Abstract
In recent years, extracellular vesicles have gained more attention. However, studies on membrane vesicles in Gram-positive bacteria were carried out relatively late because of the thick bacterial wall and the low production of membrane vesicles. Thanks to the research in recent years, the cognition of the composition and function of the membrane vesicles of Gram-positive bacteria has made significant progress. Membrane vesicles are spherical in shape comprising bilayer membranous structures with a diameter of 20–400 nm. Components of membrane vesicles are diverse, including proteins, nucleic acids, lipids, and metabolites. It also has been reported that membrane vesicles are involved in various pathophysiological processes and serve as communication tools in pathophysiological activities between the bacteria and the host. This review provided the current understanding of components and functions of membrane vesicles in Gram-positive bacteria. The findings might facilitate further research in the emerging field of membrane vesicles in Gram-positive bacteria.
Key points
• Membrane vesicles in Gram-positive bacteria contain proteins, nucleic acids, lipids, and metabolites, suggesting their biological significance.
• Membrane vesicles in Gram-positive bacteria are thought to be involved in stress response, biofilm formation, immune regulation, and so on.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, extracellular vesicles (EVs) have gained more attention (El Andaloussi et al. 2013; van Niel et al. 2018). EVs were first discovered in eukaryotes as vesicles derived from the plasma membrane carrying proteins, nucleic acids, and lipids. They are mainly divided into three categories according to their size: exosomes, microvesicles, and apoptotic bodies (Raposo and Stoorvogel 2013). Then, EVs were also found in prokaryotes such as bacteria (Brown et al. 2015). Bacteria are divided into Gram-negative and Gram-positive bacteria according to their membrane structure. The Gram-negative bacteria have two membrane layers separated by the periplasm. The membrane in Gram-positive bacteria is very different from that in Gram-negative bacteria with only one membrane and one thicker layer of peptidoglycan. The EVs in Gram-negative bacteria are called outer membrane vesicles (OMVs), while EVs in Gram-positive bacteria are called membrane vesicles (MVs) (Avila-Calderón et al. 2015). Both OMVs and MVs are subclasses of microbial EVs. Research on OMVs has been extensive, with many reviews highlighting their functions (Jan 2017; Tan et al. 2018). Many people have questioned the production of vesicles in Gram-positive bacteria in the past because it is relatively low. However, in the last decade, a great deal of research has been conducted on the role of Gram-positive MVs.
MVs have been observed in all pathogenic and nonpathogenic Gram-positive bacteria under different growth conditions and different natural environments. The secretion of MVs is a universal and widespread phenomenon (Toyofuku et al. 2019). With the increase in the number of studies on Gram-positive bacteria, such as Staphylococcus aureus, Bacillus anthracis, and Streptococcus mutans, it was found that MVs in Gram-positive bacteria had a 20- to 400-nm bilayer spherical structure (Jeon et al. 2016; Liao et al. 2014; Rivera et al. 2010).
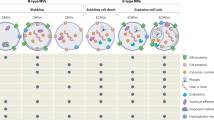
In this review, the current knowledge on content characterization and functions of MVs in Gram-positive bacteria was discussed (Fig. 1).
Composition of MVs
The mechanism of MV biogenesis in Gram-positive bacteria has been reviewed in some studies (Briaud and Carroll 2020; Brown et al. 2015). A few genes found in certain bacteria have been reported to regulate MV biogenesis. For example, the transcription factor σB regulates MV formation in Listeria monocytogenes (Lee et al. 2013). A two-component regulator CovRS impacts MV production in Group A Streptococcus (Resch et al. 2016). However, a unified gene that can be applied to all Gram-positive bacteria is still lacking. MVs in Gram-positive bacteria are produced when a section of the cytoplasmic membrane protrudes and buds off selectively encapsulating various components, and then passes through the cell wall. Therefore, the whole process of MV biogenesis is affected by many pathways. Turgor pressure, protease enzymes, and protein channels are proposed hypotheses that explain how MVs traverse thick cell walls (Brown et al. 2015). In summary, all these studies showed that MV production is regulated by a complex gene network. Whether Gram-positive bacteria share a conserved general mechanism for MV biogenesis is currently unknown.
