Abstract
Pichia pastoris is widely used as a host system for heterologous protein expression in both academia and industry. Production is typically accomplished by a fed-batch induction process that is known to have negative impacts on cell physiology that impose limits on both protein yields and quality. We have analysed recombinant protein production in chemostat cultures to understand the physiological responses associated with methanol-induced production of two human lysozyme variants with different degrees of misfolding by P. pastoris. Confounding variables associated with nutrient stress or growth-rate are minimised during steady-state growth in chemostats. Comparison of transcriptome-level data obtained during the non-inducing and inducing steady states identified changes in expression of only about 1 % of the genome during production of either an amyloidogenic human lysozyme variant prone to intracellular aggregation (I56T) or a misfolded but secretable variant (T70N), indicating near-complete acclimation to their production. A marked, but temporary, stress response involving both the unfolded protein response (UPR) and ER-associated degradation pathway was observed during the transient between steady states, particularly following induction of the T70N variant synthesis, and was accompanied by changes in expression of around 50 antisense transcripts. The results suggest that optimal heterologous protein production could best be achieved by a continuous process that minimises the number of methanol-induced transients experienced by the cultures. The processing of HAC1 mRNA required for the UPR was found to be constitutive in the culture conditions used, even in the absence of recombinant protein induction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mechanisms for achieving the efficient production of recombinant proteins (r-proteins) of therapeutic value by mammalian or microbial cell systems are the subject of continuing and intensive investigation (Carrondo and Fussenegger 2005; Gellissen 2006; Lorence 2012). Despite all these efforts, we still have only a limited understanding of the response of host networks to the perturbations caused by the high-level expression of r-protein (e.g. Carvalho et al. 2012). Mammalian cell lines should, in principle, be the best choice for the expression of human proteins with complex features that demand homologous post-translational systems for either their folding or modification (Braakman and Bulleid 2011). However, the relative low secretory capacity of mammalian cell lines, their complex and expensive nutritional requirements, and their high sensitivity to the mechanical and environmental stresses inherent to industrial processes (Walter and Ron 2011; Upton et al. 2012) means that robust microbial hosts with high secretory capacity (e.g. yeasts and filamentous fungi: Wolf et al. 2003; Mattanovich et al. 2012; Mokdad-Gargouri et al. 2012; Lorence 2012) offer an attractive alternative.
Pichia pastoris (syn. Komagataella pastoris) (e.g. GS115 haploid strain; De Schutter et al. 2009) is a host system for heterologous protein expression that has been widely used in both academia and industry. Moreover, P. pastoris has been successfully engineered, or ‘humanized’, to allow it to produce human proteins with authentic glycosylation patterns, and with biophysical and biochemical characteristics comparable to those obtained with mammalian cell lines, such as Chinese hamster ovary cells (Bollok et al. 2009; Ha et al. 2011; Liu et al. 2011; Mattanovich et al. 2012; Mokdad-Gargouri et al. 2012). Attempts to increase the yield and productivity of P. pastoris using different molecular and physiological approaches focused on the improvement of individual steps that were regarded as ‘bottlenecks’ in the pathways to r-protein production (e.g. gene dosage, promoter, growth substrates or cultivation conditions) and have had only limited success (Hohenblum et al. 2004; Resina et al. 2009; Marx et al. 2009). Therefore, increasing productivity during scale-up of processes has largely depended on trial-and-error screening, somewhat refined by the adoption of the multifactorial design of experiments (Zhao et al. 2008; Holmes et al. 2009; Jafari et al. 2011).
Until recently, an integrative, systems level approach to understand the roles of the cellular networks underlying r-protein production has largely been missing. An exception is the proteomic study carried out by Vanz et al. (2012) on the induction of the expression, in P. pastoris, of hepatitis B surface antigen (HBsAg) using the AOX1 promoter. This study revealed that induction of r-protein expression provoked two major kinds of stress response. The first was an oxidative stress response provoked by the generation of reactive oxygen species that was consequent upon the switch in the principal carbon source from glycerol to methanol in order to activate the AOX1 promoter. The second stress response related directly to the high-level production of HBsAg. This evoked the unfolded protein response (UPR), the endoplasmic reticulum-associated degradation pathway (ERAD), and the induction of vacuolar proteases and autophagy. Despite the increase in chaperone and foldase levels induced by the UPR, most of these responses will reduce the final yields of r-protein that may be achieved. Moreover, these results suggest that the fed batch fermentations used for the industrial production of r-proteins by P. pastoris will repeatedly expose the producer organism to the very stresses that prevent the achievement of high product yields.
In this work, we have made a systems level approach to understand the two stress responses associated with the high-level production of an r-protein by P. pastoris. The aim of our study was to assess the response of P. pastoris at the transcriptomic level to the production of variants of a heterologous protein (human lysozyme, HuLy) with different degrees of misfolding (Johnson et al. 2005; Kumita et al. 2006; Whyteside et al. 2011a). To minimise confounding variables, including other sorts of stresses (e.g. nutrient stress; Gutteridge et al. 2010) and growth rate differences (Castrillo et al. 2007), we studied the induction of the expression of different misfolded variants of HuLY in carbon-limited continuous culture during the transition from a non-inducing condition of a sorbitol-limited steady state to inducing conditions (sorbitol + methanol). The culture was followed through the transient provoked by the medium switch until the establishment of a new steady state. We believe this investigation complements the proteomics study of Vanz et al. (2012) and has important implications for the design of industrial processes for the production of r-proteins by P. pastoris.
Materials and methods
Strains and growth conditions
The strains used in this study are derivatives of haploid Pichia (Komagataella) pastoris GS115 (De Schutter et al. 2009) expressing variants of the human lysozyme protein as previously described by Kumita et al. (2006). The expression constructs are based on the pPIC9 vector (Invitrogen) where they are under the control of the methanol inducible AOX1 promoter and directed for secretion by fusion to the α-factor secretion signal. Vectors have been integrated into the genome at the HIS4 locus resulting in His+ Mut+ strains stably expressing a T70N misfolded lysozyme variant or an I56T amyloidogenic variant. A control strain containing only the empty pPIC9 vector sequence was also used. All strains were grown in a defined sorbitol medium when heterologous lysozyme expression was not required and in a defined sorbitol–methanol medium for induction of expression. Details of the media composition are given in Supplementary Table S1. Transition from a non-inducing medium based on sorbitol to an inducing medium containing a mixture of both sorbitol and methanol as carbon source minimises inhibitory effects that can result from using only methanol for induction, without adversely affecting the strength of expression from the AOX1 promoter via carbon source mediated repression (Ramón et al. 2007; Jungo et al. 2007a, b; Gao et al. 2012; Wang et al. 2010).
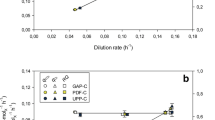
Chemostat fermentations
Chemostat fermentations, performed under carbon-limited conditions, were carried out essentially as described in Castrillo et al. (2007). Briefly, precultures of each strain were prepared by inoculating sorbitol medium (100 mL) with a single colony picked from an agar plate and incubating at 30 °C 200 rpm until reaching an OD600 between 1.0 and 1.5. Fermentors (2 L) containing sorbitol medium (900 mL) were then inoculated with the precultures and grown in batches for 24 h (30 °C, 750 rpm stirrer speed, aeration with 1 L min−1 air) before being switched to continuous mode (pH maintained at 5.0). Low dilution rates (0.014 h−1 for the control strain, and 0.005 h−1 for the T70N and I56T lysozyme variants) were used to produce growth rates comparable to fed-batch systems and to minimise issues associated with toxicity and cell lysis (Jungo et al. 2007a, b; Schenk et al. 2008). The dissolved oxygen concentration was monitored online using an Ingold polarographic oxygen electrode, and was above 40 % saturation in all cultures. Biomass was monitored offline at regular intervals by measuring UV absorbance at a wavelength of 600 nm and by dry cell weight determinations. Steady-state culture conditions were deemed to have been established once biomass and dissolved oxygen measurements had remained constant over three residence times (three vessel volume changes), at which point culture samples were taken for analysis, designated as non-induced steady-state sorbitol (SS) samples. The purity of the cultures was routinely checked by phase contrast microscopy and by plating on selective media. For induction, a pulse of methanol was added to the steady-state cultures to produce a final concentration of 1.0 g L−1 and the time of addition defined as time 0. From this time, sorbitol–methanol medium was supplied to the fermentor at the same dilution rate used prior to induction, resulting in a smooth transition to the inducible conditions. Culture samples were harvested for analysis at regular intervals following time 0, and a final sample taken after the cultures had reached a new steady state. In each case, the final samples were designated as the induced steady-state sorbitol–methanol (SM) samples.
