Abstract
Quantum mechanical calculations are used to explore the thermodynamics of possible prebiotic synthesis of the building blocks of nucleic acids. Different combinations of D-ribofuranose (Ribf) and N-(2-aminoethyl)-glycine (AEG) (trifunctional connectors (TCs)); the nature of the Ribf, its anomeric form, and its ring puckering (conformation); and the nature of the nucleobases (recognition units (RUs)) are considered. The combinatorial explosion of possible nucleosides has been drastically reduced on physicochemical grounds followed by a detailed thermodynamic evaluation of alternative synthetic pathways. The synthesis of nucleosides containing N-(2-aminoethyl)-glycine (AEG) is predicted to be thermodynamically favored suggesting a possible role of AEG as a component of an ancestral proto-RNA that may have preceded today’s nucleic acids. A new pathway for the building of free nucleotides (exemplified by 5’-uridine monophosphate (UMP)) and of AEG dipeptides is proposed. This new pathway leads to a spontaneous formation of free UMP assisted by an AEG nucleoside in an aqueous environment. This appears to be a workaround to the “water problem” that prohibits the synthesis of nucleotides in water.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The question on how the building blocks of today’s nucleic acids (NAs) emerged in the early Earth, billions of years ago, remains an intriguing open question for prebiotic chemistry (Castanedo 2024). A widely accepted theory for the origins of life considers that RNA was the first biopolymer to emerge (Cech 2012; Gesteland et al. 1999). This theory is known as the “RNA world” hypothesis and proposes that RNA came first because this biomolecule has catalytic activity in the form of ribosomal RNA (rRNA) and simultaneously can also transmit the genetic information in the form of messenger RNA (mRNA) (Ayukawa et al. 2019; Neveu et al. 2013; Orgel 2004).
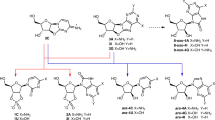
Initial attempts to synthetize nucleosides by Orgel and coworkers (Fuller et al. 1972) showed that the formation of N-glycosidic bonds between the canonical nitrogenous bases generically named as recognition units or RUs (Hud et al. 2013) and ribose, also known as the trifunctional connector or TC (Hud et al. 2013), through a condensation reaction in dehydrated conditions is thermodynamically unfavored. Orgel attempted to glycosylate ribose with adenine, guanine, inosine, and xanthine, heating the reaction at 100 °C with and without the presence of catalysts and obtained a mixture of only β- and α-ribofuranosyl adenosine with a ~ 2–10% yield.
Similar unsuccessful attempts to glycosylate uracil and cytosine with ribose were also reported (Fuller 1972). This inability of canonical bases to create nucleosides with ribose in aqueous solution has been referred as “the nucleoside problem,” a special case of the “water problem” (Hud et al. 2013; A. do Nascimento Vieira et al. 2020; Kim et al. 2016; Joyce and Orgel 1999). Alternative models have been suggested to explain how building blocks of today’s nucleic acids were selected by Nature and assembled. Amount these models we may mention the “ribose centric model” (Hud et al. 2013) and the “polymer fusion model” (Hud and Anet 2000).
Nevertheless, it is also possible that the first nucleic acids had TCs, RUs, and ionized linkers (ILs) in their building blocks different than today’s ribose (TC), the five canonical nucleobases (RUs), and the phosphate group (IL), respectively. This hypothesis proposes that the contemporary nucleic acids emerged as a product of evolutionary selection from a proto- and pre-RNAs in which non-canonical (alternative) TCs and RUs glycosylated and assembled easier in prebiotic conditions (Hud et al. 2013). These components may have then evolved to become today’s β-ribofuranose, Adenine (A), Guanine (G), Cytosine (C), Thymine (T), Uracil (U), and phosphate (Engelhart and Hud 2010).
On the search for alternative prebiotic RUs, Kolb et al. (Kolb et al. 1994) reported the prebiotic synthesis of nucleosides containing a non-canonical base (either urazole or 1,2,4-triazolidine-3,5-dione) and ribose. They obtained a mixture of α- and β-configurations of the urazole nucleosides with the ribose in the furanose (F)- and pyranose (P)-forms. From that mixture the β-ribopyranose was predominant with a 53% yield. Following this idea Cafferty and coworkers (Cafferty et al. 2018) explored a library of 91 RUs (that included the five canonical bases (A, G, C, T, U)) based on the following criteria:
-
can glycosylate with ribose in the presence of water,
-
able to create complementary base pairing with at least two hydrogen bonds,
-
capable of π-stacking in aqueous environment,
-
chemically and photo-stable,
-
with conceivable synthesis in prebiotic conditions,
-
and “good chromophores” which makes them suitable to absorb UV radiation believed to have flooded prebiotic Earth (Douglas 2017).
Three non-canonical nucleobases fulfill all these conditions simultaneously. These are the triazine melamine (MM) and the pyrimidines 2,4,6-triaminopyrimidine (TAP) and barbituric acid (BA). In addition to these three bases the triazine cyanuric acid (CA) can create hexads in aqueous solution. Hexads are defined as linear π-stacked assemblies in the form of a six-sided polygon (Cafferty et al. 2016, 2013; Cafferty and Hud 2014, 2015; Fialho 2019; Li et al. 2016). Furthermore, numerous studies have shown the ability of MM, TAP, and BA to overcome the “glycoside problem” by creating N- and C-glycosides with ribose. As an example, a mixture of β-N-, α-N-, β-C-, and α-C-nucleosides can be obtained, with a yield of ~ 33–55%, by heating at 35 °C for 24 h TAP with either ribofuranose or ribopyranose. The β-C-ribofuranose nucleoside was predominant with a 20% yield (Chen et al. 2014).
Let’s address another fundamental question: was ribose present in the first proto-nucleic acids? The most accepted synthetic route for ribose in prebiotic conditions is the formose reaction. This reaction was first proposed by Butlerow (Butlerow 1861) and includes the polymerization of formaldehyde (H2CO) to obtain sugars. However, this method faces a challenge termed “the asphalt problem” as simple sugars can also polymerize to create asphalts through a series of competitive enolization/aldol additions between the carbonyl groups. As a result of the asphalt problem, ribose is obtained only as a minor product with ~ 2% of yield. Additionally, the 5-member ring (5-MR) ribofuranose is the least of the products from the mutarotation of ribose in water with a yield of around ~ 13% for the β- and ~ 7% for the α-anomer (Sutherland 2010; Drew et al. 1998).
Given the compilation of challenges associated with considering ribose as the TC present in the first nucleic acids, alternative candidates for the first TC are sought for. Nucleic acids with a TC different from ribofuranose are named “xeno-nucleic acids” (XNA) and they, incidentally, have a wide range of applications as biomarkers for cancer, HIV, and hepatitis (Sefah et al. 2014).
Among several classes of XNA nucleic acids analogs, N-(2-aminoethyl)-glycine (AEG) peptide nucleic acids (aegPNAs)—where the sugar-phosphate is replaced by a polypeptide backbone containing repetitive units of N-(2-aminoethyl)glycine (AEG) bound through a carbonyl methylene group to the nucleobases (Frenkel-Pinter et al. 2020; Wu et al. 2017; Lee et al. 2013; Mateo-Martí and Pradie 2010; Nielsen 2007, 1996; Menchise et al. 2003)—have become attractive as a plausible candidate for a proto-nucleic acid for a number of reasons, some of which are enumerated below:
-
1.
