Abstract
Sika deer (Cervus nippon) rely on microorganisms living in the rumen to convert plant materials into chemical compounds, such as volatile fatty acids (VFAs), but how the rumen bacterial community is affected by different forages and adapt to altered diets remains poorly understood. The present study used 454-pyrosequencing of bacterial 16S ribosomal RNA (rRNA) genes to examine the relationship between rumen bacterial diversity and metabolic phenotypes using three sika deer in a 3 × 3 latin square design. Three sika deer were fed oak leaves (OL), corn stover (CS), or corn silage (CI), respectively. After a 7-day feeding period, when compared to the CS and CI groups, the OL group had a lower proportion of Prevotella spp. and a higher proportion of unclassified bacteria belonging to the families Succinivibrionaceae and Paraprevotellaceae (P < 0.05). Meanwhile, the concentration of isobutyrate was significantly lower (P < 0.05) in the OL group than in the CS and CI groups. There was no significant change of dominant bacterial genera in the OL group after 28 days of feeding. Conversely, total volatile fatty acids (TVFAs) showed an increase after 28 days of feeding, mainly due to the increasing of acetate, propionate, and valerate (P < 0.05). The interplay between bacteria and metabolism in the OL group differed from that in the CS and CI groups, especially for the interaction of TVFAs and acetate/propionate. Overall, the current study suggested that Prevotella spp. played critical roles in the fermentation of feed in the rumen of sika deer. However, the differences in interplay patterns between rumen bacterial community composition and metabolic phenotypes were altered in the native and domesticated diets indicating the changed fermentation patterns in the rumen of sika deer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The rumen harbors a complex consortium of microorganisms including bacteria (1011–1012 cells/ml), fungi (103–106 cells/ml), archaea (107–109 cells/ml), and protozoa (104–106 cells/ml) [1]. During rumen fermentation, rumen bacteria play a primary role in the conversion of feed components to peptides, ammonia, and volatile fatty acids (VFAs), such as acetate, propionate, and butyrate [2]. These end products provide the host with essential nutrients and metabolic energy (i.e., VFAs) [3], thereby affecting its performance (i.e., animal productivity) and health.
Sika deer (Cervus nippon) are ancient and primitive members of the genus Cervus because their velvet antlers are distinctive from reindeers and are prized for Chinese medicine. In addition, sika deer yield high quality meat and skin. Therefore, sika deer are farmed in China, mainly for the production of antlers. At present, the number of domesticated sika deer in China is approximately 550,000 head, most of which are distributed in northwestern China. However, very little information is known about the rumen bacterial community of sika deer, based on limited sequences from 16S ribosomal RNA (rRNA) gene clone libraries. Kobayashi et al. [4] reported that most of the bacterial species in the rumen of wild sika deer in Japan were unknown bacteria belonging to the Cytophaga–Flexibacter–Bacteroides (CFB) group. In our previous study, Prevotella spp. were the dominant bacteria in the rumen of Chinese sika deer [5]. However, the rapid advances in sequencing technologies over the past decade will allow us to comprehensively investigate the composition and densities of the rumen bacterial community of sika deer.
In nature, sika deer have been observed to prefer tannin-rich plants, such as oak leaves [6]. Additionally, in the production practice, oak leaves are beneficial to the growth of antler compared to corn silage, which is the main forage in the growing period of antler under the domesticated condition, while corn stover is the main forage of sika deer in other periods. Moreover, the hydrolysable tannin contents in oak leaves are approximately 1 %, while that in corn stover and corn silage are close to zero [7]. Thus, how forages affect the rumen bacterial community composition of sika deer and how the bacteria adapt to these diets is worthy of investigation.
Recently, several studies provided evidence for the relationship between the particular bacteria groups and fermentation products in cattle [8–12]. For example, Carberry et al. [10] identified negative associations between Prevotella spp. and isobutyrate and isovalerate concentrations in the rumen liquor. Hernandez-Sanabria et al. [11] suggested that Succinivibrio spp. were negatively correlated with valerate proportion of total VFA. Therefore, this shift in focus from determining “who is there” toward understanding “what are they doing” will help us to understand the process of the adaptation to the varied diets [13]. However, the relationship between rumen bacterial community and metabolic phenotypes of sika deer is poorly known. Moreover, the fermentation patterns in the rumen of sika deer fed native and domesticated diets may be distinct.
Using 454-pyrosequencing of bacterial 16S rRNA gene, the objectives of the present study are to (i) investigate differences of bacterial community composition in the rumen of sika deer fed native and domesticated diets; (ii) compare rumen fermentation under these different diets; (iii) explore the co-occurrence relationship between rumen bacteria and metabolic phenotypes; and (iv) test the hypothesis that the fermentation patterns in the rumen of sika deer are related to the diets.
Materials and Methods
Animals
Three rumen-cannulated adult male sika deer (C. nippon), maintained at the research farm (44.04° N, 129.09° E) of the Institute of Special Animal and Plant Sciences (ISAPS), Chinese Academy of Agricultural Sciences (CAAS), in Jilin Province, were used in this study. All animals were individually housed in pens, and all animal procedures were approved and authorized by the CAAS Animal Care and Use Committee (ACUC) and by the ISAPS Wild Animal and Plant Subcommittee.
