Abstract
Lead is an extensive contaminant. Pb-resistant bacterial strains were isolated from Saint Clair River sediments on two enrichment media with increasing concentrations of Pb (NO3)2. Bacterial strains that grew at 1.25 or 1.5 g L−1 of Pb (NO3)2 L−1) were purified and selected for further study. Ninety-seven Pb-resistant strains were screened for the ability to produce bioflocculants. The majority of the Pb-resistant strains demonstrated moderate to high flocculation activity. Metal removal assays demonstrated that the higher is the flocculation activity, the higher is the efficiency of metal removal. In the multi-metal solutions, the bacterial strain with the highest flocculation activity (R19) had the highest metal removing capability (six out of eight metals) and the highest metal removal efficiency. The highly selective affinity towards Pb2+ observed for strain R19 suggests its use for the recovery of Pb2+ from multiple metal solutions. Because they are well adapted to unfavorable conditions due to their resistance to metals (e.g., Pb) and antibiotics, these characteristics may help in developing an effective process for wastewater treatment using these strains.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Lead (Pb) is associated with many health hazards (Demayoa et al. 1982). Lead poisoning is common and leads to mental retardation in children (Moncrieff et al. 1964). It affects reproductive systems, reduces volume of sperm, increases miscarriage, increases prematurity, and decreases low birth weight (Cleveland et al. 2008). In cells, Pb decreases the efficiency of the ATPase pump and protein kinase activity and negatively affects the concentrations of Na+, K+, and Ca+. Another adverse effect of Pb in eukaryotic cells is the increase in the formation of inclusion bodies that in turn translocate metals into the nuclei and then alter gene expression (Hu et al. 1998). Lead occurs naturally in the environment although Pb found in the environments are mostly the result of human activities. The main application for lead is in the battery production industry and vehicle batteries accounting for 80% of the total within the industry. Other types of battery are produced for motorcycles, planes, tanks, trains, tractors, factories, energy, wharves, and export. It is often found in effluents discharged from industries. In freshwater environments, Pb tends to settle at the bottom of the water where it concentrates and is capable of accumulating in the tissues of aquatic biota. Even at low concentrations, the composition, distribution, and diversity benthic organisms are affected. Thus, removal of toxic heavy metals such as Pb from industrial wastewaters is essential from the standpoint of environmental pollution control (Guangyu and Thiruvenkatachari 2003).
Several methods have been used to remediate heavy metals from the environment. These methods include ion exchange, reverse osmosis, adsorption, solvent extraction, and chemical precipitation (Babel and Kurniawan 2003). However, conventional methods employ non-regenerable materials, which in turn increase the cost of these processes. Moreover, these methods are not environmental friendly as these processes generate substantial secondary waste. Hence, biosorption or adsorption by microorganisms has become a preferred alternative for heavy metal treatment, and it is cost-effective and environmental friendly. Biosorption is a mechanism that involves adsorption of metals by microorganisms (Murthy et al. 2012). In general, the cell surface of bacteria carries a net negative charge due to the presence of amine, carboxyl, hydroxyl, phosphate, and sulfhydryl groups (Naik and Dubey 2013). Thus, the cell wall can absorb appreciable quantities of positively charged cationic metals, such as Pb. Some bacteria secrete exopolysaccharides (EPS), which aid in extracellular sequestration of cations. EPS high molecular weight polyanionic polymers that have been shown to have potential to bind heavy metals with differential degree of specificity and affinity (Bhaskar and Bhosle 2006; Pal and Paul 2008) bind metals by electrostatic interaction between metal cations and negatively charged EPS (Bruins et al. 2000; Nies and Silver 1995; Silver 1996).
The biosorption capacity of bacteria is governed by factors such as physico-chemical properties of the heavy metals in question. The pH as well as the presence of other ions and charged compounds also has effects on the biosorption capacity of the microbes (El-Hendawy et al. 2009; Zouboulis et al. 2003). This paper reports on the enrichment, isolation, and characterization of Pb-tolerant bacteria from Lake Saint Clair sediments.
