Abstract
Lipids are naturally derived products that offer an attractive, renewable alternative to petroleum-based hydrocarbons. While naturally produced long-chain fatty acids can replace some petroleum analogs, medium-chain fatty acid would more closely match the desired physical and chemical properties of currently employed petroleum products. In this study, we engineered Yarrowia lipolytica, an oleaginous yeast that naturally produces lipids at high titers, to produce medium-chain fatty acids. Five different acyl-acyl carrier protein (ACP) thioesterases with specificity for medium-chain acyl-ACP molecules were expressed in Y. lipolytica, resulting in formation of either decanoic or octanoic acid. These novel fatty acid products were found to comprise up to 40 % of the total cell lipids. Furthermore, the reduction in chain length resulted in a twofold increase in specific lipid productivity in these engineered strains. The medium-chain fatty acids were found to be incorporated into all lipid classes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Renewable fuels and chemicals offer solutions to problems ranging from global warming to supply stability (Fortman et al. 2008; Liu et al. 2014; Lynd et al. 2005). Using yeast as microbial cell factories to produce valuable commodity chemicals offers a promising and sustainable alternative to the continued use of petroleum-based products (Runguphan and Keasling 2014). Fatty acids are molecules with a long alkyl chain, very similar to the chemicals found in petroleum, with a carboxylic acid attached to the end. They accumulate within the cell as energy and carbon storage molecules. Fatty acids can be further converted to other derivatives including (1) methyl and ethyl esters which can be used as biodiesel (Moser 2011); (2) fatty waxes to be used in cosmetics, food, and paper industries (Santos et al. 2014); (3) fatty alcohols with applications as surfactants and industrial solvents (Zheng et al. 2012); and (4) alkanes that can directly be used as fuel (Choi and Lee 2013).
Oleaginous yeasts naturally produce elevated amounts of lipids compared to traditional yeast strains, making them a natural choice as a microbial platform for industrial production of fatty-acid-derived products (Ageitos et al. 2011; Papanikolaou and Aggelis 2011; Beopoulos et al. 2009). Yarrowia lipolytica is the most-well studied of these yeasts, and researchers have previously engineered it for increased lipid production by a variety of rational and evolutionary approaches (Tai and Stephanopoulos 2013; Wang et al. 2014; Liu et al. 2015; Qiao et al. 2015; Zhang et al. 2014). In addition, many researchers have successfully produced novel fatty acid products in Y. lipolytica including omega-3-fatty acids, linoleic acid-derived unsaturated and polyunsaturated fatty acids, and alkanes (Xue et al. 2013; Beopoulos et al. 2014; Zhang et al. 2013; Blazeck et al. 2013; Madzak 2015). Although these yeast strains can be engineered to produce large quantities of lipids and related products, the associated alkyl chains are much longer than those found in typical petroleum-derived products, making their physical properties less attractive for industrial and commercial applications (Dwivedi and Sharma 2014; Ramos et al. 2009). Engineering oleaginous yeast for production of medium-chain fatty acids would lead to production of renewable molecules more suitable as replacements for petroleum-based products.
The mechanism of fatty acid synthesis by the multienzyme fungal type I fatty acid synthase (FAS) is illustrated in Fig. 1. Initiation of fatty acid synthesis occurs when acetyl-CoA is condensed with carbon dioxide by acetyl-CoA carboxylase to form malonyl-CoA. Malonyl-CoA and acetyl-CoA are then incorporated in the FAS complex by transacylase activity to produce malonyl-acyl carrier protein (ACP) or acetyl-ACP, respectively. The ACP will have been previously activated with phosphopantetheine at a conserved serine residue by the phosphopantetheinyl transferase subunit (Lomakin et al. 2007). Malonyl-ACP is then decarboxylated, and keto-synthase catalyzes its condensation with acetyl-ACP. Keto-reductase, dehydratase, and enoylreductase subunits act in series to catalyze removal of the remaining oxygen atom and carbon-carbon double bond to form a saturated acyl-ACP (Landriscina et al. 1972; von Wettstein‐Knowles et al. 2006). The acyl-ACP then becomes the substrate for further addition of decarboxylated malonyl-ACP. This cycle of malonyl-CoA addition to the acyl chain continues until elongation is terminated by malonyl/palmitoyl transacylase which catalyzes the transacylation of the acyl-ACP with CoA to produce a fatty acyl-CoA (Leibundgut et al. 2008; Lynen 1969; Lomakin et al. 2007). This is slightly different from fatty acid synthesis in other domains such as bacteria and plants where elongation is terminated by an acyl-ACP thioesterase which catalyzes a transesterification of fatty acyl-ACP with water to produce free fatty acid (Jing et al. 2011; Moreno-Pérez et al. 2012).