Study Design and Sampling
Three sika deer were used in a 3 × 3 latin square design (Table S1). We firstly prepared the concentrate diet (64.5 % corn, 19.7 % soybean meal, 12.8 % distiller dried grains with solubles and a 3 % mixture of vitamins and mineral salts). And then, the same concentrate diet was mixed with either corn stover (CS group), corn silage (CI group), or oak leaves (OL group), to form three different diets. The ratio of forage to concentrate was 50:50 on the base of dry matter (DM). On a DM basis, the concentrate contained 12.68 % crude protein and 3.54 % crude fat, corn stover contained 37.57 % cellulose and 4.03 % crude protein, oak leaves contained 35.58 % cellulose and 6.40 % crude protein, and corn silage contained 69.63 % cellulose and 7.50 % crude protein. All sika deer were fed twice each day at 8:00 am and 4:00 pm and had free access to water.
After one week of adaption to the diets, sika deer received each diet for 28 days. After feeding each diet for 7, 14, and 28 days, the rumen contents including solid and liquid fractions were obtained via rumen cannula before the morning. Rumen samples were stored at −80 °C for later analysis.
DNA Extraction and 454 Pyrosequencing
Total genomic DNA was extracted from the whole rumen content of each animal using the PowerSoil® DNA Isolation Kit (MOBIO, Carlsbard, CA) according to the manufacturer’s instructions. Universal bacterial primers 8F and 533R were used to amplify the V1 to V3 regions of the bacterial 16S rRNA gene [14]. The forward primer was made up of adaptor A, a seven-nucleotide barcode, and the 8F primer, and the reverse primer contained adapter B, a seven-nucleotide barcode, and the 533R primer. The following protocol was used to amplify bacterial 16S rRNA gene sequences: 94 °C for 3 min; followed by 25 cycles for 30 s at 94 °C, 30 s at 57 °C, and 30 s at 72 °C; and a final extension at 72 °C for 10 min. The purified PCR products (approximately 200 ng) were sequenced using a Roche GS-FLX 454 pyrosequencer (Roche, Mannheim, Germany). Each sample had two PCR replicates for pyrosequencing.
Analysis of Pyrosequencing Reads
Pyrosequencing data were processed and analyzed using QIIME 1.7.0 following the pipeline described by Caporaso et al. [15]. Sequences were sorted according to barcode and were examined for quality control using the following criteria: the minimum sequence length was 400 nt; the maximum sequence length was 600 nt; minimum qual score was 25; the maximum number of errors in the barcode was 0; the maximum length of homopolymer run was 6; the number of mismatches in the primer was 0; ambiguous and unassigned characters were excluded. The adaptor B and reverse primer from all assigned sequences were also removed from the raw sequences. Sequencing errors were removed from filtered sequences using denoiser 0.91 [16]. The denoised sequence was clustered into operational taxonomic units (OTUs) using Usearch61 according to the sequence identity of 97 % [17]. Representative sequences were aligned to the Greengenes database [18] using PyNAST [19] with a minimum alignment length of 400 and a minimum identity of 75 %. Potential chimera sequences were removed using Chimera Slayer [20]. OTUs that were found in at least nine samples with at least three sequences were retained for the further analysis. Principal coordinate analysis (PCoA) and partial least squares discriminant analysis (PLS-DA) were performed on a comprehensive web server for comparative metagenomics, METAGENassist [21]. All raw sequences in this study were deposited to the NCBI Sequence Read Archive (SRP033500).
Measurement of Ruminal Fermentation Parameters
Rumen pH was determined immediately using a pH meter (model PB-10/C, Sartorius, Goettingen, Germany). The concentration of ammonia nitrogen (NH3-N) was measured using a commercial kit (R-Biopharm Roche Inc., South Marshall, MI) according to the manufacturer’s instructions at a wavelength of 340 nm (SpectraMax 190; Molecular Devices, Sunnyvale, CA). Rumen fluid was centrifuged at 15,000g for 10 min at 4 °C, and 0.2 ml of 2-ethylbutyric acid (internal standard, 2 g/L) in metaphosphoric acid (25 % w/v) was added to 1 ml of clarified rumen fluid. The concentrations of VFAs in the rumen were determined by gas chromatography with a flame ionization detector and a DB-FFAP column (30 m × 0.25 μm × 0.25 μm, Agilent Technologies 6890GC, USA). The carrier gas was N2 at a flow rate of 2.2 ml/min. The analysis was a gradient oven temperature of 80–170 °C with an incremental rate of 10 °C/min for optimal separation and a detector temperature of 250 °C.
Co-occurrence Network Analyses Between Rumen Bacteria and Metabolic Phenotypes
The co-occurrence pattern was constructed between the OTUs and the rumen fermentation profiles using the method as described by Zhang et al. [22]. Briefly, Each OTU’s abundance and the concentrations of metabolic phenotypes were calculated using the mean value from all samples in each group. Statistic P values were corrected using the false discovery rate (FDR) method of the p.adjust package in R. Correlations have an absolute Pearson’s correlation above 0.70 with an FDR-corrected significance level under 0.05, and these correlations were transformed into links between two OTUs in the co-occurrence network. The co-occurrence networks were then visualized using Cytoscape 2.8.2 with a force-directed algorithm [23].