For MV composition analysis, sample concentration is critical. The separation of purified MVs is the key to mass spectrometry. MVs can be isolated from bacterial cultures or biofilms. Generally speaking, MVs can be obtained by ultracentrifugation after the supernatant culture is filtered through a filter with a pore diameter of 0.22–0.45 μm (Chutkan et al. 2013). It is worth noting that a typical filter with 0.22-μm diameter might lower the yields up to approximately 50% when the actual size of MVs is more than 200 nm under a certain environment (Gorringe et al. 2005). However, ultracentrifugation is not sufficient to purify MVs from protein aggregates or membrane fragments (Choi et al. 2013). Larger macromolecules such as fimbriae and flagella should be included in the MV pellet during the ultracentrifugation method. In addition, sucralose concentration ultracentrifugation can also be used to extract bacterial MVs, but sucrose does not fully preserve the size of MVs in the gradient. In short, the aforementioned methods cannot remove pollutants other than MVs. To characterize a pure population, density gradient ultracentrifugation is one of the best methods to isolate bacterial MVs (Chutkan et al. 2013; Klimentova and Stulik 2015). At present, a procedure for isolating MVs, consisting of multiple centrifugation, filtration, and density gradient ultracentrifugation, has replaced original methods such as gel filtration and is used to guide the separation of bacterial MVs (Klimentova and Stulik 2015).
With the development of omics technology, various contents have been discovered, indicating the diversity of vesicle functions. Studies on different bacteria under different conditions have shown that MVs in Gram-positive bacteria contain proteins, genetic material, lipids, and metabolites (Coelho et al. 2019; Rivera et al. 2010). The following sections discuss the current understanding of the components of MVs.
Proteins
Experiments have shown similarities and differences in protein components between bacteria and MVs. For example, the analysis of the protein composition of Listeria monocytogenes found that 296 proteins were present in both whole cells and 16 proteins were present only in the MVs, including PI-PLC, autolysin, uncharacterized protein yabE, competence protein ComEC/Rec2, flagellar proteins, and other uncharacterized proteins (Karthikeyan et al. 2019). Protein localization showed differences in the sources of vesicles. PSORTb 3.0 software was commonly used for protein localization, and the composition of the protein localization was found to be different from that of Gram-positive bacteria. Most MV-related proteins were predicted as cytoplasmic or plasma membrane proteins, while a few were predicted as cell wall or extracellular binding proteins (Rivera et al. 2010). The unique protein composition that MVs have proteins which were only present in MVs and that MVs can selectivity enriched with certain proteins suggested that MVs had special functions.
MVs allow the bacteria to disperse bacterial products into the surrounding environment. For pathogenic Gram-positive bacteria, many MV proteins are virulent factors and toxins, which help the bacteria exert their actions. MV-related virulence factors vary according to the bacteria. For example, the virulence factor of L. monocytogenes MVs is a toxin that can interact with host cells and contribute to the pathogenesis of L. monocytogenes during infection (Karthikeyan et al. 2019). The virulence factors of S. mutans MVs are glucosyltransferases, which help S. mutans colonize (Rainey et al. 2019). For nonpathogenic Gram-positive bacteria, for example, MV proteins have a variety of protective effects as probiotics. In the study of Lactobacillus casei, researchers identified MV proteins as mediators of probiotic effects of L. casei, which are involved in the bacteria–gastrointestinal cell interface (Dominguez Rubio et al. 2017).
The composition of MV proteins in Gram-positive bacteria is different in different strains and environments. Many researchers have performed the proteomic analysis of different clinical strains under different culture conditions and found that the composition of MV proteins had many common and unique characteristics (Wagner et al. 2018). The growth medium affects gene expression in bacteria, thereby altering the number and/or content of MVs.
Nucleic acids
In Gram-negative bacteria, DNA of OMVs can be horizontally transferred between the same and different species, leading to the transmission of bacterial resistance (Domingues and Nielsen 2017). The related studies on Gram-positive bacteria were carried out late. Previous studies showed that genetic materials, including DNA and RNA, were present in MVs isolated from Gram-positive bacteria.
DNA in Gram-positive bacteria was first reported in cellulolytic Ruminococcus spp. The study reported that MVs from wild-type bacteria could continuously “rescue” the mutants and cause them to degrade crystalline cellulose (Klieve et al. 2005). Further studies found that Clostridium perfringens also contained partial gene-fringed DNA (Jiang et al. 2014). In addition, S. mutans released MV-related eDNA to improve biofilm formation (Liao et al. 2014; Senpuku et al. 2019).
The differentially enriched intragenic RNAs in MVs, mostly rRNAs and tRNAs, were first reported in Group A Streptococcus (Resch et al. 2016). However, no studies explored the possibility of RNA-level transfer mediated by MVs in Gram-positive bacteria, which was contrary to the RNA content of mammalian exosomes.
Lipids
Lipids are important components of MVs. The identification of key lipid biomarkers in metabolic regulation can reveal the mechanism underlying various life activities.
