Abstract
Purpose
Although functional brain imaging has been used for the early and objective assessment of cognitive dysfunction, there is a lack of generalized image-based biomarker which can evaluate individual’s cognitive dysfunction in various disorders. To this end, we developed a deep learning-based cognitive signature of FDG brain PET adaptable for Parkinson’s disease (PD) as well as Alzheimer’s disease (AD).
Methods
A deep learning model for discriminating AD from normal controls (NCs) was built by a training set consisting of 636 FDG PET obtained from Alzheimer’s Disease Neuroimaging Initiative database. The model was directly transferred to images of mild cognitive impairment (MCI) patients (n = 666) for identifying who would rapidly convert to AD and another independent cohort consisting of 62 PD patients to differentiate PD patients with dementia. The model accuracy was measured by area under curve (AUC) of receiver operating characteristic (ROC) analysis. The relationship between all images was visualized by two-dimensional projection of the deep learning-based features. The model was also designed to predict cognitive score of the subjects and validated in PD patients. Cognitive dysfunction-related regions were visualized by feature maps of the deep CNN model.
Results
AUC of ROC for differentiating AD from NC was 0.94 (95% CI 0.89–0.98). The transfer of the model could differentiate MCI patients who would convert to AD (AUC = 0.82) and PD with dementia (AUC = 0.81). The two-dimensional projection mapping visualized the degree of cognitive dysfunction compared with normal brains regardless of different disease cohorts. Predicted cognitive score, an output of the model, was highly correlated with the mini-mental status exam scores. Individual cognitive dysfunction-related regions included cingulate and high frontoparietal cortices, while they showed individual variability.
Conclusion
The deep learning-based cognitive function evaluation model could be successfully transferred to multiple disease domains. We suggest that this approach might be extended to an objective cognitive signature that provides quantitative biomarker for cognitive dysfunction across various neurodegenerative disorders.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Various neurodegenerative disorders are associated with cognitive dysfunction. Alzheimer’s disease (AD) is a representative disorder characterized by abnormalities in cognition and behaviors. As another common neurodegenerative disorder, Parkinson’s disease (PD) is primarily a movement disorder, while non-motor manifestations including cognitive dysfunction have been widely recognized as well [1]. At the early stage of PD, a single domain of cognition or behavior is affected before the onset of full-blown dementia and then multiple domains such as visuospatial function and memory performance are affected [2]. The early detection of cognitive dysfunction is essential for appropriate therapeutic intervention as cognitive functions largely contributes to functional outcomes and quality of life [3, 4]. So far, the objective methods to estimate patients’ cognitive functional status considering underlying progression mechanisms of neurodegenerative disorders have not been established. The quantitative biomarker of cognitive function at the individual patient level has been required for the objective assessment of disease status during progression and therapeutic intervention for the future clinical trials.
Functional imaging studies which aim at dopaminergic neurodegeneration and neuronal activities have been used for diagnosis and evaluating the functional status of PD [5, 6]. Among the various imaging methods, FDG brain PET has been used to differentiate parkinsonian disorders as well as to define metabolic abnormal patterns related to clinical symptoms and functional properties of PD patients [7, 8]. In particular, FDG PET was also applied to find abnormal metabolic topographic patterns in cognitive dysfunction in PD [9,10,11]. In spite of the identification of cognitive function-related brain metabolic changes in PD, the quantitative score based on FDG PET has not been widely used in the clinical setting. It was due to a lack of validation and generalization for various disease spectrum as well as the difficulty in standardized image processing.
To this end, we aimed at developing a cognitive function signature based on FDG PET which can be applied to PD as well as AD and mild cognitive impairment (MCI) patients. We employed a deep convolutional neural network (CNN) for translating FDG PET images into quantitative scores reflecting cognitive dysfunction. Though deep CNN has advantages in automatically capturing discriminative features from the data, the preparation of a large dataset for the model training becomes a bottleneck of deep learning application in medicine [12]. The model transfer for relatively small datasets can be a good alternative for this data issue. The features extracted from a relatively large dataset can be transferred to other domains with relatively small data to overcome data size [13, 14]. We applied this concept to brain imaging data. Firstly, a deep CNN model to discriminate AD from normal controls (NC) was trained using a relatively large FDG brain PET dataset, and then, this model was transferred to identification of MCI patients who would rapidly convert to AD. We further investigated whether the model can be applied to an independent dataset with different disease domain, a cohort of PD patients, for identifying cognitive dysfunction. A web-based application where the uses could easily obtain the cognitive signature produced by our analysis was also developed (https://fdgbrainpet.appspot.com/). Thus, we expected deep learning-based cognitive function evaluation could provide the cognitive signature across multiple disease groups.
