Abstract
The bleomycins (BLMs) are important clinical drugs extensively used in combination chemotherapy for the treatment of various cancers. Dose-dependent lung toxicity and the development of drug resistance have restricted their wide applications. 6′-Deoxy-BLM Z, a recently engineered BLM analogue with improved antitumor activity, has the potential to be developed into the next-generation BLM anticancer drug. However, its low titer in the recombinant strain Streptomyces flavoviridis SB9026 has hampered current efforts, which require sufficient compound, to pursue preclinical studies and subsequent clinical development. Here, we report the strain improvement by combined UV mutagenesis and ribosome engineering, as well as the fermentation optimization, for enhanced 6′-deoxy-BLM production. A high producer, named S. flavoviridis G-4F12, was successfully isolated, producing 6′-deoxy-BLM at above 70 mg/L under the optimized fermentation conditions, representing a sevenfold increase in comparison with that of the original producer. These findings demonstrated the effectiveness of combined empirical breeding methods in strain improvement and set the stage for sustainable production of 6′-deoxy-BLM via pilot-scale microbial fermentation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The bleomycins (BLMs) (Umezawa et al. 1966), together with tallysomycins (TLMs) (Kawaguchi et al. 1977) and zorbamycin (ZBM) (Wang et al. 2007), are a family of anticancer glycopeptides with oxidative DNA/RNA cleavage activity (Burger 1998). The distinctive low myelo- and immunosuppression properties of BLMs make them effective broad-spectrum chemotherapeutic drugs, particularly in combination chemotherapy widely used for the treatment of malignant tumors (Galm et al. 2005), such as uterine cervical cancer (Panetta et al. 1999) and metastatic testicular cancer (Einhorn 2002). However, BLMs could impair the endothelial cells of lung vasculature and cause pneumonitis, which may further progress into lung fibrosis (Sleijfer 2001). Such dose-limiting pulmonary toxicity, as well as the continually emerged BLM resistance in tumor cells (Wang et al. 2013), has greatly restricted the clinical applications of BLMs.
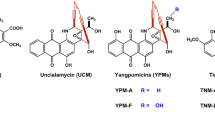
Over the past decades, substantial efforts have been devoted to the generation of BLM analogues, through congener isolation (Chen et al. 2008), organic synthesis (Leitheiser et al. 2003), and combinatorial biosynthesis (Galm et al. 2008; Hindra et al. 2017; Huang et al. 2012; Tao et al. 2010; Yang et al. 2017), aiming for the development of next-generation BLM anticancer drugs with improved efficacy and/or lower lung toxicity. Among the BLM analogues known to date, 6′-deoxy-BLM Z (6′-DO-BLM Z) (Fig. 1) exhibited potent DNA cleavage activity (Hindra et al. 2017; Huang et al. 2012) and better antitumor activity (Chen et al. 2017). Produced by the recombinant strain Streptomyces flavoviridis SB9026, 6′-DO-BLM Z structurally combines the peptide-polyketide backbone of BLMs with the C-terminal amine and the disaccharide moieties of ZBM (Fig. 1), reflecting its complex biosynthetic origin resulted from extensive cross-talks between the BLM and ZBM biogenetic machineries. As a promising lead, the low titer of 6′-DO-BLM Z (less than 10 mg/L) in the original strain SB9026 has limited its adequate supply and hampered the efforts for further preclinical studies or clinical applications.
Titer improvement is one of the key issues for the development of microbial natural products. Rational pathway engineering has great potential to boost the production of target compounds (Chen et al. 2010; Chen et al. 2011; Yu et al. 2010), but its successful application depends on a comprehensive knowledge of biosynthetic pathways and the amenability of producers to genetic modifications. Empirical medium optimization (Yang et al. 2011; Zhang et al. 2010) and mutagenesis (Zhu et al. 2010) require little prior understanding to effect strain improvement, but the processes are tedious and inefficient. On the other hand, antibiotic-induced ribosome engineering could generate structural and functional variations of ribosome and RNA polymerase, such as streptomycin and paromomycin resistance mutations at 30S ribosomal protein S12 (rpsL gene) (Okamoto-Hosoya et al. 2000; Shima et al. 1996), and rifamycin resistance mutation at RNA polymerase β-subunit (rpoB gene) (Xu et al. 2002), which lead to the transcriptional and translational modulations and consequently regulate the metabolic network to stimulate the production of secondary metabolites (Ochi et al. 2004). Hence, ribosome engineering has been regarded as an effective approach to achieve the strain improvement (Ochi 2017) and overproduction of various antibiotics (Shentu et al. 2016; Tanaka et al. 2009).
In this paper, the strain improvement by combining UV mutagenesis and ribosome engineering, as well as fermentation optimization, was conducted for the overproduction of 6′-DO-BLM Z. A high-producing strain S. flavoviridis G-4F12 was isolated, which produced 6′-DO-BLM Z with a titer of over 70 mg/L under the optimized fermentation conditions, about sevenfold increase comparing to the original titer. Our work has provided the basis for pilot-scale production of 6′-DO-BLM Z via practical microbial fermentation, and facilitating further drug development.