RNA sample preparation, cDNA library construction and SOLiD sequencing
Cells from culture samples were harvested rapidly by centrifugation and the cell pellets flash frozen in liquid nitrogen and stored at −80 °C until required. For RNA purification, frozen cells were resuspended in trizol™ (Invitrogen) and lysed mechanically at 4 °C by bead beating using a FastPrep™ homogeniser (MP Biomedicals; 6 × 1-min cycles of shaking at 5 m/s). DNA and protein were removed from the lysed samples by extraction with chloroform (×2) and total RNA precipitated using 0.7 volumes of isopropanol. RNA pellets were washed in 70 % (v/v) ethanol, air-dried and resuspended in 10 mM Tris pH 8.0. Sample integrity and absence of genomic DNA contamination was confirmed by agarose gel electrophoresis. PolyA-enriched RNA was prepared from total RNA using the MicroPoly(A)Purist™ Kit (Ambion AM1919) according to the manufacturer’s instructions. RNA sequencing libraries were generated and sequenced at an MRC-funded regional sequencing centre (EASIH, Cambridge, UK) using the SOLiD 4 system, multiplexing samples so that all were sequenced on the same run to minimise variation. Libraries were prepared from polyA-enriched RNA samples using the SOLiD RNA barcoding (ABI PN4427046) and Total RNA-seq (ABI PN4445374) kits according to the manufacturers’ recommendations. The .csfasta and .qual sequencing files generated have been deposited, together with the processed data, at the EBI ArrayExpress database (http://www.ebi.ac.uk/arrayexpress/) under accession number E-MTAB-1126.
qRT-PCR analysis
qRT-PCR analyses were performed on cDNA synthesised from total RNA (1 μg) using Superscript III (Invitrogen). Each RNA sample was first subjected to RNase-free DNaseI treatment (Invitrogen) according to the manufacturer’s recommendations prior to the cDNA synthesis step. To control for DNA contamination in the qRT-PCR, a duplicate set of cDNA synthesis reactions were performed, but with the reverse transcriptase enzyme omitted. These returned abundance values comparable to background. Following RNaseH treatment, cDNA samples were diluted 1:50 in water and 2.5-μL aliquots used in the qPCR reaction using the Quantifast Sybr green PCR master mix (Qiagen). Each 25-μL reaction contained 200 nM of forward and reverse primers. PCR cycling was performed in a Rotor-Gene 6000 (Corbett Research), typically 95 °C for 5 min followed by 40 cycles of 95 °C for 15 s and 60 °C for 30 s. Parallel reactions were performed using serial dilutions of genomic DNA to produce a standard curve for each primer pair used. All determinations were performed in triplicate, and the results analysed using the Rotor-Gene 6000 series software v1.7. The primer pair 5’-TGTTTTGCCATTTTCCAACA-3’ and 5’-TAGCGAGCTCACATCTTTCG-3’ was used to quantify expression of the human lysozyme transgene. Transcription of the internal control gene actin changes markedly during the experimental conditions, and no attempt was therefore made to normalise the data. The non-normalised lysozyme transcript abundance values from qRT-PCR were in good agreement with the normalised RNA-seq expression data (data not shown).
RT-PCR analysis of HAC1 mRNA splicing
To determine the extent to which the HAC1 gene transcript was spliced in the RNA samples, RT-PCR was performed on the cDNA aliquots synthesised for the qRT-PCR analyses above using the primer pair HAC1f2 5’-AGACAGCTAGCCCACTTCCA-3’ HAC1r2 5’-ACCTCGCGTAAAATCAATGG-3’ designed to amplify across the splice junction and produce a 1,250-bp product with the unprocessed transcript, but a 900-bp product with the mature spliced mRNA. Splicing was confirmed in the 900-bp product by sequencing using the amplification primers. A control PCR using the same primers with genomic DNA as template produced the expected 1,250-bp product (see Fig. 1).
The chemostat culture system used to explore the response of P. pastoris to inducing the production of misfolded lysozyme variants. a Expression of the I56T and T70N lysozyme variants is predicted to produce distinct stresses due to differences in their secretion and intracellular retention. b Schematic representation illustrating the establishment of a non-induced steady-state culture by growth in a defined sorbitol medium followed by transition to an induced steady state in sorbitol–methanol in which the heterologous protein is continuously expressed. The timing of the SS, T+3 and SM samples analysed in Figs. 2, 3, 4 and 5 is illustrated on the x-axis
Thioflavin T staining assay
The presence of intracellular amyloid-like protein aggregates was determined by thioflavin T staining according to the method of Summers and Cyr (2011). Yeast cells were fixed by incubation on ice for 1 h with 4 % formaldehyde in 0.1 M potassium phosphate buffer (pH 6.5). Fixed cells were washed twice in ice-cold phosphate-buffered saline and resuspended in a staining solution comprising 1 M sorbitol, 0.1 M potassium phosphate (pH 7.5), 0.1 % Triton X and 0.001 % thioflavin T. After incubation for 20 min at room temperature, stained cells were washed five times in ice-cold PST (0.1 M potassium phosphate (pH 7.5), 1 M sorbitol, 0.1 % Triton X) then viewed using an Olympus BX51 microscope. Thioflavin T bound to protein amyloid aggregates fluoresces with excitation and emission wavelengths of 450 and 482 nm, respectively, and was visualised using the CFP channel filters on the microscope. Images were processed using ImageJ (Schneider et al. 2012), scoring and counting cells as staining positively with thioflavin T if they exhibited fluorescence greater than three times higher than the average background value for the image. Results were expressed as a percentage of the total cells visible in a bright field image of the same field of view, taking the average and standard deviation values from the analysis of three different fields of view.
Assay for secreted lysozyme activity
Lysozyme activity in culture supernatants was quantified using the EnzChek Lysozyme Assay Kit (Molecular Probes) according to the manufacturer’s instructions. Values were normalised to the amount of biomass (milligramme dry cell weight) present in the equivalent culture volume.
RNAseq data analysis
Sequencing reads were mapped to the Komagataella pastoris GS115 genome sequence (Ref201010.fa; BOGAS (Sterck et al. 2012)) using TopHat v1.4.1 (Trapnell et al. 2009) employing the default settings with the following exceptions: min-intron length = 25; max-intron length = 1,500; max-multihits = 10; segment mismatches = 3; site of the non-canonical splice junction for the HAC1 transcript supplied using the raw-juncs option. For the analysis of transcripts corresponding to annotated protein encoding ORFs, sense-strand reads mapped to features in the “pichia_format.gtf” genome annotation file dated 1st March 2012 (BOGAS (Sterck et al. 2012)) were counted using HTSeq v0.5.3 (Anders and Huber 2011) in intersection non-empty mode. Transcript counts were normalised and tested for differential expression between samples using the DESeq R package v1.8.3 (Anders and Huber 2010). In the absence of genuine biological replication, differential expression tests in DESeq were realised by defining the non-induced sorbitol samples “T70N SS” and “I56T SS” as pseudo-replicates, thus allowing transcript count variances to be estimated. For the identification of antisense transcripts in the genome, reads mapped to the genome sequence were assembled de novo into transcription units for each sample using cufflinks v2.0.0 (Trapnell et al. 2010) with default settings except for: min-frags-per-transfrag = 5; min-intron-length = 25; max-intron length = 1,500; min-isoform fraction = 0.025; overlap-radius = 10. The transcripts thus identified in each sample were then merged with each other, and with the list of annotated transcripts, to produce an assembly “merged.gtf” file containing both known and novel features. Features assigned class code x correspond to antisense transcription units. This merged.gtf file was used as input to the HTSeq-DESeq pipeline outlined above to test for differential expression between antisense transcripts. Principal components analyses were performed on normalised count data which had been log2(count + 1) transformed. Principal components analysis (PCA) was realised using the R package ‘pcaMethods’ (Stacklies et al. 2007).