Materials and methods
Subjects
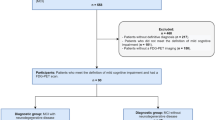
For a dataset of PD with FDG PET scans, we used FDG database of PD patients who were diagnosed as PD based on the UK PD Brain Bank criteria [15] by experienced movement disorder specialist (J.Y.L) and had been followed up at the movement disorders clinic in our institution. Among the patients, those who had history of stroke, space occupying lesions on the routine brain MRI; neurosurgical procedures including deep brain stimulation; features suggesting atypical parkinsonism or final diagnosis of atypical parkinsonism such as multiple system atrophy, progressive supranuclear palsy, and corticobasal degeneration; secondary parkinsonism related to hydrocephalus or drug; medical history of psychiatric disease which required psychiatric treatment, were excluded. Clinical information of age at PET scan, age at PD onset, Hoehn and Yahr (HY) stage, presence of PD dementia, and Korean version of mini-mental status examination scores was obtained. Diagnosis of dementia was made during the routine clinical follow-up according to the published criteria of probable dementia associated with PD suggested by the Movement Disorders Society (MDS) Task Force [2], and the diagnosis of dementia with Lewy bodies was considered as an exclusion criteria for this study following the 1-year rule. Finally, data from a total 62 PD patients (mean age ± SD, 71.0 ± 6.5; female:male, 32:30) were included in this study, and thirteen of the PD patients (20.9%) were classified as PD with dementia. The Institutional Review Board (IRB) of Seoul National University Boramae Medical Center approved this study and informed consents were waived.
In this study, as a training dataset for our model, FDG PET images acquired from subjects recruited in Alzheimer’s Disease Neuroimaging Initiative (ADNI) were used (http://adni.loni.usc.edu). The ADNI was launched in 2003 as a public-private partnership, led by Principal Investigator Michael W. Weiner, MD, VA Medical Center and University of California San Francisco. The primary goal of ADNI has been to test whether serial MRI, PET, biological markers, and clinical and neuropsychological assessment can be combined to measure the progression of MCI and early AD. ADNI has recruited more than 1600 participants from more than 50 sites across the USA and Canada. For up-to-date information, see http://www.adni-info.org. Written informed consent to cognitive testing and neuroimaging prior to participation was obtained, approved by the institutional review boards of all participating institutions. Baseline FDG PET scans of AD, MCI, and NC were used for this study (n = 1306; 243 AD, 666 MCI, and 393 NC). MCI subjects were divided into two subgroups: MCI-converters (MCI-C) and MCI-nonconverters (MCI-NC). MCI subjects who converted into AD within 3 years were defined as MCI-C (167 MCI-C and 274 MCI-NC). MCI-NC included MCI subjects whose diagnosis was still MCI at 3-year follow-up exam. Demographic and clinical characteristics of patients are summarized in Table 1.
FDG PET acquisition and processing
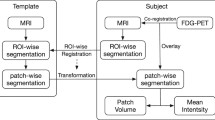
FDG PET scans were downloaded from ADNI database (http://adni.loni.usc.edu). PET images of ADNI are available for the database at various levels of preprocessing [16]. The images of dynamic frames (6 frames from 30 to 60 min after the injection) were averaged. The images were aligned and resampled to have same voxel size. The intensity was normalized by global brain FDG uptake. PET images were acquired in the 57 sites participating in ADNI; scanner-specific smoothing was additionally applied [16]. Images were downloaded at this preprocessing level. Downloaded scans were then normalized to Montreal Neurological Institute (MNI) space using statistical parametric mapping (SPM8, University College of London, London, UK).
