine learning model based on three phases are shown in Table 2.
Discussion
In this study, we established machine learning models based on single- or three-phase CT images to differentiate between low- and high-grade ccRCC. Our results showed that this machine learning model could significantly stratify patients with diverse risk assessments of ccRCC according to the Fuhrman grading system.
Currently, visual imaging interpretation based on morphological findings is the routine paradigm for evaluating renal tumors. A prior study showed that high-grade ccRCC lesions are significantly larger and have more calcifications, necrosis, collecting system infiltration, and ill-defined tumor margins than low-grade ccRCC lesions [22]. However, the value of morphological evaluation is limited by various subjective interpretations and the inability to provide quantitative indicators. Although a few previous studies sought to determine whether quantitative imaging techniques could help to grade ccRCC and found that T1 values, ADC values and metrics of diffusion kurtosis (mean kurtosis, MK; radial kurtosis, Krad; and axial kurtosis, Kax) could be valuable [23,24,25], the performances of these quantitative indicators were varied, and their repeatability needs to be validated further.
Machine learning involves an algorithm and statistical model that can evaluate invisible tumor characteristics at the pixel level; machine learning algorithms have been used to grade neurogliomas and meningiomas with high accuracy [26,27,28]. Recently, a small study (n = 53) by Bektas CT et al. showed that a machine learning classifier could accurately differentiate low- from high-grade ccRCC [29]. Nevertheless, feature extraction based on only a single portal phase CT image significantly compromised the performance and reliability of the machine learning classifier because other studies showed that features based on CMP and NP images were also helpful in differentiating low- from high-grade ccRCC [30]. In addition, the portal phase is not the optimal enhanced phase for ccRCC evaluation, which may further diminish the reliability of the machine learning classifier in the study by Bektas CT et al. Moreover, the small population could have resulted in serious overfitting. Even though dedicated algorithms, such as a naïve Bayes algorithm, were used to remedy this problem, the performance of the classifier was not sufficiently objective.
Our study used a set of three single-phase CT images to develop machine learning classifiers in a large cohort (n = 232). We found that the classifier based on three-phase CT images was superior to those based on single-phase CT images, with an increase in the AUC from 0.80 to 0.87, although the improvement was not substantial. Feature importance ranking also showed that the model including all three-phase CT images exhibited the best performance. This model also had a higher SPC, PPV, NPV, and ACC for distinguishing low- and high-grade ccRCC than the models based on other single-phase CT images. However, compared to the SPC, the TPR was relatively low for models based on each single-phase and three-phase CT images (from 64 to 67%), which is similar to the findings of a previous study [6, 31] and needs to be further improved.
Feature importance scores are common indicators that demonstrate how important a specific feature is for model performance, and a higher value indicates that the model performs better when this feature is included. However, features contribute to a model not only solely by themselves but also by interacting with other useful features, and these interactions can be computed using the CatBoost decision tree library during the training process. Feature interaction scores indicate the contribution of a combination of features, and a higher interaction score of combined features represents a greater contribution to the model [32]. In our study, feature interaction ranking analysis showed that some features, such as cmp_original_gldm_GrayLevelNonUniformity, not only contributed to the model by themselves but also by interacting with other features in other phases. This internal relationship of a combination of features from different phase images has not been mentioned in previous studies. It should be noted that these features represent algorithms [19], and most are not obvious to the human eye. Therefore, it is very difficult to associate them with traditional radiological findings on images, which is a common drawback in radiomics research.
The prognosis of patients with ccRCC is strongly associated with the Fuhrman nuclear grade [33, 34], and tumor grading prior to surgery can guide surgical planning and treatment strategies. Percutaneous biopsy is a commonly used technique to preoperatively determine the tumor grade. However, this invasive method can lead to serious complications, such as hemorrhage or infection, and cannot be used in follow-up cases. In addition, sampling bias is an unavoidable problem associated with percutaneous biopsy because only one region of the tumor can be analyzed [35], which may lead to underestimation of the actual tumor grade due to the heterogeneity of ccRCC [36]. According to our current study, machine learning-based CT texture analysis showed acceptable performance for noninvasively predicting the Fuhrman nuclear grade of ccRCC and could reduce the bias to a minimum level. Therefore, our study could have significance in potentially sparing patients from invasive techniques, such as percutaneous biopsy. In addition, positive tumor regions detected by this technique might also be good candidates for target biopsy. However, the actual benefit to patients still needs to be verified by clinical studies involving both machine learning-based CT texture and biopsy.