Other computational methods
Reporter metabolite analysis was performed using the BioMet toolbox as described in Patil and Nielsen (2005) and Cvijovix et al. (2010) using the P. pastoris metabolic model iCL915 constructed by Caspeta et al. (2012). Bayesian hierarchical clustering was performed in R using the ‘BHC’ package from Bioconductor (Savage et al. 2011). Gene ontology analyses were realised using Ontologizer (Bauer et al. 2010) and the .obo gene ontology mapping file and P. pastoris proteome annotation file (34378.P_pastoris_GS115.goa) downloaded from EBI (http://www.ebi.ac.uk) on 25th June 2012.
Results
A chemostat culture system for exploring the effects of inducing heterologous expression of misfolded proteins in P. pastoris
Secretion of variants of the human lysozyme protein expressed in batch cultures of P. pastoris has been shown to correlate with the native state stability of the variant such that less stable forms yield lower levels of the secreted product (Kumita et al. 2006). Furthermore, it has been shown that less stable variants are retained at higher levels within the P. pastoris cells and to give rise to a more pronounced up-regulation of genes involved in the UPR (Whyteside et al. 2011a). To produce a system for studying the physiological consequences of the continuous production of differently misfolded proteins in P. pastoris, we selected two of these previously characterised variants, T70N and I56T, for heterologous production in chemostat cultures (Fig. 1a). The I56T form is the least stable variant known and is prone to intracellular retention and minimal secretion, while the T70N variant is both more stable than I56T and more effectively secreted (Kumita et al. 2006: Whyteside et al. 2011a). Growth of the strains in chemostat cultures and the induction of lysozyme variant production using methanol is illustrated schematically in Fig. 1b, and described in detail in the ‘Materials and methods’. Sorbitol was retained in the medium as an additional carbon source during the induction as use of this sugar alcohol as a co-substrate reduces the metabolic stress and internal toxification, which can arise from induction with methanol alone (Celik et al. 2009; Ramón et al. 2007). Both media were designed to support carbon-limited growth (all other nutrients being in excess) and provided the equivalent of 0.326 g L−1 of carbon as either sorbitol (non-inducing conditions) or sorbitol and methanol (inducing conditions). Each culture was sampled during steady-state growth in the non-inducing conditions (designated SS), 3 h after commencing the induction by addition of methanol (designated T+3), and during steady-state growth in the induced cultures (designated SM).
Inducing expression of the I56T lysozyme variant, but not T70N, increases intracellular protein aggregation, while T70N is successfully secreted
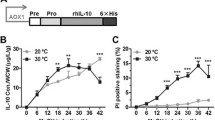
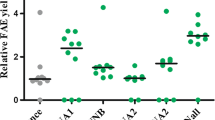
To verify that inducing expression of the lysozyme variants in the chemostat cultures was taking place and producing the expected consequences, lysozyme transcripts originating from the induction vectors were quantified using qRT-PCR and secreted lysozyme activity determined using an enzymatic assay (Fig. 2a, b). Generalised intracellular protein aggregation was also assessed via staining of cells with thioflavin T as described in the ‘Materials and methods’ (Fig. 2c). A low level of lysozyme transcription was observable in both the T70N and I56T strains prior to induction, presumably due to a basal expression of the AOX1 promoter (Fig. 2a). Methanol induction of the strain expressing the T70N variant resulted in a 6.6-fold increase in lysozyme transcript abundance, and a 35-fold increase in secreted lysozyme activity (Fig. 2a, b). No increase in the number of cells staining positive with thioflavin T was observed during the induction, but a constant 20–30 % of the population did give a positive reaction indicating a basal level of intracellular protein aggregation in these cultures (Fig. 2c). In the strain expressing the I56T lysozyme variant, transcription was induced 3.8-fold between the two steady-state conditions but peaked at 10.8-fold in the T+3 transition sample (Fig. 2a). Strikingly, the proportion of cells exhibiting protein aggregation increased approximately threefold during this time, rising from an initial 23 % in the non-induced steady-state cultures to approximately 60 % in the induced steady state (Fig. 2c). Secreted lysozyme activity was not detectable in the non-induced steady-state cultures or the T+3 transition sample, but a low level (about 9 units/mg) was present in the induced steady state (Fig. 2b). While this could result from secretion of the I56T variant, a similar profile of activity was also observed in the vector-only control culture (designated VEC in the text and “None” in the figures, as defined in Table 1), in which no lysozyme transcripts at all were detectable, suggesting that it corresponds to the induction of a small background effect. In the vector control strain, less than 6 % of cells exhibited intracellular protein aggregation in the two steady-state culture conditions (Fig. 2c).
Expression of the I56T lysozyme variant, but not T70N, increases intracellular protein aggregation in P. pastoris, while T70N is successfully secreted. a Transcription of the genes encoding the mutant lysozyme variants (T70N and I56T) during induction of heterologous expression in chemostat cultures, as compared to the control strain harbouring only the empty vector (None). RNA extracted from cultures at the non-induced steady state (SS), induced steady state (SM) and transition state (T+3) was subjected to qRT-PCR as described in the ‘Materials and methods’. Lane labels are shown below in Fig. 1d. b EnzChek lysozyme assay of culture supernatants to assess secreted lysozyme activity. Units of activity detected in the assay were normalised per milligramme of biomass. Negative values have not been normalised. c Percentage of cells giving a positive fluorescence response when stained with thioflavin T in a microscopy assay to assess intracellular protein aggregation. For each sample, images from three different fields of view were analysed and the average (± standard deviation) is shown. d RT-PCR analysis of the RNA samples from a above detects only the fully spliced 900 bp form of HAC1 mRNA, and not the unspliced 1,250-bp product readily detected by amplification of genomic DNA (G). Lanes marked M show separation of a DNA size marker ladder, and the 1,000-bp band is indicated
Splicing of the HAC1 gene transcript is constitutive and independent of heterologous lysozyme variant expression
One of the hallmarks of the UPR in eukaryotic cells is the Ire1p-mediated splicing of the HAC1 gene transcript that is required for efficient production of the Hac1p transcription factor. The extent of HAC1 mRNA splicing taking place in the different P. pastoris strains during growth and induction in the chemostat cultures was investigated by RT-PCR (Fig. 2d). The 900-bp product originating from the spliced HAC1 mRNA was readily detected in all nine samples analysed, while there was no evidence for the presence of the longer 1,250-bp unspliced transcript in any of them. Splicing of the HAC1 transcript is therefore constitutive under the culture conditions used in this study, taking place to completion not only in both the induced and non-induced conditions, but also in each of the control and lysozyme variant expression strains. Constitutive splicing of the HAC1 transcript was also observed by Guerfal et al. (2010), whereas Whyteside et al. (2011b) found that it was inducible.
Acclimation of P. pastoris cell physiology to heterologous protein production revealed at the level of transcription
Figure 2 indicates that inducing heterologous expression of the I56T lysozyme variant results in increased intracellular protein aggregation, while similar expression of the T70N variant results in lysozyme secretion and has no effect on protein aggregation. To study the impact of inducing these different cellular states in the yeast chemostat cultures, transcription profiling by RNAseq was used to determine how gene expression at the mRNA level changed following induction of the T70N and I56T variants with respect to the control strain. Polyadenylylated mRNA was purified from the total RNA samples used in the analyses shown in Fig. 2 and sequenced using SOLiD™ platform chemistry. Between ~7.2 and 11.6 million of the 50-bp reads obtained mapped to the P. pastoris genome for each sample (Table 1), and these were processed to calculate normalised transcript abundance counts for each annotated protein encoding ORF in the genome using HTSeq and DESeq as described in the ‘Materials and methods’ (Supplementary data S1). Calculation of the log2(count + 1) transform of the normalised data produced transformed values suitable for determining the Pearson correlation coefficients between each time point within each strain (Fig. 3a) and for interrogation using principal components analysis (PCA; Fig. 3b). These high-level comparisons indicate that, for each strain, transcription in the methanol induction steady-state sample (SM) is more similar to the non-induced steady-state cells (SS) than to the induced transition phase samples (T+3). In addition, while none of the T+3 transition samples clustered together in the PCA, the six SS and SM samples were only markedly separable from each other by principal component (PC) 3, which accounts for just 11.3 % of the variability in the data. We interpret this as an indication that methanol induction causes a strong transient response which is unique to each strain. Subsequent adaptations in cell physiology then return the global pattern of transcription to steady states which are both more similar to their pre-induction steady states, and more similar to each other. In the following sections, we use an analysis of the PCA loadings (Supplementary data S2–S5) together with specific pairwise differential expression tests by DESeq (Supplementary data S6–S28) to fully characterise the significant changes in gene transcription in the different lysozyme induction experiments, with particular focus on the T+3 transition states in the T70N and I56T variants. In the absence of replicate data, significantly differentially expressed genes (p < 0.05) were identified in the DESeq analyses by defining the non-induced steady-state samples T70N SS and I56T SS as pseudo-replicates, thus allowing estimates of the variance in the normalised transcript counts to be calculated (Anders and Huber 2010).