In addition to a different mechanism of termination, bacterial, plant, and mammalian FASs have widely different structures than their fungal counterpart. In bacterial and plant systems, the different catalytic subunits required for fatty acid synthesis are dissociated (White et al. 2005; Zhang et al. 2006; Wakil 1989). In many eukaryotes including mammals and fungi, however, multisubunit complexes form that contain all the necessary catalytic subunits in a single enzyme. Furthermore, the mammalian FASs consist of a hexamer of α-subunits while the fungal FAS consists of a hexamer of α-subunits that form a central core capped on either side by three β-subunits (Leibundgut et al. 2008; Lynen 1969; Lomakin et al. 2007). The dissociated nature of synthases in bacterial and plant systems makes them well suited for modification of fatty acid length by expression of heterologous proteins. Many acyl-ACP thioesterases have been identified that use medium-chain acyl-ACP molecules as their substrate (McMahon and Prather 2014; Voelker et al. 1997; Pollard et al. 1991; Dörmann et al. 1993). Numerous studies have shown that expression of a medium-chain acyl-ACP-specific acyl-ACP thioesterase leads to production of shorter fatty acids in Escherichia coli as well as several plant species (Eccleston et al. 1996; Sherkhanov et al. 2014; Liu et al. 2010; Lu et al. 2008; Steen et al. 2010; Voelker et al. 1992).
Most organisms produce only 16- and 18-carbon fatty acids, indicating that a majority of acyl-ACP-thioesterases and transacylases have specificity toward longer acyl-ACP chains (Cronan and Rock 1996; Harwood 1988). While attempts at engineering E. coli for medium-chain fatty acid production have been successful, the lack of variety in transacylase specificity combined with the bulwark-like quaternary structure of fungal FAS makes engineering yeast for medium-chain fatty acid production challenging. In fact, the first attempt at producing medium-chain fatty acids was done by induction of the β-oxidation pathway, which breaks lipids down into acetyl-CoA, along with expression of carnitine O-octanoyltransferase to terminate β-oxidation early to allow accumulation of medium-chain fatty acids. This approach was dependent on supplementation of oleic acid in the medium (Chen et al. 2014). One successful strategy for de novo short-chain fatty acid production in Saccharomyces cerevisiae required concurrent expression of a more accessible mammalian FAS with its C-terminal thioesterase domain replaced with an acyl-ACP thioesterase known to produce short-chain fatty acids (Leber and Da Silva 2014).
In this work, we tested whether a heterologous thioesterase would be able to associate with the native FAS complex in Y. lipolytica and prematurely terminate elongation to produce medium-chain fatty acids. We demonstrate that the expression of different plant and bacterial acyl-ACP-thioesterases reduces the chain length of fatty acids in Y. lipolytica. Additionally, the final length of the fatty acids produced depends on the identity of the enzyme expressed and each enzyme produces a single-length fatty acid. An increase in specific lipid yield along with a reduction in final cell mass was also observed during production of medium-chain fatty acids. Novel products were detected in all lipid classes.
Materials and methods
Strains and growth conditions
All strains and plasmids used in this study are listed in Table 1. E. coli DH5α was used for all plasmid construction and propagation. DH5α was grown in Luria Bertani (LB) medium at 37 °C with constant shaking at 250 rpm. Ampicillin was added to a concentration of 100 μg/mL for plasmid maintenance. Y. lipolytica PO1f, a leucine and uracil auxotroph variant of the W29 strain (ATCC 20460), was used as a base strain for all studies (Madzak et al. 2000). Cultivation of Y. lipolytica strains was carried out at 30 °C with shaking at 200 rpm. Two different types of media were used for cultivation of Y. lipolytica strains. Yeast Synthetic Complete (YSC) media, which was used for cultivation of the PO1f wild type, consisted of 20 g/L glucose (Sigma), 0.67 g/L Yeast Nitrogen Base (Becton, Dickinson and Company), and 0.69 g/L Complete Supplement Mixture (CSM) supplement (MP Biomedicals). YSC-LEU, which was used for cultivation of acyl-ACP thioesterase expressing derivatives of PO1f, is the same as above with CSM replaced with 0.69 g/L CSM-Leu. Lipid accumulation was achieved by cultivation in a medium with a high C/N ratio containing 50 g/L glucose, the appropriate CSM supplement, 0.17 g/L Yeast Nitrogen Base without amino acids or ammonium sulfate, 0.15 g/L yeast extract, and 1 g/L ammonium sulfate. Acyl-ACP thioesterase expressing strains of Y. lipolytica PO1f were grown in high C/N medium at 30 °C and 200 rpm and samples collected after 48 and 96 h of incubation. Agar was added to 20 g/L for solid media preparations.
