Abstract
Dry olive residue (DOR) is a waste product derived from olive oil extraction and has been proposed as an organic amendment. However, it has been demonstrated that a pre-treatment, such as its transformation by saprophytic fungi, is required before DOR soil application. A greenhouse experiment was designed where 0 and 50 g kg−1 of raw DOR (DOR), Coriolopsis floccosa-transformed DOR (CORDOR) and Fusarium oxysporum-transformed DOR (FUSDOR) were added to soil. Analyses of the soil chemical properties as well as the structure and relative abundance of bacterial and actinobacterial communities were conducted after 0, 30 and 60 days following amendment. The different amendments produced a slight decrease in soil pH and significant increases in carbon fractions, C/N ratios, phenols and K, with these increases being more significant after DOR application. Quantitative PCR assays of the 16S rRNA gene and PLFA analyses showed that all amendments favoured bacterial growth at 30 and 60 days, although actinobacterial proliferation was more evident after CORDOR and FUSDOR application at 60 days. Bacterial and actinobacterial DGGE multivariate analyses showed that the amendments produced structural changes in both communities, especially after 60 days of amendment. PLFA data analysis identified changes in soil microbial communities according to the amendment considered, with FUSDOR and CORDOR being less disruptive than DOR. Finally, integrated analysis of all data monitored in the present study enabled us to conclude that the greatest impact on soil properties was caused by DOR at 30 days and that soil showed some degree of resilience after this time.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The olive oil industry produces large quantities of waste by-products in olive-growing regions around the world [1]. Most of these residues are generated during the olive oil extraction process [2] which, in recent decades, has been performed mainly using the two- and three-phase method depending on what final result is required [3]. The two-phase olive oil extraction method is mostly used in Spain [4]. This system generates a liquid (olive oil) and an organic sludge (two-phase olive mill waste, TPOMW) [5]. Subsequently, TPOMW is revalorized by means of organic solvent and heat treatments to generate a low-quality olive oil and a by-product called dry olive residue (DOR) or “alpeorujo” [6]. In Spain alone, 5 million tons of this residue are produced annually [2]. DOR is saline, acidic and contains high levels of phenols [7]. Inappropriate disposal of DOR can therefore generate (i) negative effects on the physical, chemical and biological properties of soil, (ii) phytotoxic effects and (iii) groundwater pollution [8]. For these reasons, it is necessary to develop strategies for the correct management of this waste to avoid agro-environmental hazards. The use of DOR as an organic amendment has been proposed as a possible strategy due to its high organic matter and nutritionally relevant cation content [5, 9]. Furthermore, unlike other organic wastes, this residue is free of heavy metals and pathogenic microorganisms [6]. However, due to the aforementioned potential environmental risks that the direct application of this residue to soil may produce, a DOR pre-treatment would be required before being used as an organic amendment. One of the most effective strategies proposed for DOR bioremediation is the transformation by saprophytic fungi [10, 11]. This transformation of DOR stabilizes organic matter, decreases its C/N ratio, reduces the phenolic fraction and eliminates phytotoxic effects [12].
In the Mediterranean region, many soils are sensitive to erosion and structural deterioration due to specific ecological conditions such as aridity [13]. In this region also, the transition from traditional techniques to intensive-mechanized farming methods has produced a reduction in soil organic matter (SOM) [14]. These new practices may alter microbial community structure and composition which directly or indirectly influence the soil ecosystem, nutrient cycle activity and crop production [15]. One of the most extensive practices is the use of chemical fertilizers, which enhance crop yield but also alter soil properties and functional diversity in microbial communities [16]. Nevertheless, the maintenance of microbial functionality and composition is essential for sustainable agricultural production. In this way, it has been demonstrated that soils under an organic farming system are of higher quality and superior microbial activity than soils subjected to non-organic practices [17]. An organic amendment containing treated DOR could represent an alternative to increase the SOM content to maintain an appropriately balanced ecosystem in the Mediterranean region. However, before using treated DOR as an organic amendment, it is necessary to study the behaviour of bacterial communities in soils amended with this type of transformed residue. Within bacterial kingdom, it is also important the analysis of Actinobacteria phylum, since it has been shown to be one of the most common phyla in soil [18], playing an important role in the degradation of polymeric and xenobiotic substances [19].
Previous studies have shown the effect of raw DOR amendments on soil physical, chemical and certain biological properties, regarding soil enzymatic activity, over the long-term [6, 9]. However, to the best of our knowledge, the impact of fungi-transformed DOR on chemical soil properties or the structure and abundance of bacteria, specifically Actinobacteria, in agricultural soils has not been studied except for a preliminary study under in vitro conditions [11]. For this reason, and as it has been suggested that most potential changes in soil microbiology occur during the first weeks following application of organic amendments [20], this study aimed to assess the short-term effects of DOR transformed by the ligninolytic fungus Coriolopsis floccosa and the soil saprophytic fungus Fusarium oxysporum on selected chemical soil properties and on the abundance, structure and diversity of bacterial and actinobacterial soil communities. To obtain an integrated approach to the study’s objectives, several techniques were used: quantitative PCR (qPCR), denaturing gradient gel electrophoresis (DGGE) and phospholipid fatty acid (PLFA) analysis.
Materials and Methods
Materials
The soil used in this study was obtained from the “Cortijo Peinado” field (Granada, Spain, 37° 13′ N, 3° 45′ W). It was classified as loam (clay, 17.15 %; sand, 34.35 %; silt, 48.50 %) according to the USDA system [21]. Ten 5-kg samples were collected randomly from the Ap horizon on the plot (10,000 m2). Subsequently, the different samples were sieved (5 mm mesh) and mixed. The soil was stored for 3 days in thin mesh plastic bags at room temperature until the experiment was initiated.
DOR was supplied by an olive oil manufacturer (Sierra Sur S.A., Granada, Spain) and was frozen at −20 °C until use.
DOR Biotransformation
DOR transformation was conducted with the fungi: C. floccosa (Spanish Type Culture Collection, CECT 20449), formerly known as Coriolopsis rigida, and F. oxysporum (Mycological Culture Collection of the Department of Biological Sciences, Faculty of Exact and Natural Sciences, University of Buenos Aires, BAFC 738). For DOR transformation, polyurethane sponge (PS) cubes, 0.5 cm in width, were rinsed with water in a 1:20 (w/v) ratio and autoclaved three times prior to their use. A 1.5 g of sterilized PS cubes were placed in Erlenmeyer flasks, and 25 mL of culture medium [50 g L−1 of glucose anhydrous (Acros Organics) and 5 g L−1 of yeast extract (Fisher Chemical)] was added and again autoclaved. Subsequently, 5 mL of C. flocossa or F. oxysporum inoculum (ca. 50 mg dry weight) was aseptically added to each Erlenmeyer flask with PS and incubated at 28 °C for 7 days. After this period of time, 25 g of sterilized DOR was placed above the colonized PS. Solid-state cultures on DOR were incubated at 28 °C in the dark under stationary conditions for 30 days. Non-inoculated DOR samples were prepared and incubated as controls. DOR controls, DOR incubated with C. floccosa and DOR incubated with F. oxysporum were then autoclaved several times for complete sterilization. The different residues were sieved (2 mm) manually, homogenized and stored at 4 °C until the soil amendment experiment began.