Materials and methods
Site description and sediment sampling
Sediment samples were collected from Saint Clair River between Michigan. Saint Clair River is an outlet for Lake Huron, forming part of the boundary between Michigan, USA, and Ontario, Canada. Much of the shoreline on both sides of the St. Clair River is urbanized and heavily industrialized; therefore, the pollutants from day-to-day human activities and the industry have contributed to the impairment of the water quality in the lake and river. Pollutants in the Saint Clair River and Lake Saint Clair sediments include toxic organic compounds such as organochlorine insecticides, polychlorinated biphenyls (PCBs), chlorobenzenes, chlorotoluenes, and chlorostyrenes (Fogarty 2007). Heavy metals such as copper, nickel, zinc, lead, cadmium, chromium, and mercury have also been detected in the sediments (Table S1). Sediment samples were retrieved from three locations of Saint Clair River (SC1, SC2, and SC3). These locations are in the vicinity of known point pollution input sources (e.g., city of Saint Clair permitted wastewater discharges, marina, major river follow, and public parks) (Oest et al. 2018; Patel et al. 2018). Site SC1 (N 42° 58.393; W 82° 25.141) is located upstream where the water from Lake Huron drains into Saint Clair River; site SC2 (N 42° 54.323; W 82° 28.036) is close to the city of Marysville Wastewater Treatment Plant, City Public Park and public boat launch, and site SC3 (N 42° 49.2′ 9″; W 82° 29.2′ 0″) is situated at the mouth of Pine River, which empties into the Lake Saint Clair.
A ponar grab sampler was used to collect the sediment samples. The sampling equipment was decontaminated after each sample was processed. The decontamination procedure included rinsing the sampling pans and spoons with site water and using new sampling tube and gloves for each sample location. Sediment samples for molecular analyses were stored at − 80 °C until used. Sediment samples for enrichment of metal-resistant bacteria were stored at 4 °C and were used in the enrichment assay as soon as they were shipped in the laboratory.
Sediment analysis
The samples were analyzed for the presence of As, Ba, Cd, Cr, Cu, Fe, Pb, Mn, Ni, Se, Zn, Ag, and Hg) using inductively coupled plasma-mass spectrometry (ICP-MS) (ELAN 6000, PerkinElmer, CA, USA).
Enrichment and isolation of Pb-resistant bacteria
For the isolation of lead-resistant bacteria, the sediment sample was enriched into lead-supplemented nutrient media. Pb (NO3)2 was used at a final concentration of 10 g L−1. Luria Bertani (LB) broth (Difco and BBL 2009) was used to isolate fast-growing bacteria, while Reasoner’s 2A (R2A) (Difco and BBL 2009) broth was used to isolate slow-growing bacteria. Twenty grams of sediment sample was added to 1 L of each medium and was incubated at room temperature with shaking at 150 rpm until substantial growth was observed. During incubation, the growth of the bacteria in the enrichment media was monitored following turbidity method at 660 nm. The enrichment processes were repeated for three cycles, by transferring 20 mL of the late-exponential phase enriched culture into fresh LB and R2A broths.
Screening of Pb-resistant bacteria was carried out by inoculating 20 mL of the enriched culture in 1 L of LB or R2A media supplemented with 0 to 1.5 L−1 g Pb (NO3)2. The enrichment media were then incubated at room temperature for 48 h with shaking (150 rpm). This process was repeated for three times.
The Pb-resistant strains were obtained from the highest concentration of Pb (NO3)2 in which growth was observed (1.25 g L−1 Pb (NO3)2 for the R2A strains and 1.5 g L−1 Pb (NO3)2 for the LB strains). The enrichment cultures were diluted and aliquots spread directly on the surface of R2A and LB agar plates containing 1.25 g Pb [NO3]2 L−1 and 1.5 g Pb [NO3]2 L−1, respectively. The plates were incubated at room temperature for 1 week. Colonies arising on the plates were selected for isolation based on gross morphological features and then purified. A total of 92 Pb-resistant bacterial strains were recovered, 47 strains from the R2A enrichment and 45 strains from the LB enrichment. Morphological and biochemical characteristics of the Pb-resistant strains were determined by standard methods (Gerhardt et al. 1994).