Statistical Analysis
Statistical analysis was performed using the SigmaPlot 12.0 (Systat Software, Inc.) and R software packages. General characteristics were expressed as mean or percentages. Variations in different groups were checked for normal distribution using the Shapiro-Wilk test (significance value of P < 0.05). When normally distributed, multiple samples’ comparisons were performed using one-way analysis of variance (ANOVA) (parametric) and using Kruskal-Wallis one-way ANOVA on ranks (non-parametric) for un-normal distribution with the significant value of P < 0.05.
Results
Summary of Pyrosequencing Data
Overall, the current study obtained 661,564 raw 16S rRNA gene sequences from 54 samples. A total of 330,435 high quality sequences were obtained with an average of 6119 ± 344 (S.E) sequences for each sample. Using these sequences, we identified 2931 core OTUs based on 97 % sequence identity (equal to species level), with 783 OTUs per sample in the OL group, 815 OTUs per sample in the CS group, and 764 OTUs per sample in the CI group (Table 1).
Comparison of Bacterial Community Composition in the Three Groups
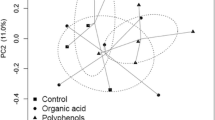
In all, these core OTUs were assigned to 12 phyla, 34 families, and 61 genera. The top three phyla were Bacteroidetes (64.9 %), Proteobacteria (17.3 %), and Firmicutes (16.1 %) (Fig.1a). Prevotella (50.69 %), Succinivibrio (11.14 %), and unclassified bacteria belonging to the family Succinivibrionaceae (6.08 %) were the top three genera in the three diet groups (Fig. 1b). Analysis of Good’s coverage showed that 93 ± 0.8 (S.E)% of bacterial species were captured in each sample (Table 1). The PCoA and PLS-DA plots showed that the bacterial community in the OL group was clearly separated from those in the CS and CI groups (Fig. 2), indicating that the native diet (OL group) potentially altered rumen bacterial community composition in sika deer.
Bacterial composition was compared between three diets across the three time points. After 7 days of feeding, there were no differences at the phylum level in the three groups (Fig. 3a). At the genus level, the OL group has a lower proportion of Prevotella spp. than the CI group (P = 0.006) and had higher abundance of unidentified bacteria belonging to the families Succinivibrionaceae and Paraprevotellaceae than the CS (P = 0.031 and P = 0.002) and CI (P = 0.043 and P = 0.002) groups (Fig. 4a). After 14 days of feeding, bacteria belonging to TM7 (OL vs CS, P = 0.018; OL vs CI, P = 0.017) and Cyanobacteria (OL vs CS, P = 0.027) were increased in the OL group (Fig. 3a). At the genus level, the unclassified bacteria belonging to the family Porphyromonadaceae (OL vs CS, P = 0.034; OL vs CI, P = 0.023) were more abundant in the OL group than the CS and CI groups, and unclassified bacteria in the family Catabacteriaceae were reduced in the OL group compared to the CI group (P = 0.043, Fig. 4a). After 28 days of feeding, bacteria belonging to the phylum Fibrobacteres were lower in the OL group than in the CI group (P = 0.029, Fig. 3a), but at the genus level, there were no differences. Moreover, for the OL group, the relative abundance of phylum Cyanobacteria was more prevalent at day 14 than at day 7 (P = 0.023) and day 28 (P = 0.027, Fig. 3b). However, there were no significant differences among the three time points.
Rumen Metabolic Phenotypes in the Three Groups Across Three Time Points
After 7 days of feeding, the concentrations of isobutyrate (OL vs CS, P < 0.001; OL vs CI, P < 0.001) and NH3-N (OL vs CS, P < 0.001; OL vs CI, P < 0.001) in the OL group were lower than in the CS and CI groups, but the ratio of acetate to propionate (AP) in the OL group was higher than in the CI group (P = 0.005, Fig. 5a).
After 14 days of feeding, the concentrations of valerate in the OL group were lower than in the CI group (P = 0.006, Fig. 5a). The amounts of NH3-N in the CS and OL groups were increased than in the CI group (CS vs CI, P < 0.001; OL vs CI, P = 0.011, Fig. 5a).
After 28 days of feeding, the concentration of NH3-N in the CS group was higher than in the OL and CI groups (CS vs OL, P = 0.007; CS vs CI, P < 0.001 Fig. 5a). In addition, for the OL group, the concentrations of total volatile fatty acids (TVFAs) (D7 vs D28, P < 0.001; D14 vs D28, P < 0.001), propionate (D7 vs D28, P < 0.001; D14 vs D28, P < 0.001), and valerate (D7 vs D28, P = 0.003; D14 vs D28, P = 0.002) at day 7 and day 14 were lower than at day 28, but the concentrations of NH3-N at day 14 (P = 0.001) and day 28 (P = 0.005) were increased compared to day 7 (Fig. 5b).
The Interplay Patterns Between Rumen Fermentation and Bacterial Community
In the OL group (Fig. 6a), TVFAs were directly positive with butyrate and acetate through 7 OTUs. Most of OTUs associated with butyrate were classified as Prevotella spp. AP was negatively correlated to propionate, and isovalerate was interactive with 16 OTUs. However, in the CS (Fig. 6b) and CI (Fig. 6c) groups, TVFAs were positively associated with acetate, propionate, butyrate, and valerate. AP was positively correlated with isovalerate and isobutyrate in the CS and CI groups, respectively (Fig. 6b, c). Across this network, many novel Prevotella spp. were positively associated with various metabolic phenotypes.