FDG PET scans of PD patients were acquired in a single institute. FDG PET scans were obtained for the subjects without stopping anti-parkinsonian medication. Images were obtained by GEMINI PET/CT machine (Philips Healthcare, Andover, MA, USA) 40 min after intravenous injection of 4.8 MBq/kg of FDG. FDG was injected after fasting for at least 6 h. Emission scans were acquired for 10 min. Images were reconstructed three-dimensional row-action maximum-likelihood algorithm (3D RAMLA) with CT-based attenuation correction. The final image matrix size was 128 × 128 × 90 and the voxel size was 2 × 2 × 2 mm3. The images of PD patients were also spatially normalized to MNI space using SPM8. The matrix size of spatially normalized PET images was 79 × 95 × 68 and the voxel size was 2 × 2 × 2 mm3.
Study design
Firstly, we built a CNN-based model that differentiates AD from NC using FDG brain PET. The model was directly transferred to differentiating MCI-C from MCI-NC. Furthermore, as another independent cohort of different disease domain, the model was tested for differentiating PD patients with dementia from those without dementia. As hidden layers of the CNN represented automatically extracted features of PET images, the features were visualized by two-dimensional projection. To predict the cognitive score, MMSE, fine tuning of the model was additionally performed. This trained model was applied to another disease group, PD patients. We tested whether the output of the model could be transferred to predicting cognitive scores of PD patients. CNN-based features of PD patients also projected to two-dimensional space for the visualization.
Deep CNN for differentiating AD from normal
FDG PET images of AD and NC subjects were used for the training of the CNN model. Among 636 PET images (243 AD 393 NC), randomly selected 64 images were used for the internal validation. The architecture of the model is represented in Fig. 1. Briefly, four 3d convolutional layers with 5 × 5 × 5 convolutional filters hierarchically extracted features of PET images. For each convolution layer, rectified linear unit (ReLU) activation was used as an activation function. These four convolutional features produced 128 feature volumes, which have 10 × 12 × 9 matrix size. A global average pooling layer summarized the feature maps into a vector for each feature map. The 128 features were finally connected to an output which represented the expectation that the input PET volume was acquired from AD. The network minimized the cross-entropy loss between the predicted diagnosis and the real diagnosis. Training was conducted by the stochastic gradient descent algorithm with Adam optimizer [17]. Batch size was 4 and iterative training was stopped by monitoring loss function and accuracy of the internal validation set.
Architecture of FDG PET-based cognitive dysfunction evaluation model. The model was based on a three-dimensional deep convolutional neural network. Four 3d convolutional layers with 5 × 5 × 5 convolutional filters followed by nonlinear activation function were hierarchically extracted image features of input images. It produced 128 feature volumes, which have 10 × 12 × 9 matrix size. A global average pooling layer was followed to summarize the feature maps into vectors. These automatically extracted features were finally connected to an output which represented likelihood of Alzheimer’s disease (AD)
Deep CNN model transfer and evaluation
The accuracy of the model was evaluated by receiver operating characteristic (ROC) curves. A ROC curve was drawn for differentiating AD from NC in the validation set. The model was directly transferred to testing accuracy for differentiating MCI-C from MCI-NC. In addition, the trained model was also transferred to the task differentiating PD patients with dementia from those without dementia. The accuracy was measured by area under curve (AUC) values of ROC curves.
Visualization of deep CNN-based features with two-dimensional projection
All PET volumes of ADNI dataset (N = 1306) were inputted into the trained CNN model. The model produced 128 features for each subject based on the CNN model. For visualizing the relationship of data according to the similarities, we employed a parametric t-distributed stochastic neighbor embedding (t-SNE) model [18]. One hundred twenty-eight features of each subject’s PET images were visualized by the parametric t-SNE by estimating the similarity. It is an unsupervised parametric dimensionality reduction technique that attempts similar samples are modeled by nearby points and dissimilar samples are modeled by distant points. More specifically, the pairwise distance of two datapoints is defined by the probabilities by centering an isotropic Gaussian over a datapoint i and computing the density of point j. It is calculated by,
where x denotes high-dimensional data, for here, 128-dimensional features. Because of the asymmetricity, the pairwise data affinity was recalculated by \( {\mathrm{p}}_{ij}=\frac{\left({p}_{j\left|i\right.}+{p}_{i\left|j\right.}\right)}{2n} \), where n is number of datapoints. Then, this datapoints move to latent feature space, where the parametric t-SNE transformed the datapoints, by minimizing a cost function defined by Kullback-Leibler divergences over all datapoints, \( \mathrm{KL}\left(\mathrm{P}\Big\Vert \mathrm{Q}\right)=\sum \limits_{i\ne j}{p}_{ij}\log \frac{p_{ij}}{q_{ij}} \). qij is the pairwise distance in the feature latent space, which has 2-dimensions in this study.