Characterization of Pb-resistant isolates
Sensitivity to antibiotics
Antibiotic sensitivity of the 92 Pb-resistant isolates recovered from the enrichment assay was determined using the disc diffusion method (Carolina Biological Supply Company, NC, USA). Overnight broth cultures of the test strains were used and the turbidity of the inoculum as adjusted in phosphate-buffered saline (pH 7.4) to a 0.5 McFarland opacity standard. These cultures (0.1 mL) were then aseptically spread onto Mueller–Hinton agar plates. Standard antibiotic-impregnated discs (6-mm diameter, Carolina Biological Supply Company, NC, USA) were placed on freshly prepared lawns of each isolate on Mueller–Hinton plates. After 24 h of incubation at 30 °C (Difco and BBL 2009), the diameter of the inhibition zones around the discs was measured after incubation and the bacterial strains were classified as resistant, intermediate, and susceptible, following the standard antibiotic disc sensitivity testing method (Difco and BBL 2009) to the following antibiotics: chloramphenicol (30 μg), erythromycin (15 μg), kanamycin (30 μg), neomycin (30 μg), novobiocin (30 μg), penicillin G (10 units), streptomycin (10 μg), and tetracycline (30 μg).
16S rRNA gene sequencing and phylogenetic analysis
The 16S rRNA gene sequences of the 92 Pb-resistant strains obtained from the enrichment experiments above were determined by direct sequencing of the PCR products. A 24-h culture was prepared for each strain for the DNA extraction. DNA extracts from each strain were prepared using DNEasy kit (Qiagen, Inc., Valencia, CA). The DNA was used as the target of PCR amplification of complete 16S rRNA fragments using bacterial primers FD1 and 1506 (Weisburg et al. 1991). The PCR products were purified and then sequenced directly using the ABI Big Dye Primer cycle sequencing kit. Sequencing was performed on an ABI 3700 DNA sequencer (Applied Biosystems, Foster, CA). All sequence fragments generated from a given template were edited against electropherograms and then assembled into contigs using SeqMan (Lasergene DNASTAR, Inc., Madison, WI). For most sequences, two to four overlapping fragments were used to assemble the contigs. Chimeric sequences were checked by Check_Chimera program available at the Ribosomal Database Project (RDP II) (Hugenholtz and Huber 2003). Complete 16S rRNA gene sequences were compared with other reference sequences as available in the NCBI database using the Basic Local Alignment Search Tool (BLAST) algorithm.
Bioflocculation assay
The Pb-resistant isolates were grown in R2A with 1.2 g L−1 Pb (NO3)2 or LB with 1.5 g L−1 Pb (NO3)2 broth for 48 h with shaking at 150 rpm. The bioflocculation activity of the Pb-resistant isolates was assessed using a kaolin clay suspension following the protocol described by Lin and Harichund (2011). The cell-free culture supernatant (1.5 mL of culture was centrifuged at 6000 rpm for 30 min) was used as crude bioflocculant. The bacterial supernatant (1 mL) was added to 9 mL of kaolin suspension (5 g L−1) in the presence of 3 mL 10 g L−1 CaCl2. The mixture was vortexed for 30 s and then left undisturbed for 5 min. The OD (optical density) of the clarifying solution was measured using a UV-visible spectrophotometer (Shimadzu UV-1250, OR, USA) at 550 nm. A control experiment was set up using the same protocol described above, except that the crude bioflocculant was replaced with uninoculated culture media. The flocculating activity was calculated based on the optical density of the control experiment at 550 nm and is the optical density of the sample experiment (Zhang et al. 2007).
