Abstract
The objective of this study was to characterise lactic acid bacteria (LAB) isolated from faecal samples of healthy Ethiopian infants, with emphasis on bacteriocin production and antibiotic susceptibility. One hundred fifty LAB were obtained from 28 healthy Ethiopian infants. The isolates belonged to Lactobacillus (81/150), Enterococcus (54/150) and Streptococcus (15/150) genera. Lactobacillus species were more abundant in the breast-fed infants while Enterococcus dominated the mixed-fed population. Bacteriocin-producing LAB species were isolated from eight of the infants. Many different bacteriocins were identified, including one new bacteriocin from Streptococcus salivarius, avicin A (class IIa) from Enterococcus avium, one class IIa bacteriocin from Enterococcus faecalis strains, one unknown bacteriocin from E. faecalis and two unknown bacteriocins from Lactobacillus fermentum strains and the two-peptide gassericin T from Lactobacillus gasseri isolate. Susceptibility tests performed for nine antibiotics suggest that some lactobacilli might have acquired resistance to erythromycin (3 %) and tetracycline (4 %) only. The streptococci were generally antibiotic sensitive except for penicillin, to which they showed intermediate resistance. All enterococci were susceptible to ampicillin while 13 % showed penicillin resistance. Only one E. faecalis isolate was vancomycin-resistant. Tetracycline (51 %) and erythromycin (26 %) resistance was prevalent among the enterococci, but multidrug resistance was confined to E. faecalis (47 %) and Enterococcus faecium (33 %). Screening of enterococcal virulence traits revealed that 2 % were β-haemolytic. The structural genes of cytolysin were detected in 28 % of the isolates in five enterococcal species, the majority being E. faecalis and Enterococcus raffinosus. This study shows that bacteriocin production and antibiotic resistance is a common trait of faecal LAB of Ethiopian infants while virulence factors occur at low levels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gut of a human being is generally considered sterile at birth, but, right from the time of birth, it is colonised by bacteria that increase fast in number and complexity and being established as a durable microflora of the growing infant [1]. The composition of the gut bacteria differs from one to another person [1]. This is because the development and composition of intestinal microflora of infants is determined by various factors [2–4], including feeding type (e.g., breast milk, formula, or other foods), delivery mode (vaginal or caesarean), environment, and hygiene conditions. It has been shown that the colonisation of the gut with lactobacilli and bifidobacteria is delayed in infants delivered by caesarean section compared with vaginally delivered infants [5, 6]. Infants in developing countries are colonised by enterobacteria (including Escherichia coli), bifidobacteria, enterococci, and lactobacilli earlier and contain a more diverse microflora than infants from developed countries [3].
The LAB play an important role as probiotics [7–9]. For safety reasons, LAB of healthy human origin, which are non-pathogenic and not resistant to antibiotics, are often preferred for selection as potential probiotics. The probiotic potential of LAB may be promoted by their ability to produce antimicrobial compounds, the most prominent being bacteriocins. Bacteriocins are ribosomally synthesised antimicrobial peptides or proteins produced by bacteria and kill other bacteria. Bacteriocin production has also been implicated in the protection of mice against the food-borne pathogen Listeria monocytogenes [10]. Thus, bacteriocin-producing LAB can be potential candidates for use as probiotics or as biological control agents.
Even though lactobacilli are generally considered non-pathogenic and clinically less important than enterococci and streptococci, it is necessary to screen them for antibiotic susceptibility. In this regard, EFSA (European Food Safety Authority) has set criteria for safety evaluation of lactobacilli intended for use in animal feeds [11]. This is because there is a concern that lactobacilli (and other normal flora) may act as potential reservoirs of several antibiotic-resistant genes, which might be transferred to pathogenic or opportunistic bacteria during passage in the GIT, thus contributing to the spread of antibiotic resistance [12, 13]. Several studies have shown that the transfer of such genes or plasmids bearing the genes from lactobacilli to pathogens is possible [14–16]. In particular, the erm(B) and tet(M) genes that confer resistance to erythromycin and tetracycline, respectively, have been identified in intestinal lactobacilli [17].
Enterococci, especially Enterococcus faecalis and Enterococcus faecium, are the major cause of nosocomial infections [18, 19]. Enterococci display both intrinsic and acquired resistance to antibiotics that entails a challenge to treatment of enterococcal infections. Tolerance to chloramphenicol, erythromycin, tetracycline and high levels of clindamycin, aminoglycosides, β-lactams and glycopeptides is acquired resistance [20, 21]. Acquired resistance is a major concern in infectious diseases not only for posing challenges in the treatment of infections but also for its transmissibility among bacteria. Vancomycin resistance also deserves particular attention since this antibiotic is considered as “a last line of defense” in the treatment of infections caused by multidrug-resistant enterococci [19, 22].
The ability of enterococci to cause diseases can be promoted by production of tissue-damaging virulence factors such as cytolysin and gelatinase [23]. Cytolysin is a two-peptide lantibiotic haemolysin whose production in enterococci is dictated by a cytolysin operon that consists of 8 genes: cylR1, cylR2, cylL L , cylL S , cylM, cylB, cylA and cylI [24]. The operon is mostly located on pheromone-responsive plasmids and sometimes on the chromosome within a pathogenicity island [24]. The occurrence of cytolysin is more prevalent in clinical isolates than in food isolates [25, 26]. Gelatinase production is more frequent in E. faecalis and is mainly associated with clinical or veterinary isolates [27, 28].
The objectives of this study were to characterise LAB from faecal samples of infants with emphasis on (1) bacteriocin production, (2) susceptibility to selected antibiotics and (3) detection of cytolysin and gelatinase production, among the enterococci.
Materials and Methods
Subjects and Sample Collection
Faecal samples were collected from 28 healthy Ethiopian infants (Dilla town, Southern Ethiopia), age between 3 and 26 weeks, of whom 16 were females and 12 males. Seventeen were breast-fed, and eleven were mixed-fed. All were vaginally delivered except one. Eight were born at home and 20 in hospital (Table 1). Informed consent of parents was obtained before samples were collected. One faecal sample was collected in sterile screw-capped plastic tubes from each of the infants.