Materials and methods
Strain and media
S. flavoviridis SB9026 (CCTCC M2011292) was inoculated on modified ISP4 (mISP4) solid medium (10 g/L soluble starch, 1 g/L MgSO4•7H2O, 1 g/L K2HPO4, 4 g/L MgCl2, 2 g/L (NH4)2SO4, 2 g/L CaCO3, 1 g/L tryptone, 0.5 g/L yeast extract, 0.001 g/L MnCl2•7H2O, 0.001 g/L FeSO4•7H2O and 15 g/L agar, pH 7.0), supplemented with 50 mg/L apramycin, and grown at 30 °C till sporulation. After 8 to 10 days growth, the collected spores were suspended in 20% glycerol, filtered, and stored at − 80 °C. Original seed and production media for fermentation of SB9026 have been described previously (Huang et al. 2012; Wang et al. 2007), while the solid fermentation medium (10 g/L soluble starch, 10 g/L glucose, 2.5 g/L soy peptone, 2.5 g/L yeast extract, 0.05 g/L CuSO4•5H2O, 0.25 g/L ZnSO4•7H2O, 3 g/L NaCl, 3 g/L CaCO3 and 10 g/L agar, pH 7.0) was used for mutant screening. Mycobacterium smegmatis ATCC607 (bought from China General Microbiological Culture Collection Center) was selected for the mutant screening bioassay and cultured with nutrient broth (NB) medium (10 g/L peptone, 3 g/L beef extract, and 5 g/L NaCl; for solid medium, 15 g/L agar was added). For homogeneous dispersion of either spores or M. smegmatis ATCC607, 5 mL soft agar (8 g/L agar) of each corresponding medium was prewarmed at 50 °C water bath and mixed with the target microbes before overlaying on a 90-mm solid plate.
Single-factor medium optimization of S. flavoviridis SB9026
Based on the original production medium, 0.1 g/L CuSO4•5H2O, 0.5 g/L ZnSO4•7H2O, and 3 g/L CaCO3 were set as the constant components, and the effects of various carbon sources and nitrogen sources on the production of 6′-DO-BLM Z by SB9026 were then evaluated. Ten common carbon sources, including the quick-acting D-fructose, D-glucose, D-galactose, lactose, mannitol, and glycerol and the slow-acting maltose, sucrose, soluble starch, and dextrin, were tested at the concentration of 20 g/L, respectively, with a constant 20 g/L soybean meal. Meanwhile, ten organic nitrogen sources, including the quick-acting peptone, tryptone, fish peptone, corn steep liquor, yeast extract, malt extract, and beef extract and the slow-acting soybean meal, pharmamedia, and corn meal, were sequentially tested at the concentration of 20 g/L, respectively, with a constant 20 g/L glucose. After the quick-acting and slow-acting carbon and nitrogen sources have been assigned, their optimal dosages have been further determined to finally formulate the optimized production medium, which has been applied in the subsequent fermentation studies.
The UV mutagenesis of S. flavoviridis SB9026
To determine the proper conditions for UV mutagenesis, 3 mL of diluted SB9026 spore suspension (106/mL) was transferred into a sterilized 60-mm glass Petri dish containing a 50-mm needle and placed on a magnetic stirrer located 12 cm beneath a UV light. After irradiation at 254 nm with constant but gentle stirring, the resulting spores were collected and resuspended in 15 mL mISP4 soft agar and then parallel overlaid onto three mISP4 plates (5 mL/plate). A series of plates containing spores treated to different irradiation time (from 3 to 8 min, with 30 s increments) was cultivated at 30 °C in the dark for 5 days to recover viable colonies. The irradiation time that could provide 10–20 colonies per plate was chose as the appropriate condition for the UV mutagenesis of 108 SB9026 spores, which was accordingly carried out on a total of 100 mISP4 plates to generate mutants.
Ribosome engineering of S. flavoviridis SB9026
To determine the minimal inhibition concentrations (MIC) of commonly used streptomycin (Str), gentamycin (Gen), and rifamycin (Rif), the unified spore suspensions of SB9026 were inoculated onto mISP4 plates (106 spores mixed with 5 mL mISP4 soft agar per plate) containing the different concentrations of target antibiotic (5, 10, 30, 50, 100 mg/L) and cultivated at 30 °C for observation. The subsequent ribosome engineering of SB9026 was carried out by using corresponding antibiotics under 1 MIC and 2 MIC concentrations, respectively, on a total of 120 mISP4 plates (20 plates per each condition) to obtain resistant mutants.