Co-occurrence network analysis between bacterial OTUs and rumen metabolic phenotypes for the OL (a), CS (b), and CI (c) groups. Circle nodes represent bacterial OTUs (or species) and rounded rectangle nodes do metabolic phenotypes. Each co-occurring pair between bacterial OTUs (or species) and metabolic phenotypes has an absolute Spearman rank correlation above 0.70 [gold line indicates positive correlation (R > 0.70); gray line indicates negative correlation (R < −0.70)] with an FDR-corrected significance level under 0.05. Circles with different color represent the OTUs from the different genera. OL oak leaves, CS corn stover, CI corn silage. Asterisk indicates unclassified bacteria taxa
Discussion
The present study used high-throughput sequencing of the 16S rRNA gene to provide a comprehensive view into understanding the alteration of bacterial compositions and explore the association between bacterial community and metabolic phenotypes in the rumen of sika deer fed three diets. The co-occurrence analyses showed diet-driven pattern of rumen microorganisms and the metabolic phenotype interaction.
This study found highly abundant bacteria belonging to the phylum Bacteroidetes, mostly Prevotella spp. Other studies on cows, goats, elk, and white-tailed deer also found that Prevotella spp. were predominant [24–29]. However, the proportion of Prevotella spp. (50.7 %) in the present study was relatively higher than previous findings on cattle (20–30 %) [30–32], moose (10 %) [33], macropods (38 %) [34], horses (0.23 %) [35], elk, and white-tailed deer (about 20 %) [28]. A possible explanation was the co-evolution between rumen bacteria and sika deer [36]. Furthermore, a recent study demonstrated that the enterotype of human gut microbiome dominated by Prevotella spp. was strongly associated with long-term diets having abundant carbohydrates [37]. Moreover, metagenomic analyses suggested that Prevotella spp. played a potential role in cellulose degradation in the foregut of the Tammar wallaby and in the rumen of Svalbard reindeer [38, 39]. Some members of the genus Prevotella contained highly active hemicellulolytic and proteolytic enzymes [40], which could degrade xylan, pectin, and starch [41–44], and comprised a large part of the genetic and metabolic diversity in rumen microbial community [45–47]. Recent studies also showed that the genus Prevotella displayed a central niche in maintaining the community structure of human gut microbiome [48] and could provide the host with the ability to adapt various diets [49]. Therefore, our findings suggested that Prevotella spp. likely played a key role in the rumen fermentation of sika deer. Interestingly, most of Prevotella spp. were identified as novel species, which was similar to the findings of Gruninger et al. [28], who also indentified a number of core OTUs that were classified as uncultured Prevotella in the rumen of elk and white-tailed deer in Canada. This suggested that the rumen of cervidae animals may harbor unique microbiota, which needed to further examine in future studies.
The distribution of bacteria, such as Prevotella spp., unclassified Succinivibrionaceae and Paraprevotellaceae, and TVFAs was changed within the OL group, as well as in comparison with the CS and CI groups (Figs. 4 and 5), suggesting that the rumen bacterial community composition and fermentation patterns were altered by the diets [50, 51]. The co-occurrence patterns further revealed the interaction between bacteria and fermentation products in the rumen of sika deer fed three diets. The present study found that different groups of Prevotella spp. took important roles in the interaction with various metabolic phenotypes, given that Prevotella spp. was the dominant bacteria in the rumen of sika deer. In addition, the results found that bacterial patterns of butyrate were considerably altered after the feed was changed from the CS and CI groups (Fig. 6b, c) to the OL group (Fig. 6a), indicating that butyrate metabolism mediated by specific rumen bacteria may play a major role in the energy acquisition of sika deer fed oak leaves. Butyrate contributed to approximately 70 % of the daily metabolic energy of ruminants [3]. Moreover, butyrate production has recently been shown to play a special role in modulating bacterial energy metabolism in the gut ecosystem [52]. The different interplays in the OL group compared to the CS and CI groups may be related to the tannins in the oak leaves, which could decrease methane emissions causing energy loss to the host [53, 54].
This study showed that many novel Prevotella spp. were positively related to butyrate. By surveying non-redundant protein sequence database in NCBI (http://www.ncbi.nlm.nih.gov/), the results showed that 24 species (31 isolates) of the Prevotella genus could express potential butyrate kinase responsible for the production of butyrate (Supplementary Fig. S1). Moreover, other butyrate producers, such as unclassified bacteria belonging to the family Lachnospiraceae [55], and Paludibacter spp. [56], were positively correlated to butyrate. These results indicated that the phylogenetically diverse butyrate-producing bacteria may contribute to host-specific butyrate biosynthesis in the rumen ecological system of sika deer [56–59]. On the other hand, the current study did not find that valerate was associated with TVFAs in the OL group, which may be caused by tannins contained in oak leaves, as tannins could affect the metabolism of proteins, and inhabit the growth of proteolytic bacteria to generate branch fatty acids, such as valerate [60–62].
The interactive patterns between the AP and other metabolic phenotypes in the OL group were different from that in the CS and CI groups. Many unclassified bacteria in the family Ruminococcaceae were higher and were positively associated with the AP in CS and CI groups (Figs. 4 and 6). The Ruminococcaceae bacteria were enriched in endo-1,4-beta-xylanase and cellulose genes, which played key roles in cleaving the cellulose and hemicellulose components of plant material [63]. Therefore, these bacteria may enhance the degradation of fiber plants in the rumen of sika deer, which could cause the increase of production of hydrogen and possibly methane. This was in agreement with the finding that these bacteria were positively associated with the AP, as the methane production was positively associated with AP [53, 64–66].
