Abstract
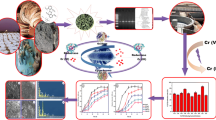
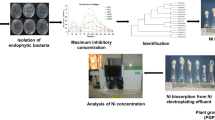
The potential of indigenous bacterial strains to accumulate three metals (Cr, Ni, Pb) was exploited here to remediate the polluted environment. In the present study, metal resistance profiles identified three most potential isolates which could tolerate 700–1000 μg/ml of Ni, 500–1000 μg/ml of Cr, and 1000–1600 μg/ml of Pb. These three bacterial strains were identified as Stenotrophomonas sp. MB339, Klebsiella pneumoniae MB361, and Staphylococcus sp. MB371. UV-Visible and atomic absorption spectrophotometric (AAS) analysis revealed gradual increase in percentage accumulation with increase in time due to increased biomass. Quantitative assessments exhibited maximum removal of Cr (83.51%) by Klebsiella pneumoniae MB361, Pb (85.30%), and Ni (48.78%) by Stenotrophomonas MB339, at neutral pH and 37 °C, whereas Staphylococcus sp. MB371 sorbed 88.33% of Pb at slightly acidic pH. The present study therefore supports the effective utilization of indigenous bacteria for comprehensive treatment of metal-rich industrial effluents.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Industrial revolution has greatly increased the heavy metals, which is a big hazard for human health. Anthropogenic activities convert such metals into various forms that are highly toxic and even persist in the environment for longer duration (Banerjee et al. 2015). Major ecosystem source of such metals is industrial effluents that are being discharged by steel manufacturing, electroplating, chemical processing, and leather tanning units into the surrounding land and water resources (Tian et al. 2012). Although small concentrations of some metals such as iron, zinc, copper, and manganese are required as essential micronutrients for rapid growth, maintenance of protein structure, and act as enzyme co-factor by various life forms, others are considered most toxic and persistent environmental pollutants causing multiple cellular alterations (Deeb and Altalhi 2009; Reis et al. 2014).
Immense release of heavy metals during the last few decades has increased the world concerns about their removal (Ren et al. 2009). Industries discharge their polluted waste rich in heavy metals such as Pb, Cr, Ni, and Cd into the nearby streams. Such harmful wastewater is being supplied to agricultural land, causing long-term effects on plants, animals, and humans health (Takahashi et al. 2012; Olaniran et al. 2013). For instance, elevated concentrations of lead and cadmium intake in human have resulted in many kidney disorders and disturbance in endocrine system (Jiménez-Ortega et al. 2012). Moreover, increased nickel amount has been linked to various respiratory and skin problems (Thyssen et al. 2013). To resolve these problems, several methods have been introduced to remove these contaminants from environmental compartments (Bibi et al. 2014; Oyewo et al. 2018).
Although various physicochemical techniques such as chemical precipitation, ion exchange, electrochemical treatment, membrane technologies, and adsorption on activated sludge (Shim et al. 2014; Eiband et al. 2014; Gupta et al. 2015) have been developed for eradicating excess amount of heavy metals (Beveridge et al. 1997; Wang and Chen 2009), these methodologies are highly expensive, less environment friendly, and sometimes ineffective at low concentrations. Moreover, these processes require many reagents but result in incomplete metal removal and generate highly toxic sludge (Sibi 2016), whereas bioremediation processes involve the use of bacteria, fungi, or plants for the removal of pollutants in a natural and economical way (Marc et al. 2012). Research reports addressing bioremediation processes reflect the potential application of living or dead biomass to remove considerable amounts of these toxic metals from different compartments of an ecosystem (OK et al. 2007; El-Gendy and El-Bondkly 2016). Similarly, metal sorption could be considered as it involves rapid surface binding of metal ions to either bacteria or any other life form (Bayramoglu and Arıca 2008; Huang et al. 2013). While bioaccumulation is an active mode of metal uptake by living cells, this process generally depends upon metal speciation and its bioavailability. The cell’s ability to bioaccumulate largely depends upon their metabolic activities and various physiological, biochemical, genetic, and structural adaptations depending upon the surrounding ecosystem (Algarra et al. 2014). Aditionally, environmental conditions such as temperature, pH, and biomass concentration always have a significant impact upon the metal ions uptake capabilities of living cells (Uslu and Tanyol 2006; Ojuederie and Babalola 2017).