Combined UV mutagenesis and ribosome engineering of S. flavoviridis SB9026
Following the same protocol, the 106 spore suspension of SB9026 was first processed via UV irradiation, in which the irradiation time that could provide 50–100 colonies per plate was used. The treated spores were mixed with 5 mL mISP4 soft agar and spread on a mISP4 plate containing a certain concentration (either 0.5 MIC or 2 MIC) of selected antibiotic and cultivated at 30 °C in the dark for 7–8 days. Similarly, the total of 120 mISP4 plates (20 plates per each condition) was cultured to obtain expected mutants.
Efficient screening of S. flavoviridis SB9026-derived mutants and HPLC analyses
Each viable colony derived from the above experiments was parallel inoculated into 96-well plates (200 μL/well) containing either solid fermentation medium or mISP4 medium (corresponding antibiotic was supplied for ribosome engineering-derived colonies) and incubated at 30 °C for 10 days. SB9026 was also inoculated as the reference. Only colonies that grew well on both plates were regarded as the potential mutants. To bioassay mutants, the well-grown agar cylinder of each mutant was picked up from the corresponding solid fermentation 96-well plate by a sterilized toothpick and appropriately placed into the punched 6-mm hole of the NB plate; then, 1 mL of M. smegmatis ATCC607 suspension (108/mL) was mixed with 4 mL NB soft agar and overlaid onto the processed NB plate. After solidification, the plates were incubated at 37 °C overnight and the inhibition zone of each agar cylinder was measured (Fig. 2). Mutants with better inhibition activity than that of SB9026 were further validated by liquid fermentation in shake flasks. Briefly, 50 μL spore suspension of the target mutant was inoculated into 30 mL seed medium and grown at 30 °C and 220 rpm for 40 h. Then, 5 mL of seed culture was transferred into a 250-mL flask containing 50 mL production medium and cultivated at the same conditions for 6–7 days.
The resulting supernatant from the fermentation broth was filtered through a 0.22-μm needle filter and analyzed on the Waters E2695 HPLC system equipped with a photo-diode array (PDA) detector and a Welch AQ-C18 column (5 μm, 250 × 4.6 mm; Welch Materials Inc., Shanghai, China). The mobile phase consisted of buffer A (ultrapure H2O containing 0.1% HCO2H) and buffer B (chromatographic grade CH3OH containing 0.1% HCO2H) at a flow rate of 1 mL/min. A gradient program (95% buffer A to 65% buffer A from 0 to 10 min; 65% buffer A to 5% buffer A from 10 to 15 min; 5% buffer A from 15 to 17 min; 5% buffer A to 95% buffer A from 17 to 23 min) was used to detect 6′-DO-BLM Z at 300 nm. The titer of 6′-DO-BLM Z in each mutant was the average from at least three parallel fermentations and quantified based on the calibration curve with authentic 6′-DO-BLM Z standard.
Genetic characterization of Gen-resistant S. flavoviridis G-4F12
The genomic DNA of SB9026 and its Gen-resistant mutant G-4F12 were prepared according to the previous report (Huang et al. 2012). On the basis of the homologous 30S ribosomal protein S12 (NCBI protein ID: NP_628819.1) and 50S ribosomal protein L6 (NCBI protein ID: NP_628876.1) from Streptomyces coelicolor A3(2), rpsL-for (5′-GTGCCTACGATCCAGCAGCTG-3′) and rpsL-rev (5′-AATGAAGAGGAAGAACCGCGG-3′) were designed to amplify the rpsL gene encoding S12; rplF-for (5′-ATGTCGCGCATCGGCAAGCT-3′) and rplF-rev (5′-AATGAATGGGCGGAAAGGCTGGA-3′) were designed to amplify the rplF gene encoding L6, while 16SrRNA-for (5′-AGAGTTTGATCCTGGCTCAG-3′) and 16SrRNA-rev (5′-ACGGCTACCTTGTTACGACTT-3′) (Hindra et al. 2014) were used to amplify the 16S ribosomal DNA. The related oligonucleotide syntheses and DNA sequencing were all performed by Tsingke Biotech Co. (Beijing, China).
Scaled-up fermentation of 6′-DO-BLM Z
The production of 6′-DO-BLM Z was further scaled up in a 15-L fermentor containing 10 L production medium via three-stage cultivation. Briefly, 500 μL of fresh spore suspension (about 108/mL) was inoculated into 50 mL seed medium and cultivated at 30 °C for 24 h. Then, this culture was transferred into 1 L fresh seed medium and cultivated at 30 °C for an additional 24 h. The resulting entire seed culture was inoculated into 10 L production medium to start the fermentation. The stirrer speed (200 to 400 rpm), aeration rate (150 to 300 L/H), and fermentor pressure (0.02 to 0.1 MPa) were coordinated to maintain the dissolved oxygen (DO) of the fermentation broth at above 20%. Samples were periodically collected and analyzed to determine the yield of 6′-DO-BLM Z. When the pH of the broth was quickly increased beyond 7.0 at the middle stage of fermentation, it was maintained in a range of 7.5 to 8.0 by the automatic addition of sterilized 20% (W/V) glucose solution. The fermentation was ended when the DO of the broth remained high (> 50%) in spite of glucose addition.