The sugar-phosphate backbone of DNA/RNA is replaced in aegPNAs by a peptide bond which is more resistant to hydrolysis in aqueous solution than the phosphodiester bond (Frenkel-Pinter et al. 2020).
-
2.
AEG may have been present in the early Earth environment and can have conceivably been incorporated in proto-nucleic acids and replaced later by 2'-deoxyribose or ribose through natural selection. Bolstering this proposition is the present-day traces of AEG which are found in certain strains of cyanobacteria (Josa et al. 2013). These bacteria are believed to have emerged around 3.5 Gyr ago and are known to be resilient in extreme environments such as geothermal vents which, in of themselves, are considered as a plausible environment for the emergence of early forms of life.
-
3.
The components of aegPNAs’ building blocks have been synthesized prebiotically. These components include AEG as TC and the N-acetic acid derivatives of adenine, guanine, cytosine, and uracil as RUs (Nelson et al. 2000). The acetic acid derivatization of the nucleobases is necessary in order to form the amide linkage between the AEG and the bases that is part of the polyamide backbone. For example, the famed Miller-Urey experiments (McCollom 2013; Miller 1955, 1953) produced, in vitro, precursors of amino acids and L-glycine in a simulated chemical composition of the early Earth that included electric discharge to simulate thunder. In a later study, Nelson and coworkers (Nelson et al. 2000) studied the prebiotic synthesis of AEGs by i) electric discharge experiments of a mixture of CH4 (g), N2(g), NH3(g), and H2O (10–5–10–6% of yield for AEG), ii) polymerization of NH4CN (10–5% of yield for AEG), and iii) polymerization of NH4CN catalyzed by H2CO (10–5% of yield for AEG). Despite the low yields of AEG from the three previous methods Nelson and coworkers hypothesize that the production of this amino acid could have been possible in the early Earth due to the high solubility of its monohydrochloride and that AEG can also be obtained through the Strecker synthesis (Strecker 1854), from the reaction of ethylenediamine with HCN and H2CO. Ethylenediamine can be obtained with a yield of 0.05% through the reaction of NH4CN and H2CO in the presence of UV radiation (Nelson et al. 2000). Nelson et al., estimate that low concentrations of the Strecker mixture (~ 10−6 M) can produce up to 33% of AEGs. Adenine-N9-acetyl (acetyl = Ac) and guanine-N9-Ac were also obtained from polymerization of HCN in the presence of glycine in 0.0062% and 0.011%, respectively. Meanwhile, cytosine-N1-Ac and uracil-N1-Ac were produced from the reaction between hydantoic acid with cyanoacetaldehyde at 100 °C in 18% and 1.8% yields, respectively.
-
4.
Nelson and coworkers (2000) also found that AEGs can polymerize in dry conditions at 100 °C better than α-amino acids at higher temperatures.
-
5.
The carbon atoms of the aegPNAs backbone lack chirality. This property may have given them an evolutionary advantage in the early Earth since, in contrast with today’s nucleic acids, they are not subjected to enantiomeric cross-inhibition (ECI). ECI refers to the exclusive relationship between DNA/RNA biochemical functions and the D-enantiomerism of the ribofuranose or 2’-deoxyribofuranose. For example, studies by Joyce and Orgel (1999) show that the template directed polymerization of DNA containing D-ribose can be inhibited when D-ribose-phosphate is replaced by the L-enantiomer in the building blocks of the D-homochiral template (Hehre et al. 1986; Rao et al. 2009).
Another important question that requires further explanation regards the adoption by the majority of contemporary DNA and RNA of a β- instead of an α-configuration at the anomeric C1’ position of the sugar D-ribofuranose (Fig. 1). But why not α?
Orientation of the nucleobase at the C1’ of the furanose with respect to the hydroxymethyl group at the C4’ for the β- (left) and α-anomers (right) of canonical ribonucleosides. RU: nucleobase or recognition unit. R: can be either H in 2’-deoxynucleosides (in DNA) or OH in ribonucleosides (in RNA) (taken from (Ni et al. 2019) and reprinted with permission of RSC Advances. © Royal Society of Chemistry, 1999)
Numerous experimental reports have suggested that α-strands of DNA can create Watson and Crick (WC) complementary α-α and α-β double stranded polynucleotides (Froeyen et al. 2001; Guesnet et al. 1990; Paoletti et al. 1989; Lancelot et al. 1989, 1987).
It is also well-known that 5-MR sugars (pentoses), e.g., ribose, adopt two preferential 3T2 (2’-exo-3’-endo) and 2T3 (2’-endo-3’-exo) conformations in equilibrium for their respective β- and α-anomers (Fig. 2). Thibaudeau and coworkers propose that the pseudorotational equilibrium between these two conformations in ribofuranose is an important factor to take into account when comparing the stability of the β- and α-anomers of the building blocks of nucleic acids (see Chapter 2 of Thibaudeau et al. (2005) and (Thibaudeau et al. 1997, 1994)).
Equilibrium between the 3T2 (2’-exo-3’-endo) and 2T3 (2’-endo-3’-exo) conformations for a nucleoside with a 5-MR sugar (see pp. 22–46 in (Thibaudeau et al. 2005) and (Thibaudeau et al. 1997, 1994)). ∆X: change in enthalpy (∆H°) or Gibbs free energy (∆G°). Red chemical groups represent atoms over, meanwhile blue atoms under the mean plane of the sugar. Modified from (Thibaudeau et al. 1997) and reprinted with permission of Elsevier © Elsevier, 1997
Castanedo and Matta (2022a, b) explore the role of thermodynamics as a driver of evolutionary selection for the chemical structure of today’s nucleic acids in implicit solvation models and in vacuum/gas phase in a study titled “[o]n the prebiotic selection of nucleotide anomers: a computational study”. C and M estimate the energetic differences between β- and α-anomers of the sugars, their monophosphates, the derived nucleosides and nucleotides and the thermodynamic feasibility of the synthesis of the five canonical nucleotides through a classic and alternative pathway in which the order of the reactants was changed. The changing of the order of addition of the components of the building blocks locks the intermediates in different local minima on the potential energy hypersurface and hence results in energetic differences of the products if those have different conformations.
Castanedo and Matta report a marginal energetic preference ~ 8 kJ/mol (or less) for the β- over the α-anomers for some nucleosides containing adenine and guanine in vacuum (Castanedo and Matta 2022a, b). The classic pathway, in which two condensation reactions give the nucleotides as product, was found to be the preferred route in vacuum. In aqueous medium, however, neither pathway is energetically favored (both are non-spontaneous). These quantum chemical results are consistent with the “water problem” prohibiting the formation of nucleotides in aqueous media from their corresponding components.
The present paper explores the thermodynamic plausibility of the synthesis of potential prebiotic nucleosides from the canonical D-ribofuranose and non-canonical AEG and the five canonical bases A, G, C, T, U, the four non-canonical bases TAP, BA, MM, and CA, and their corresponding acetyl derivatives. The influence of the two preferential sugar ring conformations 2T3 and 3T2 for the D-ribofuranose in the corresponding α- and β-anomers is also examined.
Furthermore, we wish to revisit whether the synthesis of canonical and/or non-canonical nucleosides containing ribofuranose and AEG is thermodynamically favored following the “classic model.”
Finally, a hypothetical pathway to synthesize free β-uridine 5’-monophosphate (β-UMP) as free nucleotide and an AEG dipeptide in aqueous medium is proposed by building β-UMP on an AEG-Ac-C1-C template. Part of this work has recently been published in the PhD thesis of one of the authors [L.A.M.C.] (Castanedo 2024).