On the other hand, many bacterial OTUs belonging to the family Succinivibrionaceae, which were lower in the CS and CI groups than in the OL group, were negatively associated with AP. Similarly, Pope et al. [67] found that Succinivibrionaceae bacteria were also abundant in the gut of the Tammar wallaby, which produced relatively lower methane as compared to ruminants. Bacteria belonging to the family Succinivibrionaceae depended on carbon dioxide to support its metabolism via succinate biosynthesis and, in turn, inhibited the bacterial NADH-H hydrogenases and the subsequent inhibition of methanogens producing lower methane [11, 68]. This suggested that the corn forages may cause more loss of metabolic energy of sika deer than the oak leaves mediating by rumen bacteria. Additionally, the concentrations of isobutyrate and isovalerate were positively associated with the AP in the CS and CI groups. Isobutyrate and isovalerate were derived from branched-chain amino acids such as leucine, valine, and isoleucine [69], which were required for the growth of most ruminal fiber-degrading bacteria generating lots of hydrogen during rumen fermentation [70]. In consideration of the side effects of tannins on the proteins, these results indicated that the composition and concentration of some amino acids in the diets may have a fundamental role in energy metabolism, which need to be evaluated in future study.
In conclusion, the current study suggested that Prevotella spp. played critical roles in the fermentation of feed in the rumen of sika deer. However, the interplay patterns between rumen bacterial community composition and metabolic phenotypes were altered in the native and domesticated diets, especially for the butyrate, AP, and TVFAs, indicating that the fermentation patterns were changed in the rumen of sika deer fed different diets. It is important to note that the fungi and protozoa in the rumen of sika deer were not investigated here and require future attention.
Abbreviations
- OTU:
-
Operational taxonomic units
- VFAs:
-
Volatile fatty acids
- TVFAs:
-
Total volatile fatty acids
- BLAST:
-
Basic local alignment search tool
- PCoA:
-
Principal coordinate analysis
- PLS-DA:
-
Partial least squares discriminant analysis
- AP:
-
Acetate/propionate
- FDR:
-
False discovery rate
- NH3-N:
-
Ammonia nitrogen
- ANOVA:
-
Analysis of variance
- ISAPS:
-
Institute of Special Animal and Plant Sciences
- CAAS:
-
Chinese Academy of Agricultural Sciences
- ACUC:
-
Animal Care and Use Committee
- CFB:
-
Cytophaga–Flexibacter–Bacteroides
- DM:
-
Dry matter
References
Wright ADG, Klieve AV (2011) Does the complexity of the rumen microbial ecology preclude methane mitigation? Anim Feed Sci Technol 166–167:248–253. doi:10.1016/j.anifeedsci.2011.04.015
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto J-M, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Dore J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Bork P, Ehrlich SD, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. doi:10.1038/nature08821
Bergman EN (1990) Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol Rev 70:567–590
Kobayashi Y (2006) Inclusion of novel bacteria in rumen microbiology: need for basic and applied science. Anim Sci J 77:375–385. doi:10.1111/j.1740-0929.2006.00362.x
Li Z, Liu H, Li G, Bao K, Wang K, Xu C, Yang Y, Yang F, Wright ADG (2013) Molecular diversity of rumen bacterial communities from tannin-rich and fiber-rich forage fed domestic Sika deer (Cervus nippon) in China. BMC Microbiol 13:151
Hiura T, Hashidoko Y, Kobayashi Y, Tahara S (2010) Effective degradation of tannic acid by immobilized rumen microbes of a sika deer (Cervus nippon yesoensis) in winter. Anim Feed Sci Technol 155:1–8. doi:10.1016/j.anifeedsci.2009.09.015
Yang Y (2010) Effects of tannins on the nutrition utilization and growth of young Sika deer (Cervus nippon) (in Chinese). Master, Chinese Academy of Agricultural Sciences
Wang XX, Li XB, Zhao CX, Hu P, Chen H, Liu ZX, Liu GW, Wang Z (2012) Correlation between composition of the bacterial community and concentration of volatile fatty acids in the rumen during the transition period and ketosis in dairy cows. Appl Environ Microbiol 78:2386–2392. doi:10.1128/AEM.07545-11
Hernandez-Sanabria E, Guan LL, Goonewardene LA, Li MJ, Mujibi DF, Stothard P, Moore SS, Leon-Quintero MC (2010) Correlation of particular bacterial PCR-denaturing gradient gel electrophoresis patterns with bovine ruminal fermentation parameters and feed efficiency traits. Appl Environ Microbiol 76:6338–6350. doi:10.1128/Aem.01052-10
Carberry CA, Kenny DA, Han S, McCabe MS, Waters SM (2012) Effect of phenotypic residual feed intake and dietary forage content on the rumen microbial community of beef cattle. Appl Environ Microbiol 78:4949–4958. doi:10.1128/Aem.07759-11