Results
Medium optimization of S. flavoviridis SB9026
To explore the capacity of SB9026 for 6′-DO-BLM Z production, the production medium was reformulated using single-factor optimization. The evaluation results suggested that glucose and yeast extracts were the best quick-acting carbon and nitrogen sources, while soluble starch and soybean meal were suitable slow-acting carbon and nitrogen sources, respectively (Fig. 3). After investigations of their optimal concentrations, the final production medium was simplified to contain 15 g/L soluble starch, 20 g/L glucose, 20 g/L soybean meal, 5 g/L yeast extract, 0.1 g/L CuSO4•5H2O, 0.5 g/L ZnSO4•7H2O, and 3 g/L CaCO3 (pH 7.0), and the fermentation time was shortened significantly from 12 to 7 days. Under these conditions, the average titer of 6′-DO-BLM Z from different batches of flask fermentations was improved from 8.9 ± 0.7 to 20.1 ± 2.6 mg/L.
Independent UV mutagenesis and ribosome engineering of S. flavoviridis SB9026
The preliminary experiments have determined that 6-min UV irradiation time was suitable for the mutagenesis of SB9026 (data not shown). Under this condition, a total of 672 mutants were recovered and bioassayed (Table 1). Most of generated mutants were less bioactive; only 3% of them (20 of 672) showed slightly improved bioactivity (Fig. 4A). Based on the morphological and phenotypic characteristics, 17 mutants were further investigated by liquid fermentation in shake flasks; 10 of which showed improved 6′-DO-BLM Z production. However, the highest titer of 26.0 ± 1.8 mg/L was only a 30% increase comparing to that of SB9026, whereas the production of 6′-DO-BLM Z in 7 other mutants with impaired sporulation was decreased (Fig. 4B).
To implement the ribosome engineering of SB9026, the relative MIC of streptomycin (Str), gentamycin (Gen), and rifamycin (Rif) was determined as 50, 30, and 10 mg/L, respectively. However, the antibiotic screening under various MIC conditions only led to the isolation of a total of 37 resistant mutants, and most of these were Rif-resistant (32 of 37) (Table 1). Among them, 12 mutants exhibited very poor sporulation and were not suitable for further evaluation. The exploration of the remaining 14 mutants with shake flask fermentation revealed that the production of 6′-DO-BLM Z was impaired in most mutants; only two showed comparable titers as that of SB9026 (Fig. 4C).
Combined UV mutagenesis and ribosome engineering of S. flavoviridis SB9026
Based on the data above, UV mutagenesis combined with ribosome engineering was undertaken on SB9026. A total of 384 Gen-resistant mutants were recovered, with most of them derived from 0.5 MIC Gen selection (381 of 384), while no viable colony was observed under either Str or Rif screening (Table 1). Subsequent bioassay indicated that 39 of the 384 mutants (~ 10%) were more active than SB9026; 6 of which with the highest activities were subjected to further validation (Fig. 5A). Originally, 5 of 6 mutants demonstrated titer improvement when the seed medium was supplemented with 15 mg/L Gen, with mutant G-4F12 presenting the highest yield of 26.3 ± 0.2 mg/L 6′-DO-BLM Z. In addition, when Gen concentration was increased to 30 mg/L in the seed medium, all 6 mutants showed further titer improvement at different levels, with G-4F12 still retaining the highest yield of 44.2 ± 0.4 mg/L, about twofold better than that of SB9026 (Fig. 5B). However, the attempted successive increase of Gen concentration up to 300 mg/L failed to further improve the titer of 6′-DO-BLM Z in these mutants. In fact, the cell growth was dose-dependently repressed when Gen concentration was higher than 90 mg/L (data not shown). Hence, S. flavoviridis G-4F12 with the preservation number CCTCC M2015738 was identified as a potential high producer of 6′-DO-BLM Z, and 30 mg/L Gen was supplemented to prepare its fermentation seeds.
Screening of mutants generated from combined UV mutagenesis and ribosome engineering: a bioassay (asterisk indicates the inhibition zone of SB9026) and b titer comparison of 6′-DO-BLM Z in two rounds of fermentation of six potential mutants (their seeds were respectively supplemented with 15 and 30 mg/L of Gen); and HPLC analyses of c extracts from SB9026 and G-4F12, as well as pure 6′-DO-BLM Z; and d periodically collected extracts with different pH from G-4F12 (▼, 6′-DO-BLM Z)
HPLC analyses demonstrated that the metabolite profile of G-4F12 was very similar to that of SB9026, only except the titer changes (Fig. 5C). This could be beneficial for the subsequent purification of 6′-DO-BLM Z considering about its improved titer in G-4F12. Moreover, periodical monitoring of the flask fermentation revealed that the production of 6′-DO-BLM Z preferred the neutral to slightly alkaline environment, while the pH of the broth higher than 8.0 caused the concentration decrease of 6′-DO-BLM Z (Fig. 5D). These preliminary data could provide us guidance for scaled-up production of 6′-DO-BLM Z.