The intergroup imbalance between low- and high-grade ccRCC is an inevitable issue for machine learning-based analysis due to the relatively lower incidence of high-grade ccRCC. Thus, the performance of a model based on imbalanced data will be overestimated and unreliable, and the degree of overestimation and unreliability mainly depend on the component proportions of low- and high-grade ccRCC but not on the selected texture features. This is a prevalent and critical problem in previous studies [29, 30]. The characteristic texture features of ccRCC should be retained in most slices of the tumor. Hence, every single slice can theoretically contribute to the machine learning classifier. In our study, every selected slice from high-grade ccRCC was considered as an individual sample to augment the high-grade samples and minimize the bias.
There are several limitations to our study. First, 3D radiomics features, which are features extracted across whole image slides of the tumor, were not used in this study, and in theory, these features can provide additional information. However, a prior study showed that 2D features actually exhibited better performance than 3D features [37]. Second, deep learning, which is a subset of the machine learning technique, has shown promising potential in medical imaging [38]. Unlike other machine learning methods, it is capable of discovering image features automatically without manually providing the features. Thus, this method may result in a more powerful model; however, we did not apply this technique in our study because, even though we had the largest cohort of patients among previous machine learning studies, a much larger sample size than we had was needed to obtain a stable deep learning model. Moreover, the features that deep learning detects are even more difficult to understand because they have no preexisting description or definition.
Conclusion
The results of our proof-of-concept study show that a CT-based machine learning model can be valuable for differentiating low- from high-grade ccRCC. However, further prospective studies are needed to verify its value. In addition to diagnostic accuracy, further machine learning studies could also potentially address other important clinical factors such as the survival time or genotype (BAP1 and PBRM1), which have been shown to be independent prognostic factors for tumor recurrence.
References
C. Global Burden of Disease Cancer, C. Fitzmaurice, D. Dicker, et al. The Global Burden of Cancer 2013. JAMA Oncol. 2015;1(4):505–527.
R. L. Siegel, K. D. Miller and A. Jemal. Cancer statistics, 2016. CA Cancer J Clin. 2016;66(1):7–30.
J. C. Cheville, C. M. Lohse, H. Zincke, et al. Comparisons of outcome and prognostic features among histologic subtypes of renal cell carcinoma. Am J Surg Pathol. 2003;27(5):612–624.
I. S. Gill, E. M. Remer, W. A. Hasan, et al. Renal cryoablation: outcome at 3 years. J Urol. 2005;173(6):1903–1907.
D. Jocham, A. Richter, L. Hoffmann, et al. Adjuvant autologous renal tumour cell vaccine and risk of tumour progression in patients with renal-cell carcinoma after radical nephrectomy: phase III, randomised controlled trial. Lancet. 2004;363(9409):594–599.
S. Y. Choi, D. J. Sung, K. S. Yang, et al. Small (< 4 cm) clear cell renal cell carcinoma: correlation between CT findings and histologic grade. Abdom Radiol (NY). 2016;41(6):1160–1169.
C. Chen, Q. Kang, B. Xu, et al. Differentiation of low- and high-grade clear cell renal cell carcinoma: Tumor size versus CT perfusion parameters. Clin Imaging. 2017;46:14–19.
F. Cornelis, E. Tricaud, A. S. Lasserre, et al. Multiparametric magnetic resonance imaging for the differentiation of low and high grade clear cell renal carcinoma. Eur Radiol. 2015;25(1):24–31.
G. Wu, Z. Zhao, Q. Yao, et al. The Study of Clear Cell Renal Cell Carcinoma with MR Diffusion Kurtosis Tensor Imaging and Its Histopathologic Correlation. Acad Radiol. 2018;25(4):430–438.
B. J. Erickson, P. Korfiatis, Z. Akkus, et al. Machine Learning for Medical Imaging. Radiographics. 2017;37(2):505–515.
L. Zhang, J. Tan, D. Han, et al. From machine learning to deep learning: progress in machine intelligence for rational drug discovery. Drug Discov Today. 2017;22(11):1680–1685.
R. C. Deo. Machine Learning in Medicine. Circulation. 2015;132(20):1920–1930.
B. Kocak, A. H. Yardimci, C. T. Bektas, et al. Textural differences between renal cell carcinoma subtypes: Machine learning-based quantitative computed tomography texture analysis with independent external validation. Eur J Radiol. 2018;107:149–157.
L. Yan, Z. Liu, G. Wang, et al. Angiomyolipoma with minimal fat: differentiation from clear cell renal cell carcinoma and papillary renal cell carcinoma by texture analysis on CT images. Acad Radiol. 2015;22(9):1115–1121.
H. Yu, J. Scalera, M. Khalid, et al. Texture analysis as a radiomic marker for differentiating renal tumors. Abdom Radiol (NY). 2017;42(10):2470–2478.
S. A. Fuhrman, L. C. Lasky and C. Limas. Prognostic significance of morphologic parameters in renal cell carcinoma. Am J Surg Pathol. 1982;6(7):655–663.
P. A. Yushkevich, J. Piven, H. C. Hazlett, et al. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage. 2006;31(3):1116–1128.