Isolation and Identification of Bacteria
The LAB were isolated on de Man-Rogosa-Sharpe (MRS) supplemented with cysteine–HCl (0.5 gL−1), reinforced clostridial (RC) agar and bile aesculin (BA) agar (all from Oxoid, England). Faecal samples were homogenised in broths (1 g faeces/mL) and tenfold serial dilutions were done. Samples (0.1 mL) were plated from each of the three highest dilutions (10−5 to 10−7 or 10−6 to 10−8) on MRS supplemented with cysteine–HCl (0.5 gL−1), RC and BA agar plates. The plates were incubated under anaerobic condition at 37 °C for 24–48 h. Three to four colonies from each plate were picked and inoculated in appropriate stab agar, and transported to Norway. Then, the bacteria in the transport media were streaked on appropriate agar plates to recover the isolates. Single colonies were picked and streaked on plates to confirm their purity. The resulting isolated colonies were subcultured in appropriate broths and subsequently maintained at −80 °C with 13 % glycerol-containing appropriate medium. Species identification was done by partial (about 1,440 bp) 16S rRNA gene sequence analysis which was done by BLAST search against nonredundant database.
DNA isolation, PCR and Sequencing
DNA isolation was done using the Bacterial Genomic DNA Purification Kit (Edge Biosystems, Gaithersburg, USA). The purified DNA was resuspended in 1× TE buffer pH 8.
Polymerase chain reaction (PCR) was done on genomic DNA to amplify genes for the 16S rRNA, bacteriocins and cytolysin with appropriate primers (Table 2). The PCR mixture (25 μl) contained 100–200 ng template, 0.5 unit DyNAzyme DNA polymerase (Finnzymes), 1× DyNAzyme buffer, 0.5 mM MgCl2, 200 μM dNTPs and 1 μM of each primer. PCR conditions—initial denaturation at 95 °C for 1 min and 35 cycles at 95 °C for 15 s, 50-54 °C for 30 s and 72 °C for 90 s. The PCR products were purified using QIAquick PCR purification Kit (QIAGEN, Germany). Cycle sequencing was done by using BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, USA), and the PCR products were sequenced by ABI Prism 377 DNA sequencing system (Applied Biosystems, USA).
Bacteriocin Production and Activity
Antimicrobial activity was detected by soft agar overlay assay as described previously [29]. Briefly, the isolated bacteria were spotted onto MRS or GM17 agar plates and incubated for 16 h at 37 °C. The colonies were overlaid with appropriate soft agar seeded with an overnight culture (diluted 400-fold) of the indicator strain. Five indicators (Lactobacillus sakei NCDO 2714, Lactobacillus plantarum LMGT 2003, E. faecalis LMGT 2708, Listeria innocua BL86/26 B, Staphylococcus aureus LMGT 3242) were used for detecting bacteriocin production (Table 4). After overnight incubation, the formation of growth-inhibition zones around the colonies was examined as indication of antimicrobial activity. To investigate whether the antimicrobial activity observed was caused by a non-protein compound or a bacteriocin, the antimicrobial activity was tested for sensitivity to proteinase K by adding 1 μl of 20 mgmL−1 proteinase K close to overnight colonies that produce antimicrobial substances. The colonies were then overlaid with appropriate soft agar seeded with an indicator strain and incubated overnight. The next day, the regions close to the colonies where the proteinase K was applied were checked for absence of growth inhibition which is indicative of inactivation of the antimicrobial substance by proteinase K.
In order to further characterise the bacteriocins detected in two E. faecalis strains (14M5c and 21M8a), the sensitivity of these strains to other enterococci that produce known bacteriocins was tested using soft agar assay as described above. E. faecalis LMGT 2708, which is sensitive to many bacteriocins, including pediocin-like bacteriocins, was used as a reference indicator. The producer enterococci used and their bacteriocins (in bracket) are: E. faecalis 100 (enterolysin A), E. faecalis 189 (enterolysin A and cytolysin), E. faecalis 3207 (enterocin L50 and AS-48), E. faecalis 3208 (enterocin P, L50 and AS-48), E. faecalis 26c and 116c (enterocin 1071), E. faecium L50 (enterocin P, Q, L50) and E. faecium T136 (enterocin AB) [30, 31].
Hemolytic Activity
The haemolytic activity of enterococci isolates was tested on brain heart infusion agar supplemented with 1 % (w/v) glucose, 0.03 % (w/v) l-arginine and 5 % (v/v) defibrinated horse blood [32]. The isolates were streaked on the agar, followed by anaerobic incubation at 37 °C for 24 h. E. faecalis JH2SS (PAD1) [33] and E. faecalis 111A [34] were used as positive controls while E. faecalis V583 and E. faecalis Symbioflor1 were included as negative controls. Haemolytic activity was detected by the appearance of a clear zone (β-haemolysis), greenish zone (α-haemolysis) or absence of any zone (γ-haemolysis) around the colony on the blood agar. The presence of the cytolysin structural genes (cylL L and cylL S ) was verified by PCR and DNA sequencing as described above.
Gelatinase Activity
The ability of the enterococci to produce gelatinase was tested on Todd Hewitt agar (Oxoid, England) supplemented with 3 % bovine gelatine [35]. The bacteria were spotted on the gelatine containing agar plates and incubated at 37 °C for 40 to 48 h, after which they were stored at 4 °C for 5 h. E. faecalis V583 was used as a positive control. Gelatinase production was evidenced by the formation a turbid zone around the colonies.