One hundred twenty-eight features of each subject’s PET images were used for input of the t-SNE model. Subjects of AD/NC, including training and internal validation data, were visualized. MCI subjects also visualized by the t-SNE model whether MCI-C and MCI-NC were clustered. The PD data independent from the diagnosis of AD that consist of 62 patients were projected to two-dimensional axis points using the t-SNE model. Their MMSE scores were also visualized with a colormap.
Cognitive function prediction model
To predict the cognitive score, the fine tuning of the CNN model was performed. One hundred twenty-eight features were extracted from four convolutional layers of the model. The last layer connected to output was only modified. The aim of the model was the MMSE score instead of AD/NC classification. As the minimum and maximum MMSE of the ADNI data in our study were respectively 18 and 30, MMSE scores were rescaled by (MMSEsubject − MMSEmin)/(MMSEmax − MMSEmin). So, the range of output was rescaled from 0 to 1 for the training. The CNN model was tuned to minimize mean square errors between predicted and real cognitive scores. For this tuning, all ADNI data (N = 1306) were used for the training process. Thus, MMSE scores of AD, NC, and MCI subjects were used. This fine-tuned CNN model was applied to predict the cognitive function of PD subjects. The predicted cognitive score was obtained by inputting PET images of PD subjects. The output was then rescaled to the original MMSE scale.
Individual cognitive dysfunction-related region mapping
To demonstrate that brain regions where the CNN model evaluated for AD or PD with decreased cognitive function, we employed the class activation map (CAM) visualization method [19]. This CAM-based visualization method has been widely used for indirectly explaining how deep CNN models predict the outputs [20, 21]. The idea of CAM is to generate the output map instead of a value which represents the probability of AD. Thus, simply removing the last pooling layer produces feature maps instead of feature vectors to visualize the area most indicative of the AD at individual level. More specifically, 128 feature maps with 10 × 12 × 9 matrix size were produced when the global average pooling layer was not applied. A class activation map for each subject was obtained by multiplying weighs of the last layer and 128 features. The map represented which areas were important for determining that the subject was classified into AD. We produced the class activation maps for PD subjects who showed dementia symptoms as well as AD subjects of ADNI dataset.
Results
Accuracy of the cognitive dysfunction evaluation model
A deep CNN-based model was built by the training dataset consisting of AD patients and NC (243 AD 393 NC). The model accuracy for differentiating AD from NC was evaluated by AUC of the ROC curve, which showed 0.94 (95% CI 0.89–0.98) for independent validation set (Fig. 2a). The model was directly applied to the task of differentiating MCI-C from MCI-NC. The AUC for this task was 0.82 (95% CI 0.78–0.86) for 441 baseline MCI subjects (Fig. 2b). The model was also applied to differentiating PD patients with dementia from those without dementia. The AUC was 0.81 (95% CI 0.68–0.94) (Fig. 2c).
Receiver operating characteristic curves for differentiating patients with cognitive dysfunction. The accuracy of the model was assessed by area under curves (AUC) of receiver operating characteristic curves. a The AUC for differentiating AD from normal controls (NC) was 0.94 (95% CI 0.89–0.98) for validation set. b The AUC of the model for differentiating mild cognitive impairment (MCI) patients who would convert to full-blown dementia (MCI-C) was 0.82 (95% CI 0.78–0.86). c The model was directly also applied to differentiating Parkinson’s disease (PD) patients with dementia from those without dementia and AUC was 0.81 (95% CI 0.68–0.94)
Brain metabolic features projection to two-dimensional axes
The FDG PET image features based on the deep CNN model were extracted for all subjects of our study. To visualize subjects according to the similarity of brain metabolism patterns, those were projected to 2D axes by using a parametric t-SNE embedding (Fig. 3a). Each point represents individual PET data. Subjects with similar brain metabolic features were exhibited by close points. Of note, two axes represent the two-dimensional output of the parametric t-SNE. The axes did not represent a linear quantitative value as the model visualizes the similarity between PET data in terms of nonlinear features of image patterns. Thus, the results visualized the similarities between PET data preserving local relationship, such as clusters according to diagnostic groups, however, did not reflect the distance of all data. As a result, AD and NC subjects were clustered, respectively (Fig. 3b). The location of MCI-C subjects was similar with AD and that of MCI-NC subjects was similar with NC (Fig. 3c). MMSE scores of AD, MCI, and NC subjects were additionally represented (Fig. 3d). It showed that patients with cognitive dysfunction tend to be distributed in the left lower portion of the 2D axes.