Industrial effluents served as a potential source for isolating microbes capable of tolerating and proficiently remediation contaminants. Bacterial cells have multiple surface binding sites for metal cations and possess metal-resistant phenotypes which support their use for environmental protection. For instance, Escherichia hermannii and Enterobacter cloacae uptake Cd and Ni, Bacillus thuringiensis can accumulate copper, Bacillus cereus sorbed lead proficiently, and Pseudomonas aeruginosa act as metallophore to eradicate metals Cd and Pb from the effluent (Selvi et al. 2012; Poornima et al. 2014: McFarlane et al. 2018).
In the present study, bacterial isolates were used to check their metal remediation potential. Many strains showed varied abilities to resist metals. Among these, Klebsiella pneumoniae MB361, Stenotrophomonas sp. MB339, and Staphylococcus sp. MB371 showed resistance towards higher concentration of metals which were selected for evaluating their bioaccumulation abilities.
Materials and methods
Isolation and screening of isolates
Industrial effluent samples were collected from Hattar Industrial State, Pakistan. The physicochemical analysis of samples was done using standard techniques and stored at 4 °C for isolation of bacterial strains. Isolation of bacteria was done by performing standard spread plate method of different industrial effluent samples. Initially 50 μl of each effluent sample was plated with mixed metals (Pb, Cr, and Ni) supplemented nutrient agar plates, incubated at 37 °C for 24 h. Further screening was done by plating the selected colonies on M9 minimal medium plates with different concentrations of selected metal salts, i.e., PbCl2, K2CrO4, and NiCl2. The minimal agar medium M9 of composition Na2HPO4 (6 g), KH2PO4 (3 g), NH4Cl (1 g), and NaCl (0.5 g) per liter of distilled water was prepared and the pH was adjusted to 7.04. After autoclaving, CaCl2 (0.1 M), MgSO4 (1 M), and casein hydrolysate (5 g) were added to the medium under sterilized conditions. The freshly grown culture was inoculated in M9 broth with 50, 100, 150, 200, and 250 μg/ml of each metal and incubated in a rotary shaker (150 rpm) at 30 °C for 24 h. The growth was determined by checking optical density of the medium at 600 nm using spectrophotometer. The growth of each strain declined with increasing concentration of metal salts. However, the isolated MB339, MB361, and MB371 showed maximum relative growth of more than 80% at 200 μg/ml. Thus, these three most promising strains were purified by three rounds of single colony streaking.
Morphological, physiological, and biochemical characterization
The cell and colony morphology of purified colonies grown on nutrient agar plates was studied. Furthermore, investigation of culture growth pattern at 25, 30, 37, and 45 °C and in the medium of pHs 5 to 11 was made in nutrient broth. Biochemical tests, including catalase, oxidase, and nitrate reductase, were performed using standard methods. In addition, isolates were tested for enzymes tryptophan deaminase, lysine decarboxylase, arginine dihydrolase production, and utilization of sugars glucose, sucrose, maltose etc. The biochemical tests were performed following Bergey’s manual of Descriptive Bacteriology (Holt et al. 1994).
DNA extraction and identification of bacterial strains
These isolates were then identified through 16S rDNA sequencing. The bacteria were cultured in LB broth at 30 °C in shaking incubator for 24 h. After extraction of genomic DNA, 16S rRNA gene was amplified by PCR using universal primers: 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′). The PCR product was then purified with PureLink™ Quick PCR Purification Kit (Invitrogen) and run on 0.8% agarose gel as described by Aslam et al. (2018). The 16S rRNA gene sequences obtained were compared with known sequences from the NCBI database with BLASTN algorithm. These sequences were aligned with ClustalW and phylogenetic tree was constructed by the neighbor-joining method using Molecular Evolutionary Genetic Analysis MEGA 6 software (Tamura et al. 2013).