Antibiotic Susceptibility
The sensitivity of the LAB to the following antibiotics (concentration range in micrograms per millilitre) was tested using microtitre plate twofold dilution assay: ampicillin (0.25–512), chloramphenicol (0.5–64), erythromycin (0.125–1024), gentamycin (16–2,048), kanamycin (16–8,192), penicillin G (2–8,192), streptomycin (16–8,192), tetracycline (0.25–32) and vancomycin (0.25–32). Enterococci and streptococci were grown and tested in GM17 and the lactobacilli in MRS. A twofold serial dilution (in broth) of 50 μl the antibiotic solution was prepared in a microtitre plate well containing 50 μl broth, to which 150 μl diluted overnight culture (400-fold diluted in broth) of the test strain was added. The plate was incubated for at least 16 h, after which growth inhibition was measured turbidometrically at 620 nm by a microtitre plate reader (Labsystems Acsent Reader MF, Labsystems, Helsinki, Finland). Resistance was determined by using CLSI (Clinical and Laboratory Standards Institute) breakpoints for enterococci and streptococci and EFSA breakpoints for the lactobacilli [11, 36].
Statistical Analysis
Fisher’s exact test was used to compare proportions. P value < 0.05 was considered statistically significant.
Results
Diversity of the LAB among Infants
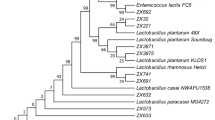
A total of 150 LAB were isolated from 28 Ethiopian infants and identified to species level by 16S rRNA gene sequencing analysis. The isolates belonged to Lactobacillus (81/150), Enterococcus (54/150) and Streptococcus (15/150) genera (Table 3). No statistically significant difference in colonisation rates (frequency of occurrence) of lactobacilli was observed between breast-fed and mixed-fed infants (82 % versus 64 %) (Table 3). The same was true for enterococci (59 % versus 82 %) and streptococci (12 % versus 36 %) (Table 3). But the differences could be ecologically significant.
Nevertheless, significant differences were observed in colonisation levels (abundance) of each of the three groups of LAB between breast-fed and mixed-fed babies. Sixty of the Lactobacillus isolates (74 %) were found in breast-fed infants which is significantly higher than the number of lactobacilli found among the mixed-fed (21 isolates, 26 %) infants (odds ratio (OR), 5.357; P value < 0.00001). Conversely, a significant enrichment of enterococci among isolates from mixed-fed infants (59 %) was observed (OR, 2.652; P value, 0.0061). The streptococci were more abundant in mixed-fed infants (13 isolates) than in breast-fed infants (two isolates) (OR, 10.057; P value, 0.0006) (Table 3).
Bacteriocin Production
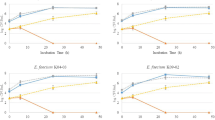
The 150 LAB isolates were screened for bacteriocin production using a universal indicator L. sakei NCDO 2714 that allows detection of most known class I and II bacteriocins [37–39]. Fourteen strains (Table 4) isolated from eight infants were found to produce antimicrobial compounds that were susceptible to proteolytic treatment. The isolates comprise six Enterococcus avium, four E. faecalis, two Lactobacillus fermentum, one L. gasseri and one Streptococcus salivarius. From one of the bacteriocinogenic E. avium isolates (XA83), a class IIa bacteriocin (avicin A) has already been purified and characterised [39]. PCR and DNA sequence analysis showed that the other 5 E. avium strains (Table 4) from two infants also contain the structural gene of avicin A. The four E. faecalis strains (Table 4) produced increased antimicrobial activity up to 6–7 h of growth in liquid GM17 medium at 37 °C, but the activity was gradually lost when incubation time was prolonged. The instability of the bacteriocin produced by E. faecalis 21A8a could be due to degradation by gelatinase [40], since this strain produces gelatinase. In order to identify class IIa bacteriocin producers, a sensitive indicator E. faecalis LMGT 2708, the resistant mutant E. faecalis LMGT 2708RA were used [39]. The faecal isolates E. faecalis 14M5a, 14A6b and 14M5c inhibited growth of E. faecalis LMGT 2708 while E. faecalis LMGT 2708RA was insensitive to the same antibacterial producers. The results strongly suggest that the bacteriocins produced by 14M5a, 14A6b and 14M5c are pediocin-like bacteriocins. PCR and its partial DNA sequence showed that the bacteriocin from L. gasseri 10M7c is most likely identical to the two-peptide bacteriocin gassericin T, which inhibits L. delbrueckii subsp. bulgaricus [41]. S. salivarius 5M6c produced a new lantibiotic that is similar to the nisins [42]. PCR did not detect salivaricin A, salivaricin B and bovicin HJ50 in this strain. We have no sequence information of the bacteriocins produced by the remaining LAB isolates (Table 4). Cross-immunity test showed that E. faecalis 14M5c and E. faecalis 21A8a are sensitive to E. faecium L50 and E. faecium T136, but they were resistant to E. faecalis 3208, 3207, 100, 189, 26c and 116c, suggesting that the bacteriocins produced by strains 14M5c and 21A8a may not be enterocins Q, A or B. PCR was not able to detect genes for AS-48, bacteriocin 31 and enterocin 1071 in strains 14M5c and 21A8a.
β-Haemolytic Activity and Gelatinase Production Are Rare in Gut Enterococci
Fifty-three enterococci were tested for their ability to haemolyse red blood cells. One E. faecalis strain (21A8a) was found to be β-haemolytic. The remaining isolates, including the enterococcal negative controls, showed varying degrees of haemolytic activities which were weaker than β-haemolysis. PCR showed that 15/53 enterococci harboured the structural genes for biosynthesis of cytolysin, and the genes were found most frequently in E. faecalis and E. raffinosus (Table 5).
Gelatinase activity has been observed earlier in faecal enterococci from infants [34]. In this study, two of the 53 tested enterococci (E. faecalis and E. maldoratus) were found to produce gelatinase. The gelatinase-producing E. faecalis (strain 21A8a) showed also bacteriocin and cytolysin activity.