Plasmid construction
Five acyl-ACP thioesterase genes were codon optimized for expression in Y. lipolytica and cloned into the pINT4 vector. This was achieved by amplifying the TEFintron region of pINT4 using primers ProF and ProR and the ACP thioesterase genes from Anaerococcus tetradius (GenBank Accession No: KR180390), Cuphea hookeriana (GenBank Accession No: KR180391), Cuphea palustris (GenBank Accession No: KR180392), Clostridium perfringens (GenBank Accession No: KR180393), and Umbellularia californica (GenBank Accession No: KR180394) (Tables S1 and S2 and Fig. S1) by primers AtetF and AtetR, ChoF and ChoR, CpaF and CpaR, CpeF an CpeR, and UcaF and UcaR, respectively. The forward primers for ACP thioesterase amplifications contained a region of homology to the 3′ region of the TEFintron. The TEFintron and individual ACP thioesterase fragments were then joined by simultaneous overlap extension polymerase chain reaction. The joined fragments were then digested by BstBI and AscI and ligated into the pINT4 vector digested by the same enzymes to generate the pINT4/ACP plasmids. PCR reactions were carried out with Phusion High-Fidelity DNA polymerase (New England Biolabs), and recommended reaction conditions were used. Ligation was carried out at room temperature for 1 h using T4 DNA ligase (New England Biolabs). Table S3 lists all primers used for plasmid construction.
Transformation
E. coli was transformed using standard electroporation protocols on a Bio-Rad Gene Pulser. Plasmids were miniprepped from E. coli using a GeneJet Kit (Thermo Scientific), digested with AatII restriction enzyme, and transformed into Y. lipolytica to get random integrants of the TEFin-ACPT-Leu2 cassette. In order to transform Y. lipolytica, cells were taken from solid media and resuspended in 100 μL of a 50 % PEG solution containing 50 mM lithium acetate and 50 mM dithiothreitol and incubated at 39 °C for 1 h before plating the whole mixture onto YSC-LEU solid medium (Gietz and Woods 2002). Colonies were picked after 3 days, genomic DNA was extracted, and integration of the acyl-ACP thioesterase gene was verified by PCR amplification.
Identification and quantification of lipids
Saponification was carried out to convert neutral and charged lipids to their respective methyl esters for analysis by gas chromatography/mass spectrometry (GC/MS). Lyophilized pellets were resuspended in 2 mL of 20:1 mixture of methanol and acetyl chloride and 2 mL of hexanes. Five microliters of 25 mg/mL tridecanoic acid dissolved in a 3:2 methanol/benzene mixture was added as an internal standard. Samples were placed in sealed glass tubes and boiled for 30 min. Upon completion, 1 mL of water was added and phases allowed to separate (Lepage and Roy 1984). The upper organic phase was collected as the total saponified cell lipid.
GC/MS was used to determine the lengths of lipid species produced and their abundance. Saponified cell lipid samples were analyzed using DB-5 ms capillary column (Agilent) on a Shimadzu GC-2010 equipped with a GCMS-QP2010. Samples (1 μL) were injected at a 10:1 split ratio using hexane as the solvent. Helium carrier gas pressure was 121.7 kPa at a flow rate of 1.0 mL/min. Injection port temperature was 250 °C. Column temperature started at 30 °C and increased to 250 °C at a rate of 10 °C/min. The eluent from the GC entered the ionization chamber at 250 °C and measured at a full scan between 15 and 250 amu (Tariq et al. 2011). Concentrations were determined using standards as a reference. Sample chromatograms and mass spectra are provided in Fig. S2.
Lipid extraction and total dry weight estimation
Whole lipid extracts were collected and weighed to determine lipid titer and total lipid content. Lyophilized cell pellets were resuspended in 1 mL of a 2:1 chloroform/methanol mixture and shaken in a FastPrep 24 (MP Biomedicals) three times for 40 s at 5 m/s. Then, 0.2 mL of water was added, and samples were vortexed. Samples were centrifuged at 15,000×g for 5 min, and the lower organic phase was removed. The organic layer was then washed with 0.1 M NaCl and centrifuged at 15,000×g for 5 min. The lower organic layer was removed and allowed to dry overnight to obtain a total lipid mass (Bligh and Dyer 1959). The same volume of culture was centrifuged and washed with one volume of phosphate-buffered saline pH 6.8, allowed to dry at 70 °C overnight, and weighed to determine the total dry cell weight (DCW).
Separation of lipid classes
Total lipids were fractionated to determine the abundance of different lipid species in strains expressing heterologous ACP thioesterase enzymes. Florisil hydrated to 7 % by incubation with shaking overnight at room temperature was used as the adsorbent for chromatography. Total lipids extracted from 50 mL of culture were loaded onto the column, and fractions were eluted in the following solvents: 95:5 hexane/ether, 85:15 hexane/ether, 75:25 hexane/ether, 50:50 hexane/ether, 98:2 ether/methanol, and 96:4 ether/acetate to elute off steryl esters, triacylglycerides, sterols, diacylglycerides, monoacylglycerides, and free fatty acids, respectively (Carroll 1961).