Metal removal assay
Cells of each culture (100 mL) selected for this assay was confined in dialysis tubing and pretreated with 0.1 M HCl in order to remove the metal ions possibly bound to the negatively charged groups of the cell envelopes. After 30 min, the confined cultures were dialysed against deionized water for 24 h, in order to remove residual HCl from the treated cultures with continuous stirring at 100 rpm. The 100 mL confined cultures were subsequently transferred in 1 L solutions containing 10 mg L−1 of Pb for 5 h at room temperature. The pH of the system (biosorbent + metal solution) was adjusted to 4.5–5.5 by addition of either 1 M HCl or 1 M NaOH to avoid hydroxide precipitation.
Samples (10 mL of the aqueous solutions) were withdrawn at different times over 5 h and filtered through a 0.2-μm Acrodisc syringe filters. The metal uptake was calculated from the difference between the concentration of the metals in solution, determined with an atomic absorption spectrometer (PinAAcle 900T, PerkinElmer, MA, USA). Pb concentrations were measured at 283.31 nm. The amount of Pb2+ removed in blanks carried out in parallel without the addition of bacterial cells, which ranged from 0.1 to 0.2 mg L−1, was subtracted from the experimental values obtained in the tests with the bacterial cultures. All the experiments were performed in triplicates per time point, and the data were reported as mean values ± standard deviation. Specific metal uptake expressed as milligrams metal removed per gram of dry biomass. The dry weight (g L−1) was determined by centrifugation 10 mL of the dialysed cultures, followed by drying 50 °C until a constant weight was reached.
For the multi-metal removal assays, 10 mg L−1 of each metal (As, Pb, Cu, Mn, Zn, Cd, Cr, and Ni) was used. The assay was carried the same way as the mono-metal system (Pb only), except that the dialysed cultures were transferred to aqueous solutions containing a combination of all eight metals. As, Pb, Cu, Mn, Zn, Cd, Cr, and Ni were determined at 193.70, 283.31, 324.75, 279.48, 213.86, 228.00, 357.87, and 232.0 nm, respectively, using the atomic absorption spectrometer (PinAAcle 900T, PerkinElmer, MA, USA).
Nucleotide sequence accession numbers
The 16S rRNA gene sequence of selected lead-tolerant bacterial isolate was deposited in the NCBI/EMBL nucleotide sequence database under the accession numbers MG022641, MG022642, MG022643, MG022644, MG022645, MG022646, MG022647, MG022648, MG022649, MG022650, MG022651, MG022652, MG022653, MG022654, MG022655, MG022656, MG022657, MG022658, MG022659, and MG022660.
Strain deposition number
Strains R19 (Klebsiella michiganensis 19), R3 (Raoultella planticola R3), L2 (Providencia rettgeri L2), and L30 (Serratia sp. L30) were deposited to the Northern Regional Research Laboratory (NRRL, Peoria, IL, USA; https://nrrl.ncaur.usda.gov) Culture Collection under the deposit numbers B-65501, B-65504, B-65502, and B-65503, respectively.
Results
Enrichment and isolation of Pb-resistant strains
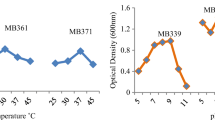
Metal analysis (As, Ba, Cd, Cr, Cu, Fe, Pb, Mn, Ni, Se, Zn, Ag, and Hg) of the sediments from the three sites demonstrated that the sediment samples retrieved from site SC1 contained slightly higher concentrations of metals (Table S1). Therefore, sediment SC1 was selected for enrichment and isolation of Pb-tolerant bacteria in R2A and LB enrichment culture media containing 0.25 to 1.5 g L−1 Pb (NO3)2 (Fig. 1). As expected, bacterial growth decreased as the concentration of Pb2+ increased, although the specific growth rates for the LB enrichments were higher than the R2A enrichments. The selected bacteria were able to grow in 1.25 or 1.5 g L−1 Pb (NO3), although these concentrations were not the optimal environments for their growth.
All 97 Pb-resistant bacterial strains were capable of both aerobic and anaerobic growth (Table S2). Overall, the LB strains were resistant to chloramphenicol, erythromycin, novobiocin, penicillin, and tetracycline (Table S2). The R2A strains on the other hand were dominated by gram-negative bacteria and were resistant to erythromycin, neomycin, novobiocin, and penicillin (Table S2).