Studies have shown that MVs contain lipids, but they also show some specific aggregation. For example, the lipids identified from Bacillus anthracis and MVs were mainly palmitic acid and stearic acid, but considerable differences were detected in the composition of secondary lipid components. MVs were rich in myristic and palmitic acids (Rivera et al. 2010). The study on Streptococcus pneumoniae found that MVs had a different fatty acid composition and were enriched in short-chain saturated fatty acids (Olaya-Abril et al. 2014). The study on L. monocytogenes found that unsaturated fatty acids such as phosphatidylethanolamine and sphingolipids were more abundant in MVs (Coelho et al. 2019). These studies showed that the lipid components of MVs did not replicate in the bacterial membrane, but had unique aggregation patterns. The result suggested that MVs were produced via certain mechanisms. One possible explanation is that short-chain saturated fatty acids and unsaturated fatty acids, which are selectively enriched in Gram-positive MVs, affect membrane curvature and membrane fluidity. Since the membrane is more fluid with shorter and more unsaturated fatty acids (Mercier et al. 2012), selective enrichment makes it easier for Gram-positive bacteria to form and release MVs. However, the specific role of these lipid components needs further exploration.
Metabolites
The changes in metabolites indicate the influence of disease, toxicity, gene modification, and environmental factors. The study on bacterial vesicle metabolites is an emerging research field (Bryant et al. 2017; Zakharzhevskaya et al. 2017). Comparative metabolomic analyses were conducted on vesicles isolated from virulent strains of Bacteroides fragilis and nonvirulent strains, and metabolic pathways of OMVs were reconstructed (Zakharzhevskaya et al. 2017).
Compared to the studies on the Gram-negative bacteria, few studies explored the metabolomics of Gram-positive bacteria. The metabolites carried by MVs include ornithine, citrate, inositol, phenylalanine, citric acid, and the key intermediate metabolite pyruvate (Coelho et al. 2019). The metabolic composition of MVs varies under different environments. Our study compared the differences in metabolites in S. mutans under acidic conditions and obtained 35 different metabolites. The upregulated MV metabolites, such as betaine, trehalose, and L-carnitine, were stress protectors, providing new information for assessing the survival and proliferation of Gram-positive bacteria under environmental pressure (Cao et al. 2020). The characteristics of metabolites in different types of bacteria are not clear, which is worth further exploration.
Function of MVs
The benefits must be sufficient considering the huge energy costs involved in producing MVs. MVs are likely to have important biological functions based on their metabolic cost and interesting contents (Brown et al. 2015). In recent years, thanks to intensive research, it was soon discovered that MVs were vital in bacterial physiology and pathogenesis. They were thought to be involved in stress response, biofilm formation, intraspecies and interspecies communication, immune regulation, and so on. In the following sections, the current understanding of the function of MVs was presented.
Stress response
Bacteria are exposed to a variety of pressures, such as malnutrition, antibiotics, and oxidative stress, under both physiological and clinical conditions. The bacteria increase the pressure response, which usually includes activation of pressure sensors, changes in transcription levels, and downstream changes in bacterial envelope composition, to counteract the effects of environmental damage on their structures (Ahn et al. 2006). Current studies suggest that vesicle involvement in pressure regulation is related to lipid involvement in changes in membrane curvature and secretion of misfolded proteins and metabolites.
OMVs are used as part of the general stress response mechanism to improve the chances of bacterial survival in Gram-negative bacteria (Atashgahi et al. 2018; Baumgarten et al. 2012). The production of OMVs is also considered as a stress response, which can eliminate bacterial misfolded proteins and provide an external bait to absorb antimicrobial agents on the cell surface (Schwechheimer and Kuehn 2015).
However, MVs in Gram-positive bacteria have been less explored. L. monocytogenes can survive under extreme environmental stress conditions, and the transcription factor sigma B is involved in this survival ability (Wiedmann et al. 1998). Studies found that a wild-type strain produces ninefold more MVs compared with sigB mutant (Lee et al. 2013). Studies found that the extrusion of the polar growth tissue at the hypha tip led to the formation of MVs, which was related to the survival of bacteria in Streptomyces venezuelae (Frojd and Flardh 2019). In the present study, acidic conditions induced S. mutans to produce more and smaller vesicles containing different proteins and metabolites. The expansion of the MVs may serve as a defensive response to acid pressure, increasing the chances of bacterial survival (Cao et al. 2020). The forementioned studies suggested that MV formation helps Gram-positive bacteria to better adapt to environmental stress and prevent the fatal effects of adverse environments. On the one hand, the release of MVs can maintain the stability of the membrane; on the other hand, MVs can carry some stress substances out of the bacteria. MV formation is a strategy for Gram-positive bacteria to respond to environmental changes. Hence, the role of MVs in the stress response of Gram-positive bacteria deserves further investigation.