Abundance of lipid in collected eluents was measured by dichromate reduction. Samples were collected and allowed to completely dry before analysis. Each dried sample was resuspended in 1 mL of a 0.2 % potassium dichromate dissolved in 96 % sulfuric acid and boiled for 15 min. Samples were allowed to cool, and an additional 1 mL of water was added. Samples were again allowed to cool, and their absorbance was measured at 440 nm to detect formation of Cr3+ (Freeman and West 1966).
Results
Production of non-native fatty acid species in Y. lipolytica by expression of heterologous acyl-ACP thioesterases
Most microorganisms produce fatty acids predominantly with acyl chains 16 or 18 carbons in length. In an attempt to produce shorter fatty acids, we expressed heterologous acyl-ACP thioesterase enzymes from five different species previously shown to have specificity toward acyl-ACP chains eight carbons in length (McMahon and Prather 2014; Sherkhanov et al. 2014). We evaluated Y. lipolytica PO1f strains with acyl-ACP-thioesterases from A. tetradius, Cuphea hookeriana, Cuphea palustris, Clostridium perfringens, and U. californica randomly integrated into the genome (referred to as PO1f-AtACPT, PO1f-AChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT, respectively) for their ability to produce medium-chain fatty acids (Fig. 2). We found that the expression of each enzyme tested was capable of terminating the elongation of fatty acid early to produce either decanoic acid or octanoic acid. PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, and PO1f-CpeACPT produced 36 ± 8, 44 ± 1, 57 ± 8, and 46 ± 8 % of their lipids as ten-carbon fatty acid species on a mass basis, respectively. PO1f-UcACPT produced 14 ± 5 % of their lipids as eight-carbon fatty acid species on a mass basis. The Y. lipolytica PO1f wild-type strain showed no production of either eight- or ten-carbon fatty acid species when grown under the same conditions. In all cases, production of shorter acids was coupled with reduction in the abundance of 18-carbon acid species. In the PO1f-UcACPT strain, 16-carbon fatty acid abundance increased to 75 ± 5 % on a mass basis compared to 34 ± 2 % in the wild-type strain.
Lipid yields from engineered strains
Total lipid titers and specific lipid production were determined for strains expressing heterologous enzyme during growth under lipid accumulation conditions (Fig. 3). Total lipid titers from engineered strains were comparable to those of the wild-type strain with PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT producing 2.9 ± 0.7, 2.3 ± 0.1, 1.7 ± 0.1, 1.8 ± 0.1, and 1.5 ± 0.6 g of lipid per liter, respectively. A lipid titer of 1.7 ± 0.6 g/L was produced by the wild-type strain (Fig. 3a). The above strains were found to grow to a final DCW of 4.5 ± 0.4, 4.1 ± 0.2, 2.4 ± 0.2, 3.9 ± 0.1, and 9.8 ± 0.9-g DCW per liter. The wild-type strain grew to 10.8 ± 1.3-g DCW per liter (Fig. 3c). The above strains produced 0.66 ± 0.23, 0.57 ± 0.32, 0.66 ± 0.06, 0.46 ± 0.02, and 0.13 ± 0.05 g lipid per gram DCW, respectively. The wild-type strain produced 0.15 ± 0.1 g lipid per gram dry cell weight, indicating that the improved specific lipid productivity was due to reduced growth of engineered strains capable of producing decanoic acid (Fig. 3b).
Incorporation of medium-chain fatty acids into different lipid classes
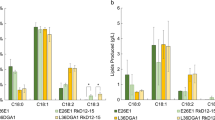
To evaluate the capacity for Y. lipolytica to incorporate non-native fatty acids into acylglycerides (mono, di, and tri), sterols, and steryl esters, total lipids were separated into different classes by chromatography and the composition with respect to carbon chain length of each fraction was determined (Fig. 4). Lipids harvested from wild-type cells were found to be 23 % triacylglyceride (TAG) while 17, 19, 17, 21, 19 and 15 % of total lipids were found in the TAG fraction from PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT, respectively. This reduction in TAG abundance coincided with an increase in free fatty acid (FFA) abundance with 17, 17, 21, 25, and 31 % of PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT lipids present as FFA, respectively, compared to 15 % of wild-type lipids present as FFA species.
Medium-chain fatty acids produced were found in all lipid classes. In the PO1f-AtACPT strain, 15 and 22 % of decanoic acid were found in the TAG and FFA fractions, respectively. In the PO1f-ChACPT strain, 20 and 19 % of decanoic acid were found in the TAG and FFA fractions, respectively. In PO1f-CpaACPT, 16 and 21 % of decanoic acid were found in the TAG and FFA fractions, respectively. In PO1f-CpeACPT, 32 and 23 % of decanoic acid were found in the TAG and FFA fractions, respectively. In PO1f-UcACPT, 20 and 28 % of octanoic acid were found in the TAG and FFA fractions, respectively (Table 2).