Computational Methods
The chemical structures of all the canonical components (TCs and RUs) from RNA were modified from the nucleotides contained in the structure of a B-DNA dodecamer (Protein Data Bank Identifier [PDB ID] # 1BNA) (Drew et al. 1981), while the structure corresponding to the PDB ID 1PNN (Betts et al. 1995) structure was used to create the initial 3D geometries of the components of aegPNA nucleosides by using the graphical interfaces UCSF Chimera (Pettersen et al. 2004), Hyperchem 7.0 (HyperChem release 7.0 2002), and GaussView 5.0 (Dennington et al. 2009).
All possible combinations between each TC and RU represented in Fig. 3 were explored. The enol form of barbituric acid (BA) which prevails in aqueous solution (Lubczak and Mendyk 2008) has been used in all calculations. For nucleosides containing D-ribofuranose (Ribf), each RU has been placed in the β- and α-configurations at the anomeric position C1’. The two preferential sugar ring conformations 2T3 or 2’-exo-3’-endo and 3T2 or 2’-endo-3’-exo for Ribf have also been considered.
Canonical and non-canonical trifunctional connectors (TCs) (top panel) and recognition units (RUs) (bottom panel) considered in the modeling of the nucleosides. (Top panel): Ribf = D-ribofuranose (β-anomer present in RNA) and AEG = N-(2-aminoethyl)-glycine (present in peptide nucleic acids (PNAs)). (Bottom panel) A = adenine, G = guanine, C = cytosine, T = thymine, U = uracil, TAP = 2, 4, 6-triaminopyrimidine, BA = barbituric acid in enol form, MM = melamine, CA = cyanuric acid. The R group is H for the RUs in non-PNAs nucleosides and acetyl (–CH2–COOH) for acetyl derivatives of RUs in the PNAs nucleosides. The blue C and green N represent reactive centers for the condensation reactions. Notice that in the case of TAP and BA they can either create N- or C5-nucleosides. Vertical lines with a dotted shadow indicate bonds that are broken during the condensation reactions and curved arrows denote rotatable bonds. In the case of Ribf-nucleosides, the leaving group is R = H while for AEG-nucleosides it is the hydroxyl (–OH) of the R = Ac
In total 110 nucleosides have been constructed. The total number of nucleosides included 88 nucleosides with a Ribf as TC {(11 RUs) × 2 environments (vacuum and implicit solvation) × 2 sugar ring conformations (3T2, 2T3) × 2 anomeric forms (β- and α-anomers, respectively) and 22 nucleosides with an AEG {(11 RUs) × 2 environments}.
Each of the TCs (Ribf been considered in the β- and α-configurations and with two sugar ring conformations) and RUs with rotatable bonds (curved arrows in Fig. 3) were subjected to a potential energy scan (PES) by reading their Z-matrices in the programme GRANADAROT (Montero 2019; Montero et al. 2000, 1998) which was used to generate, for each structure, 1,000 different conformers by randomly changing the dihedral angle around each rotatable bond. Each of these 11,000 (= 6,000 for RUs + 5,000 for TCs) conformers were optimized at the semiempirical PM7 (Parametric Method 7) (Stewart 2023) through a gradient minimization of energy until convergence (energy gradient threshold = 0.01 kcal·mol−1·Å−1) using MOPAC2016 (Stewart 2023). PM7 was chosen as it includes empirical corrections for dispersive and hydrogen bonding interactions.
Calculations were performed both in vacuum and in implicit continuum solvent modeled using the conductor-like screening model (COSMO) (Klamt 2011; Klamt and Schüümann 1993). For ribofuranose the coordinates of the carbon atoms in the furanose ring were kept frozen during the calculations at the PM7.
Some of the 1,000 optimized structures at PM7 converged without changing the conformation (only the values of the distances and angles were altered). For each set of final n unique optimized geometries, the final n’ conformers that collectively contribute at least 50% to the partition function (Z) were kept for refinement at a more accurate quantum mechanical level of theory. The rest of the conformers with minor contributions were discarded.
For each TC, RU, and acetylated RU with rotatable bonds in vacuum and in implicit aqueous medium the numbers n and n’ and their contribution to Z are summarized in Tables S1-S2 (in the Supplementary Information (SI)).
The n’ conformers were then subjected to a fully relaxed geometry optimization using the B3LYP DFT functional (Orio et al. 2009; Hertwig and Koch 1997; Becke 1988; Lee et al. 1988) with the 6–311+ +G(d, p) basis set (Hehre et al. 1986) as implemented in Gaussian 16 (Frisch et al. 2019). The B3LYP functional was chosen as it has been widely used for the computational modeling of building blocks of nucleic acids (Kaur et al. 2019, 2017; Šponer et al. 2011). The error bars for a similar level of theory, namely, DFT-B3LYP/6–31+G(d, p) have been benchmarked by Zhao and Truhlar (ZT) to be around 15 kJ/mol (Zhao and Truhlar 2008a). ZT obtained this estimate by comparing calculated and experimental thermodynamic data for 177 compounds (Zhao and Truhlar 2008a). Another study by ZT (Zhao and Truhlar 2008b) has reported a mean unassigned error of 13.8 kJ/mol in the estimation of B3LYP/6–31+G(d, p) interaction energies for 22 hydrogen-bonded complexes. Meanwhile, Rao and coworkers (Rao et al. 2009) evaluate 11 DFT functionals for their accuracy in estimating hydrogen bonding energies and relative energies of a conformational scan for 14 systems of biological interest that includes glycine, proline, and serine. In this study, the highest level of theory in predicting conformational energies is DFT-B3LYP/6–31++G(2d, 2p) with a mean absolute deviation of ~ 6 kJ/mol. Castanedo and Matta (2022a, b) estimate the uncertainty of the B3LYP/6-31G(d, p) level of theory to be within 13–17 kJ/mol, which is probably similar to the level of theory in this work.
From the final DFT-refined n’ conformers for each component the corresponding nucleoside has been designed by using an in-house bash-python script that manipulates the Z-matrices and internal coordinates to create a customized glycosidic bond between each TC and RU.
For consistency, the length of the C- and N-glycosidic bonds between each RU and TC, have been initially set to 1.52 Å while the dihedral angle that contains the glycosidic bond has been initially set to 180°.
Each of the 506 nucleoside structures was then subjected to a fully relaxed scan from 0 to 360° in 6 steps of 60° on the dihedral that involves the glycosidic bond. The lowest energy structure from each scan has been refined by subjecting it to a fully unconstrained optimization at the DFT-B3LYP/6–311++G(d, p) level of theory to obtain the final structures of the nucleosides in vacuum and in implicit solvation. A harmonic frequency calculation has been performed to ensure the absence of imaginary frequencies that indicates that the final structures are minima on the PES.
The thermodynamic feasibility for the synthesis of the canonical and non-canonical nucleosides following the “classic model” has been analyzed by estimating the ∆G° for the condensation reaction as:
for non-aegPNA nucleosides, and
for the aegPNA nucleosides.
Aqueous solvation has been accounted for in the DFT calculations using the Integral Equation Formalism variant of the “Polarizable Continuum Model” (IEFPCM) (Tomasi et al. 2010; Tomasi 2011; Barone et al. 1997; Cossi et al. 1996; Miertuš and Tomasi 1982; Miertuš et al. 1981) implemented in Gaussian 16 (Frisch et al. 2019).