Biofilm formation
Biofilm is considered as the main state of microorganisms in the environment, and the formation of biofilms is an important issue in clinic. Biofilms protect microbial growth from surface detachment and interference by antimicrobial substances (Kumar et al. 2017). The involvement of vesicles in biofilm formation is related to vesicle cargos, such as eDNA as a biofilm substrate, lipids as hydrophobic surface providers, and adhesion-related proteins as adhesive mediators. OMVs are crucial in the formation, communication, nutrition acquisition, and defense of biofilms and are the key substances of biofilms (Brown et al. 2015; Schooling and Beveridge 2006).
The effects of MVs in Gram-positive bacteria on biofilms have also been reported. On the one hand, MVs can be used as the substrate of biofilms. On the other hand, adhesion-related enzymes in MVs are actively involved in biofilm formation. In the Bacillus subtilis biofilm, MVs exist in the stroma and emerge from embedded cells in the biofilm (Brown et al. 2014). In the Mycobacterium ulceris biofilm, MVs are confined in the extracellular matrix surrounding the outer part of the bacterial community, rather than distributed throughout the complex biofilm structure. MVs isolated from planktonic L. monocytogenes contain protein components of biofilms. In S. mutans, MVs contain eDNAs and adhesive proteins that contribute to the formation of biofilms. In particular, MVs are easily dispersed and hence make local use of sucrose in the surrounding environment, facilitating the colonization of bacteria (Cao et al. 2020). MVs in S. mutans not only benefit the formation of their own biofilms but also contribute to the development of Candida albicans biofilm (Wu et al. 2020).
Immune regulation
The role of MVs in host immune regulation has been increasingly recognized because MVs contain many immune-related molecules and can be accepted by host cells. These immune-related molecules include lipoproteins and toxins which could trigger innate and adaptive immunity.
MVs in both pathogenic and nonpathogenic Gram-positive bacteria are involved in innate immunity, in which macrophages, dendritic cell, and Toll-like receptor 2 (TLR2) are the key molecules. A large number of MVs in pathogenic bacteria can activate the host innate immune response. For example, MVs in C. perfringens induce the release of inflammatory cytokine interleukin-6 (IL-6) through the TLR2 signaling pathway (Jiang et al. 2014). Staphylococcus aureus enhances the production of protein inflammatory mediators, such as tumor necrosis factor (TNF-), IL-6, and IL-12, through the expression of co-stimulatory molecules via the TLR2 signaling pathway (Choi et al. 2015; Gurung et al. 2011). MVs in Streptococcus suis also activate the nuclear factor kappa B signaling pathway in single cells and macrophages, inducing the secretion of pro-inflammatory cytokines (Haas and Grenier 2015). In a study on MVs in nonpathogenic bacteria, MVs derived from Lactobacillus sakei subsp. sakei NBRC15893 enhanced immunoglobulin A production by activating TLR2 signaling (Yamasaki-Yashiki et al. 2019). The cell wall components of Gram-positive bacteria contain large numbers of TLR2 ligands, such as lipoteichoic acid, teichoic acid, peptidoglycan, and lipoprotein (Kaji et al. 2010; Matsuguchi et al. 2003; Shida et al. 2009; Zeuthen et al. 2008). Thus, MVs in Gram-positive bacteria which also contain cell wall components could work together with bacteria to exert innate immunomodulatory effects via TLR2. To date, lipoproteins contained in MVs have been reported to be involved in immunomodulatory responses in a variety of Gram-positive bacteria (Prados-Rosales et al. 2011; Yamasaki-Yashiki et al. 2019).
MVs are also vital in adaptive immunity, which has also been confirmed by relevant studies. Mice inoculated with MVs in Streptococcus pneumoniae produced specific antibodies (Olaya-Abril et al. 2014). S. aureus induced Th1, Th17, and Th2 cells as well as IgG antibody responses (Choi et al. 2015). These findings indicated an effective vaccination efficacy of MVs in Gram-positive bacteria.
Conclusions and perspectives
The discovery of EVs is a breakthrough and provides a new mechanism for the release of components into the extracellular environment (Avila-Calderón et al. 2015; Nagakubo et al. 2019). Studies on MVs in Gram-positive bacteria are relatively few, lagging behind those on eukaryotes and Gram-negative bacteria, due to the thick cell walls and low production of vesicles (Brown et al. 2015). However, these MVs deserve further exploration owing to the importance of their functions.
The enrichment and selection of specific factors associated with Gram-positive MVs indicated a regulatory mechanism for MV cargo. Genetic and environmental factors also influence MV contents and regulate their release. The molecular mechanisms underlying MV release and cargo are still unknown, but will surely be elucidated in recent years.