Expression of heterologous enzymes changed the localization of native fatty acids with respect to lipid classes. Of the 16-carbon fatty acids produced by the wild type, 19 % were found to be FFAs while PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT strains showed 44, 25, 46, 52, and 60 % of their 16-carbon fatty acids to be present as FFAs, respectively (Table 3). Eighteen-carbon fatty acids from PO1f-AtACPT, PO1f-ChACPT, PO1f-CpaACPT, PO1f-CpeACPT, and PO1f-UcACPT were found to be 36, 17, 43, 56, and 51 % FFAs, respectively (Table 4). Wild-type strains produced 12 % of their 18-carbon fatty acids as FFAs (Fig. 5). An increase in the FFA abundance of native lipids corresponds to the increase in FFA found in total cellular lipids.
Abundance of a medium-chain lipids, b 16-carbon lipids, and c 18-carbon lipids from Y. lipolytica PO1f strains expressing different heterologous acyl-ACP thioesterase enzymes in different lipid classes. FFA free fatty acids, MAG monoacylglycerol, DAG diacylglycerol, TAG triacylglycerol, SE sterol ester. Values can be found in Tables 2, 3, and 4
Discussion
Oleaginous yeasts are attractive because they produce lipids at high titers from simple carbohydrates. Numerous researchers have engineered Y. lipolytica for increased lipid titers and productivity. Like most other microbes, Y. lipolytica produces lipids 16 carbons in length or longer. Medium- and short-chain lipids have more desirable physical properties compared to the naturally produced long-chain lipids for the production of many industrially relevant chemicals and fuels. In this study, natural lipids were shortened to medium-chain lengths of eight or ten carbons by the expression of heterologous acyl-ACP thioesterases. In addition, lipid productivity doubled in some cases, suggesting that this approach can be used to increase flux through the lipid metabolism pathway as well as introduce novel products into an oleaginous yeast strain.
Expression of four acyl-ACP thioesterases in a Y. lipolytica host resulted in production of decanoic acid as 40 % or more of the total lipids produced. These decanoic-acid-producing strains did so without drastically changing the abundance of 16-carbon lipids, and seemingly, the medium-chain lipids were produced at the expense of 18-carbon lipids. The acyl-ACP thioesterase from U. californica, when expressed in Y. lipolytica, led to production of a modest amount of octanoic acid. Additionally, the abundance of 16-carbon lipids nearly doubled in this octanoic-acid-producing strain. Again, the increase in octanoic and hexadecanoic acids was coupled to a reduction in 18-carbon lipids. These results demonstrate that an acyl-ACP thioesterase with specificity for shorter acyl-ACP molecules is capable of associating with the FAS complex and catalyzing the termination of fatty acid elongation to produce medium-chain fatty acids. It was previously shown that all enzymes tested catalyzed formation of octanoic acid in an E. coli system (McMahon and Prather 2014; Sherkhanov et al. 2014), yet decanoic acid was the product in most cases when they were expressed in Y. lipolytica.
It is unknown how a plant or bacterial acyl-ACP thioesterase is capable of associating with the fungal type I FAS, whose structure is known to orient most of the pertinent catalytic sites within a capsule-like structure (Lomakin et al. 2007; Leibundgut et al. 2008). One possibility is that the heterologous thioesterase is somehow disrupting the natural structure of the FAS in a way that allows the thioesterase catalytic access to the growing acyl-ACP molecule. This disruption may also explain why cells expressing the thioesterases exhibit reduced growth due to the FAS functioning incorrectly. We note that the change in growth also manifests itself as a change in the color of the culture from pale white to green, possibly due to the production of the shorter-chain fatty acids or altered physiology of the cell (Fig. S3). Furthermore, in most cases, the fatty acid products are longer in Y. lipolytica than when the same enzyme is expressed in E. coli suggesting impeded access of the thioesterase to acyl-ACP substrate in the fungal FAS compared to a bacterial one, indicating that the association of the thioesterase with the FAS is not optimal yet still functional.