Hernandez-Sanabria E, Goonewardene LA, Wang ZQ, Durunna ON, Moore SS, Guan LL (2012) Impact of feed efficiency and diet on adaptive variations in the bacterial community in the rumen fluid of cattle. Appl Environ Microbiol 78:1203–1214. doi:10.1128/Aem.05114-11
Mao SY, Zhang RY, Wang DS, Zhu WY (2012) The diversity of the fecal bacterial community and its relationship with the concentration of volatile fatty acids in the feces during subacute rumen acidosis in dairy cows. BMC Vet Res 8:237. doi:10.1186/1746-6148-8-237
Ursell LK, Haiser HJ, Van Treuren W, Garg N, Reddivari L, Vanamala J, Dorrestein PC, Turnbaugh PJ, Knight R (2014) The intestinal metabolome: an intersection between microbiota and host. Gastroenterology 146:1470–1476. doi:10.1053/j.gastro.2014.03.001
Mao S, Huo W, Zhu W (2013) Use of pyrosequencing to characterize the microbiota in the ileum of goats fed with increasing proportion of dietary grain. Curr Microbiol 67:341–350. doi:10.1007/s00284-013-0371-0
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Meth 7:335–336. doi:10.1038/Nmeth.F.303
Reeder J, Knight R (2010) Rapidly denoising pyrosequencing amplicon reads by exploiting rank-abundance distributions. Nat Meth 7:668–669. doi:10.1038/nmeth0910-668b
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. doi:10.1093/bioinformatics/btq461
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. doi:10.1128/AEM.03006-05
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267. doi:10.1093/bioinformatics/btp636
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G, Ciulla D, Tabbaa D, Highlander SK, Sodergren E, Methe B, DeSantis TZ, Petrosino JF, Knight R, Birren BW, Consortium HM (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21:494–504. doi:10.1101/gr.112730.110
Arndt D, Xia J, Liu Y, Zhou Y, Guo AC, Cruz JA, Sinelnikov I, Budwill K, Nesbo CL, Wishart DS (2012) METAGENassist: a comprehensive web server for comparative metagenomics. Nucleic Acids Res 40:W88–W95. doi:10.1093/nar/gks497
Zhang Z, Geng J, Tang X, Fan H, Xu J, Wen X, Ma Z, Shi P (2014) Spatial heterogeneity and co-occurrence patterns of human mucosal-associated intestinal microbiota. ISME J 8:881–893. doi:10.1038/ismej.2013.185
Smoot ME, Ono K, Ruscheinski J, Wang PL, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27:431–432. doi:10.1093/bioinformatics/btq675
Kim M, Morrison M, Yu Z (2011) Status of the phylogenetic diversity census of ruminal microbiomes. FEMS Microbiol Ecol 76:49–63. doi:10.1111/j.1574-6941.2010.01029.x
Stevenson DM, Weimer PJ (2007) Dominance of Prevotella and low abundance of classical ruminal bacterial species in the bovine rumen revealed by relative quantification real-time PCR. Appl Microbiol Biotechnol 75:165–174. doi:10.1007/s00253-006-0802-y
Wu S, Baldwin RL, Li W, Li C, Connor EE, Li RW (2012) The bacterial community domposition of the bovine rumen detected using pyrosequencing of 16S rRNA genes. Metagenomics 1:1–11. doi:10.4303/mg/235571
Jami E, Mizrahi I (2012) Composition and similarity of bovine rumen microbiota across individual animals. PLoS One 7:e33306. doi:10.1371/journal.pone.0033306
Gruninger RJ, Sensen CW, McAllister TA, Forster RJ (2014) Diversity of rumen bacteria in Canadian cervids. PLoS One 9:e89682. doi:10.1371/journal.pone.0089682
Lee HJ, Jung JY, Oh YK, Lee SS, Madsen EL, Jeon CO (2012) Comparative survey of rumen microbial communities and metabolites across one caprine and three bovine groups, using barcoded pyrosequencing and 1H nuclear magnetic resonance spectroscopy. Appl Environ Microbiol 78:5983–5993. doi:10.1128/AEM.00104-12
de Oliveira MNV, Jewell KA, Freitas FS, Benjamin LA, Tótola MR, Borges AC, Moraes CA, Suen G (2013) Characterizing the microbiota across the gastrointestinal tract of a Brazilian Nelore steer. Vet Microbiol 164:307–314. doi:10.1016/j.vetmic.2013.02.013
Kong Y, Teather R, Forster R (2010) Composition, spatial distribution, and diversity of the bacterial communities in the rumen of cows fed different forages. FEMS Microbiol Ecol 74:612–622. doi:10.1111/j.1574-6941.2010.00977.x
Zened A, Combes S, Cauquil L, Mariette J, Klopp C, Bouchez O, Troegeler-Meynadier A, Enjalbert F (2013) Microbial ecology of the rumen evaluated by 454 GS FLX pyrosequencing is affected by starch and oil supplementation of diets. FEMS Microbiol Ecol 83:504–514. doi:10.1111/1574-6941.12011
Ishaq SL, Wright ADG (2012) Insight into the bacterial gut microbiome of the North American moose (Alces alces). BMC Microbiol 12:212. doi:10.1186/1471-2180-12-212
Gulino LM, Ouwerkerk D, Kang AYH, Maguire AJ, Kienzle M, Klieve AV (2013) Shedding light on the microbial community of the Macropod foregut using 454-amplicon pyrosequencing. PLoS One 8:e61463.. doi:10.1371/journal.pone.0061463