Genetic characterization of S. flavoviridis G-4F12
To find out possible mutation sites inducing the Gen resistance, the rpsL gene and rplF gene, as well as 16S ribosomal DNA of SB9026 and G-4F12, were respectively sequenced. However, none of these genes was mutated in G-4F12. This result is consistent with other reported Gen-induced ribosome engineering cases (Shentu et al. 2016; Tanaka et al. 2017; Wang et al. 2017), in which the mutation sites could not be located although the titer of product had been improved. In addition, the genetic stability of G-4F12 was investigated by the successive five-generation propagation on mISP4 slants containing 30 mg/L Gen. Each generation was validated by liquid fermentation in shake flasks. The corresponding titer of 6′-DO-BLM Z was varied from 42.3 ± 0.9 to 51.3 ± 0.3 mg/L, which maintained at least a twofold titer increase (Fig. 6A). Furthermore, subsequent medium evaluation showed that G-4F12 shared similar nitrogen source preferences as SB9026 (Fig. 6C), while its dependences on carbon sources were distinctly changed (Fig. 6B). Glucose was still the best quick-acting carbon source for G-4F12, but maltose presented slightly better performance than soluble starch as the slow-acting carbon source. Considering about the much less impact of the slow-acting carbon source on the production of 6′-DO-BLM Z, as well as the higher cost of maltose (about five folds of the price of soluble starch), the production medium of G-4F12 was not changed.
The scaled-up production of 6′-DO-BLM Z
The production of 6′-DO-BLM Z was scaled up to a 15-L fermentor by cultivation of either SB9026 or G-4F12. A temperature-descending strategy was applied to correlate the fermentation process with the physiological behavior of microbes. The initial temperature was set at 32 °C to help relieve the stagnant cell growth during transition from shake flask to fermentor. When DO dropped after 24–30 h cultivation, the temperature was decreased to 30 °C to avoid the overgrowth of cells. At the end of the exponential growth phase (around 60 h) when the DO fluctuated and a quick rise in pH occurred, so, the temperature was adjusted to 28 °C to ensure production of 6′-DO-BLM Z until termination.
Comparing the fermentation profiles of SB9026 and G-4F12, the variation of pH and DO for SB9026 was more pronounced, especially at the later phase of fermentation (Fig. 7). In both cases, the pH of the broth rapidly increased around 60–80 h, which indicated the depletion of glucose. When the pulsed addition of 20% (W/V) glucose solution was initiated at this time point, the pH was maintained within a small range of 7.5 to 8.0, thus providing a relatively stable environment for the production of 6′-DO-BLM Z. Moreover, the concentration of residual sugar was kept at a low level (around 1 g/L) to avoid the possible re-growth of cells, which could repress or even cease the biosynthesis of 6′-DO-BLM Z. Under these rational fermentative regulations, the average titer of 6′-DO-BLM Z within the consecutive three batches of fermentation of G-4F12 was 70.7 ± 1.8 mg/L, which represented an almost threefold increase comparing to that of SB9026 (23.4 ± 0.8 mg/L) at the same conditions, and a sevenfold improvement better than the original titer (Fig. 8). In addition, the extra 40% titer improvement (from shake flask to fermentor) (Fig. 8) of G-4F12 under fine-tuned fermentative conditions was much higher than that of SB9026 (only about 10%), which further confirmed the increased productivity of G-4F12.
Discussion
The discovery of bioactive microbial natural products (Galm and Shen 2007; Shen 2015) also requires the establishment of reliable sources for sufficient supply of novel chemical entities in order to develop viable drugs. Combinatorial biosynthesis has succeeded in the production of 6′-DO-BLM Z as a promising lead for the next generation of a BLM anticancer drug, and microbial fermentation remains the only practical approach to produce such complex natural product.
The strain improvement of recombinant S. flavoviridis SB9026, using UV mutagenesis, ribosome engineering, and fermentation optimization, has been adopted to boost the production of 6′-DO-BLM Z. Although the initial medium optimization could almost double the titer of 6′-DO-BLM Z to over 20 mg/L, this increment was correlated with an increase to biomass. Hundreds of random mutants generated by UV mutagenesis were screened out by a facile bioassay system, but only a few of them demonstrated to possess enhanced 6′-DO-BLM Z production. An observed inconsistency between bioassay screening and fermentation (Fig. 4) may be due to the impaired sporulation of mutants, which led to the reduced inoculums and cell densities during fermentation. Moreover, further evaluation suggested that these mutants from UV mutagenesis may undergo strain degeneration due to the lack of selection. On the other side, several Rif- or Str-resistant mutants were derived from ribosome engineering but none of them demonstrated high-titer 6′-DO-BLM Z production.