J. J. M. van Griethuysen, A. Fedorov, C. Parmar, et al. Computational Radiomics System to Decode the Radiographic Phenotype. Cancer Res. 2017;77(21):e104–e107.
A. Zwanenburg, S. Leger, M. Vallières, et al. Image biomarker standardisation initiative. arXiv preprint arXiv:1612.07003. 2016.
A. V. Dorogush, A. Gulin, G. Gusev, et al. Fighting biases with dynamic boosting. arXiv preprint arXiv:1706.09516. 2017.
A. V. Dorogush, V. Ershov and A. Gulin. CatBoost: gradient boosting with categorical features support. arXiv preprint arXiv:1810.11363. 2018.
H. Coy, J. R. Young, M. L. Douek, et al. Association of qualitative and quantitative imaging features on multiphasic multidetector CT with tumor grade in clear cell renal cell carcinoma. Abdom Radiol (NY). 2018.
L. C. Adams, B. Ralla, P. Jurmeister, et al. Native T1 Mapping as an In Vivo Biomarker for the Identification of Higher-Grade Renal Cell Carcinoma: Correlation With Histopathological Findings. Invest Radiol. 2018.
L. Shen, L. Zhou, X. Liu, et al. Comparison of biexponential and monoexponential DWI in evaluation of Fuhrman grading of clear cell renal cell carcinoma. Diagn Interv Radiol. 2017;23(2):100–105.
Y. D. Zhang, C. J. Wu, Q. Wang, et al. Comparison of Utility of Histogram Apparent Diffusion Coefficient and R2* for Differentiation of Low-Grade From High-Grade Clear Cell Renal Cell Carcinoma. AJR Am J Roentgenol. 2015;205(2):W193–201.
A. T. Hale, D. P. Stonko, L. Wang, et al. Machine learning analyses can differentiate meningioma grade by features on magnetic resonance imaging. Neurosurg Focus. 2018;45(5):E4.
Y. W. Park, J. Oh, S. C. You, et al. Radiomics and machine learning may accurately predict the grade and histological subtype in meningiomas using conventional and diffusion tensor imaging. Eur Radiol. 2018.
Y. Li, Z. Qian, K. Xu, et al. MRI features predict p53 status in lower-grade gliomas via a machine-learning approach. Neuroimage Clin. 2018;17:306–311.
C. T. Bektas, B. Kocak, A. H. Yardimci, et al. Clear Cell Renal Cell Carcinoma: Machine Learning-Based Quantitative Computed Tomography Texture Analysis for Prediction of Fuhrman Nuclear Grade. Eur Radiol. 2018.
J. Ding, Z. Xing, Z. Jiang, et al. CT-based radiomic model predicts high grade of clear cell renal cell carcinoma. Eur J Radiol. 2018;103:51–56.
S. Oh, D. J. Sung, K. S. Yang, et al. Correlation of CT imaging features and tumor size with Fuhrman grade of clear cell renal cell carcinoma. Acta Radiol. 2017;58(3):376–384.
J. H. Friedman and B. E. Popescu. Predictive Learning via Rule Ensembles. The Annals of Applied Statistics. 2008;2(3):916–954.
I. Frank, M. L. Blute, J. C. Cheville, et al. An outcome prediction model for patients with clear cell renal cell carcinoma treated with radical nephrectomy based on tumor stage, size, grade and necrosis: the SSIGN score. J Urol. 2002;168(6):2395–2400.
T. Klatte, J. J. Patard, M. de Martino, et al. Tumor size does not predict risk of metastatic disease or prognosis of small renal cell carcinomas. J Urol. 2008;179(5):1719–1726.
L. Marconi, S. Dabestani, T. B. Lam, et al. Systematic Review and Meta-analysis of Diagnostic Accuracy of Percutaneous Renal Tumour Biopsy. Eur Urol. 2016;69(4):660–673.
R. Guarch, J. M. Cortes, C. H. Lawrie, et al. Multi-site tumor sampling (MSTS) improves the performance of histological detection of intratumor heterogeneity in clear cell renal cell carcinoma (CCRCC). F1000Res. 2016;5:2020.
C. Shen, Z. Liu, M. Guan, et al. 2D and 3D CT Radiomics Features Prognostic Performance Comparison in Non-Small Cell Lung Cancer. Transl Oncol. 2017;10(6):886–894.
J. Ker, L. Wang, J. Rao, et al. Deep learning applications in medical image analysis. IEEE Access. 2018;6:9375–9389.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare there are no conflicts of interest regarding the publication of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lin, F., Cui, EM., Lei, Y. et al. CT-based machine learning model to predict the Fuhrman nuclear grade of clear cell renal cell carcinoma. Abdom Radiol 44, 2528–2534 (2019). https://doi.org/10.1007/s00261-019-01992-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-019-01992-7