Heavy metal accumulation studies
The selected isolates MB339, MB361, and MB371 were grown in 200 μg/ml of each metal-supplemented broth at 37 °C with 150 rpm. After every 24-h incubation, bacterial cells were harvested by centrifugation and suspended in 1 ml of distilled water. The metal content was determined after acid dissolution of bacterial cell as described by Ganje and Page (1974). Concentrations of selected heavy metals accumulated were measured in μg/ml (1 ppm) units at specific wavelengths 217 nm for Pb, 232 nm for Ni, and 357.9 nm for Cr for each metal by flame atomic absorption spectrophotometer (Spectra AA 220, Australia).
Percentage bioaccumulation was then calculated as follows:
Effect of various factors on metal (Cr, Ni, and Pb) accumulation
Bioaccumulation potential of three isolates was assessed at five different temperatures 25, 30, 37, 40, and 45 °C and pHs 6, 7, and 8. Nutrient broth (10 ml) was prepared in sterilized conditions supplemented with 200 μg/ml of Cr, Ni, and Pb and inoculated with 50 μl of fresh bacterial cultures.
Effects of six different inoculum sizes (50, 100, 200, 300, 400, and 500 μl) and five concentrations between 100 and 500 μg/ml of each metal were also used. Furthermore, impact of incubation time (24, 48, and 72 h) in shaking incubator at 150 rpm at 37 °C was also studied. The percentage of metal accumulated was analyzed through AAS.
Statistical analysis
All experiments were performed in triplicate. The results are expressed as standard error of mean (±SEM).
Results and discussion
The industrial effluent samples collected were apparently and chemically different from one another. The color observed was blank and yellow with pH value 7.1 and 9.4 having electrical conductivity 653 and 1311 μS/s. Whereas the total hardness, dissolved, and suspended solid concentrations were a little above the permissible limits.
Isolation and characterization of metal-resistant bacteria
In the present study, bacterial isolates that were obtained using selected mixed metals (Pb, Cr, and Cd) supplemented nutrient agar medium were further tested for tolerance against twelve different metals. The bacterial cultures were purified by several rounds of single colony streaking and maintained on nutrient agar medium. Strains MB339 and MB361 were gram-negative, while MB371 was gram-positive. Isolates showed optimal growth between 30 and 37 °C. Isolates MB361 and MB371 were neutrophilic whereas MB339 exhibited slightly alkalophilic nature (Fig. 1). After the screening, isolates MB339, MB361, and MB371 tolerating more than 1000 μg/ml of Cr, Ni, and Pb were finally selected for carrying out bioaccumulation studies. The lethal concentration LC50 values for each metal (Pb, Cr, Ni) were assessed using eight different concentrations from 100 to 800 μg/ml. With increasing concentration of each metal, bacterial growth was gradually decreasing. In the case of strain MB339, the LC50 calculated for metal (Pb, Cr) was 400 μg/ml and 300 μg/ml for Ni. For strain MB361, 300 μg/ml for Cr and Ni, while 400 μg/ml for Pb was found lethal concentration. While LC50 value for bacterial strain MB371 was observed at 400 μg/ml for Pb and 300 μg/ml for Ni and Cr, research reports suggested that gram-negative bacteria always had higher metal uptake capacity than gram-positive because active metal-binding groups such as carboxy, sulphuryl, phosphorous, and amino are present in the cell wall (Thomas and Rice 2014). A previous study revealed that gram-negative Escherichia coli K-12 and Pseudomonas aeruginosa more efficiently biosorbed Cu, Cr, and Ni as compared with gram-positive strains Micrococcus luteus (Churchill et al. 1995). However, Kaewehai and Prasertson (2002) demonstrated that Enterobacter agglomerrans SM 38, Bacillus subtilis SM 29, and Bacillus subtilis WD 90 proficiently accumulate cadmium and nickel.