The transcript profiles of steady-state methanol-induced samples (SM) are more similar to the steady-state samples prior to induction (SS) than to the transition samples (T+3). a Pearson correlation matrices of normalised transformed transcript abundances, where a value of 1 signifies perfect correlation and 0 represents no correlation. b PCA of normalised transformed transcript abundances: variation explained in PC1 = 40.3 %; PC2 = 21.2 %; PC3 = 11.3 % and PC4 = 10.1 %
Expression of KAR2 and PDI1 is constitutively high in the steady-state chemostat culture conditions
Transcription of the KAR2 (PAS_chr2-1_0140) and PDI1 (PAS_chr4_0844) genes is activated by the Hac1p transcription factor and has been used as an indicator to assess the occurrence and strength of the UPR in yeast cells (Whyteside et al. 2011a, b; Guerfal et al. 2010). Neither KAR2 nor PDI1 were identified as being significantly differentially expressed at the 5 % probability level in comparisons between the pre- and post-induction steady-state cultures expressing the lysozyme variants (Supplementary data S7, S9). However, both were among the top 4 % most highly expressed genes in these samples, including the pre-induction steady state of the vector control strain where they ranked 125th (PDI1) and 126th (KAR2) out of the 5,320 genes. This suggests the stimulation of a generically strong UPR by the culture conditions used and is consistent with the observed constitutive splicing of the HAC1 mRNA transcript reported above.
Induction of T70N lysozyme synthesis causes a temporary up-regulation of genes encoding respiratory enzymes that is associated with a transient up-regulation of the UPR and ERAD
PCA showed a clear separation of the T70N T+3 transition phase sample from the rest of the data by PC 2 (Fig. 3b). Genes significantly (p < 0.05) changing in expression between the T70N SS and T+3 samples were therefore identified using DESeq (Supplementary data S6). Discarding those genes also found to be significantly altered by induction in the control strain identified 240 genes that were up-regulated in the 3 h following induction of T70N lysozyme synthesis (Supplementary data S10) and 237 genes which were down-regulated (Supplementary data S11). Gene ontology (GO) analysis indicated that genes associated with respiration were significantly over-represented in the list of up-regulated genes (GO004533 cellular respiration; p value of 1.19 × 10−8) together with DNA replication (GO0006260 DNA replication; p value of 7.82 × 10−6), the septin cytoskeleton (GO0032156; p value of 7.46 × 10−3), and protein O-linked glycosylation (GO0006493; p value of 3.49 × 10−2; Supplementary data S10). Strikingly, genes in all four significant categories highlighted exhibited a strong peak in their expression in the T+3 time point of the T70N strain but not in the vector-only control strain (Fig. 4a, i–iv) and were also identified as significantly differentially expressed in a direct comparison between the T70N T+3 and VEC T+3 samples (Supplementary data S14-15). In contrast, genes from rRNA processing and ribosome biogenesis GO categories, GO0006364 and GO0042254 respectively, were significantly over-represented only in the list of genes down-regulated in the T70N T+3 sample (Fig. 4a, v; Supplementary data S11). This indicates a decrease in production of the translational apparatus at this time and suggests that the observed up-regulation of genes associated with respiration and DNA synthesis does not simply reflect an increase in growth rate. This is supported by a comparison with the transcription profiles of genes in the ribosomal subunit GO category GO0044391 where a markedly higher peak in expression is observed in the VEC T+3 sample than the T70N T+3 (Fig. 4a, vi). Interestingly, of the GO0044391 genes exhibiting an increased expression in the T70N strain, many encode subunits of the mitochondrial ribosome (see Supplementary data S29). The effect on genes involved in DNA synthesis could be explained by a longer residence in the S phase of the cell cycle or an increased demand for either DNA repair or the synthesis of mitochondrial DNA. The latter is supported by the observed up-regulation of the PAS_FRAGB_0014 gene encoding a protein required for mitochondrial DNA replication and PAS_chr2-1_0500 encoding a subunit of the β DNA polymerase complex implicated in DNA repair (Supplementary data S6). However, it is also interesting to note that four genes encoding cyclins (PAS_chr1-4_0440, PAS_chr2-1_0992, PAS_FRAGD_0025, PAS_chr1-1_0097) were significantly and transiently up-regulated in the T+3 sample (see Fig. 4b, v). To highlight regions of cellular metabolism that would be significantly affected by the induction of T70N variant synthesis at the T+3 time point, reporter metabolite methods were used to summarise the significant changes in expression occurring around specific metabolites (Patil and Nielsen 2005). Up to the top ten most significant metabolites (p < 0.05) are shown in Table 2 for the up- or down-regulated genes (full data is available in Supplementary data S30). Consistent with the results from the GO analyses, the reporter metabolites identified indicate that the pathways perturbed are largely involved in central carbon metabolism, particularly the TCA and glyoxylate cycles but also acyl group shuttling mediated via acetylcarnitine.
Functional characterisation of genes identified as being significantly differentially expressed in the T+3 transition phase sample following induction of the synthesis of the misfolded soluble lysozyme variant T70N. a Comparison of the expression profiles of genes in GO categories that are significantly over-represented in the up-regulated (i–iv) or down-regulated (v) genes with those for the ribosomal subunit category (vi). *Only genes significantly down-regulated in GO0042254 are shown whereas, in the other GO categories, all genes are shown. b Transcription profiles of significantly up-regulated genes that are discussed in the text. c Expression profiles of genes encoding enzymes in the glycerol metabolism and fatty acid synthase GO categories which were identified as being significantly enriched in a comparison between the T70N T+3 and VEC T+3 samples. Expression profiles for the genes included in this Figure are presented individually in Supplementary data S29
Expression of the KAR2 and PDI1 genes, although already constitutively very high, was significantly further up-regulated by around twofold in the T70N T+3 transition phase sample (Fig. 4b, i and Supplementary data S10). Following Bayesian hierarchical clustering (BHC) of the significantly up-regulated genes, the transcription profiles of both genes clustered together along with many of the genes from the respiration, DNA synthesis and protein glycosylation GO categories (BHC cluster 11 in Supplementary data S10). GO analysis of all the genes present in cluster 11 also revealed a significant enrichment for the GO0048475 membrane coat category and a total of seven genes encoding components of the COPI or COPII vesicles were identified as being significantly up-regulated in the T70N T+3 sample (Fig. 4b, ii). This indicates an increase in both retrograde and anterograde protein transport between the ER and the Golgi apparatus that is not observed during induction of the control strain and occurs only to a markedly reduced extent when expression of the I56T lysozyme variant is induced. A further nine genes encoding functions associated with the ER or Golgi were identified among the genes whose expression was transiently up-regulated following induction of T70N lysozyme synthesis (see Fig. 4b, vi) providing additional evidence for activation of an adaptive response in these organelles. Interestingly, these include PAS_chr1-3_0202 encoding the major subunit of the Sec61 complex which forms a channel in the ER membrane and mediates translocation of secretory and membrane proteins into the ER and also retrograde transport of misfolded proteins to the cytoplasm for degradation (reviewed in Römisch 1999). Two genes encoding protein chaperones were also induced, PAS_chr1-3_0174 and PAS_chr2-1_0322, the latter encoding calnexin which is a resident in the ER and promotes correct folding of proteins entering the secretory pathway and also the targeting of misfolded proteins for degradation by the proteasome in ERAD (reviewed in Määttänen et al. 2010). PAS_chr2-1_0753 encoding an α-1,2-mannosidase involved in quality control of protein N-glycosylation in the ER was about twofold up-regulated, together with an OST3 homologue (PAS_chr4_0610) predicted to catalyse the asparagine-linked N-glycosylation of newly synthesised proteins (Schwarz et al. 2005). Consistent with the suggestion that protein glycosylation is up-regulated following the induction of T70N lysozyme synthesis, a GDA1 homologue PAS_chr4_0021 encoding a guanosine diphosphatase located in the Golgi was up-regulated approximately twofold. In Saccharomyces cerevisiae, Gda1p is involved in the transport of GDP-mannose to the Golgi lumen for protein mannosylation (Berninsone et al. 1994). A search for additional genes encoding functions associated with protein folding or mannosylation present in the list of significantly up-regulated transcripts identified four more that were at least twofold induced in the T70N T+3 sample, including two encoding GDP-mannose pyrophoshorylases (PAS_chr2-1_0093 and PAS_chr3_0870) and a co-chaperone (PAS_chr2-1_0323) to the heat shock protein Hsp90p (Fig. 4b, vii–viii). Twelve genes encoding proteases or components of the ubiquitin/proteasome proteolytic machinery were also significantly up-regulated (Fig. 4b, iii–iv). These data are all consistent with a coordinated transient up-regulation of the UPR, protein trafficking and ERAD with respiratory activity occurring in response to the increased synthesis of the misfolded T70N lysozyme variant. Interestingly, this appears to occur independently of regulation resulting from differential splicing of HAC1 mRNA (see Fig. 2), and also of HAC1 expression which was not significantly altered in the time points analysed. Five genes encoding tRNA synthetase enzymes also showed a similar pattern of up-regulation in the T70N strain (Fig. 4b, ix).