The taxonomic identity of all the isolated Pb-resistant bacterial strains was ascertained through 16S rRNA gene sequence analysis. Sequence similarity search in NCBI Genbank and RDP database revealed that the Pb-resistant strains were from different genera falling under the phylum Proteobacteria. The strains composed mainly of 16S rRNA genes related to the β and γ subdivisions of Proteobacteria. The R2A strains grouped together near the top of the tree, while the LB isolates clustered together near the bottom of the tree indicating a distant relationship between two groups of enrichment cultures (Table S3). The LB strains were dominated by Providencia spp. and Enterococcus spp. (Fig. 2a), whereas the R2A strains were dominated by Raoultella spp. and Klebsiella (Fig. 2b).
Flocculation activities of the Pb-resistant strains
The flocculation activity of the Pb-resistant strains was assayed by monitoring their ability to flocculate kaolin clay. The flocculation activity was expressed in terms of percentage activity. Only three LB strains showed high flocculation activity, and five strains showed no evidence of activity (Fig. 3a). Most of the R2A strains showed either moderate to high flocculation activity. The maximum flocculation activity (79%) was recorded for strain R19 strain (Fig. 3b). In general, the flocculation activity was higher in R2A strains than the LB strains. Among the 97 Pb-resistant strains, 20 demonstrated high (> 50%) flocculation activity (Fig. 3b).
The majority of the R2A strains were similar to the 16S rRNA gene sequence of Rauotella spp.; a few have similarities with 16S rRNA gene sequence of other genera (Table S3).
Pb removal assay in mono-metal (Pb) and multi-metal solutions
The specific metal removal [expressed as (mg metal removed) (g dry biomass)−1] of the selected Pb-resistant strains was determined in mono- and multi-metal cation solutions. The strains with the highest flocculation activity from the R2A (R2 and R19 strains) and LB (L2 and L30) enrichments (Fig. 3b) were used in the Pb2+ removal assay. The four strains were capable of removing Pb2+ from aqueous solution (Fig. 4). In general, Pb removal was higher for the R2A isolates than the LB isolates. In mono-metal (Pb only) solutions, the Pb2+ removal was highest for strain R19 and lowest for L30 (Fig. 4a). In the mixed metal solutions, the highest Pb removal was found in strain R19, followed by strain R3 and L2. No Pb2+ removal was found for strain R19 (Fig. 4b) with multiple metal solutions. Sorption of Pb2+ by the bacterial cultures was gradual. The saturation of the metal removal capacity of each strain was achieved within the first 4 h in the single metal solution and after 5-h measurements in the multi-metal solution (Fig. 4). Metal cation removal was generally higher for the bacterial strains in the mono-metal solution than in the multi-metal solutions. With the exception of strain R19, the simultaneous presence of eight metals reduced the removal of Pb2+ in the aqueous solution (Fig. 5). Surprisingly, the presence of multiple metals had a stimulatory effect for R19. The Pb2+ removal for R19 increased significantly by 31% (Figs. 4 and 5).
The R2A strains (R3 and R19) were able to remove more metal cations than the LB strains in multi-metal cation solutions (Fig. 6). Metal uptake was the highest for strain R19, in particular Pb2+ and Cu2+, which generally removed larger amounts than Mn2+, Zn2+, Cd2+, and Ni2+ (Fig. 6b). The metal accumulation of strain R19 decreased in the order Pb2+ > Cu2+ > Cd2+ > Ni2+ > Mn2+ > Zn2+. Strain L2 only removed three of the eight metals, which decreased in the order Cu2+ > Ni2+ > Pb2+ (Fig. 6c). None of the strains removed Cr6+ nor As3+ from the multi-metal solution (Fig. 6).