Antibiotic Resistance Is Common among Faecal LAB from Ethiopian Infants
The antibiotic resistance pattern is different for the three LAB groups (Table 6). The lactobacilli were generally susceptible to chloramphenicol, erythromycin and tetracycline but resistant to aminoglycosides and glycopeptides. Even though many lactobacilli showed sensitivity to ampicillin, the majority of L. gasseri did not. Kanamycin resistance was highly prevalent (96 %). One L. oris, one L. vaginalis and one L. salivarius strain were susceptible to kanamycin. Among the Lactobacillus species, L. gasseri and L. johnsonii were sensitive to vancomycin; the rest (83 %) were intrinsically resistant [43].
Most of the Streptococcus isolates appeared to be more susceptible to the nine antibiotics than the enterococci (Table 6). All streptococci were susceptible to ampicillin. All streptococci showed intermediate resistance to penicillin (MIC = 0.25–2 μg mL−1) while all were sensitive to ampicillin, chloramphenicol, erythromycin and vancomycin. One Streptococcus isolate was resistant to tetracycline (MIC > 8 μg mL−1).
E. faecalis and E. faecium Constitute a Reservoir for Antibiotic Resistance among the Gut Enterococci
Out of the 53 enterococci isolates tested for antibiotic sensitivity, 34 were resistant to one or more antibiotics. Resistance includes both full resistance and intermediate resistance, unless otherwise stated. Tetracycline resistance was the most prevalent (51 %), followed by erythromycin (26 %) and kanamycin (21 %) and was present in five species. Resistance to ampicillin was not observed. Only one Enterococcus isolate showed high vancomycin resistance (MIC > 32 μg mL−1), and this E. faecalis strain was also resistant to penicillin and erythromycin (MIC = 32 μg mL−1 for both). In addition, two E. gallinarum isolates showed intermediate resistance to vancomycin (MIC = 8–16 μg mL−1).
A high level of aminoglycoside resistance was observed in E. faecalis and E. faecium isolates. Overall, the prevalence of high level of aminoglycoside resistance was observed in five of fourteen E. faecalis and five of nine E. faecium isolates. Among these, two E. faecalis isolates were resistant to the three aminoglycosides (streptomycin, gentamycin and kanamycin).
Ten isolates (19 %) of the enterococci showed multidrug (more than two antibiotics) resistance, and they included only E. faecalis and E. faecium isolates. So, multidrug resistance is significantly more prevalent in E. faecalis and E. faecium than in other enterococci (P value, 0.0001). The multidrug resistance observed combined for the most part tetracycline, aminoglycosides and erythromycin resistance.
Discussion
In the present study, 150 faecal LAB from 28 healthy Ethiopian infants were isolated and identified to species level. Their ability to produce bacteriocins and their susceptibility to antibiotics was investigated. Moreover, haemolytic and gelatinase activities were determined. Our study indicates that lactobacilli and enterococci dominate the faecal LAB microflora of Ethiopian infants, consistent with studies of infant gut microflora in other parts of the world [44]. Our results indicate that the lactobacilli are slightly more predominant in the breast-fed infants, but they also constitute a significant part of the faecal LAB flora among the mixed-fed infants. The gut microbiota of the infants is partly acquired during delivery from the vaginal microflora [45], but the mother’s milk has also been shown to be an important source of the lactobacilli [46]. A previous study showed that Ethiopian infants contain more enterococci and lactobacilli during the first 2 weeks of life as compared with infants from Sweden [47]. This observation is also in agreement with our result, indicating that this trend extends beyond 2 weeks after birth. Furthermore, two previous studies have shown that enterococci and lactobacilli are the prevailing LAB in breast-fed Norwegian infants [30, 31], providing more support to our finding. The higher level of occurrence of enterococci in the faeces of mixed-fed than breast-fed infants has previously been attributed to foods other than breast milk [48, 49].
A previous study showed that, in 45 % of Swedish breast-fed infants, lactobacilli reached their highest number at the age of 6 months, the most common being L. rhamnosus and L. gasseri [50]. L. rhamnosus, L. johnsonii and L. paracasei prevail in the intestinal flora of Greek neonates, but no infants contained more than one species at a time [5]. In our study, these lactobacilli species were also common in the faecal samples of Ethiopian infants; however, the most prevailing species was L. fermentum. The most common enterococci isolated from infants are E. faecalis and E. faecium [3, 51, 52]. Also, E. avium, E. raffinosus and S. salivarius have been identified from neonates [30, 53]. Consistent with these studies, we isolated E. avium and E. faecalis from many infants and E. faecium, E. raffinosus and E. gallinarum from several samples (Table 3). In this study, the number of infants colonised with enterococci (68 %, Table 3) is higher than what has been reported in a study from Germany in which 23 % of the neonates were colonised with enterococci [52].
Recently, a large number of bacteriocin-producing LAB strains have been isolated from the gut of mammals, including man [54]. Bacteriocin-producing LAB were also found frequently in faecal samples of Norwegian infants [30, 31]. This study also shows that potential bacteriocin producers are commonly found in faecal LAB of Ethiopian infants (10 %) (Table 4). Identification of 14 bacteriocin-like compounds from 150 LAB producers obtained from 28 infants is a relatively high number when compared with isolation of bacteriocin-producing LAB from different environments [55, 56]. Most bacteriocin production was found in enterococci, and this is in line with a similar study of faecal samples from Norwegian infants [30, 31]. It is interesting to note that enterococci which occasionally occur as pathogens are also among the most frequent bacteriocin producers among the gut bacteria of infants [30, 31]. This suggests that enterococci may be better targets (sources) for bacteriocin screening compared with other LAB. In some infants, at least three to six bacteriocin-producing strains belonging to same species were found, suggesting that bacteriocin producers may predominate in their resident gut microflora.