Considering the high mutation rate of UV mutagenesis and traceable antibiotic resistance of ribosome engineering, a straightforward strategy combining both two methods was applied for strain improvement. With the significantly increased positive mutation rate other than that of UV mutagenesis alone, the resulting mutants were all Gen-resistant in contrast to that from ribosome engineering alone. A 6′-DO-BLM Z high-producing strain S. flavoviridis G-4F12 has been successfully identified. Unlike streptomycin and rifamycin, the origin of gentamycin resistance is still obscure; mutations at 16S-rRNA (Pfister et al. 2003), or ribosomal protein S12 (Vila-Sanjurjo et al. 2007), or ribosomal protein L6 (rplF gene) (Buckel et al. 1977; Davies et al. 1998) all could confer such resistance in Escherichia coli. However, our attempts have so far failed to locate any possible mutation site in the genes of G-4F12, which is consistent with other published Gen resistance-related ribosome engineering results (Shentu et al. 2016; Tanaka et al. 2017; Wang et al. 2017). Although the Gen-induced mutation could not be clarified in G-4F12, variations in utilization of different carbon sources suggested its metabolic pathway changes are distinct to that of SB9026. In addition, the genetic stability and productivity of G-4F12 have also been shown to provide a stable yield of over 50 mg/L at shake flask scale (Fig. 6), which indicates a practical improvement for the next scaled-up production of 6′-DO-BLM Z.
Compared to the flask fermentation, the cultivation process in the fermentor could be controlled by temperature regulation and fed-batch strategies. As a result, the stationary phase of fermentation for secondary metabolite production was prolonged. Moreover, the comparison of fermentation profiles revealed that G-4F12 grew faster and in a more controllable fashion than its parental strain SB9026. It also exhibited better tolerance to environmental changes at the later phase of fermentation (Fig. 7). Hence, the better physiological performance and optimized fermentation conditions have both contributed to the final titer improvement of 6′-DO-BLM Z in G-4F12.
Overall, by combining the phenotype-oriented strain improvement via ribosome engineering and UV mutagenesis, we have succeeded in the isolation of a 6′-DO-BLM Z high-producing strain of S. flavoviridis G-4F12. Our efforts have demonstrated the efficacy of combined empirical strain-breeding approaches to effect a rapid improvement in titer, which could be complementary to rational pathway engineering, especially for recombinant strains that may be limited to further genetic manipulations. In addition, the obtained high producer and established fermentation process have set the stage to produce gram quantities of 6′-DO-BLM Z for subsequent drug development.
References
Buckel P, Buchberger A, Bock A, Wittmann HG (1977) Alteration of ribosomal protein L6 in mutants of Escherichia coli resistant to gentamicin. Mol Gen Genet 158(1):47–54. https://doi.org/10.1007/BF00455118
Burger RM (1998) Cleavage of nucleic acids by bleomycin. Chem Rev 98(3):1153–1170. https://doi.org/10.1021/cr960438a
Chen C, Si S, He Q, Xu H, Lu M, Xie Y, Wang Y, Chen R (2008) Isolation and characterization of antibiotic NC0604, a new analogue of bleomycin. J Antibiot (Tokyo) 61(12):747–751. https://doi.org/10.1038/ja.2008.88
Chen Y, Smanski MJ, Shen B (2010) Improvement of secondary metabolite production in Streptomyces by manipulating pathway regulation. Appl Microbiol Biotechnol 86(1):19–25. https://doi.org/10.1007/s00253-009-2428-3
Chen Y, Yin M, Horsman GP, Shen B (2011) Improvement of the enediyne antitumor antibiotic C-1027 production by manipulating its biosynthetic pathway regulation in Streptomyces globisporus. J Nat Prod 74(3):420–424. https://doi.org/10.1021/np100825y
Chen JK, Yang D, Shen B, Murray V (2017) Bleomycin analogues preferentially cleave at the transcription start sites of actively transcribed genes in human cells. Int J Biochem Cell Biol 85:56–65. https://doi.org/10.1016/j.biocel.2017.02.001
Davies C, Bussiere DE, Golden BL, Porter SJ, Ramakrishnan V, White SW (1998) Ribosomal proteins S5 and L6: high-resolution crystal structures and roles in protein synthesis and antibiotic resistance. J Mol Biol 279(4):873–888. https://doi.org/10.1006/jmbi.1998.1780