Discussion
The use of bacteria specifically adapted to high concentrations of heavy metals has potential in the remediation of heavy metal-contaminated wastewaters. Since Pb is one of the most extensive environmental contaminants, the isolation of robust bacterial strains having potential for Pb resistance is necessary. In the present study, enrichment of sediment samples from the Saint Clair River led to the isolation of bacteria having significant tolerance for Pb2+. While the high concentration of Pb (NO3)2 in the enrichment culture media made the isolation procedure highly selective, many of the isolates were inhibited by and unable to grow on high Pb (NO3)2 concentrations. The Pb tolerance of the bacterial strains isolated in this study is significantly higher than the Pb tolerance potential of Pseudomonas spp. reported by Lin and Harichand (2011).
In this study, the ability of the Pb-resistant bacterial strains to absorb Pb2+ was examined by their flocculation activity. Twenty of the 97 Pb-resistant strains showed high flocculation activity. It is interesting to note that while the bacterial strains from the LB enrichment culture had higher specific growth rate than bacterial strains from the R2A enrichment cultures (Fig. 1), higher specific growth rate did not reflect higher flocculation activity (Fig. 5). Four strains exhibiting the highest flocculating activity (R3, R19, L2, and L30) were selected. Based on 16S rRNA gene sequence analysis, R3 was identified as Raoultella planticola, R19 as Klebsiella michiganensis, L2 as Providencia rettgeri, and L30 as Serratia sp. (Table S3).
Metal cation removal assays demonstrated that the higher is the flocculation activity of the strain, the higher is the efficiency of metal cation removal. The Pb2+ removal of strain R19 was higher than the other three strains (R3, L2, and L30). The higher efficiency of Pb removal maybe due to the fact that the EPS produced by R19 possess higher number and/or more accessible carboxyl and amide functional groups, which are the most important metal binding sites (as demonstrated by Micheletti et al. (2008) in cyanobacteria).
In solutions that simultaneously contain eight different metal cations, strain R19, the bacterial strain with the highest flocculation activity, had the highest efficiency of Pb removal. While the two R2A strains (R3 and R19) were able to remove six (Pb2+, Cu2+, Mn2+, Zn2+, Cd2+, and Ni2+) of the eight metal cations from the multiple metal solution, the efficiency of metal removal was higher for strain R19 than for strain R3.
In the presence of eight metal cations, the process of cation binding to the strains was in general inhibitory rather than stimulatory, which was expected, as these metal cations need to compete for the binding sites on the cell. Surprisingly, the presence of multiple metal cations had stimulatory effect on the Pb2+ removal efficiency of strain R19. The Pb2+ removed by R19 in the multiple metal solution was 31% higher than in solution containing only Pb2+. The greatest inhibition in Pb2+ removal was observed for strain L2. In general, the bacterial strains had higher selectivity and affinity for Pb2+ both in single- and in multi-metal cation solutions.
The values of the electronegativity of the two metals are 2.18 (for As3+) and 1.66 (for Cr6+) based on Pauling scale. Conferring to these values, the affinities should have followed the order Pb2+ > As3+ > Ni2+ > Cu2+ > Cd2+ > Cr6+ > Zn2+ > Mn2+. It is possible that As3+ and Cr6+ may have generated very stable aqueous complexes or precipitates in the multiple metal solutions, and thus, they became poorly exchangeable with the protons bound to the active sites of the cells and the EPS.
It is clear in this study that the long-term effects of pollutants have led to the emergence of metal and antibiotic-resistant bacteria in the sediments of the Saint Clair River. Although bacterial exposure to metals predates human history, anthropogenic-derived sources of metals represent a major source of contamination in the environment. Notably, a substantial number of reports suggest that metal contamination in natural environments could have an important role in the maintenance and proliferation of antibiotic resistance (Alonso et al. 2001; Summers 2002; Summers et al. 1993). The Pb-resistant strains isolated in this study were found to be resistant to penicillin, tetracycline, novobiocin, neomycin, erythromycin, and chloramphenicol. It is known that many bacterial isolates are resistant to antibiotics (Novick and Roth, 1968). For example, the penicillase plasmid of Staphylococcus aureus carries genes determining resistance to several metal ions including Pb2+, As3+, Cd2+, Zn2+, and Hg2+ (Novick and Roth 1963), indicating that the resistance of bacteria to metals may be mediated by the same plasmid mediating drug resistance. It is assumed that the Pb-resistant strains isolated in this study are thus well adapted to unfavorable conditions by their resistance metals (e.g., Pb) and antibiotics, and these may help in developing an effective process for wastewater treatment by using these strains.