Soil Amendment
The experiment was performed in 0.5-L pots. Untransformed DOR (DOR), DOR transformed by C. floccosa (CORDOR) and DOR transformed by F. oxysporum (FUSDOR) were added to soil pots at concentrations of 50 g kg−1. Soil samples without the residue (C) were also prepared. One sorghum plant (Sorghum bicolor) was planted in each pot. The experiment was conducted in a greenhouse with supplementary light at 25/19 °C and 50 % relative humidity. Regular manual watering was provided during the experiment. The soil watering ensured that water content of the samples was 15–20 %.
The soil without the residue and amended with DOR, CORDOR and FUSDOR was analysed after 0, 30 and 60 days of treatment. The experiment consisted of five pots of each treatment at all sampling times. In each soil sampling, the soil from the five pots was mixed, homogenized and sieved (2 mm mesh). Subsequently, three 100-g soil subsamples for each treatment were placed in sterile Falcon™ tubes and stored at −80 °C until sample analysis was initiated.
The sorghum plants were harvested at 30 and 60 days, and shoot dry weight was measured after the plants were kept for 48 h in a dried oven.
DOR and Soil Chemical Analyses
The pH and electrical conductivity (EC) of DOR, CORDOR and FUSDOR as well as the soil samples were determined in a 1:5 (w/v) amendment/water extract and in a 1:10 (w/v) soil/water extract, respectively [9, 22]. The phenolic content of amendments (1 g) and the different soil samples (0.5 g) was determined by extraction with a 10 mL distilled water/acetone mixture (50:50, v/v) for 24 h under orbital shaking (200 rpm). Total phenolic content was estimated according to Sampedro et al. [23], using tannin acid as the standard. Total concentrations of K, Ca, Mg, Na and P of amendments and soil samples were determined by digestion with HNO3 and H2O2, followed by analysis using inductively coupled plasma optical emission spectrometry (ICP-OES) (ICP 720-ES, Agilent, Santa Clara, USA). The analyses were performed by the Instrumental Technical Services of EEZ-CSIC, Granada, Spain. The dissolved organic carbon (DOC) of soil was extracted with de-ionized water at 1:10 (soil/water) and determined by the wet oxidation method [24]. The reaction was conducted with 3 mL K2Cr2O7 and 6 mL H2SO4, and the Cr3+ resulting from organic C oxidation were determined using spectrophotometry (590 nm). The measurements of organic carbon (Corg), total carbon (Ctot) and N (Ntot) from amendments and soil samples were determined using the Leco TruSpec® CN system (Leco Corporation, St. Joseph, USA) after dry combustion of the samples. The C/N ratio was calculated as Corg/Ntot. Colour, chemical oxygen demand (COD) and ergosterol content of DOR, CORDOR and FUSDOR were determined according to Sampedro et al. [23], Brozzoli et al. [25] and Šnajdr et al. [26], respectively.
DNA Extraction
DNA was extracted from 0.25 g of the different soil treatments using the MoBio UltraClean Soil DNA Isolation Kit (MoBio Laboratories Inc., Solana Beach, CA, USA) following the manufacturer’s instructions. Three different DNA extractions were performed for each treatment. Subsequently, all DNAs were quantified using the QuantiFluor™ dsDNA System (Promega, Madison, USA). The DNA concentration for each extraction was standardized to a final concentration of 5 ng μL−1 and stored at −20 °C.
Quantitative PCR
qPCR was executed on the iCycler iQ5 (Bio-Rad, Hercules, CA, USA). The 16S rRNA gene amplification reactions were performed with the set of primers Eub338/Eub518 for bacteria and Actino235/Eub518 for Actinobacteria [27]. Each 25 μL reaction contained 12.5 μL iQ™ SYBR® Green Supermix (Bio-Rad, Hercules, USA), 0.5 μL per primer (10 μM) (Sigma-Aldrich Co., St. Louis, USA), 1 μL template DNA (5 ng) and 10.5 μL H2O. All the samples were analysed in triplicate on polypropylene 96-well plates under the quantitative PCR conditions described by Fierer et al. [27]. Melting curve analysis of the PCR products was conducted to ensure amplification of a single product.
The bacterial and actinobacterial standard curves were obtained by serial dilutions (ranging from 102 to 104) of genomic DNA from Enterobacter cloacae (HF954380) and Streptomyces pilosus (HF954395), respectively. The curve was obtained by plotting the Ct value as a function of the log of the copy number of the tenfold serial dilutions of genomic DNA. The relationship between Ct and the gene copy number of targets and standards was calculated as described by Yun et al. [28] using the data on the 16S rRNA gene copy number provided by Vetrovsky and Baldrian [29].
PCR-DGGE
The bacterial and actinobacterial communities in the different samples were analysed by means of PCR-DGGE. For the bacteria, the V6-V8 region of the 16S rRNA gene was amplified through the primers 968F + GC and 1401R [30]. A 40-bp GC clamp at the end 5′ of primer 968F was used. Each PCR, consisting of 2.5 μL dNTPs (2 mM), 2.5 μL NH4 buffer (10×), 1 μL MgCl2 (50 mM), 0.5 μL per primer (10 μM) (Sigma-Aldrich Co., St. Louis, USA), 0.5 μL Taq DNA Polymerase (5 U μL−1) (BIOTAQ™ DNA pol, Bioline, London, UK) and 1 μL template DNA (5 ng), was completed with H2O up to 50 μL. The PCR program was performed according to Brons and van Elsas [30]. For the Actinobacteria, the V3 and V4 region of the 16S rRNA gene was amplified with the primers 341F + GC and Act704R [31]. The PCR mixtures were the same as those for bacterial amplification, and the PCR program was performed as described by Xiao et al. [31]. All the amplifications were executed in a Mastercycler® Personal (Eppendorf, Applied Biosystems, Foster City, USA), and all PCR products were tested by electrophoresis in 1.5 % agarose gels stained with SYBR® Gold Nucleic Acid Gel Stain (Life Technologies™, Carlsbad, USA).
For the bacteria and Actinobacteria, DGGE analyses were conducted on an INGENYphorU-2 × 2 system (Ingeny International BV, Goes, The Netherlands). Polyacrilamide gels (9 %) were prepared in 1× TAE buffer (40 mM Tris base, 20 mM acetic acid and 1 mM disodium EDTA, pH 8.2). Ten microlitres of a mixture from the three different PCR products of each sample were used in the DGGE analyses. The polyacrylamide gels were made with a denaturing gradient ranging from 40 to 60 %. Gel electrophoresis was run for 16 h at 60 °C and 85 V. After completion of electrophoresis, the gels were stained with SYBR® Gold Nucleic Acid Gel Stain. The stained gel was captured using a digital camera. The image was then analysed using InfoQuest FP software (Bio-Rad Laboratories, Inc., Hercules, USA).