Besides the analysis of individual omics, the combined analysis of multiple omics has become more important. In previous studies, the analysis of single omics data was often carried out; the relationship between the omics data and biological processes was explored through the independent analysis of each omics to explain the single information of some species. However, single omics is far from enough for the regulation of complex biological processes. Researchers began to integrate and analyze multiple sets of data (Biagini et al. 2015; Coelho et al. 2019; Olaya-Abril et al. 2014). In the field of MV research, multiomics is beginning to be applied and some important conclusions have been drawn.
The diversity of MV composition indicates the diversity of their functions. This review briefly introduced the role of MVs in stress resistance, biofilm formation, and immunity, but more roles are worth exploring. In particular, as OMVs in Gram-negative bacteria have been used as vaccines (Jiang et al. 2019; Pastor et al. 2021), there is great potential for the application of MVs in Gram-positive bacteria, too.
The findings of this review might stimulate the exploration of MVs in Gram-positive bacteria.
References
Ahn SJ, Wen ZZT, Burne RA (2006) Multilevel control of competence development and stress tolerance in Streptococcus mutans UA159. Infect Immun 74(3):1631–1642. https://doi.org/10.1128/iai.74.3.1631-1642.2006
Atashgahi S, Sanchez-Andrea I, Heipieper HJ, van der Meer JR, Stams AJM, Smidt H (2018) Prospects for harnessing biocide resistance for bioremediation and detoxification. Science 360(6390):743–746. https://doi.org/10.1126/science.aar3778
Avila-Calderón ED, Araiza-Villanueva MG, Cancino-Diaz JC, López-Villegas EO, Sriranganathan N, Boyle SM, Contreras-Rodríguez A (2015) Roles of bacterial membrane vesicles. Arch Microbiol 197(1):1–10. https://doi.org/10.1007/s00203-014-1042-7
Baumgarten T, Sperling S, Seifert J, von Bergen M, Steiniger F, Wick LY, Heipieper HJ (2012) Membrane vesicle formation as a multiple-stress response mechanism enhances Pseudomonas putida DOT-T1E cell surface hydrophobicity and biofilm formation. Appl Environ Microbiol 78(17):6217–6224. https://doi.org/10.1128/aem.01525-12
Biagini M, Garibaldi M, Aprea S, Pezzicoli A, Doro F, Becherelli M, Taddei AR, Tani C, Tavarini S, Mora M, Teti G, D’Oro U, Nuti S, Soriani M, Margarit I, Rappuoli R, Grandi G, Norais N (2015) The human pathogen Streptococcus pyogenes releases lipoproteins as lipoprotein-rich membrane vesicles. Mol Cell Proteomics 14(8):2138–2149. https://doi.org/10.1074/mcp.M114.045880
Briaud P, Carroll RK (2020) Extracellular vesicle biogenesis and functions in gram-positive bacteria. Infect Immun 88(12):e00433–e00420. https://doi.org/10.1128/iai.00433-20
Brown L, Kessler A, Cabezas-Sanchez P, Luque-Garcia JL, Casadevall A (2014) Extracellular vesicles produced by the Gram-positive bacterium Bacillus subtilis are disrupted by the lipopeptide surfactin. Mol Microbiol 93(1):183–198. https://doi.org/10.1111/mmi.12650
Brown L, Wolf JM, Prados-Rosales R, Casadevall A (2015) Through the wall: extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat Rev Microbiol 13(10):620–630. https://doi.org/10.1038/nrmicro3480
Bryant WA, Stentz R, Le Gall G, Sternberg MJE, Carding SR, Wilhelm T (2017) In silico analysis of the small molecule content of outer membrane vesicles produced by Bacteroides thetaiotaomicron indicates an extensive metabolic link between microbe and host. Front Microbiol 8:2440. https://doi.org/10.3389/fmicb.2017.02440
Cao Y, Zhou Y, Chen D, Wu R, Guo L, Lin H (2020) Proteomic and metabolic characterization of membrane vesicles derived from Streptococcus mutans at different pH values. Appl Microbiol Biotechnol 104(22):9733–9748. https://doi.org/10.1007/s00253-020-10563-6
Choi DS, Kim DK, Kim YK, Gho YS (2013) Proteomics, transcriptomics and lipidomics of exosomes and ectosomes. Proteomics 13(10-11):1554–1571. https://doi.org/10.1002/pmic.201200329