On the basis of biochemical characterization, bacterial strains were distinctly different from one another. Both MB361 and MB371 showed positive Voges-Proskauer test, nitrate reduction, catalase, and oxidase activity. These two strains were able to utilize citrate with the ability to produce urease and tryptophan deaminase. These could not hydrolyze gelatin or produce indole, but could utilize all types of sugars including glucose, mannitol, inositol, sorbitol, rhamnose, saccharose, and melibiose.
On the basis of 16S rRNA, strains were identified as Stenotrophomonas sp. MB339 (KP723528), Klebsiella sp. MB361 (KP723532), and Staphylococcus sp. MB371 (KP723530). Phylogenetically, neighbor-joining algorithm showed high similarity of strain MB339 with Stenotrophomonas sp. as described in Aslam et al. (2018). Similarly, in the case of Klebsiella pneumoniae MB361, the strain showed 99% similarity with other Klebsiella pneumoniae MB369, Klebsiella pneumoniae nucleomorph CEES16, and others given in Fig. 2. The strain Staphylococcus sp. MB371 was found to be highly similar (99%) to Staphylococcus sp. SQ7-5-9, A9, BAB-4167 and different strains of Staphylococcus sciuri.
Effect of incubation temperature on metal bioaccumulation
The results obtained from experiments performed at different incubation temperatures showed that all the three strains accumulated Cr maximally, ranging from 37 to 40 °C (Fig. 3). Most effectively Stenotrophomonas sp. MB339 accumulated 65.85% Cr at 37 °C. In the case of Ni, all three isolates achieved optimum metal uptake at 30 °C which gradually decreased with increasing temperature up to 45 °C. Same trend of percentage Pb accumulation was recorded by MB339, MB361, and MB371 strains. Temperature is known to affect the stability of the cell wall and its configuration. Bacterial strains normally remove maximum metal at 30 °C like Acromobacter sp. and Pseudomonas stutzeri could uptake 82% copper effectively (Olmezoglu et al. 2012). On contrary to this, higher temperatures generally lead to increase in metabolic activity along with an increase in energy of the system that would result in active uptake of metals. For instance, the rate of Cr (VI) removal by S.cerevisiae was very high at 45 °C (Goyal et al. 2003; Banerjee et al. 2015).
Effect of pH on metal bioaccumulation
Under natural conditions, the presence of heavy metals in wastewater greatly affects its pH (Abourached et al. 2014). Therefore, experiments were performed in order to evaluate the effect of varying pH on metal uptake. The outcomes of this study demonstrated that percentage bioaccumulation of Cr and Ni was increased with increasing pH (Fig. 4). Strain MB339 showed maximum 85.3% of Pb being sorbed at pH 7, while acidic pH significantly reduced Ni accumulation. Potential of cells to uptake metal ions varies with the type of adsorbent (biomass) and adsorbate (metal ions). At low pH (below 7), bioaccumulation percentage decrease due to hydronium ions occupying the binding sites offering repulsion in contrast to higher pH values where more ligands carrying negative charge are exposed providing space for metal ions to biosorb on cell surfaces (Dexilin et al. 2007). The decrease in removal of Pb(II) in alkaline medium is owing to the formation of Pb(OH)2 precipitates (Colak et al. 2011; Sahu et al. 2015). Bacillus thuringiensis OSM29 had greater sorption of Cd and Cu at low pH values (Oves et al. 2013). On contrary to this, Masood and Malik (2011) observed more Cr removal by Bacillus sp. FM1 because of increasing metal anions as in the case of Cr(VI) in the form of chromate ion (CrO4−2).