We observed that production of decanoic acid resulted in more than double the specific yield of lipids on a dry cell weight basis. An increase in lipid titer during the fermentation, however, was not observed for any engineered strains. This increase in specific yield was not observed in the strain expressing the U. californica acyl-ACP thioesterase. Feedback inhibition of several enzymes in the fatty acid metabolism pathway by long-chain fatty acyl-CoA is known to occur (Jiang and Cronan 1994; Ohlrogge and Jaworski 1997). Decanoic-acid-producing strains show drastically reduced levels of long-chain fatty acids compared to the wild type, potentially removing feedback inhibition and increasing productivity in these engineered strains. This effect is not observed in the octanoic-acid-producing strain, because it produces high amounts of long-chain fatty acids. As mentioned above, the disruption of the native FAS along with reduction in native fatty acids (16 and 18) and production of potentially toxic medium-chain fatty acids retards cell growth (Viegas et al. 1989). Despite the slower growth of these engineered strains, final lipid titers remain unchanged relative to the wild type and indicate that fermentation by engineered strains results in a lower final density of more productive cells.
Fatty acids naturally segregate into several classes of lipids. This segregation requires the activity of a variety of enzymes that may or may not have activity toward these newly created medium-chain fatty acids. We found non-native fatty acids in all the different lipid classes including triacylglycerides, diacylglycerides, and monoacylglycerides; sterol; steryl esters; and FFAs. This indicates that the native machinery in Y. lipolytica is capable of recognizing and modifying fatty acids as short as octanoic acid. Some engineered strains also showed altered abundances of different lipid classes as compared to the wild type. Typically, the abundance of FFAs was increased in the engineered strains with a corresponding decrease in sterol and TCA abundance. A corresponding change was not observed in the medium-chain fatty acid lipid class abundances but rather in the modification of the native long-chain fatty acids. An increase in the amount of hexadecanoic and octadecanoic acids present as FFAs was increased in all engineered strains.
In this study, we demonstrated that expression of a single acyl-ACP thioesterase with activity toward short acyl-ACP substrates in Y. lipolytica results in alteration of the length of fatty acids produced. Enzymes previously shown to produce fatty acids as short as octanoic acid in E. coli (McMahon and Prather 2014; Sherkhanov et al. 2014) were able to produce either octanoic or decanoic acid in Y. lipolytica. The production of novel medium-chain fatty acids accompanied reduction in native long-chain fatty acid production, which resulted in a reduction of cell growth and an increase in lipid yields. Finally, we found that the medium-chain fatty acids were found to incorporate in all different lipid classes. Taken together, this data shows that oleaginous yeasts can be engineered to produce medium-chain fatty acids which, in turn, results in improved flux through the fatty acid metabolism pathway. The ability of Y. lipolytica to further modify medium-chain fatty acids makes it a promising host for production of molecules derived from fatty acid precursors.
References
Ageitos JM, Vallejo JA, Veiga-Crespo P, Villa TG (2011) Oily yeasts as oleaginous cell factories. Appl Microbiol Biotechnol 90(4):1219–1227
Beopoulos A, Chardot T, Nicaud JM (2009) Yarrowia lipolytica: a model and a tool to understand the mechanisms implicated in lipid accumulation. Biochimie 91:692–696
Beopoulos A, Verbeke J, Bordes F, Guicherd M, Bressy M, Marty A, Nicaud JM (2014) Metabolic engineering for ricinoleic acid production in the oleaginous yeast Yarrowia lipolytica. Appl Microbiol Biotechnol 98(1):251–262
Blazeck J, Liu L, Knight R, Alper HS (2013) Heterologous production of pentane in the oleaginous yeast Yarrowia lipolytica. J Biotechnol 165(3):184–194
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37(8):911–917
Carroll KK (1961) Separation of lipid classes by chromatography on Florisil. J Lipid Res 2(2):135–141
Chandler DS, Qi J, Mattox W (2003) Direct repression of Splicing by transformer-2. Mol Cell Biol 23(15):5174–5185
Chen L, Zhang J, Chen WN (2014) Engineering the Saccharomyces cerevisiae β-oxidation pathway to increase medium chain fatty acid production as potential biofuel. PLoS One 9(1)