PLFA Analysis
Microbial lipids from soil were extracted using a mixture of chloroform-methanol-phosphate buffer (1:2:0.8, v/v/v) according to Bligh and Dyer [32]. Phospholipids were then separated using solid-phase extraction cartridges (LiChrolut Si-60, Merck, Whitehouse Station, USA), and the samples were subjected to mild alkaline methanolysis as described by Šnajdr et al. [26]. The free methyl esters of phospholipid fatty acids were analysed by gas chromatography mass spectrometry (450-GC, 240-MS ion trap detector, Varian, Walnut Creek, USA) according to Sampedro et al. [11].
Bacterial biomass (PLFAbac) was quantified as a sum of i14:0, i15:0, a15:0, 16:1ω7t, 16:1ω9, 16:1ω7, 10Me-16:0, i17:0, a17:0, cy17:0, 17:0, 10Me-17:0, 10Me-18:0 and cy19:0. Fungal biomass (PLFAfun) was estimated on the basis of 18:2ω6,9 content. Actinobacteria biomass (PLFAact) was determined according to 10Me-16:0, 10Me-17:0 and 10Me-18:0. The fatty acids found in both bacteria and fungi, such as 15:0, 16:0 and 18:1ω7, were excluded from the analysis [33]. The total content of PLFA molecules was used as a measure of total microbial biomass (PLFAtot). Several microbial ratios [G+/G− (Gram-positive bacteria/Gram-negative bacteria), F/B (PLFAfun/PLFAbac)] and stress indicators [cy/pre ((cy17:0 + cy19:0)/(16:1ω7/18:1ω7)), S/M (saturated PLFAs/monosaturated PLFAs)] were calculated [34].
Data Analysis
Statistical differences between the four treatments replicated three times at a given sampling time were analysed by ANOVA, and Tukey’s honest significance difference (HSD) test was used for multiple comparison of means at a 95 % confidence interval within treatments.
The number and area of bands from bacterial and actinobacterial DGGE analyses were used to calculate different diversity indices: species richness (S), the Shannon index (H) and evenness (J) using the PAST software package [35]. Significant differences (p ≤ 0.05) in the Shannon diversity index between each amended sample and its respective control at every sampling time were checked by using the Shannon diversity t test [36].
Principal component analysis (PCA) was employed to assess the similarity between samples and, in some cases, to find the variables which determined a specific sample ordination. PCA was firstly used to evaluate bacteria and actinobacteria DGGEs as well as PLFA data. Finally, a PCA was performed which included all the chemical and biological characteristics examined in the present study.
Results
DOR Transformation
DOR incubation with C. floccosa and F. oxysporum produced important changes in most of the parameters measured (Table 1). While the transformation caused by the fungi increased pH, colour (less dark) and Ntot in the residue, EC, phenols, Corg, C/N rate and COD decreased. According to ergosterol measurements, both fungi were able to grow using DOR as a culture medium, although F. oxysporum colonized the residue more extensively. With regard to the different mineral elements evaluated, DOR biotransformation with both fungi caused an increase in Mg content, a decrease in P and no changes in K, Ca and Na levels.
Effects of Amendments on Soil Chemical Properties and Plant Growth
The results showed that pH decreased significantly in the samples amended with DOR, CORDOR and FUSDOR at the initial sampling time with respect to the control treatment. However, at the other sampling times, only untreated DOR produced a significant reduction in soil pH (Table 2). EC only increased after addition of amendments at time 0 day with respect 30 and 60 days. Conversely, all the amendments applied caused a significant increase in soil phenol content at initial sampling time. At the other times, untransformed DOR also generated a rise in phenol concentrations. Nevertheless, it was not possible to detect significant differences in phenol concentration between the control samples and soils amended with CORDOR and FUSDOR, especially at 60 days. Regarding Ntot concentration, no significant changes were observed in the different treatments through time (Table 2). Nevertheless, the application of DOR, CORDOR and FUSDOR significantly increased levels of Ctot, Corg and DOC with respect to the control samples. The increases in C fractions were more evident in the soil amended with DOR at all sampling times (Table 2). The constant Ntot values for all the treatments at all sampling times and the increment in Corg for the amended samples produced an increment in the C/N ratio in the amended soils, an increase which was more evident in the soils amended with DOR. The concentrations of several agronomically important mineral elements (K, Ca, Mg, Na and P) were analysed, of which, only K concentration varied among treatments (Table 2).
DOR application to soil generally had a phytotoxic effect on plant growth. In particular, the growth inhibition of shoot sorghum plants grown in the presence of DOR for 30 and 60 days was approximately 74 % and 93 %, respectively. However, the CORDOR and FUSDOR amendments did not result in any significant changes in sorghum shoot dry weight with respect to the plants grown in the unamended samples (data not shown).
Effects of Amendments on Soil Bacterial Communities
Quantitative PCR
Relative abundance of bacterial 16S rRNA gene determined by qPCR did not change between treatments at initial sampling time (Fig. 1a). At 30 and 60 days, an increment in the number of 16S rRNA gene copies was observed in all the amended treatments, with the increases being amendment-type dependent at both sampling times, meaning that DOR produced greater bacterial proliferation than FUSDOR or CORDOR.
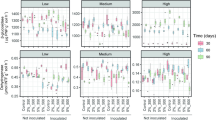
Quantification of bacterial (a) and actinobacterial (b) 16S rRNA gene copy number by means of qPCR in unamended soil (C) and soil amended with untransformed DOR (DOR), C. floccosa-transformed DOR (CORDOR) or F. oxysporum-transformed DOR (FUSDOR) at 0, 30 and 60 days. Error bars represent standard deviation. For each sampling time, data followed by different letters are significantly different according to Tukey’s HSD test (p ≤ 0.05)
In actinobacterial communities, at initial sampling time, the number of 16S rRNA gene copies did not vary among samples (Fig. 1b). At 30 days, the highest levels of actinobacterial abundance were observed in the samples amended with DOR and FUSDOR. After 60 days of residue applications, control soil and soil amended with DOR did not differ in terms of the number of 16S rRNA gene copies. However, actinobacterial proliferation was detected in the samples amended with CORDOR and FUSDOR.
DGGE Analysis
Changes in the structure and diversity of bacteria and actinobacteria communities due to the application of the different amendments at all sampling times were assessed by means of PCR-DGGE. Pre-testing showed that there were no differences in the banding patterns obtained from replicates of the same treatment. For this reason, only one profile per sample was included in the study to simplify the analysis. Bacterial DGGE analysis detected a complex band pattern with a large number of bands for each sample (Fig. 2a). Several diversity indices (S, H and J) were calculated for each sample (Supplementary Table 1). On the whole, no drastic changes in any of the indices for the samples were observed. However, the Shannon diversity t test detected significant differences in bacterial community diversity between control soil and soil amended with DOR (p < 0.01), CORDOR (p < 0.01) and FUSDOR (p < 0.01) at 30 days. PCA of DGGE profiles (Fig. 2b) showed that 37 % of the variance can be explained by two principal components. The PCA ordination of the samples did not detect any major differences in the bacterial structure between the unamended and amended samples at 0 and 30 days. In contrast, the first axis separated the treatments at 60 days; the amended samples were grouped in a cluster which differed from the control sample (Fig. 2b).