Haemolytic phenotype and genotype have been studied in enterococci obtained from several sources. One study that investigated enterococci from poultry showed that 1 % was β-haemolytic, but 39 % contained the cylL S gene which was most prevalent among the E. faecalis isolates [27], and this is similar to the result we obtained. In another study, both cytolysin production and cytolysin operon were detected in 13 % of clinical enterococci [57]. On the contrary, enterococci isolated from faecal samples of wild boars did not show cytolysin activity or presence of the complete cytolysin operon [58]. Still in another study, β-haemolysis was observed in 33 % and 6 % of clinical and food enterococci, respectively, with high prevalence of cylL gene in E. faecalis (88 %) and E. faecium (70 %) [26]. In E. faecalis isolated from healthy Norwegian infants, the prevalence of cytolysin production and cytolysin structural genes was 9/31 and 15/31, respectively [34]. In contrast to the above-mentioned studies, we found very low frequency (2 %) of β-haemolysis activity after 24 h growth on blood plates.
Cytolysin production is common in E. faecalis but rare among other enterococcal species. In our study, PCR detected cytolysin structural genes (cylL L and cylL S ) also in E. faecium, E. avium, E. maldoratus and E. raffinosus (Table 5). Interestingly, one previous study [26], which examined 164 enterococci representing 20 species, detected β-haemolysis under anaerobic incubation in 16 % of the isolates. However, the structural genes of cytolysin were detected only in 9 % of the haemolytic strains, compared with our study which detected the structural genes in 28 % of the enterococci. Cytolysin structural genes were not detected in some haemolytic enterococci. This can be explained by variation in the sequence of the structural genes of cytolysin, or it might be conferred by different haemolysins other than cytolysin, as seen in another study [26]. The detection of structural genes in some non-haemolytic strains (e.g., E. avium) might be due to mutations and/or non-functional genes or gene products. It could also be due to loss of some genes from the cytolysin operon.
Gelatinase production is more common in E. faecalis [27, 28], although it has been reported in other species such as E. faecium, E. durans and E. hirae [59]. In one study, almost all E. faecalis isolated from poultry produced gelatinase [27]. About half of the 31 E. faecalis strains isolated from the Norwegian infants produced gelatinase [34]. In the current study, only one out of 15 E. faecalis produced gelatinase. These results show that gelatinase production is highly variable among faecal enterococci. In contrast, there are several reports that show that gelatinase production in E. faecalis is most prevalent among clinical isolates [28, 60].
Many studies have reported high prevalence of tetracycline resistance among enterococci [61, 62]. This was conferred by acquired resistance mainly due to tetracycline resistant genes tetL, tetM, tetS or tetO [61, 63] that is more pronounced in poultry isolates. In one study, it has been shown that tetracycline resistance occurred in 24 % of food isolates [63]. In enterococci isolates from healthy Portuguese children, 29 % of the isolates were resistant to tetracycline, 22 % were resistant to erythromycin, and 9 % were resistant to kanamycin [64]. In another study, a very high level of resistance to tetracycline and erythromycin was present in poultry and pet animals compared with isolates from faeces of healthy humans [62]. In the same study, no ampicillin resistance was found in human isolates and no vancomycin resistance in any of the isolates [62]. Consistent with these studies, we found high prevalence of tetracycline resistance in the enterococci, followed by erythromycin resistance. Also, tetracycline resistance was the most prevalent (55 %) among E. faecalis isolated from healthy Norwegian infants [34], comparable to the present study (53 %). It has been shown in several studies that E. faecium is more resistant to penicillin than E. faecalis [65, 66]. Our result is in agreement with these studies, since 44 % of the E. faecium were resistant to penicillin compared with 13 % of E. faecalis. Although we found no enterococci resistant to ampicillin, one E. faecium isolate has a MIC value of 4 μg mL−1, and according to EFSA criteria, this isolate is not safe for use in animal feeds [67].
Higher prevalence of multidrug resistance (57 %) was reported in enterococci isolated from German neonates [52], as compared with our study (19 %) (P value, 0.0001). The former study also reported absence of vancomycin resistance. Recently, it has been shown that multidrug resistance was frequent in E. faecalis clinical isolates [25]. E. faecalis was also the most frequent and showed the highest levels of multidrug resistance in the present study. High-level aminoglycoside resistance was also observed among the E. faecalis and E. faecium isolates, and this might be due to aminoglycoside-modifying enzymes [68]. In contrast to our study, a high level of aminoglycoside resistance occurred in all species of enterococci isolated from poultry but with high prevalence among E. faecium and E. faecalis.
Many studies have documented the resistance of most lactobacilli, including L. casei [69, 70], L. salivarius, L. acidophilus, L. plantarum, L. delbreuckii, L. paracasei and L. brevis [43, 71], to vancomycin which is due to intrinsic resistance. However, some lactobacilli belonging to the L. acidophilus group (in our case L. gasseri and L. johnsonii) were sensitive to vancomycin. This phenotypic characteristic can be used to differentiate the latter group from the rest of lactobacilli [43]. Similarly, the high prevalence of aminoglycoside resistance in the lactobacilli in this study might be due to intrinsic resistance [70]. Eleven L. casei strains isolated from Italian hard cheeses were all susceptible to ampicillin, penicillin G, tetracycline and vancomycin, but 1 and 8 were resistant to erythromycin and gentamycin, respectively [72]. In our study, all the 12 L. casei were susceptible to tetracycline, chloramphenicol, erythromycin and ampicillin (except one isolate), but all were resistant to vancomycin, gentamycin, kanamycin and most to streptomycin.
A study done on antibiotic susceptibility of intestinal lactobacilli of healthy children showed that all were susceptible to ampicillin, erythromycin and gentamycin and the majority were susceptible to tetracycline while 73 % were resistant to vancomycin [71]. Except for gentamycin, the study is generally in agreement with our study.
We did not define penicillin-resistant lactobacilli because the relevant breakpoints are not available in either EFSA or CLSI documents, but the MIC values we obtained are generally higher than those found by other studies, especially for L. fermentum, L. gasseri, L. salivarius and L. vaginalis [73, 74].