Direct comparison of the T70N T+3 and VEC T+3 samples identified 363 genes, which were significantly up-regulated in the strain expressing the lysozyme variant and 647 down-regulated genes (Supplementary data S14-S15). In addition to the processes discussed above, GO analysis of BHC cluster members from the up-regulated gene set provided evidence for an enrichment of genes involved in glycerol-3-phosphate metabolism (GO:0006072 in cluster 14; p = 0.00947) and fatty acid biosynthesis (GO:0004312 in cluster 7; p = 0.00424). Transcripts encoding both the alpha- and beta-subunits of fatty acid synthase (PAS_1-4_0295, PAS_1-1_0008) were 4.5-fold up-regulated, while those for glycerol kinase (PAS_4_0783) and the mitochondrial glycerol-3-phosphate dehydrogenase (PAS_3_0579) were approximately tenfold more abundant (Fig. 4c). In S. cerevisiae, the UPR is associated with an up-regulation of lipid biosynthesis to increase the physical capacity of the ER and Golgi (Schuck et al. 2009), and the results in Fig. 4c are consistent with this also transiently occurring in the P. pastoris strain expressing the T70N lysozyme variant.
Inducing synthesis of the insoluble I56T lysozyme variant transiently up-regulates the transcription of genes encoding functions associated with the peroxisome, nucleosome, cytoskeleton, DNA synthesis and cell cycle progression
Figures 3 and 4 illustrate that the transcriptional response induced following synthesis of the amyloidogenic I56T lysozyme variant was markedly different to that produced following a similar activation of T70N lysozyme synthesis. In particular, there was no evidence for an up-regulation of respiration or KAR2 and PDI1 gene expression, and the genes associated with the ER, Golgi and protein degradation were generally induced only to a markedly reduced extent. PCA, however, showed a clear separation of the I56T T+3 transition phase sample from the rest of the data by PC 4 (see Fig. 3b), and 224 genes whose transcript values make the most important contribution to the variability observed in this component were identified by taking those with loadings >0.03 or less than −0.03 in PC 4 (Supplementary data S5). GO analysis indicates an enrichment for genes associated with the nucleosome (GO0000786; p value of 1.43 × 10−6), peroxisome (GO00044439; p value of 8.67 × 10−6), microtubules (GO0005874; p value of 7.42 × 10−4) and the cytoskeleton (GO0044430; p value of 3.49 × 10−2) in this list of genes (Supplementary data S5), and Fig. 5a shows that expression of the genes in these categories is generally strongly but transiently up-regulated in the T+3 sample. DESeq analysis identified a total of 191 genes (147 up-regulated and 44 down-regulated) significantly (p < 0.05) changing in expression between the I56T SS and T+3 samples that did not also change in the control induction using the vector only strain (Supplementary data S12–S13). GO analysis of the down-regulated genes did not identify any significantly enriched functional categories, but confirmed that cytoskeletal and microtubule functions were over-represented among the up-regulated genes. The peroxisome and nucleosome categories were not identified, the former because gene members were also significantly up-regulated in the control strain (although to a lesser extent (see Fig. 5)), while the latter was presumably due to gene members failing the differential expression test criteria. As suggested from the data in Fig. 4a however, the DNA replication category (GO0006261; p value of 2.61 × 10−5) was found to be significantly over-represented, along with two describing the cell cycle (GO0007049 (p value 1.43 × 10−4) and GO0022402 (p value 3.63 × 10−2). This is again suggestive of a delay during the DNA synthetic S phase of the cell cycle, while the observed up-regulation of genes encoding cytoskeletal and microtubule functions may be associated with a role for cytokinesis in the clearance of aggregated proteins.
Functional characterisation of genes identified as being differentially expressed in the T+3 transition phase sample following induction of the synthesis of the misfolded amyloidogenic lysozyme variant I56T. Expression profiles of genes in GO categories that are significantly over-represented in the most influential loadings in PC4 of the PCA (a) or in the list of up-regulated genes identified in the DESeq comparison of the I56T SS and I56T T+3 samples (b). Expression profiles for the genes included in this figure are presented individually in Supplementary data S29
Only about 1 % of genes are significantly differently expressed after the chemostat cultures induced for lysozyme variant production achieve steady-state growth
Consistent with the PCA (Fig. 3b), DESeq analysis comparing the transcriptome data for the T70N SM and I56T SM samples with the corresponding sample from the vector only control (VEC SM) identified only 40 and 50 genes, respectively, that were differentially expressed at the 5 % probability level (Supplementary data S7 and S23–26). In the strain expressing the I56T lysozyme variant, 30 transcripts were down-regulated that are enriched for functions associated with the hydrolysis of O-glycosyl bonds (GO:0004553 p value 1.274 × 10−3 (and also therefore carbohydrate metabolism (GO:0005975 p value 1.107 × 10−3)) and cell wall organisation (GO:0070822 p value 1.501 × 10−2). PAS_chr1-4_0416 encoding the cell wall assembly regulator SMI1 is similarly down-regulated consistent with a generalised decrease in cell wall biosynthesis. Reporter metabolite analysis of the significantly differentially expressed genes in the I56T SM sample also highlighted metabolites associated with cell wall precursor biosynthesis as being significantly affected in the data for the down-regulated genes (Table 3 and Supplementary data S31). Yeast cell walls are extracellular structures that are dependent on a functional protein secretory pathway for their construction and maintenance, and the cell wall and ER stress responses are co-ordinately regulated in S. cerevisiae (Krysan 2009). The down-regulation of cell wall-associated processes in the induced I56T SM sample could result from a persistent inhibition of protein secretion arising from the accumulation of intracellular protein aggregates. GO analysis did not identify any significantly over-represented functional categories in the 20 up-regulated genes in this sample, nor anything of more than marginal significance in the down- and up-regulated genes (16 and 24, respectively) in the T70N SM sample.