The results of this study show the potential applicability of the recently isolated Pb-tolerant strain R19 in the treatment of Pb-metal-containing solutions or solutions containing single and multiple metals. Because efficient metal removal and growth over a range of metal concentrations under aerobic conditions are advantageous, this organism may be employed for metal remediation in simple reactors or even in situ. The highly selective affinity towards Pb2+ observed for strain R19 suggests its use for the recovery of Pb2+ from multiple metal solutions.
References
Alonso A, Sánchez P, Martínez JL (2001) Environmental selection of antibiotic resistance genes. Environ Microbiol 3:1–9
Babel S, Kurniawan TA (2003) Low-cost adsorbents for heavy metals uptake from contaminated water: a review. J Hazard Mat 97(1–3):219–243. https://doi.org/10.1016/S0304-3894(02)00263-7
Bhaskar PV, Bhosle NB (2006) Bacterial extracellular polymeric substances (EPS) a carrier of heavy metals in the marine food-chain. Environ Int 32:192–198
Bruins MR, Kapil S, Oehme FW (2000) Microbial resistance to metals in the environment. Ecotoxicol Environ Saf 45:198–207
Cleveland LM, Minter ML, Cobb KA, Scott AA, German VF (2008) Lead hazards for pregnant women and children: part 1: immigrants and the poor shoulder most of the burden of lead exposure in this country. Am J Nurs 108:40–49
Demayoa A, Taylora MC, Taylora KW, Hodsonb PV, Hammond PB (1982) Toxic effects of lead and lead compounds on human health, aquatic life, wildlife plants, and livestock. Crit Rev Environ Contr 12(4):257–305. https://doi.org/10.1080/10643388209381698
Difco and BBL (2009) Difco and BBL manual of microbiological media, 2nd edn. BD Diagnostics-Diagnostic System, Sparks 700 p
El-Hendawy HH, Ali Dena A, El-Shatoury EH, Ghanem SM (2009) Bioaccumulation of heavy metals by Vibrio alginolyticus isolated from wastes of iron and steel factory, Helwan, Egypt. Egypt Acad J Biol Sci 1:23–28
Fogarty LR (2007) Bacteria and emerging chemical contaminants in the St. Clair River/Lake St. Clair Basin, Michigan: U.S. Geological Survey Open-File Report 2007–1083, 10
Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Kreig NR, Phillips GB. Manual of Methods for General Bacteriology (1994) Washington (DC): American Society for Microbiology. 791 p
Guangyu Y, Thiruvenkatachari V (2003) Heavy metals removal from aqueous solution by fungus Mucor rouxii. Water Res 37:4486–4496
Hu H, Rabinowitz M, Smith D (1998) Bone lead as a biological marker in epidemiologic studies of chronic toxicity: conceptual paradigms. Environ Health Perspect 106(1):1–8. https://doi.org/10.1289/ehp.981061
Hugenholtz P, Huber T (2003) Chimeric 16S rDNA sequences of diverse origin are accumulating in public databases. Inter J Syst Evol Microbiol 53(1):289–293. https://doi.org/10.1099/ijs.0.02441-0
Lin J, Harichund C (2011) Isolation and characterization of heavy metal removing bacterial bioflocculants. Afr J Microbiol Res 5:559–607
Micheletti E, Colica G, Viti C, Tamagnini P, De Philippis R (2008) Selectivity in the heavy metal removal by exopolysaccharide-producing cyanobacteria. J Appl Microbiol 105:88–94
Moncrieff AA, Koumides OP, Clayton BE, Patrick AD, Renwick AGC, Roberts GE (1964) Lead poisoning in children. Arch Dis Child 39(203):1–13. https://doi.org/10.1136/adc.39.203.1
Murthy S, Bali G, Sarangi SK (2012) Biosorption of lead by Bacillus cereus isolated from industrial effluents. British Biotechnology Journal 2:73–84