Visualization of brain metabolic features by two-dimensional projection mapping. a Deep learning-based features were extracted for all subjects and they were visualized according to the similarity of brain metabolism patterns. The individual features were projected to 2D axes by using parametric t-distributed stochastic neighbor embedding (t-SNE). b For training data, AD and NC subjects were distinctively clustered. c PET images of MCI patients were embedded to 2D axes. MCI-C and MCI-nonconverters (MCI-NC) were distinctively located as AD and NC subjects. d MMSE scores of subjects of ADNI cohort were simultaneously plotted. Subjects with lower MMSE score were relatively located on left lower portion of the 2D axes. e PD patients were projected to the 2D projection map and overlaid with AD and NC subjects. PD patients with low MMSE score were located on the left lower portion
The FDG PET images of PD patients were also projected to 2D axes (Fig. 3e). The color scale represents MMSE score. Note that the points of PD patients were overlaid with AD and NC subjects. As AD, MCI, and NC subjects, PD patients with cognitive dysfunction also tend to be distributed in left lower portion, which suggested that FDG PET images of PD patients with cognitive dysfunction share similar patterns with those of AD and MCI-C subjects.
Predicting cognitive score
Fine tuning of the model was performed to predict individual cognitive score using FDG PET images of all AD, MCI, and NC subjects (Fig. 4a). The output of the model, predicted cognitive score, was plotted with MMSE score (Fig. 4b). The model was applied to FDG PET images of PD patients. The predicted cognitive scores of PD patients were significantly correlated with MMSE scores (r = 0.55, p < 1 × 10−4, Fig. 4c). Notably, the predicted MMSE scores of some subjects were larger than 30, which was caused by the training with normalized value.
Fine tuning of the model for predicting cognitive score. a The last layer of the model was tuned to predict individual MMSE score using FDG PET images. b For ADNI cohort, the output of fine-tuned model was plotted with MMSE score. c This model was applied to the independent cohort, PD patients. The output of the model was significantly correlated with MMSE score (r = 0.55, p < 1 × 10−4)
Cognitive dysfunction-related region mapping
We visualized the regions related to cognitive dysfunction by the deep CNN model. Brain metabolic features related to the cognitive dysfunction were localized on individual FDG PET images. The cognitive dysfunction-related regions of individuals were partly different to each other. As shown in Fig. 5a, cognitive dysfunction-related regions were obtained for three PD patients with dementia. The regions mainly included cingulate and high frontal/parietal cortices. However, cognitive dysfunction-related regions of the first patient were posterior cingulate area while those of the second patient were the superior frontal and parietal cortices. Those of the third patient were the cingulate and superior frontoparietal cortices. Cognitive dysfunction-related regions were averaged across AD patients and PD patients with dementia, respectively (Fig. 5b). The averaged regions represented cingulate and superior frontal/parietal cortices.
Cognitive dysfunction-related region mapping. The feature map of deep CNN could visualize the cognitive dysfunction-related regions for each individual. This mapping represented what regions the model had seen to determine that the subject had cognitive dysfunction. a The cognitive dysfunction-related regions differed among individuals. b These regions were averaged across AD patients and PD patients with dementia. They included cingulate and superior frontal/parietal cortices
Discussion
In this study, we developed an objective cognitive function signature using FDG PET which can be applied to PD as well as AD. The deep CNN model trained by AD and NC could discriminate PD patients with dementia. In addition, each individual’s cognitive status was intuitively visualized according to the similarity of FDG PET patterns with other subjects by mapping into two-dimensional space. We also found each individual represented similar but different cognitive dysfunction-related regions.