If we accept as true the “water problem” (Joyce and Orgel 1999) this implies that the complex prebiotic chemistry that may have produced the first building blocks of ancestral NAs took place in a non-aqueous environment or at least that the reactants had controlled exposure to water (see for example Ref. (A. do Nascimento Vieira et al. 2020) and literature cited therein). Thus, the effect of aqueous solvation has been included in this study for a complete evaluation of this “problem” as well.
The 2, 4, 6-triaminopyrimidine (TAP) and the barbituric acid (BA) can create either N- or C-glycosidic bonds (Fialho 2019; Fialho et al. 2020), hence in the next sections TAP-C5 or BA-C5 and TAP-N or BA-N refer to the C5- and N-glycosides, respectively.
For the modeling of the new prebiotic pathway to obtain dipeptides and free nucleotides the base uracil was selected for the nucleosides since due to the “water problem” it is not possible to glycosylate it with ribose in water.
The n’ conformers from β-Ribf with the 2T3 sugar ring conformation, AEG, uracil (U), and cytosine (C)-N1-Ac were refined at the M05-2X/6–311++G(d, p).
The meta-generalized hybrid functional M05-2X has been selected since it has been widely used in the modeling of non-covalent interactions (hydrogen bonds and dispersive interactions) in molecular systems of biological interest predicting results in terms of energies and geometries comparable to experimental data (Zhao and Truhlar 2008b, 2007; Jissy et al. 2011; Johnson et al. 2009). Some of these studies include the optimization of π-stacking systems containing DNA nitrogenous bases (Zhao and Truhlar 2008a, 2007; Josa et al. 2013).
For each newly created bond a fully relaxed torsion scan was performed at the M05-2X/6–31G(d, p) level by performing 6 steps of 60°. The local minimum from each PES was then reoptimized at the higher M05-2X/6–311++G(d, p) level of theory.
The ∆G° for each reaction step in this synthetic sequence is estimated as follows:
Results and Discussion
Thermodynamics of the Formation of Canonical and Non-Canonical Nucleosides
The prebiotic synthesis of (non-) canonical nucleosides, that is, the condensation of an electrophile {TC} and a nucleophile {RU}, is explored thermodynamically on the basis of theoretical calculations. The broad question addressed is whether the emergence of proto-nucleosides that may have preceded the building blocks of today’s DNA and RNA is consistent with this simple modeling from a thermodynamical standpoint. The order of the synthetic steps is unimportant since the free energy is a state function and hence depends only on the two endpoints of the reaction except if the intermediate reactants are trapped in different potential energy wells on the conformational potential energy hypersurface.
It is well documented that using the classic model implies facing the “nucleoside problem” as a specific case of the “water problem” (Hud et al. 2013; A. do Nascimento Vieira et al. 2020; Kim et al. 2016; Joyce and Orgel 1999). This challenge refers to the inability of canonical bases to N-glycosylate with ribose in an aqueous environment due to the thermodynamic instability of the glycosidic bond. The “nucleoside problem” was first reported by Orgel et al. in 1972 (Fuller et al. 1972). Despite these challenges, Orgel et al. obtained in their study adenosine albeit in low yield (1–5%). Interestingly, these authors also obtained N-nucleosides with the adenine exocyclic NH2 in 50–70% yield.
To overcome the “nucleoside problem” different strategies have been proposed in the literature that include using different synthetic routes and non-canonical components (where non-canonical refers to components non present in the building blocks of today’s NAs). In this paper we use non-canonical components to glycosylate the TCs to RUs.
We searched the literature for TCs that can create non-conventional xeno-NAs backbones, selecting AEG for its highlighted advantages and RUs that have been proven to be successful in circumventing the “glycoside problem.” These RUs are 2,4,6-triaminopyrimidine (TAP), barbituric acid (BA), and melamine (MM).
Additionally, the base cyanuric acid (CA), is also tested since its nucleotides can create hydrogels (stacking hexads) through complementary base pairing with BA nucleotides (Cafferty et al. 2016), adenosine 5’-monophosphate (AMP), and 2,6-diaminopurine (DAP) (Li et al. 2016), similarly to the complementary base pairing observed in today’s DNA double helixes.
TAP is a pyrimidine that contains three exocyclic amino groups. This base exhibits four nucleophilic sites: The three exocyclic NH2 groups in positions 2, 4, 6 and the endocyclic C5. The amino groups in position 4 and 6 are equivalent by symmetry and exhibit a better nucleophilic character than the one in position 2 because of its depleted negative charge due to the electron withdrawing effect of the two ortho NH2 in position 4 and 6. Additionally, the prebiotic synthesis of TAP has been previously proposed (Pérez-Fernández et al. 2022; Salván et al. 2020; Trinks 1987).
TAP can glycosylate with different TCs depending on the reaction conditions (Fialho 2019; Fialho et al. 2018). This property is enhanced by its higher solubility in water compared to canonical bases (Chen et al. 2014). For example, Chen and coworkers (Chen et al. 2014) were the first to test the glycosylation of TAP with ribose by heating the mixture in aqueous solution at 35–95 ºC and pH = 8. A complex mixture of products was obtained. An important result from this study is that after 10 days of reaction a combined yield of 60% was obtained for the mixture of TAP-ribose and 90% was obtained for the same mixture at a higher temperature. The β-ribofuranose-C5-TAP nucleoside was detected as majoritarian product.
Studies by Cafferty and coworkers (Cafferty et al. 2018) reproduced these results obtaining in similar conditions (35 ºC and 10 days of reaction) a complex mixture of TAP-nucleosides that included pyranose and furanose rings for the ribose, β- and α-anomers and C5- and N-glycosides identified by EM-HPLC after basic hydrolysis in NH4OH for 4, 20, and 44 h. This mixture represented a combined yield of 60%. The presence of N-nucleosides with exocyclic NH2 of TAP is somehow related with the findings by Fuller et al. (Fuller 1972) of N-glycosides of adenosine.
Even as N-nucleosides are predominant in today’s DNA and RNA, C-nucleosides can also be found in Nature, e.g., pseudouridine which is a product from the post-translation of RNA. For example, rRNA from mammals contains around 100 pseudouridines per ribosome (Chen et al. 2014; Clercq 2015; Wellinton and Benner 2006). Several of these C-nucleosides have anticancer and antibiotic activity (Clercq 2015; Wellinton and Benner 2006).
Fialho expanded on the work by Cafferty et al. (Cafferty et al. 2018) by testing the glycosylation of TAP with 17 different sugars that included hexoses, pentoses, and tetroses. They heated at 85 °C a mixture of TAP and sugar in a 1:1 ratio for 24 h at pH = 1 or 7. The products were analyzed by UV-LC/MS and 1H-NMR and it was proven that all glycosylation reactions proceeded to a certain extent. The sugar that exhibited the higher yield of glycosylation was the pentose arabinose with 61% and 55% at pH = 1 and 7, respectively. Ribonucleosides were obtained in a 28% and 31% of yield at pH = 1 and 7, respectively (Fialho 2019; Fialho et al. 2020, 2018).
Barbituric acid (BA) is a strong acid (pKa = 4) that can react with ribose in aqueous solution due to the nucleophilic character of its barbiturate ion (Fialho 2019). The glycosylation of BA has been tested with glucosamine at pH = 7 in a 1:2 ratio of BA:glucosamine, for 10 h at 50 °C. The majoritarian C-β-nucleoside has been produced in basic catalysis (Fialho 2019; Gonzalez et al. 1986). The C-glycoside of the enol form of BA can create stacked hexads with MM in aqueous solution (Cafferty et al. 2016).