L. casei and L. johnsonii in this study were kanamycin-resistant which is in agreement with a previous work of probiotic L. casei and L. johnsonii isolates that were also shown to be resistant to kanamycin [70]. The same work also showed that the L. johnsonii strains were susceptible to chloramphenicol, penicillin and tetracycline, which is also consistent with our study. Also, the high prevalence of kanamycin resistance among the lactobacilli was observed in both studies (81 % in the published probiotic study [70] and 98 % in our study). Lactobacilli are generally more susceptible to chloramphenicol, erythromycin and tetracycline but more resistant to the aminoglycosides [75]. Our result is in agreement with this since nearly all strains were susceptible to chloramphenicol, tetracycline and erythromycin (see Table 4). The only exceptions were the three tetracycline-resistant L. ruminis isolates, and the one L. vaginalis isolate and one of the L. gasseri isolates that were erythromycin-resistant.
It has been shown that nearly 70 % of lactobacilli, including L. fermentum, L. gasseri, L. johnsonii and L. casei, isolated from human faeces were found to harbour tet(M) and or erm(B) genes [17]. Also, L. fermentum, L. salivarius, L. casei and L. vaginalis from food were found to contain these genes, and they were resistant to erythromycin and tetracycline [74]. Therefore, erythromycin resistance observed in our L. gasseri and L. vaginalis isolates could be due to presence of erm(B) genes in these bacteria. In contrast to a study by Delgado et al. [76], we found high prevalence of multidrug resistance in human faecal lactobacilli. However, this multidrug resistance refers to aminoglycosides and vancomycin and should not impose a big concern, since most of these resistances are most likely intrinsic and not transferrable [70, 76].
This work demonstrates that probiotic properties such as bacteriocin production is a major feature among faecal LAB. Some of the isolates shall be evaluated for their potential as probiotics. Also, putative virulence properties like gelatinase and cytolysin activities are rarely found among faecal LAB. The present study also shows that antibiotic resistance is frequently found among most of the faecal LAB.
References
Palmer C, Bik EM, DiGiulio DB et al (2007) Development of the human infant intestinal microbiota. PLoS Biol 5:e177
Penders J, Thijs C, Vink C et al (2006) Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118:511–521
Adlerberth I, Wold AE (2009) Establishment of the gut microbiota in Western infants. Acta Paediatr 98:229–238
Orrhage K, Nord CE (1999) Factors controlling the bacterial colonization of the intestine in breastfed infants. Acta Paediatr Suppl 88:47–57
Mitsou EK, Kirtzalidou E, Oikonomou I et al (2008) Fecal microflora of Greek healthy neonates. Anaerobe 14:94–101
Gronlund MM, Lehtonen OP, Eerola E et al (1999) Fecal microflora in healthy infants born by different methods of delivery: permanent changes in intestinal flora after cesarean delivery. J Pediatr Gastroenterol Nutr 28:19–25
Ljungh A, Wadstrom T (2006) Lactic acid bacteria as probiotics. Curr Issues Intest Microbiol 7:73–89
Reid G, Jass J, Sebulsky MT (2003) Potential uses of probiotics in clinical practice. Clin Microbiol Rev 16:658–672
Thomas DW, Greer FR (2010) Probiotics and prebiotics in pediatrics. American Academy of Pediatrics 126:1217–1231
Corr SC, Li Y, Riedel CU et al (2007) Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc Natl Acad Sci U S A 104:7617–7621
EFSA (2012) EFSA panel on additives and products or substances used in animal feed (FEEDAP); guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA Journal 10(6):2740
Salyers AA, Gupta A, Wang Y (2004) Human intestinal bacteria as reservoirs for antibiotic resistance genes. Trends Microbiol 12:412–416
Teuber M, Meile L, Schwarz F (1999) Acquired antibiotic resistance in lactic acid bacteria from food. Antonie Van Leeuwenhoek 76:115–137
Gevers D, Huys G, Swings J (2003) In vitro conjugal transfer of tetracycline resistance from Lactobacillus isolates to other gram-positive bacteria. FEMS Microbiol Lett 225:125–130
Toomey N, Monaghan A, Fanning S et al (2009) Assessment of antimicrobial resistance transfer between lactic acid bacteria and potential foodborne pathogens using in vitro methods and mating in a food matrix. Foodborne Pathog Dis 6:925–933
Jacobsen L, Wilcks A, Hammer K et al (2007) Horizontal transfer of tet(M) and erm(B) resistance plasmids from food strains of Lactobacillus plantarum to Enterococcus faecalis JH2-2 in the gastrointestinal tract of gnotobiotic rats. FEMS Microbiol Ecol 59:158–166
Cataloluk O, Gogebakan B (2004) Presence of drug resistance in intestinal lactobacilli of dairy and human origin in Turkey. FEMS Microbiol Lett 236:7–12
Giraffa G (2002) Enterococci from foods. FEMS Microbiol Rev 26:163–171
Franz CM, Stiles ME, Schleifer KH et al (2003) Enterococci in foods—a conundrum for food safety. Int J Food Microbiol 88:105–122
Murray BE (1990) The life and times of the Enterococcus. Clin Microbiol Rev 3:46–65
Murray BE (1998) Diversity among multidrug-resistant enterococci. Emerg Infect Dis 4:37–47
Moellering RC Jr (1998) Vancomycin-resistant enterococci. Clin Infect Dis 26:1196–1199
Jett BD, Huycke MM, Gilmore MS (1994) Virulence of enterococci. Clin Microbiol Rev 7:462–478
Shankar N, Coburn P, Pillar C et al (2004) Enterococcal cytolysin: activities and association with other virulence traits in a pathogenicity island. Int J Med Microbiol 293:609–618
Abriouel H, Omar NB, Molinos AC et al (2008) Comparative analysis of genetic diversity and incidence of virulence factors and antibiotic resistance among enterococcal populations from raw fruit and vegetable foods, water and soil, and clinical samples. Int J Food Microbiol 123:38–49
Semedo T, Almeida Santos M, Martins P (2003) Comparative study using type strains and clinical and food isolates to examine hemolytic activity and occurrence of the cyl operon in enterococci. J Clin Microbiol 41:2569–2576
Poeta P, Costa D, Klibi N et al (2006) Phenotypic and genotypic study of gelatinase and beta-haemolysis activities in faecal enterococci of poultry in Portugal. J Vet Med B Infect Dis Vet Public Health 53:203–208
Eaton TJ, Gasson MJ (2001) Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl Environ Microbiol 67:1628–1635
Mortvedt CI (1990) IF Nes: plasmid-associated bacteriocin production by a Lactobacillus sake strain. J Gen Microbiol 136:1601–1607
Forberg T.(2005) Acid bacteria of different origin, production of antimicrobial substances and distribution of bacteriocin genes. Master Thesis. Norwegian University of Life Sciences, Ås.