Effect of initial metal concentration on bioaccumulation
Pb, Ni, and Cr accumulation of bacterial isolates was observed at five different metal concentrations, i.e., 100, 200, 300, 400, and 500 μg/ml. From the results, it can be inferred that percentage of metal accumulation of selected bacterial isolates decreased with increasing concentration of respective metals (Fig. 5). Staphylococcus sp. MB371 could efficiently accumulate 71.45% of Cr; however, a little less than that (68.54% and 65.98%) was observed in Stenotrophomonas sp. MB339 and Klebsiella pneumoniae MB361 respectively, when 100 μg/ml concentration of Cr was provided in the medium. This percentage decreased in a regular manner with increasing amount of Cr. More than 50% of 100 μg/ml Ni and Pb were accumulated by three strains which then reduced to less than 10% when the provided concentration was raised to 500 μg/ml. The decrease in bioaccumulation with increasing metal concentration may be attributed to saturation of adsorption sites (Huang et al. 2014). At a lower concentration, all the metal ions in solution could interact with the available binding sites resulting in higher bioaccumulation (Pandiyan and Mahendradas 2011; Ergul-Ulger et al. 2014).
Effect of inoculum size on metal bioaccumulation
All bacterial strains exhibited lowest chromium accumulation at 50 μl, whereas highest chromium accumulation was observed at 500 μl. Klebsiella pneumoniae MB361 exhibited 18.24 and 83.51% accumulation in 50 and 500 μl inoculum size, respectively (Fig. 6). Ni uptake was almost doubled when inoculum was increased from 50 (13.97%) to 500 μl (26.97%) by Stenotrophomonas sp. MB339 after 24-h incubation. A maximum of 56.7% Pb was removed by Stenotrophomonas whereas 48.17% and 38.46% were accumulated by Klebsiella pneumoniae MB361 and Staphylococcus sp. MB371 respectively, with 500 μl of inoculum. Effect of biomass on the biosorption process was investigated by other researchers for both B. cereus and B. pumilus and it was found that Pb(II) uptake increased as the inoculum size increased (Colak et al. 2011). Previous studies revealed that augmented biomass resulted in an increase in biosorption due to enlarged surface area of the biosorbent, which in turn enhances the number of binding sites (Devika et al. 2014).
Effect of incubation time on bioaccumulation
Contact time is of immense significance in biosorption process (Arivalagan et al. 2014). Results depicted that as the contact time increased, there was a drastic increase in capability of isolates to bioaccumulate metals (Cr, Ni, and Pb). Highest potential to remove 200 μg/ml Cr from the medium was shown by Stenotrophomonas MB339 after 72 h. More than 30% of Cr removal took place by other two strains Klebsiella pneumoniae MB361 and Staphyloccocus sp. MB371 after 24 h and 72 h that rose to 64.52 and 65.24%, respectively (Fig. 7). A similar pattern was observed in all three strains for Ni with maximum accumulation 63.19% by Stenotrophomonas sp. MB339. Strain MB339 accumulated 27% of initial concentration of Pb after 24 h which increased to 72% with an increase in incubation time, i.e., up to 72 h.
The high rate of bioaccumulation was due to the greater affinity of free metal ions to the binding sites available at adsorbent and after attaining equilibrium, the percentage biosorption started to decrease. In this context, Gabr et al. (2008) concluded that initial short contact time is relatively more important to attain a higher rate of biosorption as in the case of Ni and Cd. Odokuma and Series 2012) reported a rise in metal uptake with increase in contact time for Bacillus sp., Pseudomonas sp., and Aeromonas sp. Initially the cells are not dividing rapidly or in lag phase, hence exhibit less binding sites that is why metal uptake by isolates was of no consequence (Hossain and Aditya 2013). Following this period, metal uptake as well as cell growth is amplified depending upon the availability of active binding sites. Similarly, slower rate of biosorption at short incubation time become doubled with increase in contact time up to certain limit (Abd-Alla et al. 2012; Barka et al. 2013).