Melamine (MM) contains a heterocyclic ring with three nitrogen atoms that makes it more electron withdrawing, hence it is less nucleophilic than TAP and BA. Additionally, MM is less soluble in water at pH \(\ge\) 5. Its solubility at 65 ºC is 0.2 M (Fialho 2019). MM can co-precipitate in aqueous solution with phosphates, sulfates, and carboxylates (Fialho 2019).
Nevertheless, Cafferty and coworkers (Cafferty et al. 2016) showed that MM can glycosylate with ribose in aqueous solution when the reaction lasts for 24 h at 20 ºC despite the MM disadvantages described above. Both β- and α-anomers of the MM ribosides with the 5-MR and 6-MR of ribose were detected by AFM-HPLC and by measuring the 1H-NMR coupling constants for the anomeric hydrogens.
Fialho (Fialho 2019) explored the reactivity of 80 reactions between 8 nucleophilic bases that included adenine, uracil, TAP, BA, CA, and MM with a set of 10 electrophiles that included the aldehydes D-, L-glyceraldehyde, D-ribose, one anhydride, one imide, three esters, two thioesters, and one Michael acceptor. All nucleobases tested, apart from uracil and CA, were able to react with D-ribose.
Table 1 and Figs. 4, 5, represent the values and the bar graphs for the ∆G° of reaction for all 110 nucleosides. ∆G° < 0 suggests that the condensation reaction is thermodynamically favored meanwhile ∆G° > 0 indicates otherwise.
Comparison of Gibbs energies of reaction (∆G°) at 298 K for the classic synthesis, leading to the 5 canonical and 6 non-canonical β- and α-counterparts of Ribf nucleosides. Each colored bar represents the initial puckering conformations for the 5-MR sugar. (Top): ∆G° obtained at B3LYP/6–311++G(d, p) in vacuum, (bottom): ∆G° at B3LYP/6–311++G(d,p) using the IEFPCM model to include aqueous implicit solvation
Comparison of the Gibbs energies of reaction (∆G°) at 298 K for the classic synthesis, leading to the 11 aegPNA nucleosides between AEG and the five canonical acetylated bases (RUs-Ac): A = adenine-N9-Ac, G = guanine-N9-Ac, C = cytosine-N1-Ac, T = thymine-N1-Ac, U = uracil-N1-Ac, and the six non-canonical RUs-Ac: TAP-C5 = 2,4,6-triaminopyrimidine-C5-Ac, TAP-N = 2,4,6-triaminopyrimidine-N4-Ac, BA-C5 = barbituric acid-C5-Ac, BA-N = barbituric acid-N1-Ac, CA = cyanuric acid-N5-Ac, and MM = melamine-N-Ac. (Top): ∆G° obtained at B3LYP/6–311++G(d, p) in vacuum, (bottom): ∆G° at B3LYP/6–311++G(d, p) using the IEFPCM model for the aqueous solvation
Figure 4 represents the ∆G° values for the classic formation of the Ribf nucleosides. For the case of the canonical nucleosides in vacuum the formation of both β-2T3 and β-3T2 is estimated as more favorable over the α-anomers. The cytidine has the more negative values for the ∆G° with − 16.5 and − 17.8 kJ/mol for the 2T3 and β-3T2, respectively. A similar picture is observed in the case of the canonical nucleosides in aqueous environment with the exceptions of the thymidine and uridine for which no reaction is favored. The nucleoside that is more favored in aqueous solution is the β-2T3 adenosine with a ∆G° = −18.6 kJ/mol.
In the case of the synthesis of the non-canonical nucleosides containing Ribf in vacuum even when the β-2T3 and β-3T2 are more favored to be obtained for the TAP-C5 and the BA-C5, in the case of TAP-C5 the energies are within the error of the method. All BA-C5 anomers in all sugar ring conformations are predicted to be synthesized with values that overcome the error of the DFT method: (∆G° = −32.7 kJ/mol for the α-2T3; ∆G° = −50.7 kJ/mol for the β-2T3; ∆G° = −36.7 kJ/mol for the α-3T2 and ∆G° = −48.1 kJ/mol for the β-3T2). In contrast, the formation of BA-N and CA Ribf nucleosides is estimated to be unfavorable with values beyond the DFT intrinsic error (BA-N: \(\approx\) 11 to 18 kJ/mol and CA: \(\approx\) 10 to 23 kJ/mol).
For the non-canonical Ribf nucleosides in implicit solvation a similar trend is observed. In this case the β-anomer of both F-form conformations is predicted to be thermodynamically favored while the α-anomer is not for the TAP-C5, BA-C5, BA-N, and MM. For TAP-C5 the lowest ∆G° value is for β-3T2 with a ∆G° = −15.2 kJ/mol. The BA-C5 nucleoside has the lowest ∆G° values across all non-canonical nucleosides in water for the β-anomers: (∆G° = −26.5 kJ/mol for the β-2T3 and ∆G° = −22.1 kJ/mol for the β-3T2). Again, the formation of BA-N and CA nucleosides was estimated as non favored with ∆G° ≈ 2.6–26.8 kJ/mol. The formation of MM nucleosides was favored but the energies were within the error for the estimation of ∆G° at the B3LYP/6–311++G(d, p) level.
Since the condensation reaction of nucleobases with ribose has been widely reported in the literature, we can compare our results with some of these reports. For the case of the canonical nucleosides, it can be noticed that our results are in agreement with the studies by Fuller et al. (Fuller et al. 1972; Fuller 1972) on the condensation reaction between different bases and ribose in aqueous solution. In these two papers the authors obtained β-glycosides in a low yield for the case of the reaction between A or G and ribose. No product was observed when reacting a mixture of pyrimidines with the ribose in aqueous solution. Similarly in our study the formation of β-nucleosides for A and G is predicted to be thermodynamically favored in both aqueous and non-polar environments. Our results are consistent with the well-known “nucleoside problem” in the case of pyrimidine bases (Fuller et al. 1972; Fuller 1972; Benner et al. 2012).
In the case of the non-canonical nucleosides our results are in agreement with: (1) the results obtained by Chen et al. (Chen et al. 2014) for the glycosylation of TAP with ribose where the β-ribofuranose-C5-TAP was the majoritarian product, (2) the studies by Cafferty and coworkers (Cafferty et al. 2018, 2016; Cafferty and Hud 2014) for the synthesis of TAP, BA, CA, and MM ribosides, and (3) the work by Fialho (Fialho 2019; Fialho et al. 2020, 2018) in which the CA does not to react with the ribose, meanwhile β-nucleosides were obtained for TAP, BA, and MM in different yields depending on the reaction conditions.
In our case the β-anomers of the TAP-C5, TAP-N, BA-C5, and MM nucleosides with Ribf with the initial sugar ring conformations 2T3 and 3T2 were favored in implicit solvation. This result suggests that at least TAP, BA, and MM can react with ribose in water and circumvent the “nucleoside problem.”
In a previous paper, Castanedo and Matta (2022a, b) analyze the plausibility of a similar classic synthesis for the β- and α-anomers of canonical 2’-deoxyribofuranose (2dRibf) and Ribf nucleosides but at a lower DFT-B3LYP/6-31G(d, p) level in vacuum and in implicit aqueous medium using IFPCM. In this paper it was found that, overall, the synthesis of neither of the canonical nucleosides could be predicted as thermodynamically favored/unfavored in either environment since most of the energies were within the intrinsic error of the method used (3–4 kcal/mol). Meanwhile, in this study the formation of some A, G, and C nucleosides is predicted to be favored through the condensation reaction of their components when considering different puckering conformations of the sugar and a higher DFT level of calculation.