Bacterial DGGE profiles (a) and PCA based on DGGE banding patterns (b) from unamended soil (C) and soil amended with untransformed DOR (DOR), C. floccosa-transformed DOR (CORDOR) or F. oxysporum-transformed DOR (FUSDOR) at 0 (T0), 30 (T1) and 60 days (T2). Percent variability explained by each principal component is shown in parentheses after each axis legend
Actinobacteria DGGE also showed a complex band pattern, and a large number of bands could be observed in all the samples (Fig. 3a). It was not possible to detect any significant changes in actinobacteria diversity characteristics among the unamended and amended soils at the different sampling times (Supplementary Table 2). PCA showed that the two principal components accounted for 39 % of the variance (Fig. 3b). PCA grouped the samples in three clusters, with one cluster consisting of all the treatments at initial sampling time and another cluster made up of all the samples at 30 days as well as control soil and soil amended with FUSDOR at 60 days. The last group contained the remaining amended soils treated with DOR and CORDOR for 60 days.
Actinobacterial DGGE profiles (a) and PCA based on DGGE banding patterns (b) from unamended soil (C) and soil amended with untransformed DOR (DOR), C. floccosa-transformed DOR (CORDOR) or F. oxysporum-transformed DOR (FUSDOR) at 0 (T0), 30 (T1) and 60 days (T2). Percent variability explained by each principal component is shown in parentheses after each axis legend
PLFA Analysis
PLFAtot significantly increased after application of amendments at 30 and 60 days (Table 3). The different amendments also caused an increase in PLFAbac after 30 and 60 days of treatment. At 30 days, the highest bacterial proliferation levels were found in the soil amended with DOR. Conversely, PLFAbac was significantly greater in all amended soils compared to the control soil at 60 days (Table 3). With respect to the measurement of PLFAact at 30 days, only DOR produced significant proliferation levels of actinobacterial community with respect to the other treatments. However, at last sampling time, the highest actinobacteria proliferation levels were observed for the treatments with CORDOR and FUSDOR (Table 3). No change in the G+/G− ratio was observed between the unamended and amended soils at any of the sampling times. A significant increase in the F/B ratio was detected in the treatments amended with DOR and CORDOR at 30 days and soil amended with DOR after 60 days (Table 3). On the other hand, the PLFA stress indicators cy/pre and S/M were greatly affected by the application of amendments to soil, especially at 30 and 60 days. In all the treatments, the application of the different amendments produced a diminution of both ratios in relation to their respective controls (Table 3).
The PCA of the PLFA profiles showed that around 94 % of variability was explained by the first two principal components (Fig. 4). It was possible to establish three different sample groups in the PCA, thus highlighting the strong impact of the different amendments on soil microbiology. One group, in the lower left quadrant was made up of all the samples at 0 day and control samples at 30 and 60 days. Another group consisted of the soil treated with DOR at 30 and 60 days and the treatment with CORDOR at 30 days. The last group was made up of the control samples at 30 and 60 days, the soil amended with CORDOR at 60 days and soil amended with FUSDOR at 30 and 60 days.
PCA of PLFA dataset for unamended soil (C) and soil amended with untransformed DOR (DOR), C. floccosa-transformed DOR (CORDOR) or F. oxysporum-transformed DOR (FUSDOR) at 0 (T0), 30 (T1) and 60 days (T2). Percent variability explained by each principal component is shown in parentheses after each axis legend
Integrated Multivariate Analysis
The PCA of the 45 variables analysed in each soil sample in the present study (all the chemical properties of soil, diversity characteristics from bacterial and actinobacterial communities determined by means of DGGE, qPCR data for both communities and PLFA marker dataset) showed that around 83 % of the variability of the data was explained by two first principal components (71.06 and 11.68 % respectively) (Fig. 5). The 12 soil treatments assayed were grouped into four clearly defined clusters. All the control samples were part of a group located in the lower-left quadrant (Fig. 5a), which was closely related to the actinobacterial community (Fig. 5b). The amended samples at initial sampling time were located in the upper-left quadrant (Fig. 5a). This group of samples was characterised by some PLFA indices (G+/G− and S/M), the PLFA biomarker (10Me-17:0) and several chemical parameters (DOC, Corg, EC, phenols and C/N ratio), which changed drastically after the amended application (Fig. 5b). All the amended samples after 30 and 60 days of treatment were situated to the right of PC1 and PC2 separated according to sampling time. Amended 30-day samples were thus situated in the upper-right quadrant which was highly related with several PLFA biomarkers (18:1ω9, i14:0, i16:0) and PLFAtot. Amended 60-day samples were clustered in the lower-right quadrant (Fig. 5a), which was strongly influenced by several PLFA markers (18:2ω6,9, 16:1ω7, 16:1ω5, 10-Me-16:0) and the relative abundance of the bacterial 16S rRNA genes (Fig. 5b). PC1 was related to the application of amendments to the soil, and PC2 probably indicated the analysis time of the samples.
PCA scores from the different samples (a) and loadings of the chemical and biological variables (b) measured in unamended soil (C) and soil amended with untransformed DOR (DOR), C. floccosa-transformed DOR (CORDOR) or F. oxysporum-transformed DOR (FUSDOR) at 0 (T0), 30 (T1) and 60 days (T2). Percent variability explained by each principal component is shown in parentheses after each axis legend
Discussion
DOR Transformation by Saprophytic Fungi
The DOR transformation by C. floccosa and F. oxysporum produced changes in chemical waste properties, as previous works have demonstrated [10, 12]. The choice criteria of these fungi were based on the contrasting degradation behaviour of these strains towards phenols as well as on their capacity to degrade lignin [37, 38]. The phenol content reduction in the residue after incubation has been related to the production of oxidoreductases by fungi (laccases in the case of C. floccosa while Mn peroxidases and Mn-independent peroxidase activities in the case of F. oxysporum), which produce enzymatic oxidation and polymerization processes of simple phenols [39, 38]. As a result, phenolic compounds with high molecular mass are generated which are unable to pass through the plant cell membranes [40, 41]. For this reason, the soil amendment with CORDOR and FUSDOR did not produce phytotoxic effect on sorghum plants. Likewise, these high molecular mass compounds were responsible for the increment of colour in the transformed residues [42]. Alternatively, the increment in residue’s pH after the transformation with both fungi may be related to the degradation of acidic substances as well as the mineralization of organic compounds [43]. This mineralization of organic matter produced a diminution of Corg in the residue. This reduction, together with the increment in Ntot content, led to a decrease in C/N rate in CORDOR and FUSDOR, which is line with previous studies [10]. Regarding the different mineral elements evaluated, it is worth noting the drastic reduction of P in the residue after fungal transformation, which may be a consequence of the mineralization of organic P by fungi [44].