One of the major contribution of this study was to develop a cognitive signature that covered multiple neurodegenerative disorders. Cognitive decline is a common symptom of neurodegeneration disorder. Among PD patients, 80% would eventually develop cognitive dysfunction in the late stage of disease progression [22]. The prediction of future cognitive decline in MCI subjects is crucial for managing dementia. So far, imaging biomarkers for cognitive function evaluation have been developed for each type of disorders. These studies have aimed at identifying a common imaging pattern associated with cognitive decline based on disease group analysis. Specifically, hypometabolic brain regions related to MCI-C [23] and those related to PD with dementia [24] were independently investigated. Although these analyses have revealed neural correlates of cognitive impairment, they could hardly be used for individual assessment of cognitive function in multiple types of diseases. Our model showed a single model-based stratification of cognitive function in PD as well as MCI patients. The model learned FDG PET patterns of cognitive dysfunction in AD subjects compared with NC, and then, they could identify MCI-C and PD patients with dementia without any modification. It suggested a feasibility of the usage of deep CNN-based imaging biomarker for cognitive function evaluation in multiple types of neurodegenerative disorders.
This approach could explicitly visualize the status based on an individual’s brain metabolism. It is important to identify individual’s status based on brain metabolism compared with other subjects for therapeutic management and further clinical decisions. The parametric t-SNE embedding intuitively visualized the cognitive status of each individual compared with other patients. Moreover, our approach could provide individual cognitive biomarker by capturing individual cognitive dysfunction-related regions. This individualized approach was resulted from the hierarchical feature discovery of the deep CNN model. As presented in Fig. 5, the cognitive dysfunction-related regions of individual subjects were quite different. It suggested that the model could capture unique FDG PET patterns of each subject to estimate cognitive function. This individual pattern recognition is one of the different factor compared with previous methods of image-based cognitive function evaluation. Previous study used a group-wise regional distribution pattern of FDG uptake and individual cognitive scores were obtained [10]. On the other hand, our results showed the individual variability in the location of cognitive dysfunction-related regions. It suggests that brain regions affecting cognitive dysfunction could be varied in PD patients; thus, individualized evaluation might be needed for subtyping and further clinical trials. The cognitive dysfunction-related regions were identified by CAM, which has been widely used for explaining the decision of CNN models [20, 21]. However, the interpretation of the decision of CNN model should be cautious when we try to use it in the clinical setting. Since this approach does not mean decreased cortical metabolism, but represent the location of patterns, the model could find patterns of relative FDG distribution in non-cortical area compared with near gray matter. It is a limitation of the CNN model to use in the clinical setting if the patterns located on atypical brain areas cannot be explained by our knowledge of the FDG PET of cognitive dysfunction. In our approach, the visualization of cognitive dysfunction-related regions could indirectly provide whether the deep CNN model captured appropriate brain regions for the decision; thus, we may refer these patterns in the comprehensive interpretation of the results. According to our representative figure (Fig. 5a), the cognitive dysfunction-related regions were varied according to cases; they included frontal and parietal cortices as well as the cingulate cortex. As these regions commonly show abnormal metabolism on FFG PET of AD, the output of deep learning model could be in line with the clinical situation and visual interpretation. However, if the regions include atypical brain areas and the output of the model is different from visual interpretations, we should be cautious to refer to the output of the CNN model. Because of this limitation in the explainable model using CAM, further visualization and explainable methods will be needed as future work. Furthermore, clinical manifestation and symptomatic features correlated with various cognitive dysfunction-related regions can be deeply investigated, which will enrich the application of deep CNN to classify subtypes of cognitive dysfunction.