The only other computational modeling studies of TAP, MM, BA, and CA nucleosides(tides) that we could find are the works by Kaur et al. (2017, 2019). In the publication of 2019, Kaur and coworkers studied, using molecular dynamics (MD), molecular mechanics (MM), and DFT-B3LYP, the β- and α-ribonucleosides of the non-canonical TAP-C5 and CA, their complementary base pairing TAP-C5:CA, stacking energies and deglycosylation barriers in vacuum and implicit solvation using IEFPCM. Kaur et al. found from the deglycosylation profiles obtained for the most stable β- and α-anomers of both TAP ribonucleosides anomers at the B3LYP/6-31G(d, p) that the glycosidic bond in the TAP nucleosides (relative dissociation energy (Erel.) ≈ 350–447 kJ/mol) is stronger than the one in the canonical ribonucleosides (Erel. ≈ 222–258 kJ/mol) in both environments. Meanwhile, an opposite result was obtained for both CA ribonucleosides anomers (Erel. ≈ 155–193 kJ/mol {vacuum} and 207–267 kJ/mol {IFPCM}), leading the authors to propose that maybe TAP-C5 ribonucleosides were present in more hydrolytic environments, while CA ribonucleosides may have been in less hostile conditions.
Another study by Kaur et al. was published in 2017 for the BA-C5 in its keto form and MM ribonucleosides. When the authors analyzed the energetic barriers of dissociation of the glycosidic bond between the two non-canonical nucleobases and D-ribofuranose in the β- and α-configurations using the same DFT levels from the paper in 2019 they found that for BA-C5 (Erel. ≈ 317–451 kJ/mol at a glycosidic bond length of 3.0–4.1 Å) the cleavage of the glycosidic bond was less energetically favored than for canonical ribonucleosides. The glycosidic bond in MM ribonucleoside is predicted to be more resistant to dissociation when in the β-configuration and, overall, even when less stable than the one in BA-C5 ribonucleosides it can be stronger than the glycosidic bond in canonical ribonucleosides. This could make both non-canonical ribonucleosides suitable for hydrolytic prebiotic scenarios.
The present study complements the results obtained by Kaur et al. (2017, 2019). In our case we expand the analysis by considering the sugar puckering of the furanose form of D-ribose and also analyze the C- and N-glycosides of the TAP and BA bases. In this study instead of analyzing the stability of the glycosidic bond we estimated the thermodynamic feasibility for the formation of the given ribonucleosides and compared them with the formation of the canonical nucleosides, predicting that, overall, the formation of the β-anomer of the BA-C5 ribonucleosides is the most favored in both environments compared to the canonical nucleosides. For the case of the TAP-C5 and MM a similar trend is observed even when in some cases the energies are within the error of the calculations. In general, the classic synthesis of the CA ribonucleosides is estimated as the most unfavored, even more unfavored than the one for some canonical nucleosides. These results are in agreement with the studies by Kaur et al. (Kaur et al. 2017, 2019).
When analyzing the thermodynamic plausibility for the formation of canonical and non-canonical nucleosides with AEG it is observed that the condensation reaction of the acetylated derivatives of canonical and non-canonical nucleobases with AEG (Fig. 5) is thermodynamically favored in aqueous environment (but not in vacuum) in all cases. The most favored reactions are between AEG and A-acetylated (−19.4 kJ/mol), C-acetylated (−22.3 kJ/mol) and BA-N1-acetylated (−17.9 kJ/mol) nucleosides.
These observations suggest the possibility that maybe the TCs present in the first proto-NAs were different from ribose and that these were not necessarily sugar-based. A “ribonucleoprotein (RNP) world” instead of an “RNA world” has been proposed in the literature (Lehman 2015). This topic was widely discussed in a special issue of Life with 17 papers dedicated to discuss the origins and evolution of RNA and the “RNA world” hypothesis. Carter and coworkers (Carter 2015) discussed this hypothesis on the basis of a coexistence between both polynucleotides (as storage of information (Gatlin 1972; Volkenstein 1979, 2009; Hooft et al. 2024)) and polypeptides (as cofactors with catalytic activity) to complement the formation of RNA in the evolutionary process.
Also, in support of the RNP theory Smith et al. (Smith et al. 2014) addresses the fact that some proteins can store information (Gatlin 1972; Volkenstein 1979, 2009; Hooft et al. 2024) and catalyze the synthesis of other proteins.
Wächtershäuser (Wächtershäuser 2014) argued against the “Strong RNA World” hypothesis analyzing reactions that require metals as catalyzers and hypothesizes that until this type of chemistry was well defined in the prebiotic world none of the processes invoked by the “RNA world” theory could have happened.
Taking into consideration these observations, our predicted ∆G° of reaction for the synthesis of AEG nucleosides further support the idea that the building blocks of aegPNAs could have been precursors of today’s nucleosides if these molecules emerged in aqueous environment as a way to circumvent the “water problem.”
If this hypothesis can be proven experimentally, then an obvious next question emerges: how the building blocks of PNA transitioned to today’s nucleosides (tides)? One accepted theory is that their strands can participate in template-directed reactions since they both can create hybrid double stranded helixes with DNA and/or RNA (Frenkel-Pinter et al. 2020; Meggers and Zhang 2010; Zhang et al. 2005; Nielsen et al. 1994, 1991; Nielsen 1993; Egholm et al. 1992).
Another possibility could be that single AEG nucleosides(tides) could have assisted the glycosylation and phosphorylation of Ribf in today’s nucleotides. Supporting this idea, Nature has left us a clue hidden in our own genetic material: the “transfer-RNA (tRNA)”. tRNA can create polypeptides by transferring an amino acid bonded through an ester bond to the C3'-terminal of one of the tRNA strands to the nascent polypeptide chain in the peptidyl transferase center of ribosomal RNA (rRNA) (Bose et al. 2022; Massa et al. 2010; Simonović and Steitz 2009; Gindulyte et al. 2007).
In addition, a recent study by Suárez-Marina and coworkers (Suárez-Marina et al. 2019) has shown that canonical nucleosides and nucleotides can be synthesized through condensation reactions of their components in dehydrating conditions in the presence of glycine. This study showed that glycine can react with the canonical bases and direct the glycosylation to the correct site with the ribose-5’-monophosphate.
Hirakawa et al. (2022) were able to synthesize ribose-5’-phosphate in the presence of urea, borate (BO3−3) and phosphate ions in heating conditions at 80 ºC for 24 h. The product was obtained in a 22% yield after removing by acidic hydrolysis the excess of reactants, the borate and urea (contains two amine groups and a secondary carbonyl with certain similarity to amino acids). Hirakawa et al. cleverly combined the use of borate to overcome the “asphalt problem” (Neveu et al. 2013) derived from the formose reaction for the synthesis of ribose. The use of BO3−3 was initially proposed by Ricardo and coworkers (Neveu et al. 2013; Ricardo et al. 2004) as a way to synthesize and stabilize pentoses in the presence of borates.
What if it is possible to use AEG nucleoside as scaffold to further assist the condensation reactions between ribose, canonical bases, and phosphate, helped by possible π-π stacking interactions between the existing base in the AEG nucleoside and the base been glycosylated to Ribf?