Choi SJ, Kim MH, Jeon J, Kim OY, Choi Y, Seo J, Hong SW, Lee WH, Jeon SG, Gho YS, Jee YK, Kim YK (2015) Active immunization with extracellular vesicles derived from Staphylococcus aureus effectively protects against staphylococcal lung infections, mainly via th1 cell-mediated immunity. PLoS One 10(9):e0136021. https://doi.org/10.1371/journal.pone.0136021
Chutkan H, Macdonald I, Manning A, Kuehn MJ (2013) Quantitative and qualitative preparations of bacterial outer membrane vesicles. Methods Mol Biol 966:259–272. https://doi.org/10.1007/978-1-62703-245-2_16
Coelho C, Brown L, Maryam M, Vij R, Smith DFQ, Burnet MC, Kyle JE, Heyman HM, Ramirez J, Prados-Rosales R, Lauvau G, Nakayasu ES, Brady NR, Hamacher-Brady A, Coppens I, Casadevall A (2019) Listeria monocytogenes virulence factors, including listeriolysin O, are secreted in biologically active extracellular vesicles. J Biol Chem 294(4):1202–1217. https://doi.org/10.1074/jbc.RA118.006472
Domingues S, Nielsen KM (2017) Membrane vesicles and horizontal gene transfer in prokaryotes. Curr Opin Microbiol 38:16–21. https://doi.org/10.1016/j.mib.2017.03.012
Dominguez Rubio AP, Martinez JH, Martinez Casillas DC, Coluccio Leskow F, Piuri M, Perez OE (2017) Lactobacillus casei BL23 produces microvesicles carrying proteins that have been associated with its probiotic effect. Front Microbiol 8:1783–1783. https://doi.org/10.3389/fmicb.2017.01783
El Andaloussi S, Maeger I, Breakefield XO, Wood MJA (2013) Extracellular vesicles: biology and emerging therapeutic opportunities. Nat Rev Drug Discov 12(5):348–358. https://doi.org/10.1038/nrd3978
Frojd MJ, Flardh K (2019) Extrusion of extracellular membrane vesicles from hyphal tips of Streptomyces venezuelae coupled to cell-wall stress. Microbiology (Reading) 165(12):1295–1305. https://doi.org/10.1099/mic.0.000836
Gorringe A, Halliwell D, Matheson M, Reddin K, Finney M, Hudson M (2005) The development of a meningococcal disease vaccine based on Neisseria lactamica outer membrane vesicles. Vaccine 23(17-18):2210–2213. https://doi.org/10.1016/j.vaccine.2005.01.055
Gurung M, Moon DC, Choi CW, Lee JH, Bae YC, Kim J, Lee YC, Seol SY, Cho DT, Kim SI, Lee JC (2011) Staphylococcus aureus produces membrane-derived vesicles that induce host cell death. PLoS One 6(11):e27958. https://doi.org/10.1371/journal.pone.0027958
Haas B, Grenier D (2015) Isolation, characterization and biological properties of membrane vesicles produced by the swine pathogen Streptococcus suis. PLoS One 10(6):e0130528. https://doi.org/10.1371/journal.pone.0130528
Jan AT (2017) Outer membrane vesicles (OMVs) of Gram-negative bacteria: a perspective update. Front Microbiol 8:1053. https://doi.org/10.3389/fmicb.2017.01053
Jeon H, Oh MH, Jun SH, Kim SI, Choi CW, Kwon HI, Na SH, Kim YJ, Nicholas A, Selasi GN, Lee JC (2016) Variation among Staphylococcus aureus membrane vesicle proteomes affects cytotoxicity of host cells. Microb Pathog 93:185–193. https://doi.org/10.1016/j.micpath.2016.02.014
Jiang Y, Kong Q, Roland KL, Curtiss R 3rd (2014) Membrane vesicles of Clostridium perfringens type A strains induce innate and adaptive immunity. Int J Med Microbiol 304(3-4):431–443. https://doi.org/10.1016/j.ijmm.2014.02.006
Jiang L, Schinkel M, van Essen M, Schiffelers RM (2019) Bacterial membrane vesicles as promising vaccine candidates. Eur J Pharm Biopharm 145:1–6. https://doi.org/10.1016/j.ejpb.2019.09.021
Kaji R, Kiyoshima-Shibata J, Nagaoka M, Nanno M, Shida K (2010) Bacterial teichoic acids reverse predominant IL-12 production induced by certain lactobacillus strains into predominant IL-10 production via TLR2-dependent ERK activation in macrophages. J Immunol 184(7):3505–3513. https://doi.org/10.4049/jimmunol.0901569