Shepherd ML, Swecker WS Jr, Jensen RV, Ponder MA (2012) Characterization of the fecal bacteria communities of forage-fed horses by pyrosequencing of 16S rRNA V4 gene amplicons. FEMS Microbiol Lett 326:62–68. doi:10.1111/j.1574-6968.2011.02434.x
Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS, Schlegel ML, Tucker TA, Schrenzel MD, Knight R, Gordon JI (2008) Evolution of mammals and their gut microbes. Science 320:1647–1651. doi:10.1126/science.1155725
Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R, Sinha R, Gilroy E, Gupta K, Baldassano R, Nessel L, Li H, Bushman FD, Lewis JD (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108. doi:10.1126/science.1208344
Pope PB, Denman SE, Jones M, Tringe SG, Barry K, Malfatti SA, McHardy AC, Cheng JF, Hugenholtz P, McSweeney CS, Morrison M (2010) Adaptation to herbivory by the Tammar wallaby includes bacterial and glycoside hydrolase profiles different from other herbivores. Proc Natl Acad Sci U S A 107:14793–14798. doi:10.1073/pnas.1005297107
Pope PB, Mackenzie AK, Gregor I, Smith W, Sundset MA, McHardy AC, Morrison M, Eijsink VG (2012) Metagenomics of the Svalbard reindeer rumen microbiome reveals abundance of polysaccharide utilization loci. PLoS One 7:e38571. doi:10.1371/journal.pone.0038571
Matsui H, Ogata K, Tajima K, Nakamura M, Nagamine T, Aminov RI, Benno Y (2000) Phenotypic characterization of polysaccharidases produced by four Prevotella type strains. Curr Microbiol 41:45–49. doi:10.1007/s002840010089
Kabel MA, Yeoman CJ, Han Y, Dodd D, Abbas CA, de Bont JA, Morrison M, Cann IK, Mackie RI (2011) Biochemical characterization and relative expression levels of multiple carbohydrate esterases of the xylanolytic rumen bacterium Prevotella ruminicola 23 grown on an ester-enriched substrate. Appl Environ Microbiol 77:5671–5681. doi:10.1128/AEM.05321-11
Cotta MA (1992) Interaction of ruminal bacteria in the production and utilization of maltooligosaccharides from starch. Appl Environ Microbiol 58:48–54
Gardner RG, Wells JE, Russell JB, Wilson DB (1995) The cellular location of Prevotella ruminicola beta-1,4-D-endoglucanase and its occurrence in other strains of ruminal bacteria. Appl Environ Microbiol 61:3288–3292
Krause DO, Denman SE, Mackie RI, Morrison M, Rae AL, Attwood GT, McSweeney CS (2003) Opportunities to improve fiber degradation in the rumen: microbiology, ecology, and genomics. FEMS Microbiol Rev 27:663–693. doi:10.1016/S0168-6445(03)00072-X
Bekele AZ, Koike S, Kobayashi Y (2010) Genetic diversity and diet specificity of ruminal Prevotella revealed by 16S rRNA gene-based analysis. FEMS Microbiol Lett 305:49–57. doi:10.1111/j.1574-6968.2010.01911.x
Ramsak A, Peterka M, Tajima K, Martin JC, Wood J, Johnston MEA, Aminov RI, Flint HJ, Avgustin G (2000) Unravelling the genetic diversity of ruminal bacteria belonging to the CFB phylum. FEMS Microbiol Ecol 33:69–79. doi:10.1016/S0168-6496(00)00049-0
Purushe J, Fouts D, Morrison M, White B, Mackie R, Coutinho P, Henrissat B, Nelson K (2010) Comparative genome analysis of Prevotella ruminicola and Prevotella bryantii: insights into their environmental niche. Microb Ecol 60:721–729. doi:10.1007/s00248-010-9692-8
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Baldassano RN, Anokhin AP, Heath AC, Warner B, Reeder J, Kuczynski J, Caporaso JG, Lozupone CA, Lauber C, Clemente JC, Knights D, Knight R, Gordon JI (2012) Human gut microbiome viewed across age and geography. Nature 486:222–227. doi:10.1038/nature11053
Schnorr SL, Candela M, Rampelli S, Centanni M, Consolandi C, Basaglia G, Turroni S, Biagi E, Peano C, Severgnini M, Fiori J, Gotti R, De Bellis G, Luiselli D, Brigidi P, Mabulla A, Marlowe F, Henry AG, Crittenden AN (2014) Gut microbiome of the Hadza hunter-gatherers. Nat Commun 5:3654. doi:10.1038/ncomms4654
Lettat A, Benchaar C (2013) Diet-induced alterations in total and metabolically active microbes within the rumen of dairy cows. PLoS One 8:E60978. doi:10.1371/journal.pone.0060978
Belanche A, Doreau M, Edwards JE, Moorby JM, Pinloche E, Newbold CJ (2012) Shifts in the rumen microbiota due to the type of carbohydrate and level of protein ingested by dairy cattle are associated with changes in rumen fermentation. J Nutr 142:1684–1692. doi:10.3945/jn.112.159574
Li F, Hinderberger J, Seedorf H, Zhang J, Buckel W, Thauer RK (2008) Coupled ferredoxin and crotonyl coenzyme a (CoA) reduction with NADH catalyzed by the butyryl-CoA dehydrogenase/Etf complex from Clostridium kluyveri. J Bacteriol 190:843–850. doi:10.1128/JB.01417-07
Johnson KA, Johnson DE (1995) Methane emissions from cattle. J Anim Sci 73:2483–2492