Choi YJ, Lee SY (2013) Microbial production of short-chain alkanes. Nature 502(7472):571–574
Cronan JE Jr, Rock CO (1996) Biosynthesis of membrane lipids Escherichia coli and Salmonella: cellular and molecular biology, 2nd edn. ASM Press, Washington, DC, pp 612–636
Dörmann P, Spener F, Ohlrogge JB (1993) Characterization of two acyl-acyl carrier protein thioesterases from developing Cuphea seeds specific for medium-chain-and oleoyl-acyl carrier protein. Planta 189(3):425–432
Dwivedi G, Sharma M (2014) Impact of cold flow properties of biodiesel on engine performance. Renew Sustain Energy Rev 31:650–656
Eccleston VS, Cranmer AM, Voelker TA, Ohlrogge JB (1996) Medium-chain fatty acid biosynthesis and utilization in Brassica napus plants expressing lauroyl-acyl carrier protein thioesterase. Planta 198(1):46–53
Fortman J, Chhabra S, Mukhopadhyay A, Chou H, Lee TS, Steen E, Keasling JD (2008) Biofuel alternatives to ethanol: pumping the microbial well. Trends Biotechnol 26(7):375–381
Freeman C, West D (1966) Complete separation of lipid classes on a single thin-layer plate. J Lipid Res 7(2):324–327
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Harwood JL (1988) Fatty acid metabolism. Annu Rev Plant Physiol Plant Mol Biol 39(1):101–138
Jiang P, Cronan J (1994) Inhibition of fatty acid synthesis in Escherichia coli in the absence of phospholipid synthesis and release of inhibition by thioesterase action. J Bacteriol 176(10):2814–2821
Jing F, Cantu DC, Tvaruzkova J, Chipman JP, Nikolau BJ, Yandeau-Nelson MD, Reilly PJ (2011) Phylogenetic and experimental characterization of an acyl-ACP thioesterase family reveals significant diversity in enzymatic specificity and activity. BMC Biochem 12(1):44
Landriscina C, Gnoni GV, Quagliariello E (1972) Fatty‐acid biosynthesis. Eur J Biochem 29(1):188–196
Leber C, Da Silva N (2014) Engineering of Saccharomyces cerevisiae for the synthesis of short chain fatty acids. Biotechnol Bioeng 111(2):347–358
Leibundgut M, Maier T, Jenni S, Ban N (2008) The multienzyme architecture of eukaryotic fatty acid synthases. Curr Opin Struct Biol 18:714–725
Lepage G, Roy CC (1984) Improved recovery of fatty acid through direct transesterification without prior extraction or purification. J Lipid Res 25(12):1391–1396
Liu T, Vora H, Khosla C (2010) Quantitative analysis and engineering of fatty acid biosynthesis in E. coli. Metab Eng 12(4):378–386
Liu H, Cheng T, Xian M, Cao Y, Fang F, Zou H (2014) Fatty acid from the renewable sources: a promising feedstock for the production of biofuels and biobased chemicals. Biotechnol Adv 32(2):382–389
Liu L, Pan A, Spofford C, Zhou N, Alper HS (2015) An evolutionary metabolic engineering approach for enhancing lipogenesis in Yarrowia lipolytica. Metab Eng 29:36–45
Lomakin IB, Xiong Y, Steitz TA (2007) The crystal structure of yeast fatty acid synthase, a cellular machine with eight active sites working together. Cell 129(2):319–332
Lu X, Vora H, Khosla C (2008) Overproduction of free fatty acids in E. coli: implications for biodiesel production. Metab Eng 10(6):333–339
Lynd LR, van Zyl WH, McBride JE, Laser M (2005) Consolidated bioprocessing of cellulosic biomass: an update. Curr Opin Biotechnol 16(5):577–583
Lynen F (1969) Yeast fatty acid synthase. Methods Enzymol 14:17–33
Madzak C (2015) Yarrowia lipolytica: recent achievements in heterologous protein expression and pathway engineering. Appl Microbiol Biotechnol 99(11):4559–4577
Madzak C, Tréton B, Blanchin-Roland S (2000) Strong hybrid promoters and integrative expression/secretion vectors for quasi-constitutive expression of heterologous proteins in the yeast Yarrowia lipolytica. J Mol Microbiol Biotechnol 2(2):207–216
McMahon MD, Prather KL (2014) Functional screening and in vitro analysis reveal thioesterases with enhanced substrate specificity profiles that improve short-chain fatty acid production in Escherichia coli. Appl Environ Microbiol 80:1042–1050
Moreno-Pérez AJ, Venegas-Caleron M, Vaistij FE, Salas JJ, Larson TJ, Garces R, Graham IA, Martinez-Force E (2012) Reduced expression of FatA thioesterases in Arabidopsis affects the oil content and fatty acid composition of the seeds. Planta 235(3):629–639
Moser BR (2011) Biodiesel production, properties, and feedstocks. In: Biofuels. Springer, pp 285–347
Ohlrogge JB, Jaworski JG (1997) Regulation of fatty acid synthesis. Annu Rev Plant Biol 48:109–136
Papanikolaou S, Aggelis G (2011) Lipids of oleaginous yeasts. Part II: technology and potential applications. Eur J Lipid Sci Technol 113(8):1052–1073