Antisense transcription may play a role in the UPR
Construction of the cDNA libraries from the RNA samples for sequencing preserved the information on the forward or reverse strand origin of the sequenced reads, allowing previously un-annotated antisense transcripts to be identified and analysed. A total of 1,722 regions of antisense transcription were identified from a consideration of all nine samples sequenced, and 200 of these were identified as being significantly differently expressed in DESeq comparisons between the samples (Supplementary data S32). In the strain expressing the T70N lysozyme variant (discounting transcripts also changing in the vector-only control), 11 transcripts were up-regulated, and 35 down-regulated, in the T+3 transition sample compared to the non-induced steady-state sample SS (Supplementary data S31). While their functional significance is yet to be ascertained, it is interesting to note that several are antisense to ORFs with predicted functions related to those summarised in Fig. 4 (Table 4). TCONS00001972 is more than twofold down-regulated in the T+3 sample and is antisense to the 3’ end of PAS_chr1-1_0215 (Fig. 6). This gene is predicted to encode a mannosyl-oligosaccharide glucosidase orthologous to CWH41 in S. cerevisiae which is essential for the degradation of mal-folded proteins in the ER (Hitt and Wolf 2004). Down-regulation of the antisense transcript may play a role in up-regulating synthesis of this enzyme in P. pastoris. A transcript antisense to a putative glycopeptidase-encoding gene PAS_chr4_0590 is also similarly down-regulated, along with TCONS00007242 which is antisense to the YAK1 homologue PAS_chr3_0873. In S. cerevisiae Yak1p is a protein kinase that forms part of a glucose sensing system for controlling growth in response to glucose availability. Interestingly, the antisense transcripts significantly up-regulated in the T+3 sample include one against PAS_chr1-1_0300 predicted to encode a plasma membrane localised sugar sensor, and a full-length antisense transcript to 6-phosphofructo-2-kinase PAS_chr2-1_0624. These changes may consequently play a role in the reorganisation of carbon metabolism accompanying the observed up-regulation of genes encoding respiratory enzymes in the T70N T+3 cultures (see above). Significant changes in the expression of three transcripts (TCONS00003504, TCONS00003515 and TCONS00005855) antisense to genes predicted to encode functions associated with mRNA processing or translation may reflect generalised changes in protein synthesis while recovering from UPR stress, as suggested by the observed down-regulation of genes associated with ribosome biogenesis in Fig. 4a, v. Up-regulation of the TCONS00000499 transcript antisense to PAS_chr1-1_0203 encoding the general amino acid permease may similarly reflect a decreased demand for substrates for the translational machinery.
An antisense transcript to the 3’ end of PAS_chr1-1_0215 (a) is significantly down-regulated in T70N T+3. PAS_chr1-1_0215 encodes a putative mannosyl-oligosaccharide glucosidase orthologous to CWH41 in S. cerevisiae which is essential for the degradation of mal-folded proteins in the ER (Hitt and Wolf 2004)
In the I56T SM sample, where cells exhibit the maximum protein aggregation, five antisense transcripts were significantly up-regulated compared to the control sample VEC SM, and ten down-regulated (Supplementary data S34). Antisense transcripts to ORFs with an annotated function are listed in Table 5. Several are antisense to genes putatively associated with cell envelope biosynthesis and metabolism (PAS_chr1-3_0229, PAS_chr1-1_0203, PAS_FragB_0067, PAS_chr1-1_0130, PAS_chr2-1_0807), supporting the results from the analysis of sense transcription where cell wall-associated processes were identified as being significantly affected. A transcript (TCONS_00005654) antisense to the putative peptidyl-prolyl cis-trans isomerase gene PAS_chr3_0378 is also down-regulated in the I56T SM sample and may serve to increase translation of this enzyme to assist in protein folding.
Discussion
The production of recombinant proteins by P. pastoris exploits the powerful AOX1 promoter and, in normal industrial practice, the expression of a recombinant protein is repeatedly induced by methanol addition in a fed-batch process. In this paper, we have presented an integrated analysis, using ‘omic technologies and continuous culture that followed the induction process during the transition from growth on sorbitol to that on methanol/sorbitol. We have studied the expression of two unfolded variants of human lysozyme in P. pastoris and have found that the amyloidogenic I56T version of lysozyme invokes intracellular protein aggregation, while the less extreme lysozyme variant, T70N, is successfully secreted from the cell. The expression of heterologous proteins in either their native or unfolded state is known to induce the unfolded protein response (UPR) in P. pastoris (Gasser et al. 2008, 2012). The UPR is dependent on the splicing of the transcript of the HAC1 gene (Whyteside et al. 2011b), and we found such processing of HAC1 mRNA to be constitutive under the culture conditions used in this study. Consitutively spliced HAC1 mRNA was also a feature described previously in P. pastoris under batch culture conditions and further induction of the UPR was dependent on transcriptional regulation of the HAC1 gene (Guerfal et al. 2010). Thus splicing was carried through to completion not only in both inducing (methanol/sorbitol) and non-inducing (sorbitol) conditions, but was also independent of whether, or which, lysozyme variant was expressed. It is not clear whether this constitutive splicing phenotype is a consequence of growth under nutrient limiting conditions, or whether it is characteristic of a true steady state. Interestingly, further up-regulation in the expression of genes associated with UPR and ERAD was found to be possible even from a baseline of constitutive HAC1 mRNA splicing.
In addition to valuable insights into the role of intron splicing in the UPR, our RNAseq data also revealed a possible role for untranslated RNA molecules in regulating gene action in response to the production of misfolded proteins. For example, during steady-state growth of the strain expressing the I56T lysozyme variant under inducing (methanol/sorbitol) conditions, the cells exhibited their maximum level of protein aggregation, and five antisense transcripts were significantly up-regulated and ten down-regulated compared to the empty vector control grown under similar condition. Several of these transcripts were found to be antisense to genes associated with cell envelope biosynthesis and metabolism. This confirmed the inference from the ‘sense’ transcriptome that the overproduction of a misfolded protein had a significant impact on cell wall-associated processes. Moreover, a transcript antisense to a putative peptidyl-prolyl cis-trans isomerase mRNA was also down-regulated in cells expressing the I56T variant at steady state under inducing conditions; this may serve to increase translation of this enzyme to assist in protein folding.
The most important implication of our study is that it has revealed just how unsuitable the fed-batch process, based on methanol induction, is for the production of r-proteins in their native state. We found that stress responses, including unfolded protein response, were induced within 3 h of the switch from sorbitol to methanol/sorbitol. A stress response was observed even in a strain bearing only empty vector and thus not producing r-protein, but was more extreme when an r-protein with a mild folding defect was expressed. Most strikingly, we found that once steady-state growth had been established on the new substrate combination (methanol/sorbitol), then the cells no longer exhibited stress, although they continued to produce the r-protein at high titre. All of this indicates that processes that exploit repeated methanol feeds in batch culture (even with the co-substrate sorbitol) are doomed never to produce optimal yields of r-proteins since the cells are repeatedly subjected to a stress-inducing transient which both reduces yields and promotes protein misfolding.
On the basis of our results, we would propose that optimal production of r-proteins by P. pastoris would best be achieved by an integrated continuous process for both production and recovery in which product formation is growth-linked. The use of continuous fermentation processes is rare in industrial practice, being largely confined to the production of beer in New Zealand (Davies 1988) and Quorn® mycoprotein in the UK (Trinci 1992). There are two main reasons for this: the first is the genetic instability of production microbes and the second is that the efficiency gains inherent to continuous production are lost if subsequent downstream processing steps are performed in batch mode. We believe not only that these problems are soluble, but also that continuous production has some advantages in the field of biopharmaceuticals manufacture that have been overlooked hitherto.
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106. doi:10.1186/gb-2010-11-10-r106
Anders S, Huber W (2011) HTSeq: analysing high-throughput sequencing data with Python (http://www-huber.embl.de/users/anders/HTSeq/).