Herrera CB. (2006) Faecal bifidobacteria and lactic acid bacteria from breast-fed infants with emphasis on the antimicrobial properties of lactic acid bacteria. Master Thesis. Norwegian University of Life Sciences, Ås.
Booth MC, Bogie CP, Sahl HG et al (1996) Structural analysis and proteolytic activation of Enterococcus faecalis cytolysin, a novel lantibiotic. Mol Microbiol 21:1175–1184
Jett BD, Jensen HG, Nordquist RE et al (1992) Contribution of the pAD1-encoded cytolysin to the severity of experimental Enterococcus faecalis endophthalmitis. Infect Immun 60:2445–2452
Solheim M, Aakra A, Snipen LG et al (2009) Comparative genomics of Enterococcus faecalis from healthy Norwegian infants. BMC Genomics 10:194
Qin X, Singh KV, Weinstock GM et al (2000) Effects of Enterococcus faecalis fsr genes on production of gelatinase and a serine protease and virulence. Infect Immun 68:2579–2586
CLSI: Performance standards for antimicrobial susceptibility testing; twenty-second informational supplement. CLSI document M100-S22. Clinical and Laboratory Standards Institute, Wayne, PA (2012)
Faye T, Brede DA, Langsrud T et al (2002) An antimicrobial peptide is produced by extracellular processing of a protein from Propionibacterium jensenii. J Bacteriol 184:3649–3656
Bogovic-Matijasic B, Rogelj I, Nes IF et al (1998) Isolation and characterization of two bacteriocins of Lactobacillus acidophilus LF221. Appl Microbiol Biotechnol 49:606–612
Birri DJ, Brede DA, Forberg T et al (2010) Molecular and genetic characterization of a novel bacteriocin locus in Enterococcus avium isolates from infants. Appl Environ Microbiol 76:483–492
Sedgley CM, Clewell DB, Flannagan SE (2009) Plasmid pAMS1-encoded, bacteriocin-related ″Siblicide″ in Enterococcus faecalis. J Bacteriol 191:3183–3188
Kawai Y, Saitoh B, Takahashi O (2000) Primary amino acid and DNA sequences of gassericin T, a lactacin F-family bacteriocin produced by Lactobacillus gasseri SBT2055. Biosci Biotechnol Biochem 64:2201–2208
Birri DJ, Brede DA, Nes IF (2012) Salivaricin D, a novel intrinsically trypsin-resistant lantibiotic from Streptococcus salivarius 5M6c isolated from a healthy infant. Appl Environ Microbiol 78:402–410
Hamilton-Miller JM, Shah S (1998) Vancomycin susceptibility as an aid to the identification of lactobacilli. Lett Appl Microbiol 26:153–154
Edwards CA, Parrett AM (2002) Intestinal flora during the first months of life: new perspectives. Br J Nutr 88(Suppl 1):S11–S18
Dominguez-Bello MG, Costello EK, Contreras M et al (2010) Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 107:11971–11975
Martin R, Heilig GH, Zoetendal EG et al (2007) Diversity of the Lactobacillus group in breast milk and vagina of healthy women and potential role in the colonization of the infant gut. J Appl Microbiol 103:2638–2644
Bennet R, Eriksson M, Tafari N et al (1991) Intestinal bacteria of newborn Ethiopian infants in relation to antibiotic treatment and colonisation by potentially pathogenic gram-negative bacteria. Scand J Infect Dis 23:63–69
Stark PL, Lee A (1982) The microbial ecology of the large bowel of breast-fed and formula-fed infants during the first year of life. J Med Microbiol 15:189–203
Yoshioka H, Iseki K, Fujita K (1983) Development and differences of intestinal flora in the neonatal period in breast-fed and bottle-fed infants. Pediatrics 72:317–321
Ahrne S, Lonnermark E, Wold AE et al (2005) Lactobacilli in the intestinal microbiota of Swedish infants. Microbes Infect 7:1256–1262
Adlerberth I (2008) Factors influencing the establishment of the intestinal microbiota in infancy. Nestle Nutr Workshop Ser Pediatr Program 62:13–29, discussion 29–33
Hufnagel M, Liese C, Loescher C et al (2007) Enterococcal colonization of infants in a neonatal intensive care unit: associated predictors, risk factors and seasonal patterns. BMC Infect Dis 7:107
Favier CF, Vaughan EE, De Vos WM et al (2002) Molecular monitoring of succession of bacterial communities in human neonates. Appl Environ Microbiol 68:219–226
O’Shea EF, Gardiner GE, O’Connor PM et al (2009) Characterization of enterocin- and salivaricin-producing lactic acid bacteria from the mammalian gastrointestinal tract. FEMS Microbiol Lett 291:24–34
Garver KI, Muriana PM (1993) Detection, identification and characterization of bacteriocin-producing lactic acid bacteria from retail food products. Int J Food Microbiol 19:241–258
Sedgley CM, Lennan SL, Clewell DB (2004) Prevalence, phenotype and genotype of oral enterococci. Oral Microbiol Immunol 19:95–101
Hallgren A, Claesson C, Saeedi B et al (2009) Molecular detection of aggregation substance, enterococcal surface protein, and cytolysin genes and in vitro adhesion to urinary catheters of Enterococcus faecalis and E. faecium of clinical origin. Int J Med Microbiol 299:323–332
Poeta P, Igrejas G, Costa D et al (2008) Virulence factors and bacteriocins in faecal enterococci of wild boars. J Basic Microbiol 48:385–392
Lopes Mde F, Simoes AP, Tenreiro R (2006) Activity and expression of a virulence factor, gelatinase, in dairy enterococci. Int J Food Microbiol 112:208–214