Effects of Amendments on Soil Chemical Properties
The different amendments (DOR, CORDOR and FUSDOR) tested in the present survey resulted in changes in soil chemical properties. As other studies have previously reported, soil amended with DOR produced a slight decrease in soil pH due to its high concentration of organic acids [6, 8]. However, the transformed residues did not produce changes in pH, except immediately after the application of amendments. This may be an important finding, as soil bacterial dynamics have been demonstrated to be highly sensitive to pH [45]. The application of amendments to soil produced an increase in phenol content, which was more marked for treatments with DOR. It is worth noting that we did not find significant differences in phenol concentrations between the control soil and soil amended with CORDOR and FUSDOR at 60 days. According to Piotrowska et al. [46], this is a remarkable result as soil responses to olive mill wastewater (OMW), a residue obtained from three-phase olive oil extraction system with a similar chemical composition to that of DOR [7, 47], was mainly determined by phenolic fractions in the waste. In our study, the application of the different amendments to soil did not increase Ntot content. Instead, previous studies have reported an increment in Ntot rates after long-term DOR amendments [6, 9, 48] and short-term OMW treatments [49–51]. These discrepancies in relation to our data could be due to differences in the composition of the residues, the doses used or the different type of soils tested. Conversely, we detected a sharp increase in organic carbon in all the amended treatments at each sampling time. This finding represents one of the most important advantages of using olive waste as an organic amendment in zones with degraded soils such as Mediterranean countries [4]. These improvements in organic carbon content and the constant proportion of Ntot led to a rise in C/N ratio in amended treatments, especially in soil containing DOR. The increment in C/N ratio may affect soil functionality which can involve a slowdown in the rate of organic matter mineralization [52, 53]. For this reason, some authors have suggested that nitrogen fertilization is required when olive wastes are applied to soil to reduce these ratios [54, 55]. The transformation of DOR by C. floccosa and F. oxysporum prior to its use as an amendment could therefore solve this problem as a lower C/N ratio was obtained in the samples amended with CORDOR and FUSDOR.
The application of the different amendments to soil did not produce changes in the concentrations of Ca, Mg and Na. Similar findings have also been reported in other studies following OMW soil treatments [8, 56]. On the other hand, in the present study, we did not detect any changes in P concentrations among unamended and amended soils at the different sampling times. By contrast, an increment in this mineral in soils following short- and long-time olive wastes application has previously been reported [6, 57]. These discrepancies in relation to our results could be due to differences in soil characteristics or the waste doses used. Our survey demonstrated that soil K content increased after application of the different amendments, a result which concurs with other studies [9, 58]. This rise may be beneficial for plant status as this mineral plays an important role in the stress tolerance of plants [59].
Effects of Amendments on Soil Bacterial Communities
On the whole, the soil amendment with DOR, CORDOR and FUSDOR caused an increase in bacterial biomass. This finding is in line with other studies where microorganism abundance increased after short-term olive waste treatment [49, 60]. In our study, an amendment-type-dependent rise in bacterial biomass was observed at 30 days using qPCR and PLFA techniques. Nevertheless, some discrepancies were found between data from both techniques at 60 days. Previous studies have indicated that PLFA data are more reliable than findings based on DNA as the phosphate group is rapidly hydrolysed when a cell dies [61, 62]. However, the qPCR data obtained in our experiment are in line with total viable cells and CFU counts reported in a parallel experiment [60]. In this sense, according to Drenovsky et al. [63], the best way of obtaining a reliable estimation of microbial biomass in soil is through a combination of several methodologies.
The exponential growth of bacteria in the amended treatments is related to the input of easily decomposable C sources, which favours r-strategist bacteria capable of using these nutrients to multiply to the detriment of K-strategist bacteria [64]. The microbial PLFA stress indicator ratios cy/pre and S/M decreased after amendment. High values for these indices have been explained by reductions in bacterial growth rates due to nutrient limitations [62, 65]. Thus, these findings demonstrate the beneficial effect of these amendments on bacterial growth. However, it has been widely reported that certain olive waste components such as phenols have a toxic effect on a wide variety of microorganisms [1, 57, 66] and nematodes [22]. Many authors have suggested that when raw olive wastes are applied to soil, the changes observed in microbial communities are due to complex, sometimes conflicting effects, depending on the relative amounts of beneficial, toxic organic and inorganic compounds/ions added to the residue [8, 50, 55]. On the basis of these explanations, the impact of DOR, CORDOR and FUSDOR amendments on soil bacterial communities should differ due to their different chemical compositions. Thus, although PLFA analysis showed that CORDOR and FUSDOR produced a different impact on microbial structure from that of DOR, this was detected by DGGE analysis only slightly. This could be because of the complex nature of the fingerprints obtained by this technique probably due to the high efficiency of the primers selected [30]. Indeed, we cannot be sure that the number and volume of bands for each soil sample were accurately determined and thus that changes in microbial communities caused by the different treatments were precisely gauged. Sampedro et al. [11] and Montecchia et al. [67] have actually reported that DGGE does not enable the full determination of the system complexity.
Actinobacteria play a significant role in the soil organic matter cycling due to their ability to degrade highly recalcitrant substances [50]. In addition, some microorganisms of this group have an intensive secondary metabolism which may strongly determine the dynamic of soil microbial community structure [68]. In our survey, qPCR and PLFA results have shown that DOR soil treatments cause an increase in actinobacterial biomass at 30 days with respect to the control soil. Conversely, CORDOR and FUSDOR produced this increase at 60 days, probably due to certain substances that are favourable to actinobacteria growth which were not previously available to microorganisms due to the transformation of residue caused by fungi. Mekki et al. [49] and Di Serio et al. [52] have reported a rise in actinobacteria CFU after the application of untreated and treated OMW at all sampling times. Mechri et al. [50] have also reported an increment in actinobacteria PLFA biomarkers following the application of raw OMW to soil. On the contrary, Sampedro et al. [11] showed a diminution in actinobacteria PLFA biomarkers after the incubation of soil with untransformed and transformed DOR under in vitro conditions. Thus, despite the different techniques used in these studies, there is no consensus concerning the effect of olive wastes on actinobacterial community abundance. Regarding the impact of the different amendments on actinobacterial diversity and structure, DGGE did not detect changes in Actinobacteria diversity after the addition of amendments. In contrast, we found that actinobacterial structures experienced changes depending on amendment type, with FUSDOR being the least disruptive. Karpouzas et al. [19] also detected changes in soil actinobacteria communities from two different soils following raw OMW treatments. These authors suggested that Actinobacteria are less sensitive to olive waste phenols than other groups of bacteria. However, Siles et al. [60], in a culture-dependent study, have demonstrated that the response of Actinobacteria to DOR and CORDOR depends on the taxonomic group considered.