Alternative Pathway for the Formation of Nucleotides from an AEG Nucleoside Template
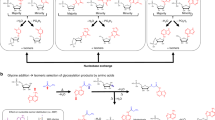
As a preliminary test of the previous hypothesis, we have modeled the synthesis of uridine-5’-monophosphate uridine (UMP) and AEG-Ac-N1-C dipeptides by following the reaction sequence represented in Fig. 6. The corresponding molecular structures are visualized in Fig. 7.
The proposed prebiotic pathway in aqueous medium includes the condensation reaction between N-(2-aminoethyl)-glycine (AEG) and Ac-N1-C to obtain the AEG-N2-Ac-N1-C derivative (product 1, {∆G°1 = −36.8 kJ/mol}). This reaction is followed by the condensation reaction between AEG-N2-Ac-N1-C and the OH in the C3′ position of β-Ribf in the 2T3 puckering conformation to obtain the β-Ribf-C3′-O-AEG-N2-Ac-N1-C (product 2, {∆G°2 = 5.9 kJ/mol}). Product 2 condensates then with uracil to obtain the uridine-C3′-O-AEG-N2-Ac-N1-C (product 3, {∆G°3 = −6.6 kJ/mol}). The phosphorylation of the OH-C5′ in product 3 with the di-hydrogen phosphate ion gives the β-UMP-C3′-O-AEG-N2-Ac-N1-C derivative (product 4, {∆G°4 = −8.7 kJ/mol}). Finally, product 4 condensates with a molecule of AEG-N2-Ac-N1-C to obtain a UMP, the AEG-N2-Ac-N1-C dipeptide (product 5, {∆G°5 = −36.4 kJ/mol}) and 4 water molecules are generated in total. While step 2 has a slightly positive ∆G°2 ≈ + 6 kJ/mol and step 3 is borderline spontaneous with ∆G°3 = −6.6 kJ/mol, these reactions are driven by the overall negative ∆G° of the reaction pathway whereby ∆G°(pathway) = \(\sum_{i=1}^{5}{\Delta G^\circ }_{i}=-82.6 \text{kJ}/\text{mol}\).
This reaction pathway that builds free UMP on an AEG-N2-Ac-N1-C template is, thus, overall favored by -82.6 kJ/mol in implicit solvation. Notice that this reaction pathway goes through a β-Ribf-C3′-O-AEG-N2-Ac-N1-C reactant which is similar to the C3′ end of the transfer RNA (tRNA) for the catalysis of the peptide bond formation in the ribosomal RNA (rRNA).
These results open the door to new prebiotic routes to overcome the “water problem” that could have used other potential prebiotic molecules as reagents to stabilize the building blocks of today’s nucleic acids. This pathway offers a way to obtain AEG dipeptides and free UMP nucleotides Fig. 7.
Discussion and Closing Remarks
The thermodynamic plausibility for the selection of the β- over the α-anomers of canonical and non-canonical nucleosides has been studied at the DFT-B3LYP/6–311++G(d, p) level of theory. A “classic” prebiotic synthesis for ribofuranose-containing nucleosides favors the formation of the β-anomer of canonical nucleosides containing adenine, guanine, and cytosine in both vacuum and implicit solvation and most non-canonical nucleosides containing N- and C4-glycosilated 2, 4, 6-triaminopyrimidine in aqueous environment and C4-glycosilated barbituric acid in both environments.
The formation of (2-aminoethyl)-glycine (AEG) containing nucleoside is favorable across the board in implicit solvation. This reinforces the hypothesis that an “RNP world” may have preceded RNA and DNA and constitutes theoretical evidence on how thermodynamics could justify the possible existence of a proto-RNA with a TC different from ribofuranose or 2’-deoxyribofuranose.
A novel hypothetical pathway was tested for the synthesis of β-uridine 2’-monophosphate (UMP) and a dipeptide containing the glycoside of AEG and the acetyl-N1 derivative of cytosine. This pathway includes the formation of some subproducts that are similar to existing aminoacyl derivatives of RNA present in today’s transfer RNA. The overall synthesis was predicted to be thermodynamically favored with a total ∆G° = − 82.6 kJ/mol. This represents a workaround by Nature to overcome the “water problem” for the glycosylation and phosphorylation of Ribf in the correct β configuration by building the free nucleotide on an amino acid derivative template. Hence, considering the possible coexistence of the components of nucleotides and ancestor amino acids in a prebiotic soup open numerous possibilities for the synthesis of both proteins and nucleic acids.
The results presented in this paper complement the theoretical studies previously published and constitute a step in the direction of understanding the drivers that may have led to the chemical nature of today’s nucleic acids. Nevertheless, we cannot disregard the relevance of other important factors. Some of these may include kinetic considerations (Jeilani and Nguyen 2020; Singh et al. 2014; Sheng et al. 2009). Singh and coworkers (Singh et al. 2014), for instance, describe a hydrogen-bond self-activated glycosylation mechanism for adenine and cytidine with D-ribose. After 8 days of heating at 60–70 °C β-adenosine was obtained in 15% of yield while β-cytidine was obtained in a 12%. The proposed reaction mechanism supports a hydrogen bond-assisted interaction of the base with the α-anomer of the sugar leading to a more stable transition state that goes through an oxonium intermediate that eventually interconverts into the β-nucleosides. This mechanism is suggested to happen on the surface of water.
Another factor that was not discussed here is pH which can have dramatic consequences since it determines the dominant protonation and tautomerization states of the nucleobases (Krishnamurthy 2012). Thibaudeau and coworkers (pp. 22–46 in Thibaudeau et al. (2005) and (Thibaudeau et al. 1994, 1997)) propose that the preference for one specific β-anomer over the α-counterpart for canonical nucleosides is related to its higher flexibility (lower differences between the ΔG° for the 2T3 / 3T2 pseudorotational equilibrium) in the β-configuration when transitioning from an acidic to an alkaline pH. Additionally, and related to the formation of non-canonical nucleosides containing TAP, BA, CA, and MM, Fialho (Fialho 2019) refers to the difference between the ionization states of the four non-canonical bases TAP, BA, CA, and MM. Due to this difference it can be inferred that only the pairs MM:BA and TAP:CA can be found at pH = 4–5 and pH = 6–7, respectively. Considering this if the first proto-NAs had these RUs and depending on the pH of the environment they may have only contained either the pair BA, MM or TAP, CA.
Ions and minerals would also have a significant effect on the prebiotic selection of building blocks of nucleic acid precursors. This is expected, at least, on the basis of the well-known ion-based catalysis especially for ions like Mg+ (Gull et al. 2017) or BO3−3 (Neveu et al. 2013; Fuller et al. 1972; Fuller 1972; Ricardo et al. 2004; Šponer et al. 2016; Holm 2012). Divalent cations, e.g., Mg2+, can assist in the folding of nucleic acids (Holm 2012). Possibly an early example of the use of Mg2+ as a catalyst in this context is the study of Fuller et al. (Fuller 1972) showing the marked increase in the synthetic yields of nucleosides in prebiotic conditions upon inclusion of Mg2+ in the reaction mixture.