Einhorn LH (2002) Curing metastatic testicular cancer. Proc Natl Acad Sci U S A 99(7):4592–4595. https://doi.org/10.1073/pnas.072067999
Galm U, Shen B (2007) Natural product drug discovery: the times have never been better. Chem Biol 14(10):1098–1104. https://doi.org/10.1016/j.chembiol.2007.10.004
Galm U, Hager MH, Van Lanen SG, Ju J, Thorson JS, Shen B (2005) Antitumor antibiotics: bleomycin, enediynes, and mitomycin. Chem Rev 105(2):739–758. https://doi.org/10.1021/cr030117g
Galm U, Wang L, Wendt-Pienkowski E, Yang R, Liu W, Tao M, Coughlin JM, Shen B (2008) In vivo manipulation of the bleomycin biosynthetic gene cluster in Streptomyces verticillus ATCC15003 revealing new insights into its biosynthetic pathway. J Biol Chem 283(42):28236–28245. https://doi.org/10.1074/jbc.M804971200
Hindra HT, Yang D, Rudolf JD, Xie P, Xie G, Teng Q, Lohman JR, Zhu X, Huang Y, Zhao LX, Jiang Y, Duan Y, Shen B (2014) Strain prioritization for natural product discovery by a high-throughput real-time PCR method. J Nat Prod 77(10):2296–2303. https://doi.org/10.1021/np5006168
Hindra YD, Teng Q, Dong LB, Crnovcic I, Huang T, Ge H, Shen B (2017) Genome mining of Streptomyces mobaraensis DSM40847 as a bleomycin producer providing a biotechnology platform to engineer designer bleomycin analogues. Org Lett 19(6):1386–1389. https://doi.org/10.1021/acs.orglett.7b00283
Huang SX, Feng Z, Wang L, Galm U, Wendt-Pienkowski E, Yang D, Tao M, Coughlin JM, Duan Y, Shen B (2012) A designer bleomycin with significantly improved DNA cleavage activity. J Am Chem Soc 134(32):13501–13509. https://doi.org/10.1021/ja3056535
Kawaguchi H, Tsukiura H, Tomita K, Konishi M, Saito K (1977) Tallysomycin, a new antitumor antibiotic complex related to bleomycin. I. Production, isolation and properties. J Antibiot (Tokyo) 30(10):779–788. https://doi.org/10.7164/antibiotics.30.779
Leitheiser CJ, Smith KL, Rishel MJ, Hashimoto S, Konishi K, Thomas CJ, Li C, McCormick MM, Hecht SM (2003) Solid-phase synthesis of bleomycin group antibiotics. Construction of a 108-member deglycobleomycin library. J Am Chem Soc 125(27):8218–8227. https://doi.org/10.1021/ja021388w
Ochi K (2017) Insights into microbial cryptic gene activation and strain improvement: principle, application and technical aspects. J Antibiot (Tokyo) 70(1):25–40. https://doi.org/10.1038/ja.2016.82
Ochi K, Okamoto S, Tozawa Y, Inaoka T, Hosaka T, Xu J, Kurosawa K (2004) Ribosome engineering and secondary metabolite production. Adv Appl Microbiol 56:155–184. https://doi.org/10.1016/S0065-2164(04)56005-7
Okamoto-Hosoya Y, Sato TA, Ochi K (2000) Resistance to paromomycin is conferred by rpsL mutations, accompanied by an enhanced antibiotic production in Streptomyces coelicolor A3(2). J Antibiot (Tokyo) 53(12):1424–1427. https://doi.org/10.7164/antibiotics.53.1424
Panetta A, Angelelli B, Martoni A (1999) Pilot study on induction chemotherapy with cisplatin, epirubicin, etoposide and bleomycin in cervical cancer stage Ib, IIa and IIb. Anticancer Res 19(1B):765–768
Pfister P, Hobbie S, Vicens Q, Bottger EC, Westhof E (2003) The molecular basis for A-site mutations conferring aminoglycoside resistance: relationship between ribosomal susceptibility and X-ray crystal structures. Chembiochem 4(10):1078–1088. https://doi.org/10.1002/cbic.200300657
Shen B (2015) A new golden age of natural products drug discovery. Cell 163(6):1297–1300. https://doi.org/10.1016/j.cell.2015.11.031
Shentu X, Liu N, Tang G, Tanaka Y, Ochi K, Xu J, Yu X (2016) Improved antibiotic production and silent gene activation in Streptomyces diastatochromogenes by ribosome engineering. J Antibiot (Tokyo) 69(5):406–410. https://doi.org/10.1038/ja.2015.123
Shima J, Hesketh A, Okamoto S, Kawamoto S, Ochi K (1996) Induction of actinorhodin production by rpsL (encoding ribosomal protein S12) mutations that confer streptomycin resistance in Streptomyces lividans and Streptomyces coelicolor A3(2). J Bacteriol 178(24):7276–7284. https://doi.org/10.1128/jb.178.24.7276-7284.1996
Sleijfer S (2001) Bleomycin-induced pneumonitis. Chest 120(2):617–624. https://doi.org/10.1378/chest.120.2.617