Ross EM, Moate PJ, Marett L, Cocks BG, Hayes BJ (2013) Investigating the effect of two methane-mitigating diets on the rumen microbiome using massively parallel sequencing. J Dairy Sci 96:6030–6046. doi:10.3168/jds.2013-6766
Cotta M, Forster R (2006) The family Lachnospiraceae, including the genera Butyrivibrio, Lachnospira and Roseburia. In: Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) Dworkin, M. The Prokaryotes, Springer US, pp 1002–1021
Li RW, Wu S, Baldwin RLT, Li W, Li C (2012) Perturbation dynamics of the rumen microbiota in response to exogenous butyrate. PLoS One 7:e29392. doi:10.1371/journal.pone.0029392
Barcenilla A, Pryde SE, Martin JC, Duncan SH, Stewart CS, Henderson C, Flint HJ (2000) Phylogenetic relationships of butyrate-producing bacteria from the human gut. Appl Environ Microbiol 66:1654–1661. doi:10.1128/AEM.66.4.1654-1661.2000
Pryde SE, Duncan SH, Hold GL, Stewart CS, Flint HJ (2002) The microbiology of butyrate formation in the human colon. FEMS Microbiol Lett 217:133–139. doi:10.1111/j.1574-6968.2002.tb11467.x
Louis P, Flint HJ (2009) Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol Lett 294:1–8. doi:10.1111/j.1574-6968.2009.01514.x
McSweeney CS, Palmer B, McNeill DM, Krause DO (2001) Microbial interactions with tannins: nutritional consequences for ruminants. Anim Feed Sci Technol 91:83–93. doi:10.1016/S0377-8401(01)00232-2
Min B, Attwood G, McNabb W, Molan A, Barry T (2005) The effect of condensed tannins from Lotus corniculatus on the proteolytic activities and growth of rumen bacteria. Anim Feed Sci Technol 121:45–58
Andries JI, Buysse FX, De Brabander DL, Cottyn BG (1987) Isoacids in ruminant nutrition: their role in ruminal and intermediary metabolism and possible influences on performances—a review. Anim Feed Sci Technol 18:169–180. doi:10.1016/0377-8401(87)90069-1
Biddle A, Stewart L, Blanchard J, Leschine S (2013) Untangling the genetic basis of fibrolytic specialization by Lachnospiraceae and Ruminococcaceae in diverse gut communities. Diversity 5:627–640. doi:10.3390/d5030627
Newbold CJ, Lopez S, Nelson N, Ouda JO, Wallace RJ, Moss AR (2005) Propionate precursors and other metabolic intermediates as possible alternative electron acceptors to methanogenesis in ruminal fermentation in vitro. Br J Nutr 94:27–35. doi:10.1079/BJN20051445
Wang YY, Zhang YL, Wang JB, Meng L (2009) Effects of volatile fatty acid concentrations on methane yield and methanogenic bacteria. Biomass Bioenergy 33:848–853. doi:10.1016/j.biombioe.2009.01.007
Christophersen CT, Wright ADG, Vercoe PE (2008) In vitro methane emission and acetate: propionate ratio are decreased when artificial stimulation of the rumen wall is combined with increasing grain diets in sheep. J Anim Sci 86:384–389. doi:10.2527/jas.2007-0373
Pope PB, Smith W, Denman SE, Tringe SG, Barry K, Hugenholtz P, McSweeney CS, McHardy AC, Morrison M (2011) Isolation of Succinivibrionaceae implicated in low methane emissions from Tammar wallabies. Science 333:646–648. doi:10.1126/science.1205760
Stackebrandt E, Hespell R (2006) The family Succinivibrionaceae. In: Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E, Dworkin M (eds) The Prokaryotes. Springer, New York, pp 419–429
Allison MJ (1978) Production of branched-chain volatile fatty acids by certain anaerobic bacteria. Appl Environ Microbiol 35:872–877
Gorosito AR, Russell JB, Vansoest PJ (1985) Effect of carbon-4 and carbon-5 volatile fatty acids on digestion of plant-cell wall in vitro. J Dairy Sci 68:840–847
Acknowledgments
The authors would like to thank Kunming Biological Diversity Regional Center of Large Apparatus and Equipments, Kunming Institute of Zoology, and Chinese Academy of Sciences for their super technical assistances. This work is supported by a leading talent and creative team project (20121810), natural science foundation (20130101104JC) and major scientific research projects (20140203018NY) from Jilin province, key projects in the national science and technology pillar program (2011BAI03B02), and the Foundation of State Key Laboratory of Genetic Resources and Evolution Kunming Institute of Zoology Chinese Academy of Sciences (GREKF14-14).
Conflict of Interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
The phylogenetic tree of butyrate kinases using amino acid sequences from 31 isolates of genus Prevotella. Bootstrap value larger than 70 were shown. (PDF 163 kb)
Table S1
The study design for the 3×3 latin square using three sika deer.(PDF 10 kb)
Rights and permissions
About this article
Cite this article
Li, Z., Wright, AD.G., Liu, H. et al. Bacterial Community Composition and Fermentation Patterns in the Rumen of Sika Deer (Cervus nippon) Fed Three Different Diets. Microb Ecol 69, 307–318 (2015). https://doi.org/10.1007/s00248-014-0497-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0497-z