Karthikeyan R, Gayathri P, Gunasekaran P, Jagannadham MV, Rajendhran J (2019) Comprehensive proteomic analysis and pathogenic role of membrane vesicles of Listeria monocytogenes serotype 4b reveals proteins associated with virulence and their possible interaction with host. Int J Med Microbiol 309(3-4):199–212. https://doi.org/10.1016/j.ijmm.2019.03.008
Klieve AV, Yokoyama MT, Forster RJ, Ouwerkerk D, Bain PA, Mawhinney EL (2005) Naturally occurring DNA transfer system associated with membrane vesicles in cellulolytic Ruminococcus spp. of ruminal origin. Appl Environ Microbiol 71(8):4248–4253. https://doi.org/10.1128/aem.71.8.4248-4253.2005
Klimentova J, Stulik J (2015) Methods of isolation and purification of outer membrane vesicles from gram-negative bacteria. Microbiol Res 170:1–9. https://doi.org/10.1016/j.micres.2014.09.006
Kumar A, Alam A, Rani M, Ehtesham NZ, Hasnain SE (2017) Biofilms: survival and defense strategy for pathogens. Int J Med Microbiol 307(8):481–489. https://doi.org/10.1016/j.ijmm.2017.09.016
Lee JH, Choi CW, Lee T, Kim SI, Lee JC, Shin JH (2013) Transcription factor sigmaB plays an important role in the production of extracellular membrane-derived vesicles in Listeria monocytogenes. PLoS One 8(8):e73196. https://doi.org/10.1371/journal.pone.0073196
Liao S, Klein MI, Heim KP, Fan Y, Bitoun JP, Ahn SJ, Burne RA, Koo H, Brady LJ, Wen ZT (2014) Streptococcus mutans extracellular DNA is upregulated during growth in biofilms, actively released via membrane vesicles, and influenced by components of the protein secretion machinery. J Bacteriol 196(13):2355–2366. https://doi.org/10.1128/jb.01493-14
Matsuguchi T, Takagi A, Matsuzaki T, Nagaoka M, Ishikawa K, Yokokura T, Yoshikai Y (2003) Lipoteichoic acids from Lactobacillus strains elicit strong tumor necrosis factor alpha-inducing activities in macrophages through Toll-like receptor 2. Clin Diagn Lab Immunol 10(2):259–266. https://doi.org/10.1128/cdli.10.2.259-266.2003
Mercier R, Domínguez-Cuevas P, Errington J (2012) Crucial role for membrane fluidity in proliferation of primitive cells. Cell Rep 1(5):417–423. https://doi.org/10.1016/j.celrep.2012.03.008
Nagakubo T, Nomura N, Toyofuku M (2019) Cracking open bacterial membrane vesicles. Front Microbiol 17(10):3026. https://doi.org/10.3389/fmicb.2019.03026
Olaya-Abril A, Prados-Rosales R, McConnell MJ, Martin-Pena R, Gonzalez-Reyes JA, Jimenez-Munguia I, Gomez-Gascon L, Fernandez J, Luque-Garcia JL, Garcia-Lidon C, Estevez H, Pachon J, Obando I, Casadevall A, Pirofski L-A, Rodriguez-Ortega MJ (2014) Characterization of protective extracellular membrane-derived vesicles produced by Streptococcus pneumoniae. J Proteome 106:46–60. https://doi.org/10.1016/j.jprot.2014.04.023
Pastor Y, Ting I, Berzosa M, Irache JM, Gamazo C (2021) Vaccine based on outer membrane vesicles using hydrogels as vaccine delivery system. Methods Mol Biol 2182:153–160. https://doi.org/10.1007/978-1-0716-0791-6_14
Prados-Rosales R, Baena A, Martinez LR, Luque-Garcia J, Kalscheuer R, Veeraraghavan U, Camara C, Nosanchuk JD, Besra GS, Chen B, Jimenez J, Glatman-Freedman A, Jacobs WR Jr, Porcelli SA, Casadevall A (2011) Mycobacteria release active membrane vesicles that modulate immune responses in a TLR2-dependent manner in mice. J Clin Invest 121(4):1471–1483. https://doi.org/10.1172/jci44261
Rainey K, Michalek SM, Wen ZT, Wu H (2019) Glycosyltransferase-mediated biofilm matrix dynamics and virulence of Streptococcus mutans. Appl Environ Microbiol 85(5):e02247–e02218. https://doi.org/10.1128/aem.02247-18
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200(4):373–383. https://doi.org/10.1083/jcb.201211138
Resch U, Tsatsaronis JA, Le Rhun A, Stubiger G, Rohde M, Kasvandik S, Holzmeister S, Tinnefeld P, Wai SN, Charpentier E (2016) A two-component regulatory system impacts extracellular membrane-derived vesicle production in group A Streptococcus. mBio 7(6):e00207–e00216. https://doi.org/10.1128/mBio.00207-16