Pollard MR, Anderson L, Fan C, Hawkins DJ, Davies HM (1991) A specific acyl-ACP thioesterase implicated in medium-chain fatty acid production in immature cotyledons of Umbellularia californica. Arch Biochem Biophys 284(2):306–312
Qiao K, Abidi SHI, Liu H, Zhang H, Chakraborty S, Watson N, Stephanopoulos G (2015) Engineering lipid overproduction in the oleaginous yeast Yarrowia lipolytica. Metab Eng 29:56–65
Ramos MJ, Fernández CM, Casas A, Rodríguez L, Pérez Á (2009) Influence of fatty acid composition of raw materials on biodiesel properties. Bioresour Technol 100(1):261–268
Runguphan W, Keasling JD (2014) Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid-derived biofuels and chemicals. Metab Eng 21:103–113
Santos TM, Pinto A, Oliveira AV, Ribeiro HL, Caceres CA, Ito EN, Azeredo H (2014) Physical properties of cassava starch–carnauba wax emulsion films as affected by component proportions. Int J Food Sci Technol 49(9):2045–2051
Sherkhanov S, Korman TP, Bowie JU (2014) Improving the tolerance of Escherichia coli to medium-chain fatty acid production. Metab Eng 25:1–7
Steen EJ, Kang Y, Bokinsky G, Hu Z, Schirmer A, McClure A, Keasling JD (2010) Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 463(7280):559–562
Tai M, Stephanopoulos G (2013) Engineering the push and pull of lipid biosynthesis in oleaginous yeas Yarrowia lipolytica for biofuel production. Metab Eng 15:1–9
Tariq M, Ali S, Ahmad F, Ahmad M, Zafar M, Khalid N, Khan MA (2011) Identification, FT-IR, NMR (1 H and 13 C) and GC/MS studies of fatty acid methyl esters in biodiesel from rocket seed oil. Fuel Process Technol 92(3):336–341
Viegas CA, Rosa MF, Sá-Correia I, Novais JM (1989) Inhibition of yeast growth by octanoic and decanoic acids produced during ethanolic fermentation. Appl Environ Microbiol 55(1):21–28
Voelker TA, Worrell AC, Anderson L, Bleibaum J, Fan C, Hawkins DJ, Davies HM (1992) Fatty acid biosynthesis redirected to medium chains in transgenic oilseed plants. Science 257(5066):72–74
Voelker TA, Jones A, Cranmer AM, Davies HM, Knutzon DS (1997) Broad-range and binary-range acyl-acyl-carrier-protein thioesterases suggest an alternative mechanism for medium-chain production in seeds. Plant Physiol 114(2):669–677
von Wettstein‐Knowles P, Olsen JG, McGuire KA, Henriksen A (2006) Fatty acid synthesis. FEBS J 273(4):695–710
Wakil SJ (1989) Fatty acid synthase, a proficient multifunctional enzyme. Biochemistry 28(11):4523–4530
Wang G-Y, Zhang Y, Chi Z, Liu G-L, Wang Z-P, Chi Z-M (2014) Role of pyruvate carboxylase in accumulation of intracellular lipid of the oleaginous yeast Yarrowia lipolytica ACA-DC 50109. Appl Microbiol Biotechnol 1–9
White SW, Zheng J, Zhang YM, Rock CO (2005) The structural biology of type II fatty acid biosynthesis. Annu Rev Biochem 74:791–831
Xue Z, Sharpe PL, Hong SP, Yadav NS, Xie D, Short DR, Damude HG, Ruper RA, Seip JE, Wang J, Pollak DW, Bostick MW, Bosak MD, Macool DJ, Hollerbach DH, Zhang H, Arcilla DM, Bledsoe SA, Croker K, McCord EF, Tyreus BD, Jackson EN, Zhu Q (2013) Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat Biotechnol 31(8):734–740
Zhang YM, White SW, Rock CO (2006) Inhibiting bacterial fatty acid synthesis. J Biol Chem 281(26):17541–17544
Zhang B, Chen H, Li M, Gu Z, Song Y, Ratledge C, Chen W (2013) Genetic engineering of Yarrowia lipolytica for enhanced production of trans-10, cis-12 conjugated linoleic acid. Microb Cell Factories 12(1):70
Zhang H, Zhang L, Chen H, Chen YQ, Chen W, Song Y, Ratledge C (2014) Enhanced lipid accumulation in the yeast Yarrowia lipolytica by over-expression of ATP: citrate lyase from Mus musculus. J Biotechnol 192:78–84
Zheng Y-N, Li L-L, Liu Q, Yang J-M, Wang Z-W, Liu W, Xu X, Liu H, Zhao G, Xian M (2012) Optimization of fatty alcohol biosynthesis pathway for selectively enhanced production of C12/14 and C16/18 fatty alcohols in engineered Escherichia coli. Microb Cell Factories 11:65
Acknowledgments
This work was funded by the Energy Biosciences Institute. We thank Jeffrey Skerker for his help during the early stages of this project and Chris Somerville for his help with lipid analysis.
Conflict of interest
The authors declare no conflict of interest
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1144 kb)
Rights and permissions
About this article
Cite this article
Rutter, C.D., Zhang, S. & Rao, C.V. Engineering Yarrowia lipolytica for production of medium-chain fatty acids. Appl Microbiol Biotechnol 99, 7359–7368 (2015). https://doi.org/10.1007/s00253-015-6764-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6764-1