Naik MM, Dubey SK (2013) Lead resistant bacteria: lead resistance mechanisms, their applications in lead bioremediation and biomonitoring. Ecotox Environm Saf 98:1–7. https://doi.org/10.1016/j.ecoenv.2013.09.039
Nies DH, Silver S (1995) Ion efflux systems involved in bacterial metal resistances. J Ind Microbiol 14(2):186–199. https://doi.org/10.1007/BF01569902
Novick RP, Roth C (1968) Plasmid-linked resistance to inorganic salts in Staphylococcus aureus. J Bacteriol 95(4):1335–1342
Oest A, Alsaffar A, Fenner M, Azzopardi D, Tiquia-Arashiro SM (2018) Patterns of change in metabolic capabilities of sediment microbial communities along pollution gradients in river and lake ecosystems. Environmental Technology. In review
Pal A, Paul AK (2008) Microbial extracellular polymeric substances: central elements in heavy metal bioremediation. Indian J Microbiol 48(1):49–64. https://doi.org/10.1007/s12088-008-0006-5
Patel D, Gismondi R, Alsaffar A, Tiquia-Arashiro SM (2018) Extracellular enzyme activities in sediments of St. Clair River and Lake Saint Clair, Michigan. Journal of Microbiology. In review
Silver S (1996) Bacteria resistances to toxic metals. Gene 179(1):9–19. https://doi.org/10.1016/S0378-1119(96)00323-X
Summers AO (2002) Generally overlooked fundamentals of bacterial genetics and ecology. Clin Infect Dis 34(s3):S85–S92. https://doi.org/10.1086/340245
Summers AO, Wireman J, Vimy MJ, Lorscheider FL, Marshall B, Levy SB, Bennett S, Billard L (1993) Mercury released from dental silver fillings provokes an increase in mercury-resistant and antibiotic resistant bacteria in oral and intestinal floras of primates. Antimicrob Agents Chemother 37(4):825–834. https://doi.org/10.1128/AAC.37.4.825
Weisburg WW, Barns SM, Pelletier DA, Lane DJ (1991) SSU ribosomal DNA amplification for phylogenetic study. J Bacteriol 173(2):697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Zhang ZQ, Lin B, Xia SQ, Wang XJ, Yang AM (2007) Production and application of a novel bioflocculant by multiple microorganism consortia using brewery wastewater as carbon source. J Environ Sci 19(6):667–673. https://doi.org/10.1016/S1001-0742(07)60112-0
Zouboulis AI, Loukidou MX, Matis KA (2003) Biosorption of toxic heavy metals from aqueous solutions by bacteria strains isolated from metal-polluted soils. Process Biochem 39:909–916
Acknowledgements
This research was funded by the University of Michigan-Dearborn Office of Sponsored Programs. We thanked the anonymous reviewers for critically reading the manuscript and suggesting substantial improvements.
Author information
Authors and Affiliations
Contributions
Tiquia-Arashiro conceived and designed the study. Bowman performed the experiments and analyzed the data with Patel, Sanchez, Xu, and Alsaffar. Alsaffar collected the sediment samples and field data. Tiquia-Arashiro, Bowman, Patel, and Sanchez wrote the manuscript, which was completed with input from Xu and Alsaffar
Corresponding author
Ethics declarations
This article does not contain any experiments involving human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 114 kb)
Rights and permissions
About this article
Cite this article
Bowman, N., Patel, D., Sanchez, A. et al. Lead-resistant bacteria from Saint Clair River sediments and Pb removal in aqueous solutions. Appl Microbiol Biotechnol 102, 2391–2398 (2018). https://doi.org/10.1007/s00253-018-8772-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8772-4