Rivera J, Cordero RJ, Nakouzi AS, Frases S, Nicola A, Casadevall A (2010) Bacillus anthracis produces membrane-derived vesicles containing biologically active toxins. Proc Natl Acad Sci U S A 107(44):19002–19007. https://doi.org/10.1073/pnas.1008843107
Schooling SR, Beveridge TJ (2006) Membrane vesicles: an overlooked component of the matrices of biofilms. J Bacteriol 188(16):5945–5957. https://doi.org/10.1128/jb.00257-06
Schwechheimer C, Kuehn MJ (2015) Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat Rev Microbiol 13(10):605–619. https://doi.org/10.1038/nrmicro3525
Senpuku H, Nakamura T, Iwabuchi Y, Hirayama S, Nakao R, Ohnishi M (2019) Effects of complex DNA and MVs with GTF extracted from Streptococcus mutans on the oral biofilm. Molecules 24(17):3131. https://doi.org/10.3390/molecules24173131
Shida K, Kiyoshima-Shibata J, Kaji R, Nagaoka M, Nanno M (2009) Peptidoglycan from lactobacilli inhibits interleukin-12 production by macrophages induced by Lactobacillus casei through Toll-like receptor 2-dependent and independent mechanisms. Immunology 128(1 Suppl):e858–e869. https://doi.org/10.1111/j.1365-2567.2009.03095.x
Tan K, Li R, Huang X, Liu Q (2018) Outer membrane vesicles: current status and future direction of these novel vaccine adjuvants. Front Microbiol 9:783. https://doi.org/10.3389/fmicb.2018.00783
Toyofuku M, Nomura N, Eberl L (2019) Types and origins of bacterial membrane vesicles. Nat Rev Microbiol 17(1):13–24. https://doi.org/10.1038/s41579-018-0112-2
van Niel G, D'Angelo G, Raposo G (2018) Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19(4):213–228. https://doi.org/10.1038/nrm.2017.125
Wagner T, Joshi B, Janice J, Askarian F, Skalko-Basnet N, Hagestad OC, Mekhlif A, Wai SN, Hegstad K, Johannessen M (2018) Enterococcus faecium produces membrane vesicles containing virulence factors and antimicrobial resistance related proteins. J Proteome 187:28–38. https://doi.org/10.1016/j.jprot.2018.05.017
Wiedmann M, Arvik TJ, Hurley RJ, Boor KJ (1998) General stress transcription factor sigmaB and its role in acid tolerance and virulence of Listeria monocytogenes. J Bacteriol 180(14):3650–3656. https://doi.org/10.1128/jb.180.14.3650-3656.1998
Wu R, Tao Y, Cao Y, Zhou Y, Lin H (2020) Streptococcus mutans membrane vesicles harboring glucosyltransferases augment Candida albicans biofilm development. Front Microbiol 11:581184. https://doi.org/10.3389/fmicb.2020.581184
Yamasaki-Yashiki S, Miyoshi Y, Nakayama T, Kunisawa J, Katakura Y (2019) IgA-enhancing effects of membrane vesicles derived from Lactobacillus sakei subsp. sakei NBRC15893. Biosci Microbiota Food Health 38(1):23–29. https://doi.org/10.12938/bmfh.18-015
Zakharzhevskaya NB, Vanyushkina AA, Altukhov IA, Shavarda AL, Butenko IO, Rakitina DV, Nikitina AS, Manolov AI, Egorova AN, Kulikov EE, Vishnyakov IE, Fisunov GY, Govorun VM (2017) Outer membrane vesicles secreted by pathogenic and nonpathogenic Bacteroides fragilis represent different metabolic activities. Sci Rep 7(1):5008. https://doi.org/10.1038/s41598-017-05264-6
Zeuthen LH, Fink LN, Frøkiaer H (2008) Toll-like receptor 2 and nucleotide-binding oligomerization domain-2 play divergent roles in the recognition of gut-derived lactobacilli and bifidobacteria in dendritic cells. Immunology 124(4):489–502. https://doi.org/10.1111/j.1365-2567.2007.02800.x
Funding
This work was funded by the National Natural Science Foundation of China (No. 81970928).
Author information
Authors and Affiliations
Contributions
CY wrote and reviewed the main manuscript text and prepared the figures. LH reviewed and revised the main manuscript. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
None required.
Conflict of interest
The author declares no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cao, Y., Lin, H. Characterization and function of membrane vesicles in Gram-positive bacteria. Appl Microbiol Biotechnol 105, 1795–1801 (2021). https://doi.org/10.1007/s00253-021-11140-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11140-1