Integrated Multivariate Analysis
In general, it has been shown that organic amendments applied to soil lead to an improvement in soil health by raising nutrient levels, increasing aggregation, reducing bulk density and increasing biological activity [69]. This is achieved directly through the intrinsic properties of the organic amendments themselves or indirectly by modifying physical, biological and chemical soil properties [70]. Nevertheless, the application of organic amendments may also introduce heavy metals, salts or recalcitrant compounds into the ecosystem, possibly leading to reduced soil functionality and affecting yield [71]. Integrated multivariate analysis (Fig. 5) has demonstrated that the chemical and biological properties analysed in the present study appear to be more sensitive to the different amendments after 30 days than at 60 days. At the latter sampling time, DOR, CORDOR and FUSDOR had a similar impact on the soil system, and PCA showed that these treatments were close to the control samples. This can be explained by the soil’s resilience following olive mill waste amendments, which has been demonstrated by other studies [72]. However, although the soil appeared to show some capacity to return to the initial properties at 60 days, the changes caused by DOR at 30 days may endanger soil functionality. Moreover, there was a phytotoxic effect of DOR on sorghum plants. In other words, the lack of phytotoxic activity of CORDOR and FUSDOR, their lower impact on soil microbial communities and their potential beneficial effect on some chemical properties demonstrate the advantages of using this type of biotransformed olive waste as an organic amendment, especially in systems for seasonal crops.
Conclusions
The soil amendment with raw DOR, C. floccosa-transformed DOR and F. oxysporum-transformed DOR led to an increase not only in organic matter and K content in the soil but also in other potentially toxic compounds such as phenols, especially when raw waste was applied. PLFA and qPCR analyses demonstrated that the incorporation of easily decomposable materials caused an increase in bacterial and actinobacterial biomass. Raw DOR favoured more bacterial multiplication than both types of transformed DOR, while C. floccosa-transformed DOR and F. oxysporum-transformed DOR generated more actinobacterial proliferation than DOR at final sampling times. On the other hand, after the application of amendments, important changes in soil bacterial and actinobacterial community structures were detected by means of DGGE and PLFA, which were probably due to alterations in the nutritional status of the soil ecosystem as well as the addition of toxic substances, although no drastic changes in the diversity of either community were detected. Integrated multivariate soil analysis showed that soil experienced the greatest chemical and biological changes following the addition of DOR at 30 days, which may alter soil functionality. Therefore, due to the lack of phytotoxic effects after CORDOR and FUSDOR application and their more limited impact on the soil properties analysed, these biotransformed wastes could be an appropriate organic amendment.
References
Justino CI, Pereira R, Freitas AC, Rocha-Santos TA, Panteleitchouk TS, Duarte AC (2012) Olive oil mill wastewaters before and after treatment: a critical review from the ecotoxicological point of view. Ecotoxicology 21:615–629. doi:10.1007/s10646-011-0806-y
Tortosa G, Alburquerque JA, Ait-Baddi G, Cegarra J (2012) The production of commercial organic amendments and fertilisers by composting of two-phase olive mill waste (“alperujo”). J Clean Prod 26:48–55. doi:10.1016/j.jclepro.2011.12.008
Alburquerque JA, Gonzalvez J, Garcia D, Cegarra J (2004) Agrochemical characterisation of “alperujo”, a solid by-product of the two-phase centrifugation method for olive oil extraction. Bioresour Technol 91:195–200. doi:10.1016/S0960-8524(03)00177-9
Roig A, Cayuela ML, Sanchez-Monedero MA (2006) An overview on olive mill wastes and their valorisation methods. Waste Manag 26:960–969. doi:10.1016/j.wasman.2005.07.024
Alburquerque JA, Gonzalvez J, Tortosa G, Baddi GA, Cegarra J (2009) Evaluation of “alperujo” composting based on organic matter degradation, humification and compost quality. Biodegradation 20:257–270. doi:10.1007/s10532-008-9218-y
López-Piñeiro A, Albarran A, Nunes JM, Barreto C (2008) Short and medium-term effects of two-phase olive mill waste application on olive grove production and soil properties under semiarid mediterranean conditions. Bioresour Technol 99:7982–7987. doi:10.1016/j.biortech.2008.03.051
Ntougias S, Bourtzis K, Tsiamis G (2013) The microbiology of olive mill wastes. BioMed Res Int 2013:784591. doi:10.1155/2013/784591
Barbera AC, Maucieri C, Cavallaro V, Ioppolo A, Spagna G (2013) Effects of spreading olive mill wastewater on soil properties and crops, a review. Agr Water Manag 119:43–53. doi:10.1016/j.agwat.2012.12.009
López-Piñeiro A, Albarrán A, Rato Nunes JM, Peña D, Cabrera D (2011) Long-term impacts of de-oiled two-phase olive mill waste on soil chemical properties, enzyme activities and productivity in an olive grove. Soil Till Res 114:175–182. doi:10.1016/j.still.2011.05.002
Sampedro I, Marinari S, D’Annibale A, Grego S, Ocampo JA, García-Romera I (2007) Organic matter evolution and partial detoxification in two-phase olive mill waste colonized by white-rot fungi. Int Biodeter Biodegr 60:116–125. doi:10.1016/j.ibiod.2007.02.001
Sampedro I, Giubilei M, Cajthaml T, Federici E, Federici F, Petruccioli M, D’Annibale A (2009) Short-term impact of dry olive mill residue addition to soil on the resident microbiota. Bioresour Technol 100:6098–6106. doi:10.1016/j.biortech.2009.06.026
Sampedro I, Cajthaml T, Marinari S, Petruccioli M, Grego S, D’Annibale A (2009) Organic matter transformation and detoxification in dry olive mill residue by the saprophytic fungus Paecilomyces farinosus. Process Biochem 44:216–225. doi:10.1016/j.procbio.2008.10.016
Toscano P, Casacchia T, Diacono M, Montemurro F (2013) Composted olive mill by-products: compost characterization and application on olive orchards. J Agric Sci Technol 15:627–638
Lal R (2006) Enhancing crop yields in the developing countries through restoration of the soil organic carbon pool in agricultural lands. Land Degrad Dev 17:197–209. doi:10.1002/ldr.696
Kibblewhite MG, Ritz K, Swift MJ (2008) Soil health in agricultural systems. Philos T Roy Soc B 363:685–701. doi:10.1098/rstb.2007.2178
Chaudhry V, Rehman A, Mishra A, Chauhan PS, Nautiyal CS (2012) Changes in bacterial community structure of agricultural land due to long-term organic and chemical amendments. Microbial Ecol 64:450–460. doi:10.1007/s00248-012-0025-y