Conclusion
Use of microorganisms for removal of toxic contaminants from environment is a method of potential application. In this study, the ability of bacteria to remediate heavy metals from aqueous solution was determined. Klebsiella pneumoniae MB361 exhibited the highest accumulation (83.51%) with initial concentration of 200 μg/ml Cr at 37 °C for 24 h, while 85.30% of Pb and 48.78% of Ni were accumulated by Stenotrophomonas sp. MB339 at neutral pH, 37 °C and with maximum inoculum size. Present investigation also highlighted the significance of various factors in the process of bioaccumulation. Bacteria could be efficient, promising, and prospective microorganisms for the removal of Ni, Cr, and Pb from aqueous solution, owing to their significant biosorption capacity and environmental friendly nature. These findings serve as a foundation for further exploitation of bacterial ability to remediate metals.
References
Abd-Alla MH, Morsy FM, El-Enany AWE, Ohyama T (2012) Isolation and characterization of a heavy-metal-resistant isolate of Rhizobium leguminosarum bv. viciae potentially applicable for biosorption of Cd2+ and Co2+. Int Biodeterior Biodegrad 67:48–55
Abourached C, Catal T, Liu H (2014) Efficacy of single-chamber microbial fuel cells for removal of cadmium and zinc with simultaneous electricity production. Water Res 51:228–233
Algarra M, Vázquez MI, Alonso B, Casado CM, Casado J, Benavente J (2014) Characterization of an engineered cellulose based membrane by thiol dendrimer for heavy metals removal. Chem Eng J 253:472–477
Arivalagan P, Singaraj D, Haridass V, Kaliannan T (2014) Removal of cadmium from aqueous solution by batch studies using Bacillus cereus. Ecol Eng 71:728–735
Aslam F, Yasmin A, Thomas T (2018) Essential gene clusters identified in Stenotrophomonas MB339 for multiple metal/antibiotic resistance and xenobiotic degradation. Curr Microbiol 75(11):1484–1492
Banerjee G, Pandey S, Ray AK, Kumar R (2015) Bioremediation of heavy metals by a novel bacterial strain Enterobacter cloacae and its antioxidant enzyme activity, flocculants production, and protein expression in presence of lead, cadmium, and nickel. Water Air Soil Pollut 226:91
Barka N, Abdennouri M, El Makhfouk M, Qourzal S (2013) Biosorption characteristics of cadmium and lead onto eco-friendly dried cactus (Opuntia ficus indica). J Environ Chem Eng 1:144–149
Bayramoglu G, Arıca MY (2008) Removal of heavy mercury (II), cadmium (II) and zinc (II) metal ions by live and heat inactivated Lentinus edodes pellets. Chem Eng J 143:133–140
Beveridge T, Hughes M, Lee H (1997) Metal-microbe interactions: contemporary approaches. Adv Microb Physiol 38:177–243
Bibi A, Farooq U, Mirza N, Khan A, Sarwar R, Alam A, Mahmood Q (2014) Excessive chromium may cause dietary toxicity in parsley (Petroselinum crispum). Toxicol Environ Chem 96(2):287–295
Churchill S, Walters J, Churchill P (1995) Sorption of heavy metals by prepared bacterial cell surfaces. J Environ Eng 121:706–711
Colak F, Atar N, Olgun A (2011) Biosorption of lead from aqueous solutions by Bacillus strains possessing heavy-metal resistance. Chem Eng J 173:422–428
Deeb BE, Altalhi AD (2009) Degradative plasmid and heavy metal resistance plasmid naturally coexist in phenol and cyanide assimilating bacteria. Am J Biochem Biotechnol 5(2):84–93
Devika MV, Thatheyus AJ, Ramya D (2014) Bioremoval of nickel using Pseudomonas aeruginosa. Ann Res Rev Biol 4(3):538–546
Dexilin M, Congeevaram S, Dhanarani S (2007) Biosorption of chromium and nickel by heavy metal resistant fungal and bacterial isolates. J Hazard Mater 146:270–277