One of the contributions of this study was to show clinical feasibility of deep CNN-based biomarker by validating the model in an independent cohort that consists of PD patients. A critical issue in application of deep learning to medical imaging is the limited number of data [12, 25]. One of the strategies to overcome this challenge was to transfer a model trained on a huge number of other data. Several studies have used deep learning models trained by natural image database, ImageNet, to various types of medical images to overcome the issue of data size [26,27,28,29]. However, as the brain PET is three-dimensional and has unique image textures distinct from natural images, it was difficult to transfer model trained by natural images. Therefore, we trained a 3D CNN model trained by relatively large data consisting of AD and NC and then validated in PD patents. Some previous studies showed transfer of the model trained by AD and NC to MCI patients, while this domain transfer was based on the knowledge that metabolic topographic patterns of MCI-C shares AD patients [30,31,32]. Considering the model could measure cognitive function in the independent PD cohort, we suggest this transfer strategy could facilitate the application of deep learning in relatively small cohorts. Furthermore, it enables clinical trials to prove clinical utility of deep learning-based biomarker.
The neural substrates of cognitive impairment in neurodegenerative disorders are varied in accordance with types of diseases and still remain poorly understood. For an objective assessment of cognitive impairment, there is a need to establish imaging biomarkers which could aid in the identification of patients at risk of cognitive decline, for early diagnosis and for therapeutic trials. Our suggested deep CNN-based cognitive signature could accurately and objectively assess cognitive dysfunction in PD as well as MCI patients. The model also intuitively visualized individual status compared with population by 2D projection map and provided cognitive dysfunction-related regions at the individual level. As a proof-of-concept study, we expect that the deep learning-based biomarker might be extended to a generalized cognitive function assessment model for various neurodegenerative disorders. The cognitive signature based on FDG PET can be estimated by a web-based application (https://fdgbrainpet.appspot.com/) (Supplementary Fig. 1). It will facilitate clinical trials of therapeutic candidates as it may play a role in disease status monitoring and surrogates for therapeutic targets.
Data availability
The imaging data can be found in ADNI database (http://adni.loni.usc.edu/). The application of our method is developed for a web-based resource (https://fdgbrainpet.appspot.com/).
References
Chaudhuri KR, Healy DG, Schapira AH. Non-motor symptoms of Parkinson’s disease: diagnosis and management. Lancet Neurol. 2006;5(3):235–45.
Emre M, et al. Clinical diagnostic criteria for dementia associated with Parkinson’s disease. Mov Disord. 2007;22(12):1689–707.
McKinlay A, et al. A profile of neuropsychiatric problems and their relationship to quality of life for Parkinson’s disease patients without dementia. Parkinsonism Relat Disord. 2008;14(1):37–42.
Schrag A, Jahanshahi M, Quinn N. What contributes to quality of life in patients with Parkinson’s disease? J Neurol Neurosurg Psychiatry. 2000;69(3):308–12.
Ravina B, et al. The role of radiotracer imaging in Parkinson disease. Neurology. 2005;64(2):208–15.
Brooks DJ. Imaging approaches to Parkinson disease. J Nucl Med. 2010;51(4):596–609.
Eidelberg D. Metabolic brain networks in neurodegenerative disorders: a functional imaging approach. Trends Neurosci. 2009;32(10):548–57.
Eckert T, et al. FDG PET in the differential diagnosis of parkinsonian disorders. Neuroimage. 2005;26(3):912–21.
Huang C, et al. Metabolic abnormalities associated with mild cognitive impairment in Parkinson disease. Neurology. 2008;70(16 Part 2):1470–7.
Huang C, et al. Metabolic brain networks associated with cognitive function in Parkinson’s disease. Neuroimage. 2007;34(2):714–23.
Svenningsson P, Westman E, Ballard C, Aarsland D. Cognitive impairment in patients with Parkinson’s disease: diagnosis, biomarkers, and treatment. Lancet Neurol. 2012;11(8):697–707.
Choi H. Deep learning in nuclear medicine and molecular imaging: current perspectives and future directions. Nucl Med Mol Imaging. 2017:1–10.
Oquab M, Bottou L, Laptev I, & Sivic J. Learning and transferring mid-level image representations using convolutional neural networks. Proceedings of the IEEE conference on computer vision and pattern recognition. 2014; pp 1717–1724.
Yosinski J, Clune J, Bengio Y, & Lipson H. How transferable are features in deep neural networks? Advances in neural information processing systems. 2014 pp 3320–3328.
Gibb W, Lees A. The relevance of the Lewy body to the pathogenesis of idiopathic Parkinson’s disease. J Neurol Neurosurg Psychiatry. 1988;51(6):745–52.