Bauer S, Gagneur J, Robinson PN (2010) GOing Bayesian: model-based gene set analysis of genome-scale data. Nucleic Acids Res 38:3523–3532. doi:10.1093/nar/gkq045
Berninsone P, Miret JJ, Hirschberg CB (1994) The Golgi guanosine diphosphatase is required for transport of GDP-mannose into the lumen of Saccharomyces cerevisiae Golgi vesicles. J Biol Chem 269:207–211
Bollok M, Resina D, Valero F, Ferrer P (2009) Recent patents on the Pichia pastoris expression system: expanding the toolbox for recombinant protein production. Recent Pat Biotechnol 3:192–201. doi:10.2174/187220809789389126
Braakman I, Bulleid NJ (2011) Protein folding and modification in the mammalian endoplasmic reticulum. Annu Rev Biochem 80:71–99. doi:10.1146/annurev-biochem-062209-093836
Carrondo M, Fussenegger M (2005) Advances in heterologous gene expression. Elsevier, New York
Carvalho ND, Jørgensen TR, Arentshorst M, Nitsche BM, van den Hondel CA, Archer DB, Ram AF (2012) Genome-wide expression analysis upon constitutive activation of the HacA bZIP transcription factor in Aspergillus niger reveals a coordinated cellular response to counteract ER stress. BMC Genomics 13:350. doi:10.1186/1471-2164-13-350
Caspeta L, Shoaie S, Agren R, Nookaew I, Nielsen J (2012) Genome-scale metabolic reconstructions of Pichia stipitis and Pichia pastoris and in silico evaluation of their potentials. BMC Syst Biol 6:24. doi:10.1186/1752-0509-6-24
Castrillo JI, Zeef LA, Hoyle DC, Zhang N, Hayes A, Gardner DC, Cornell MJ, Petty J, Hakes L, Wardleworth L, Rash B, Brown M, Dunn WB, Broadhurst D, O'Donoghue K, Hester SS, Dunkley TP, Hart SR, Swainston N, Li P, Gaskell SJ, Paton NW, Lilley KS, Kell DB, Oliver SG (2007) Growth control of the eukaryote cell: a systems biology study in yeast. J Biol 6:4. doi:10.1186/jbiol54
Celik E, Calik P, Oliver SG (2009) Fed-batch methanol feeding strategy for recombinant protein production by Pichia pastoris in the presence of co-substrate sorbitol. Yeast 26:473–484. doi:10.1002/yea.1679
Cvijovic M et al (2010) BioMet Toolbox: genome-wide analysis of metabolism. Nucleic Acids Res 38(Web Server issue):W144–W149. doi:10.1093/nar/gkq404
Davies AW (1988) Continuous fermentation—30 years on. Proc 20th Conv of the Inst of Brew (Australia and New Zealand section), Brisbane, pp 165–168
De Schutter K, Lin YC, Tiels P, Van Hecke A, Glinka S, Weber-Lehmann J, Rouzé P, Van de Peer Y, Callewaert N (2009) Genome sequence of the recombinant protein production host Pichia pastoris. Nat Biotechnol 27:561–566. doi:10.1038/nbt.1544
Gao MJ, Li Z, Yu RS, Wu JR, Zheng ZY, Shi ZP, Zhan XB, Lin CC (2012) Methanol/sorbitol co-feeding induction enhanced porcine interferon-α production by P. pastoris associated with energy metabolism shift. Bioprocess Biosyst Eng 35:1125–1136. doi:10.1007/s00449-012-0697-1
Gasser B, Saloheimo M, Rinas U, Dragosits M, Rodríguez-Carmona E, Baumann K, Giuliani M, Parrilli E, Branduardi P, Lang C, Porro D, Ferrer P, Tutino ML, Mattanovich D, Villaverde A (2008) Protein folding and conformational stress in microbial cells producing recombinant proteins: a host comparative overview. Microb Cell Fact 7:11. doi:10.1186/1475-2859-7-11
Gasser B, Maurer M, Rautio J, Sauer M, Bhattacharyya A, Saloheimo M, Penttilä M, Mattanovich D (2012) Monitoring of transcriptional regulation in Pichia pastoris under protein production conditions. BMC Genomics 8:179. doi:10.1186/1471-2164-8-179
Gellissen G (2006) Production of recombinant proteins. Novel microbial and eukaryotic expression systems. Wiley, Weinhem
Guerfal M, Ryckaert S, Jacobs PP, Ameloot P, Van Craenenbroeck K, Derycke R, Callewaert N (2010) The HAC1 gene from Pichia pastoris: characterization and effect of its overexpression on the production of secreted, surface displayed and membrane proteins. Microb Cell Fact 9:49. doi:10.1186/1475-2859-9-49
Gutteridge A, Pir P, Castrillo JI, Charles PD, Lilley KS, Oliver SG (2010) Nutrient control of eukaryote cell growth: a systems biology study in yeast. BMC Biol 8:68. doi:10.1186/1741-7007-8-68
Ha S, Wang Y, Rustandi RR (2011) Biochemical and biophysical characterization of humanized IgG1 produced in Pichia pastoris. MAbs 3453–460. doi: 10.4161/mabs.3.5.16891
Hitt R, Wolf DH (2004) DER7, encoding alpha-glucosidase I is essential for degradation of malfolded glycoproteins of the endoplasmic reticulum. FEMS Yeast Res 4:815–820. doi:10.1016/j.femsyr.2004.04.002
Hohenblum H, Gasser B, Maurer M, Borth N, Mattanovich D (2004) Effects of gene dosage, promoters, and substrates on unfolded protein stress of recombinant Pichia pastoris. Biotechnol Bioeng 85:367–375. doi:10.1002/bit.10904
Holmes WJ, Darby RA, Wilks MD, Smith R, Bill RM (2009) Developing a scalable model of recombinant protein yield from Pichia pastoris: the influence of culture conditions, biomass and induction regime. Microb Cell Fact 8:35. doi:10.1186/1475-2859-8-35
Jafari R, Sundström BE, Holm P (2011) Optimization of production of the anti-keratin 8 single-chain Fv TS1-218 in Pichia pastoris using design of experiments. Microb Cell Fact 10:34. doi:10.1186/1475-2859-10-34
Johnson RJ et al (2005) Rationalising lysozyme amyloidosis: insights from the structure and solution dynamics of T70N lysozyme. J Mol Biol 352(4):823–836. doi:10.1016/j.jmb.2005.07.040
Jungo C, Marison I, von Stockar U (2007a) Regulation of alcohol oxidase of a recombinant Pichia pastoris Mut+ strain in transient continuous cultures. J Biotechnol 130:236–246. doi:10.1016/j.jbiotec.2007.04.004
Jungo C, Schenk J, Pasquier M, Marison IW, von Stockar U (2007b) A quantitative analysis of the benefits of mixed feeds of sorbitol and methanol for the production of recombinant avidin with Pichia pastoris. J Biotechnol 131:57–66. doi:10.1016/j.jbiotec.2007.05.019
Krysan DJ (2009) The cell wall and endoplasmic reticulum stress responses are coordinately regulated in Saccharomyces cerevisiae. Commun Integr Biol 2:233–235. doi:10.4161/cib.2.3.8097
Kumita JR, Johnson RJ, Alcocer MJ, Dumoulin M, Holmqvist F, McCammon MG, Robinson CV, Archer DB, Dobson CM (2006) Impact of the native-state stability of human lysozyme variants on protein secretion by Pichia pastoris. FEBS J 273:711–720. doi:10.1111/j.1742-4658.2005.05099.x
Liu L, Stadheim A, Hamuro L, Pittman T, Wang W, Zha D, Hochman J, Prueksaritanont T (2011) Pharmacokinetics of IgG1 monoclonal antibodies produced in humanized Pichia pastoris with specific glycoforms: a comparative study with CHO produced materials. Biologicals 39:205–210. doi:10.1016/j.biologicals.2011.06.002
Lorence A (2012) Recombinant gene expression. reviews and protocols. Methods in molecular biology 824, 3rd edn. Humana Press, New York
Määttänen P, Gehring K, Bergeron JJ, Thomas DY (2010) Protein quality control in the ER: the recognition of misfolded proteins. Semin Cell Dev Biol 21:500–511. doi:10.1016/j.semcdb.2010.03.006