Nautiyal CS, Chauhan PS, Bhatia CR (2010) Changes in soil physico-chemical properties and microbial functional diversity due to 14 years of conversion of grassland to organic agriculture in semi-arid agroecosystem. Soil Till Res 109:55–60. doi:10.1016/j.still.2010.04.008
Janssen PH (2006) Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72:1719–1728. doi:10.1128/AEM. 72.3.1719-1728.2006
Karpouzas DG, Ntougias S, Iskidou E, Rousidou C, Papadopoulou KK, Zervakis GI, Ehaliotis C (2010) Olive mill wastewater affects the structure of soil bacterial communities. Appl Soil Eco 45:101–111. doi:10.1016/j.apsoil.2010.03.002
Blagodatskaya Е, Kuzyakov Y (2008) Mechanisms of real and apparent priming effects and their dependence on soil microbial biomass and community structure: critical review. Biol Fert Soils 45:115–131. doi:10.1007/s00374-008-0334-y
USDA-NRCS (1996) Soil survey laboratory methods manual. Soil Survey Investigations Report N. 42, Version 3.0. USDA, Washington, DC
Cayuela ML, Millner PD, Meyer SL, Roig A (2008) Potential of olive mill waste and compost as biobased pesticides against weeds, fungi, and nematodes. Sci Total Environ 399:11–18. doi:10.1016/j.scitotenv.2008.03.031
Sampedro I, Aranda E, Martín J, García-Garrido JM, García-Romera I, Ocampo JA (2004) Saprobic fungi decrease plant toxicity caused by olive mill residues. Appl Soil Ecol 26:149–156. doi:10.1016/j.apsoil.2003.10.011
Mingorance MD, Barahona E, Fernández-Gálvez J (2007) Guidelines for improving organic carbon recovery by the wet oxidation method. Chemosphere 68:409–413. doi:10.1016/j.chemosphere.2007.01.021
Brozzoli V, Crognale S, Sampedro I, Federici F, D’Annibale A, Petruccioli M (2009) Assessment of olive-mill wastewater as a growth medium for lipase production by Candida cylindracea in bench-top reactor. Bioresour Technol 100:3395–3402. doi:10.1016/j.biortech.2009.02.022
Šnajdr J, Valášková V, Merhautová V, Cajthaml T, Baldrian P (2008) Activity and spatial distribution of lignocellulose-degrading enzymes during forest soil colonization by saprotrophic basidiomycetes. Enzyme Microb Tech 43:186–192. doi:10.1016/j.enzmictec.2007.11.008
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120. doi:10.1128/AEM. 71.7.4117-4120.2005
Yun JJ, Heisler LE, Hwang II, Wilkins O, Lau SK, Hyrcza M, Jayabalasingham B, Jin J, McLaurin J, Tsao MS, Der SD (2006) Genomic DNA functions as a universal external standard in quantitative real-time PCR. Nucleic Acids Res 34:e85. doi:10.1093/nar/gkl400
Vetrovsky T, Baldrian P (2013) The variability of the 16S rRNA gene in bacterial genomes and its consequences for bacterial community analyses. PLoS One 8:e57923. doi:10.1371/journal.pone.0057923
Brons JK, van Elsas JD (2008) Analysis of bacterial communities in soil by use of denaturing gradient gel electrophoresis and clone libraries, as influenced by different reverse primers. Appl Environ Microbiol 74:2717–2727. doi:10.1128/AEM. 02195-07
Xiao Y, Zeng GM, Yang ZH, Ma YH, Huang C, Xu ZY, Huang J, Fan CZ (2011) Changes in the actinomycetal communities during continuous thermophilic composting as revealed by denaturing gradient gel electrophoresis and quantitative PCR. Bioresour Technol 102:1383–1388. doi:10.1016/j.biortech.2010.09.034
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Phys 37:911–917. doi:10.1139/o59-099
Tornberg K, Bååth E, Olsson S (2003) Fungal growth and effects of different wood decomposing fungi on the indigenous bacterial community of polluted and unpolluted soils. Biol Fert Soils 37:190–197. doi:10.1007/s00374-002-0574-1
Moore-Kucera J, Dick RP (2008) PLFA profiling of microbial community structure and seasonal shifts in soils of a Douglas-fir chronosequence. Microbial Ecol 55:500–511. doi:10.1007/s00248-007-9295-1
Lv X-C, Weng X, Zhang W, Rao P-F, Ni L (2012) Microbial diversity of traditional fermentation starters for Hong Qu glutinous rice wine as determined by PCR-mediated DGGE. Food Control 28:426–434. doi:10.1016/j.foodcont.2012.05.025
Magurran A (1988) Ecological diversity and its measurement. Princeton University Press, Princeton
Saparrat MCN, Balatti PA, Arambarri AM, Martínez MJ (2014) Coriolopsis rigida, a potential model of white-rot fungi that produce extracellular laccases. J Ind Microbiol Biot 41:607–617. doi:10.1007/s10295-014-1408-5
Sampedro I, D’Annibale A, Ocampo JA, Stazi SR, García-Romera I (2007) Solid-state cultures of Fusarium oxysporum transform aromatic components of olive-mill dry residue and reduce its phytotoxicity. Bioresour Technol 98:3547–3554. doi:10.1016/j.biortech.2006.11.015
Aranda E, Sampedro I, Ocampo JA, García-Romera I (2006) Phenolic removal of olive-mill dry residues by laccase activity of white-rot fungi and its impact on tomato plant growth. Int Biodeter Biodegr 58:176–179. doi:10.1016/j.ibiod.2006.06.006
Hulzebos EM, Adema DMM, Dirven-van Breemen EM, Henzen L, van Gestel CAM (1991) QSARs in phytotoxicity. Sci Total Environ 109–110:493–497. doi:10.1016/0048-9697(91)90203-Q
Reina R, Liers C, Ocampo JA, García-Romera I, Aranda E (2013) Solid state fermentation of olive mill residues by wood- and dung-dwelling Agaricomycetes: effects on peroxidase production, biomass development and phenol phytotoxicity. Chemosphere 93:1406–1412. doi:10.1016/j.chemosphere.2013.07.006
Sayadi S, Allouche N, Jaoua M, Aloui F (2000) Detrimental effects of high molecular-mass polyphenols on olive mill wastewater biotreatment. Process Biochem 35:725–773. doi:10.1016/S0032-9592(99)00134-X
Paredes C, Roig A, Bernal MP, Sánchez-Monedero MA, Cegarra J (2000) Evolution of organic matter and nitrogen during co-composting of olive mill wastewater with solid organic wastes. Biol Fert Soils 32:222–227. doi:10.1007/s003740000239
Jarboui R, Sellami F, Azri C, Gharsallah N, Ammar E (2010) Olive mill wastewater evaporation management using PCA method: case study of natural degradation in stabilization ponds (Sfax, Tunisia). J Hard Mater 176:992–1005. doi:10.1016/j.jhazmat.2009.11.140
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631. doi:10.1073/pnas.0507535103
Piotrowska A, Iamarino G, Rao MA, Gianfreda L (2006) Short-term effects of olive mill waste water (OMW) on chemical and biochemical properties of a semiarid Mediterranean soil. Soil Biol Biochem 38:600–610. doi:10.1016/j.soilbio.2005.06.012
Morillo JA, Antizar-Ladislao B, Monteoliva-Sánchez M, Ramos-Cormenzana A, Russell NJ (2009) Bioremediation and biovalorisation of olive-mill wastes. Appl Microbiol Biot 82:25–39. doi:10.1007/s00253-008-1801-y