Jagust WJ, et al. The Alzheimer’s disease neuroimaging initiative 2 PET Core: 2015. Alzheimer’s & dementia: the journal of the Alzheimer’s Association. 2015;11(7):757–71.
Kingma D & Ba J. Adam: a method for stochastic optimization. arXiv preprint arXiv. 2014;1412.6980.
van der Maaten L. Learning a parametric embedding by preserving local structure. RBM. 2009;500(500):26.
Zhou B, Khosla A, Lapedriza A, Oliva A, & Torralba A. Learning deep features for discriminative localization. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 2016; pp 2921–2929.
Zintgraf LM, Cohen TS, Adel T, & Welling M. Visualizing deep neural network decisions: prediction difference analysis. arXiv preprint arXiv. 2017;1702.04595.
Rajpurkar P, et al. Chexnet: radiologist-level pneumonia detection on chest x-rays with deep learning. arXiv preprint arXiv. 2017;1711.05225.
Aarsland D, Andersen K, Larsen JP, Lolk A. Prevalence and characteristics of dementia in Parkinson disease: an 8-year prospective study. Arch Neurol. 2003;60(3):387–92.
Schroeter ML, Stein T, Maslowski N, Neumann J. Neural correlates of Alzheimer’s disease and mild cognitive impairment: a systematic and quantitative meta-analysis involving 1351 patients. Neuroimage. 2009;47(4):1196–206.
Bohnen NI, et al. Cerebral glucose metabolic features of Parkinson disease and incident dementia: longitudinal study. J Nucl Med. 2011;52(6):848–55.
Greenspan H, van Ginneken B, Summers RM. Guest editorial deep learning in medical imaging: overview and future promise of an exciting new technique. IEEE Trans Med Imaging. 2016;35(5):1153–9.
Tajbakhsh N, et al. Convolutional neural networks for medical image analysis: full training or fine tuning? IEEE Trans Med Imaging. 2016;35(5):1299–312.
Gulshan V, et al. Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. Jama. 2016;316(22):2402–10.
Esteva A, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115.
Coudray N, et al. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat Med. 2018:1.
Cheng B, Zhang D, & Shen D. Domain transfer learning for MCI conversion prediction. Medical Image Computing and Computer-Assisted Intervention–MICCAI. 2012;2012:82–90
Choi H, Jin KH, Initiative AsDN. Predicting cognitive decline with deep learning of brain metabolism and amyloid imaging. Behav Brain Res. 2018;344:103–9.
Choi H, et al. Deep learning only by normal brain PET identify unheralded brain anomalies. EBioMedicine. 2019. 2019;43:447-53.
Funding
This work was supported by a clinical research grant-in-aid from the Seoul Metropolitan Government Seoul National University Boramae Medical Center (02-2017-5) and National Research Foundation grant funded by the Ministry of Education, Science, and Technology in Korea (NRF-2016R1D1A1B03936159, NRF-2018R1C1B3008971, and NRF-2018R1A5A2025964).
Data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.;Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd. and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Disease Cooperative Study at the University of California, San Diego. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Consortia
Contributions
Y.K.K., C.H. and J.Y.L. designed the study. C.H. developed the deep learning model and analyzed the data. Y.K.K. and E.J.Y. collected the data. J.Y.K. performed clinical assessment and supported the analysis. D.S.L supervised the study. All authors interpreted data results, drafted, and edited manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent to clinical testing and neuroimaging prior to participation of the ADNI cohort was obtained, approved by the institutional review boards (IRB) of all participating institutions. The Institutional Review Board of Seoul National University Boramae Hospital approved this study and informed consents were waived for a retrospective cohort of Parkinson’s disease patients.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection on Advanced Image Analyses (Radiomics and Artificial Intelligence)
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
Electronic supplementary material
ESM 1
(DOCX 432 kb)
Rights and permissions
About this article
Cite this article
Choi, H., Kim, Y.K., Yoon, E.J. et al. Cognitive signature of brain FDG PET based on deep learning: domain transfer from Alzheimer’s disease to Parkinson’s disease. Eur J Nucl Med Mol Imaging 47, 403–412 (2020). https://doi.org/10.1007/s00259-019-04538-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-019-04538-7