Eiband MM, Kamélia CDA, Gama K, de Melo JV, Martínez-Huitle CA, Ferro S (2014) Elimination of Pb2+ through electrocoagulation: applicability of adsorptive stripping voltammetry for monitoring the lead concentration during its elimination. J Electroanal Chem 717:213–218
El-Gendy MM, El-Bondkly AM (2016) Evaluation and enhancement of heavy metals bioremediation in aqueous solutions by Nocardiopsis sp. MORSY1948, and Nocardia sp. MORSY2014. Braz J Microbiol 47:571–586
Ergul-Ulger Z, Ozkan AD, Tunca E, Atasagun S, Tekinay T (2014) Chromium (VI) biosorption and bioaccumulation by live and acid-modified biomass of a novel Morganella morganii isolate. Sep Sci Technol 49(6):907–914
Gabr R, Hassan S, Shoreit A (2008) Biosorption of lead and nickel by living and non-living cells of Pseudomonas aeruginosa ASU 6a. Int Biodeterior Biodegrad 62:195–203
Ganje TJ, Page AL (1974) Rapid acid dissolution of plant tissue for cadmium determination by atomic absorption spectrophotometry. Atom Absorpt Newslett 13(6):131–134
Goyal N, Jain SC, Banerjee UC (2003) Comparative studies on the microbial adsorption of heavy metals. Adv Environ Res 7(2):311–319
Gupta VK, Nayak A, Agarwal S (2015) Bioadsorbents for remediation of heavy metals: current status and their future prospects. Environ Eng Res 20(1):1–18
Holt JG, Krieg NR, Sneath PH, Staley JT, Williams ST (1994) Group 19. Regular, nonsporing Gram-positive rods. Bergey’s manual of determinative bacteriology, 9th edn. Williams & Wilkins Co., Baltimore, p 566
Hossain A, Aditya G (2013) Cadmium biosorption potential of shell dust of the fresh water invasive snail Physa acuta. J Environ Chem Eng 1:574–580
Huang F, Dang Z, Guo CL, Lu GN, Gu RR, Liu HJ, Zhang H (2013) Biosorption of Cd (II) by live and dead cells of Bacillus cereus RC-1 isolated from cadmium contaminated soil. Colloids Surf B 107:11–18
Huang F, Guo CL, Lu GN, Yi XY, Zhu LD, Dang Z (2014) Bioaccumulation characterization of cadmium by growing Bacillus cereus RC-1 and its mechanism. Chemosphere 109:134–142
Jiménez-Ortega V, Barquilla PC, Fernández-Mateos P, Cardinali DP, Esquifino AI (2012) Cadmium as an endocrine disruptor: correlation with anterior pituitary redox and circadian clock mechanisms and prevention by melatonin. Free Radic Biol Med 53(12):2287–2297
Kaewehai S, Prasertson P (2002) Biosorption of heavy metals. J Sci Technol 24(3):422–430
Marc M, Burton JP, Reid G (2012) Bioremediation and tolerance of humans to heavy metals through microbial processes. Appl Environ Microbiol 78(18):6397–6404
Masood F, Malik A (2011) Biosorption of metal ions from aqueous solution and tannery effluent by Bacillus sp. FM1. J Environ Sci Health A 46:1667–1674
McFarlane JS, Davis CL, Lamb AL (2018) Staphylopine, pseudopaline and yersinopine dehydrogenases: a structural and kinetic analysis of a new functional class of opine dehydrogenase. J Biol Chem 293:8009–8019
Odokuma LO and Series IL (2012) The genius in microbes: an indispensible for the management of xenobiotic mediated environmental flux. University of Port Harcourt Inaugral Lecture, pp 87
Ojuederie O, Babalola O (2017) Microbial and plant-assisted bioremediation of heavy metal polluted environments. A review. Int J Environ Res Public Health 14(12):1504
Ok Y, Yang J, Zhang Y, Kim S, Chung D (2007) Heavy metal adsorption by a formulated zeolite-Portland cement mixture. J Hazard Mater 147:91–96
Olaniran AO, Balgobind A, Pillay B (2013) Bioavailability of heavy metals in soil: impact on microbial biodegradation of organic compounds and possible improvement strategies. Int J Mol Sci 14:10197–10228