Coque TM, Patterson JE, Steckelberg JM et al (1995) Incidence of hemolysin, gelatinase, and aggregation substance among enterococci isolated from patients with endocarditis and other infections and from feces of hospitalized and community-based persons. J Infect Dis 171:1223–1229
Cauwerts K, Decostere A, De Graef EM et al (2007) High prevalence of tetracycline resistance in Enterococcus isolates from broilers carrying the erm(B) gene. Avian Pathol 36:395–399
Poeta P, Costa D, Rodrigues J et al (2006) Antimicrobial resistance and the mechanisms implicated in faecal enterococci from healthy humans, poultry and pets in Portugal. Int J Antimicrob Agents 27:131–137
Huys G, D’Haene K, Collard JM et al (2004) Prevalence and molecular characterization of tetracycline resistance in Enterococcus isolates from food. Appl Environ Microbiol 70:1555–1562
Barreto A, Guimaraes B, Radhouani H et al (2009) Detection of antibiotic resistant E. coli and Enterococcus spp. in stool of healthy growing children in Portugal. J Basic Microbiol 49:503–512
Fontana R, Ligozzi M, Pittaluga F et al (1996) Intrinsic penicillin resistance in enterococci. Microb Drug Resist 2:209–213
Hayes JR, English LL, Carr LE et al (2004) Multiple-antibiotic resistance of Enterococcus spp. isolated from commercial poultry production environments. Appl Environ Microbiol 70:6005–6011
EFSA (2012) EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP); guidance on the safety assessment of Enterococcus faecium in animal nutrition. EFSA Journal 10(5):2682
Chow JW (2000) Aminoglycoside resistance in enterococci. Clin Infect Dis 31: 586–589
D’Aimmo MR, Modesto M, Biavati B (2007) Antibiotic resistance of lactic acid bacteria and Bifidobacterium spp. isolated from dairy and pharmaceutical products. Int J Food Microbiol 115:35–42
Temmerman R, Pot B, Huys G et al (2003) Identification and antibiotic susceptibility of bacterial isolates from probiotic products. Int J Food Microbiol 81:1–10
Mandar R, Lijvukene K, Huftt P et al (2001) Antibacterial susceptibility of intestinal lactobacilli of healthy children. Scand J Infect Dis 33:344–349
Belletti N, Gatti M, Bottari B et al (2009) Antibiotic resistance of lactobacilli isolated from two Italian hard cheeses. J Food Prot 72:2162–2169
Klare I, Konstabel C, Werner G (2007) Antimicrobial susceptibilities of Lactobacillus, Pediococcus and Lactococcus human isolates and cultures intended for probiotic or nutritional use. J Antimicrob Chemother 59:900–912
Nawaz M, Wang J, Zhou A et al (2011) Characterization and transfer of antibiotic resistance in lactic acid bacteria from fermented food products. Current microbiol 62:1081–1089
Charteris WP, Kelly PM, Morelli L et al (1998) Antibiotic susceptibility of potentially probiotic Lactobacillus species. J Food Prot 61:1636–1643
Delgado S, Florez AB, Mayo B (2005) Antibiotic susceptibility of Lactobacillus and Bifidobacterium species from the human gastrointestinal tract. Curr Microbiol 50:202–207
Weisburg WG, Barns SM, Pelletier DA et al (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Edwards U, Rogall T, Blocker H et al (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
Stackebrandt E, Charfreitag O (1990) Partial 16S rRNA primary structure of five Actinomyces species: phylogenetic implications and development of an Actinomyces israelii-specific oligonucleotide probe. J Gen Microbiol 136:37–43
Hyink O, Wescombe PA, Upton M et al (2007) Salivaricin A2 and the novel lantibiotic salivaricin B are encoded at adjacent loci on a 190-kilobase transmissible megaplasmid in the oral probiotic strain Streptococcus salivarius K12. Appl Environ Microbiol 73:1107–1113
Xiao H, Chen X, Chen M et al (2004) Bovicin HJ50, a novel lantibiotic produced by Streptococcus bovis HJ50. Microbiology 150:103–108
Shankar N, Baghdayan AS, Gilmore MS (2002) Modulation of virulence within a pathogenicity island in vancomycin-resistant Enterococcus faecalis. Nature 417:746–750
Martinez-Bueno M, Maqueda M, Galvez A et al (1994) Determination of the gene sequence and the molecular structure of the enterococcal peptide antibiotic AS-48. J Bacteriol 176:6334–6339
Tomita H, Fujimoto S, Tanimoto K et al (1996) Cloning and genetic organization of the bacteriocin 31 determinant encoded on the Enterococcus faecalis pheromone-responsive conjugative plasmid pYI17. J Bacteriol 178:3585–3593
Balla E, Dicks LM, Du Toit M et al (2000) Characterization and cloning of the genes encoding enterocin 1071A and enterocin 1071B, two antimicrobial peptides produced by Enterococcus faecalis BFE 1071. Appl Environ Microbiol 66:1298–1304
Acknowledgements
Grants from Norwegian Research Council have supported this study. We thank Zhian Salehian for technical assistance. We acknowledge parents for allowing us to collect samples from their infants.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Birri, D.J., Brede, D.A., Tessema, G.T. et al. Bacteriocin Production, Antibiotic Susceptibility and Prevalence of Haemolytic and Gelatinase Activity in Faecal Lactic Acid Bacteria Isolated from Healthy Ethiopian Infants. Microb Ecol 65, 504–516 (2013). https://doi.org/10.1007/s00248-012-0134-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-012-0134-7