Tanaka Y, Komatsu M, Okamoto S, Tokuyama S, Kaji A, Ikeda H, Ochi K (2009) Antibiotic overproduction by rpsL and rsmG mutants of various actinomycetes. Appl Environ Microbiol 75(14):4919–4922. https://doi.org/10.1128/AEM.00681-09
Tanaka Y, Kasahara K, Izawa M, Ochi K (2017) Applicability of ribosome engineering to vitamin B12 production by Propionibacterium shermanii. Biosci Biotechnol Biochem 81(8):1636–1641. https://doi.org/10.1080/09168451.2017.1329619
Tao M, Wang L, Wendt-Pienkowski E, Zhang N, Yang D, Galm U, Coughlin JM, Xu Z, Shen B (2010) Functional characterization of tlmH in Streptoalloteichus hindustanus E465-94 ATCC 31158 unveiling new insight into tallysomycin biosynthesis and affording a novel bleomycin analog. Mol BioSyst 6(2):349–356. https://doi.org/10.1039/b918106g
Umezawa H, Maeda K, Takeuchi T, Okami Y (1966) New antibiotics, bleomycin A and B. J Antibiot (Tokyo) 19(5):200–209
Vila-Sanjurjo A, Lu Y, Aragonez JL, Starkweather RE, Sasikumar M, O’Connor M (2007) Modulation of 16S rRNA function by ribosomal protein S12. Biochim Biophys Acta 1769(7–8):462–471. https://doi.org/10.1016/j.bbaexp.2007.04.004
Wang L, Yun BS, George NP, Wendt-Pienkowski E, Galm U, TJ O, Coughlin JM, Zhang G, Tao M, Shen B (2007) Glycopeptide antitumor antibiotic zorbamycin from Streptomyces flavoviridis ATCC 21892: strain improvement and structure elucidation. J Nat Prod 70(3):402–406. https://doi.org/10.1021/np060592k
Wang Q, Cui K, Espin-Garcia O, Cheng D, Qiu X, Chen Z, Moore M, Bristow RG, Xu W, Der S, Liu G (2013) Resistance to bleomycin in cancer cell lines is characterized by prolonged doubling time, reduced DNA damage and evasion of G2/M arrest and apoptosis. PLoS One 8(12):e82363. https://doi.org/10.1371/journal.pone.0082363
Wang L, Chen X, Wu G, Li S, Zeng X, Ren X, Tang L, Mao Z (2017) Enhanced epsilon-poly-L-lysine production by inducing double antibiotic-resistant mutations in Streptomyces albulus. Bioprocess Biosyst Eng 40(2):271–283. https://doi.org/10.1007/s00449-016-1695-5
Xu J, Tozawa Y, Lai C, Hayashi H, Ochi K (2002) A rifampicin resistance mutation in the rpoB gene confers ppGpp-independent antibiotic production in Streptomyces coelicolor A3(2). Mol Gen Genomics 268(2):179–189. https://doi.org/10.1007/s00438-002-0730-1
Yang D, Zhu X, Wu X, Feng Z, Huang L, Shen B, Xu Z (2011) Titer improvement of iso-migrastatin in selected heterologous Streptomyces hosts and related analysis of mRNA expression by quantitative RT-PCR. Appl Microbiol Biotechnol 89(6):1709–1719. https://doi.org/10.1007/s00253-010-3025-1
Yang D, Hindra DLB, Crnovcic I, Shen B (2017) Engineered production and evaluation of 6′-deoxy-tallysomycin H-1 revealing new insights into the structure-activity relationship of the anticancer drug bleomycin. J Antibiot (Tokyo). https://doi.org/10.1038/ja.2017.93
Yu Z, Smanski MJ, Peterson RM, Marchillo K, Andes D, Rajski SR, Shen B (2010) Engineering of Streptomyces platensis MA7339 for overproduction of platencin and congeners. Org Lett 12(8):1744–1747. https://doi.org/10.1021/ol100342m
Zhang N, Zhu X, Yang D, Cai J, Tao M, Wang L, Duan Y, Shen B, Xu Z (2010) Improved production of the tallysomycin H-1 in Streptoalloteichus hindustanus SB8005 strain by fermentation optimization. Appl Microbiol Biotechnol 86(5):1345–1353. https://doi.org/10.1007/s00253-009-2406-9
Zhu X, Zhang W, Chen X, Wu H, Duan Y, Xu Z (2010) Generation of high rapamycin producing strain via rational metabolic pathway-based mutagenesis and further titer improvement with fed-batch bioprocess optimization. Biotechnol Bioeng 107(3):506–515. https://doi.org/10.1002/bit.22819
Funding
This work was supported in part by grants from the National High Technology Research and Development Program of China 2012AA02A705, the Chinese Ministry of Education 111 Project B0803420, National Major Scientific and Technological Special Project 2011ZX09401-001, and the Natural Products Library Initiative at TSRI.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhu, X., Kong, J., Yang, H. et al. Strain improvement by combined UV mutagenesis and ribosome engineering and subsequent fermentation optimization for enhanced 6′-deoxy-bleomycin Z production. Appl Microbiol Biotechnol 102, 1651–1661 (2018). https://doi.org/10.1007/s00253-017-8705-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8705-7