Olmezoglu E, Herand B, Oncel M, Tunc K, Ozkan M (2012) Copper bioremoval by novel bacterial isolates and their identification by 16S rRNA gene sequence analysis. Turk J Biol 36:469–476
Oves M, Khan M, Zaidi A (2013) Biosorption of heavy metals by Bacillus thuringiensis strain OSM29 originating from industrial effluent contaminated north Indian soil. Saudi J Biol Sci 20:121–129
Oyewo OA, Agboola O, Onyango MS, Popoola P and Bobape MF (2018) Current methods for the remediation of acid mine drainage including continuous removal of metal from wastewater and mine dump. In Bio-Geotechnologies for mine site rehabilitation. Elsevier, pp 103–114
Pandiyan S, Mahendradas D (2011) Application of bacteria to remove Ni (II) ions from aqueous solution. Eur J Sci Res 52:345–358
Poornima M, Kumar RS, Thomas PD (2014) Isolation and molecular characterization of bacterial strains from tannery effluent and reduction of chromium. Int J Curr Microbiol App Sci 3(4):530–538
Reis KC, Silva CF, Duarte WF, Schwan RF (2014) Bioaccumulation of Fe3+ by bacteria isolated from soil and fermented foods for use in bioremediation processes. Afr J Microbiol Res 8(26):2513–2521
Ren WX, Li PJ, Geng Y, Li XJ (2009) Biological leaching of heavy metals from a contaminated soil by Aspergillus niger. J Hazard Mater 167:164–169
Sahu S, Roy B, Varghese TA (2015) Bioremediation of lead using indigenous bacterial consortium isolated from tannery effluent-a bioreactor based approach. Int J Curr Microbiol App Sci 4(7):919–927
Selvi AT, Anjugam E, Devi RA, Madhan B, Kannappan S, Chandrasekaran B (2012) Isolation and characterization of bacteria from tannery effluent treatment plant and their tolerance to heavy metals and antibiotics. Asian J Exp Biol Sci 3(1):34–41
Shim HY, Lee KS, Lee DS, Jeon DS, Park MS, Shin JS, Chung DY (2014) Application of electrocoagulation and electrolysis on the precipitation of heavy metals and particulate solids in washwater from the soil washing. J Agri Chem Environ 3(4):130
Sibi G (2016) Biosorption of chromium from electroplating and galvanizing industrial effluents under extreme conditions using Chlorella vulgaris. Green Energy Environ 1(2):172–177
Takahashi C, Turner A, Millward G, Glegg G (2012) Persistence and metallic composition of paint particles in sediments from a tidal inlet. Marine Poll Bull 64:133–137
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30(12):2725–2729
Thomas KJ III, Rice CV (2014) Revised model of calcium and magnesium binding to the bacterial cell wall. Biometals 27(6):1361–1370
Thyssen JP, Gawkrodger DJ, White IR et al (2013) Coin exposure may cause allergic nickel dermatitis: a review. Contact Dermatitis 68:148–157
Tian H, Lu L, Cheng K, Hao J, Zhao D, Wang Y, Jia W, Qiu P (2012) Anthropogenic atmospheric nickel emissions and its distribution characteristics in China. Sci Total Environ 417–418:148–157
Uslu G, Tanyol M (2006) Equilibrium and thermodynamic parameters of single and binary mixture biosorption of lead (II) and copper (II) ions onto Pseudomonas putida: effect of temperature. J Hazard Mater B 135:87–93
Wang J, Chen C (2009) Biosorbents for heavy metals removal and their future. Biotechnol Adv 27:195–226
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aslam, F., Yasmin, A. & Sohail, S. Bioaccumulation of lead, chromium, and nickel by bacteria from three different genera isolated from industrial effluent. Int Microbiol 23, 253–261 (2020). https://doi.org/10.1007/s10123-019-00098-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-019-00098-w