Lozano-García B, Parras-Alcántara L (2013) Short-term effects of olive mill by-products on soil organic carbon, total N, C:N ratio and stratification ratios in a Mediterranean olive grove. Agr Ecosyst Environ 165:68–73. doi:10.1016/j.agee.2012.12.007
Mekki A, Dhouib A, Sayadi S (2006) Changes in microbial and soil properties following amendment with treated and untreated olive mill wastewater. Microbiol Res 161:93–101. doi:10.1016/j.micres.2005.06.001
Mechri B, Echbili A, Issaoui M, Braham M, Elhadj SB, Hammami M (2007) Short-term effects in soil microbial community following agronomic application of olive mill wastewaters in a field of olive trees. Appl Soil Ecol 36:216–223. doi:10.1016/j.apsoil.2007.03.005
Sierra J, Marti E, Garau MA, Cruanas R (2007) Effects of the agronomic use of olive oil mill wastewater: field experiment. Sci Total Environ 378:90–94. doi:10.1016/j.scitotenv.2007.01.009
Di Serio MG, Lanza B, Mucciarella MR, Russi F, Iannucci E, Marfisi P, Madeo A (2008) Effects of olive mill wastewater spreading on the physico-chemical and microbiological characteristics of soil. Int Biodeter Biodegr 62:403–407. doi:10.1016/j.ibiod.2008.03.006
Mekki A, Dhouib A, Sayadi S (2009) Evolution of several soil properties following amendment with olive mill wastewater. Prog Nat Sci 19:1515–1521. doi:10.1016/j.pnsc.2009.04.014
Karpouzas DG, Rousidou C, Papadopoulou KK, Bekris F, Zervakis GI, Singh BK, Ehaliotis C (2009) Effect of continuous olive mill wastewater applications, in the presence and absence of nitrogen fertilization, on the structure of rhizosphere-soil fungal communities. FEMS Microbiol Ecol 70:388–401. doi:10.1111/j.1574-6941.2009.00779.x
Rousidou C, Papadopoulou K, Zervakis G, Singh BK, Ehaliotis C, Karpouzas DG (2010) Repeated application of diluted olive mill wastewater induces changes in the structure of the soil microbial community. Eur J Soil Biol 46:34–40. doi:10.1016/j.ejsobi.2009.10.004
Magdich S, Jarboui R, Rouina BB, Boukhris M, Ammar E (2012) A yearly spraying of olive mill wastewater on agricultural soil over six successive years: impact of different application rates on olive production, phenolic compounds, phytotoxicity and microbial counts. Sci Total Environ 430:209–216. doi:10.1016/j.scitotenv.2012.05.004
Di Bene C, Pellegrino E, Debolini M, Silvestri N, Bonari E (2013) Short- and long-term effects of olive mill wastewater land spreading on soil chemical and biological properties. Soil Biol Biochem 56:21–30. doi:10.1016/j.soilbio.2012.02.019
Montemurro F, Convertini G, Ferri D (2004) Mill wastewater and olive pomace compost as amendments for rye-grass. Agronomie 24:481–486. doi:10.1051/agro:2004044
López-Piñeiro A, Albarrán A, Rato Nunes JM, Peña D, Cabrera D (2011) Cumulative and residual effects of two-phase olive mill waste on olive grove production and soil properties. Soil Sci Soc Am J 75:1061–1069. doi:10.2136/sssaj2010.0230
Siles JA, Pascual J, González-Menéndez V, Sampedro I, García-Romera I, Bills GF (2014) Short-term dynamics of culturable bacteria in a soil amended with biotransformed dry olive residue. Syst Appl Microbiol 37:113–120. doi:10.1016/j.syapm.2013.08.005
Bossio DA, Scow KM (1998) Impacts of carbon and flooding on soil microbial communities: phospholipid fatty acid profiles and substrate utilization patterns. Microbial Ecol 35:265–278. doi:10.1007/s002489900082
Wixon DL, Balser TC (2013) Toward conceptual clarity: PLFA in warmed soils. Soil Biol Biochem 57:769–774. doi:10.1016/j.soilbio.2012.08.016
Drenovsky RE, Feris KP, Batten KM, Hristova K (2008) New and current microbiological tools for ecosystem ecologists: towards a goal of linking structure and function. Amer Midl Nat 160:140–159. doi:10.1674/0003-0031(2008)160[140:NACMTF]2.0.CO;2
Kotsou M, Mari I, Lasaridi K, Chatzipavlidis I, Balis C, Kyriacou A (2004) The effect of olive oil mill wastewater (OMW) on soil microbial communities and suppressiveness against Rhizoctonia solani. Appl Soil Ecol 26:113–121. doi:10.1016/j.apsoil.2003.12.001
Bach EM, Baer SG, Meyer CK, Six J (2010) Soil texture affects soil microbial and structural recovery during grassland restoration. Soil Biol Biochem 42(12):2182–2191. doi:10.1016/j.soilbio.2010.08.014
Medina E, Romero C, de Los SB, de Castro A, Garcia A, Romero F, Brenes M (2011) Antimicrobial activity of olive solutions from stored Alpeorujo against plant pathogenic microorganisms. J Agr Food Chem 59:6927–6932. doi:10.1021/jf2010386
Montecchia MS, Correa OS, Soria MA, Frey SD, García AF, Garland JL (2011) Multivariate approach to characterizing soil microbial communities in pristine and agricultural sites in Northwest Argentina. Appl Soil Ecol 47:176–183. doi:10.1016/j.apsoil.2010.12.008
Bontemps C, Toussaint M, Revol PV, Hotel L, Jeanbille M, Uroz S, Turpault MP, Blaudez D, Leblond P (2013) Taxonomic and functional diversity of Streptomyces in a forest soil. FEMS Microbiol Lett 342:157–167. doi:10.1111/1574-6968.12126
Tella M, Doelsch E, Letourmy P, Chataing S, Cuoq F, Bravin MN, Saint Macary H (2013) Investigation of potentially toxic heavy metals in different organic wastes used to fertilize market garden crops. Waste Manag 33:184–192. doi:10.1016/j.wasman.2012.07.021
Larney FJ, Angers DA (2012) The role of organic amendments in soil reclamation: a review. Can J Soil Sci 92:19–38. doi:10.4141/cjss2010-064
Cardoso EJBN, Vasconcellos RLF, Bini D, Miyauchi MYH, Santos CA, Alves PRL, Paula AM, Nakatani AS, Pereira JM, Nogueira MA (2013) Soil health: looking for suitable indicators. What should be considered to assess the effects of use and management on soil health? Sci Agric 70:274–289. doi:10.1590/S0103-90162013000400009
Piotrowska A, Rao MA, Scotti R, Gianfreda L (2011) Changes in soil chemical and biochemical properties following amendment with crude and dephenolized olive mill waste water (OMW). Geoderma 161:8–17. doi:10.1016/j.geoderma.2010.11.011
Acknowledgments
This study has been funded by the Spanish Ministry of Science and Innovation (Project AGL2008–572) and by a grant from the Competence Center TE01020218 of the Czech Technology Agency. J.A. Siles, D. Pérez-Mendoza and I. Sampedro gratefully acknowledge assistance from the JAE program, which is co-financed by the Consejo Superior de Investigaciones Científicas (CSIC) and the European Social Fund.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 81 kb)
Rights and permissions
About this article
Cite this article
Siles, J.A., Cajthaml, T., Hernández, P. et al. Shifts in Soil Chemical Properties and Bacterial Communities Responding to Biotransformed Dry Olive Residue Used as Organic Amendment. Microb Ecol 70, 231–243 (2015). https://doi.org/10.1007/s00248-014-0552-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0552-9