Marx H, Mecklenbräuker A, Gasser B, Sauer M, Mattanovich D (2009) Directed gene copy number amplification in Pichia pastoris by vector integration into the ribosomal DNA locus. FEMS Yeast Res 9:1260–1270. doi:10.1111/j.1567-1364.2009.00561.x
Mattanovich D, Branduardi P, Dato L, Gasser B, Sauer M, Porro D (2012) Recombinant protein production in yeasts. Methods Mol Biol 824:329–358. doi:10.1007/978-1-61779-433-9_17
Mokdad-Gargouri R, Abdelmoula-Soussi S, Hadiji-Abbès N, Amor IY, Borchani-Chabchoub I, Gargouri A (2012) Yeasts as a tool for heterologous gene expression. Methods Mol Biol 824:359–370. doi:10.1007/978-1-61779-433-9_18
Patil KR, Nielsen J (2005) Uncovering transcriptional regulation of metabolism by using metabolic network topology. Proc Natl Acad Sci U S A 102:2685–2689. doi:10.1073/pnas.0406811102
Ramón R, Ferrer P, Valero F (2007) Sorbitol co-feeding reduces metabolic burden caused by the overexpression of a Rhizopus oryzae lipase in Pichia pastoris. J Biotechnol 130:39–46. doi:10.1016/j.jbiotec.2007.02.025
Resina D, Maurer M, Cos O, Arnau C, Carnicer M, Marx H, Gasser B, Valero F, Mattanovich D, Ferrer P (2009) Engineering of bottlenecks in Rhizopus oryzae lipase production in Pichia pastoris using the nitrogen source-regulated FLD1 promoter. N Biotechnol 25:396–403. doi:10.1016/j.nbt.2009.01.008
Römisch K (1999) Surfing the Sec61 channel: bidirectional protein translocation across the ER membrane. J Cell Sci 112:4185–4191
Savage R, Cooke E, Darkins R and Xu Y (2011) BHC: Bayesian hierarchical clustering. R package version 1.8.0. http://www.bioconductor.org/packages/release/bioc/html/BHC.html?pagewanted=all
Schenk J, Balazs K, Jungo C, Urfer J, Wegmann C, Zocchi A, Marison IW, von Stockar U (2008) Influence of specific growth rate on specific productivity and glycosylation of a recombinant avidin produced by a Pichia pastoris Mut+ strain. Biotechnol Bioeng 99:368–377. doi:10.1002/bit.21565
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. doi:10.1038/nmeth.2089
Schuck S, Prinz WA, Thorn KS, Voss C, Walter P (2009) Membrane expansion alleviates endoplasmic reticulum stress independently of the unfolded protein response. J Cell Biol 187:525–536. doi:10.1083/jcb.200907074
Schwarz M, Knauer R, Lehle L (2005) Yeast oligosaccharyltransferase consists of two functionally distinct sub-complexes, specified by either the Ost3p or Ost6p subunit. FEBS Lett 579:6564–6568. doi:10.1016/j.febslet.2005.10.063
Stacklies W, Redestig H, Scholz M, Walther D, Selbig J (2007) pcaMethods—a bioconductor package providing PCA methods for incomplete data. Bioinformatics 23:1164–1167. doi:10.1093/bioinformatics/btm069
Sterck L, Billiau K, Abeel T, Rouzé P, Van de Peer, Y (2012) ORCAE: online resource for community annotation of eukaryotes. Nat Methods 9: 1041. doi:10.1038/nmeth.2242. Online Resource: Pichia pastoris: http://bioinformatics.psb.ugent.be/orcae/overview/Picpa and https://bioinformatics.psb.ugent.be/gdb/pichia/
Summers DW, Cyr DM (2011) Use of yeast as a system to study amyloid toxicity. Methods 53:226–231. doi:10.1016/j.ymeth.2010.11.007
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25:1105–1111. doi:10.1093/bioinformatics/btp120
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515. doi:10.1038/nbt.1621
Trinci APJ (1992) Myco-protein—a 20-year overnight success story. Mycol Res 96:1–13. doi:10.1016/S0953-7562(09)80989-1
Upton JP et al (2012) IRE1α cleaves select microRNAs during ER stress to derepress translation of proapoptotic Caspase-2. Science 338(6108):818–822. doi:10.1126/science.1226191
Vanz AL, Lünsdorf H, Adnan A, Nimtz M, Gurramkonda C, Khanna N, Rinas U (2012) Physiological response of Pichia pastoris GS115 to methanol-induced high level production of the hepatitis B surface antigen: catabolic adaptation, stress responses, and autophagic processes. Microb Cell Fact 11:103. doi:10.1186/1475-2859-11-103
Walter P, Ron D (2011) The unfolded protein response: from stress pathway to homeostatic regulation. Science 334:1081–1086. doi:10.1126/science.1209038
Wang Z, Wang Y, Zhang D, Li J, Hua Z, Du G, Chen J (2010) Enhancement of cell viability and alkaline polygalacturonate lyase production by sorbitol co-feeding with methanol in Pichia pastoris fermentation. Bioresour Technol 101:1318–1323. doi:10.1016/j.biortech.2009.09.025
Whyteside G, Alcocer MJ, Kumita JR, Dobson CM, Lazarou M, Pleass RJ, Archer DB (2011a) Native-state stability determines the extent of degradation relative to secretion of protein variants from Pichia pastoris. PLoS One 6:e22692. doi:10.1371/journal.pone.0022692
Whyteside G, Nor RM, Alcocer MJ, Archer DB (2011b) Activation of the unfolded protein response in Pichia pastoris requires splicing of a HAC1 mRNA intron and retention of the C-terminal tail of Hac1p. FEBS Lett 585:1037–1041. doi:10.1016/j.febslet.2011.02.036
Wolf K, Breunig K, Barth G (2003) Non-conventional Yeasts in Genetics, Biochemistry and, Biotechnology. Springer-Verlag, Heidelberg
Zhao HL, Xue C, Wang Y, Yao XQ, Liu ZM (2008) Increasing the cell viability and heterologous protein expression of Pichia pastoris mutant deficient in PMR1 gene by culture condition optimization. Appl Microbiol Biotechnol 81:235–241. doi:10.1007/s00253-008-1666-0
Acknowledgments
This work was supported by grant BB/F004389/1 from the BBSRC’s Bioprocessing Research Industry Club to SGO and DA. We thank the Eastern Sequencing and Informatics Hub (University of Cambridge; supported by MRC grant MR/K00199X/1) for the SOLiD sequencing, and for helpful discussions about data processing. We also thank Graham Whyteside for the supply of validated strains, Janet Kumita for helpful discussions on the analysis of lysozyme aggregation and Mikko Arvas for access to a script implementing the BHC cluster analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors Andrew R. Hesketh and Juan I. Castrillo contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLSX 1471 kb)
ESM 2
(XLSX 699 kb)
ESM 3
(XLSX 639 kb)
ESM 4
(XLSX 534 kb)
ESM 5
(XLSX 671 kb)
ESM 6
(XLSX 571 kb)
ESM 7
(XLSX 91 kb)
ESM 8
(XLSX 581 kb)
ESM 9
(XLSX 61 kb)
ESM 10
(XLSX 820 kb)
ESM 11
(XLSX 622 kb)
ESM 12
(XLSX 403 kb)
ESM 13
(XLSX 135 kb)
ESM 14
(XLSX 1067 kb)
ESM 15
(XLSX 1741 kb)
ESM 16
(XLSX 796 kb)
ESM 17
(XLSX 1165 kb)
ESM 18
(XLSX 472 kb)
ESM 19
(XLSX 237 kb)
ESM 20
(XLSX 300 kb)
ESM 21
(XLSX 154 kb)
ESM 22
(XLSX 101 kb)
ESM 23
(XLSX 105 kb)
ESM 24
(XLSX 91 kb)
ESM 25
(XLSX 117 kb)
ESM 26
(XLSX 127 kb)
ESM 27
(XLSX 136 kb)
ESM 28
(XLSX 151 kb)
ESM 29
(XLSX 1340 kb)
ESM 30
(XLSX 148 kb)
ESM 31
(XLSX 90 kb)
ESM 32
(XLSX 565 kb)
ESM 33
(XLSX 3013 kb)
ESM 34
(XLSX 970 kb)
ESM 35
(PDF 85 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Hesketh, A.R., Castrillo, J.I., Sawyer, T. et al. Investigating the physiological response of Pichia (Komagataella) pastoris GS115 to the heterologous expression of misfolded proteins using chemostat cultures. Appl Microbiol Biotechnol 97, 9747–9762 (2013). https://doi.org/10.1007/s00253-013-5186-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5186-1