Formamide (H2NCOH) has received special attention as a potential prebiotic solvent, since it can be synthesized from different routes, and it is present in interstellar space (Saladino et al. 2012). Nucleobases and some other molecules of life can be synthesized from H2NCOH (Saladino et al. 2012). Furthermore, the phosphorylation of different nucleosides in the presence of H2NCOH is possible with different yields going from 6%-59% (Gull 2014). There are still, however, issues that may cast doubt on formamide as the solvent that eventually led to modern day nucleic acid since this solvent inhibits base pair complementary (Hud et al. 2013). This latter point provided the impetus for the search of other prebiotic solvents. Polar organic solvents, e.g., ionic liquids and deep eutectic solvents have been considered (Mamajanov et al. 2010). For instance, choline chloride:urea mixture assists the folding of several nucleic acids imparting a higher stability to some than in water itself (Cafferty and Hud 2014). Other solvents may include urea (Salván et al. 2020) and deep eutectic solvents. In closing, the conclusions reached in this work are provisional despite being novel and suggestive and do call for more investigation.
Supporting Information
All the scripts written in bash and python to generate the nucleosides, post-process/analyze all the results and generate the diagrams and tables can be accessed through https://github.com/mattas-research-group/scripts_PhD_thesis_Lazaro/. For more information check the Supplementary Information.
References
Ayukawa S, Enomoto T, Kiga D (2019) RNA world. In: Astrobiology: from the origins of life to the search for extraterrestrial intelligence. Springer Nature, Singapore, pp 77–90. https://doi.org/10.1007/978-981-13-3639-3_6
Barone V, Cossi M, Tomasi J (1997) A new definition of cavities for the computation of solvation free energies by the polarizable continuum model. J Chem Phys 107:3210–3221
Becke AD (1988) Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A 38:3098–3100
Benner SA, Kim HJ, Carringa MA (2012) Asphalt, water, and the prebiotic synthesis of ribose, ribonucleosides and RNA. Acc Chem Res 45:2025–2034
Betts L, Josey JA, Veal JM, Jordan SR (1995) A nucleic acid triple helix formed by a peptide nucleic acid-DNA complex. Science 270:1838–1841
Bose T, Fridkin G, Davidovich C, Krupkin M, Dinger N, Falkovich AH, Peleg Y, Agmon I, Bashan A, Yonath A (2022) Origin of life: protoribosome forms peptide bonds and links RNA and protein dominated worlds. Nucleic Acids Res 28:1815–1828
Butlerow A (1861) Formation synthetique d’une substance sucrée. Comp Rend Acad Sci 63:145–147
Cafferty BJ, Fialho DM, Hud NV (2018) Searching for possible ancestors of RNA: The self-assembly hypothesis for the origin of proto-RNA. In: Menor-Salván C (ed) Prebiotic chemistry and chemical evolution of nucleic acids, nucleic acids and molecular biology. Springer, Philadelphia, Pennsylvania, pp 143–174
Cafferty BJ, Fialho DM, Khanam J, Krishnamurthy R, Hud NV (2016) Spontaneous formation and base pairing of plausible prebiotic nucleotides in water. Nat Commun 7:11328
Cafferty BJ, Gállego I, Chen MC, Farley KI, Eritja R, Hud NV (2013) Efficient self-assembly in water of long noncovalent polymers by nucleobase analogues. J Am Chem Soc 135:2447–2450
Cafferty BJ, Hud NV (2014) Abiotic synthesis of RNA in water: a common goal of prebiotic chemistry and bottom-up synthetic biology. Curr Opin Chem Biol 22:146–157
Cafferty BJ, Hud NV (2015) Was a pyrimidine-pyrimidine base pair the ancestor of watson-crick base pairs? Insights from a systematic approach to the origin of RNA. Isr J Chem 55:891–905
Carter CWJ (2015) What RNA world? Why a peptide/RNA partnership merits renewed experimental attention. Life 5:294–320
Castanedo LAM (2024) Computational biochemical study of the prebiotic selection of nucleic acids, PhD Thesis, Saint Mary’s University, Halifax
Castanedo LAM, Matta CF (2022a) On the prebiotic selection of nucleotide anomers: a computational study. Heliyon 8:e09657
Castanedo LAM, Matta CF (2022b) On the prebiotic selection of nucleotide anomers: computational data. Mendeley Data, version 2. https://doi.org/10.17632/khxvtshbs2.2
Cech TR (2012) The RNA worlds in context. Cold Spring Harbor Perspect Biol 4:a006742
Chen MC, Cafferty BJ, Mamajanov I, Gállego I, Khanam J, Krishnamurthy R, Hud NV (2014) Spontaneous prebiotic formation of a β-ribofuranoside that self assembles with a complementary heterocycle. J Am Chem Soc 136:5640–5646
Clercq ED (2015) C-nucleosides to be revisited. J Med Chem 59:2301–2311
Cossi M, Barone V, Cammi R, Tomasi J (1996) Ab initio study of solvated molecules: a new implementation of the polarizable continuum model. Chem Phys Lett 255:327–335
Dennington R, Keith TA, Millam JM (2009) GaussView, version 5. Semichem Inc, Shawnee Mission
Do Vieira AN, Do Kleinermanns K, MartinPreiner WFM (2020) The ambivalent role of water at the origins of life. FEBS Lett 594:2717–2733
Douglas K (2017) Nanoscience DNA: from prebiotic origins to emerging nanotechnology. CRC Press, Boca Ratón
Drew HR, Wing RM, Takano T, Broka C, Tanaka S, Itakura K, Dickerson RE (1981) Structure of a B-DNA dodecamer: conformation and dynamics. Proc Natl Acad Sci USA 78:2179–2183
Drew KN, Zajicek J, Bondo G, Bose B, Serianni AS (1998) 13C-labeled aldopentoses: detection and quantitation of cyclic and acyclic forms by heteronuclear 1D and 2D NMR spectroscopy. Carbohyd Res 307:199–209
Egholm M, Buchardt O, Nielsen PE, Berg R (1992) Peptide nucleic acids (PNA). oligonucleotide analogs with an achiral peptide backbone. J Am Chem Soc 114:1895–1897
Engelhart AE, Hud NV (2010) Primitive genetic polymers. Perspect Biol 2:a002196
Fialho DM (2019) Physical organic principles governing the spontaneous prebiotic emergence of proto-nucleic acids, PhD Thesis, Georgia Tech, Atlanta, pp 1–3
Fialho DM, Clarke KC, Moore MK, Schuster GB, Krishnamurthy R, Hud NV (2018) Glycosylation of a model proto-RNA nucleobase with non-ribose sugars: implications for the prebiotic synthesis of nucleosides. Org Biomol Chem 16:1263–1271
Fialho DM, Roche TP, Hud NV (2020) Prebiotic syntheses of noncanonical nucleosides and nucleotides. Chem Rev 120:4806–4830
Frenkel-Pinter M, Samanta M, Ashkenasy G, Leman LJ (2020) Prebiotic peptides: molecular hubs in the origin of life. Chem Rev 120:4707–4765
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Petersson GA, Nakatsuji H, Li X, Caricato M, Marenich AV, Bloino J, Janesko BG, Gomperts R, Mennucci B, Hratchian HP, Ortiz JV, Izmaylov AF, Sonnenberg JL, Williams-Young D, Ding F, Lipparini F, Egidi F, Goings J, Peng B, Petrone A, Henderson T, Ranasinghe D, Zakrzewski VG, Gao J, Rega N, Zheng G, Liang W, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Throssell K, Montgomery JAJ, Peralta JE, Ogliaro F, Bearpark MJ, Heyd JJ, Brothers EN, Kudin KN, Staroverov VN, Keith TA, Kobayashi R, Normand J, Raghavachari K, Rendell AP, Burant JC, Iyengar SS, Tomasi J, Cossi M, Millam JM, Klene M, Adamo C, Cammi R, Ochterski JW, Martin RL, Morokuma K, Farkas O, Foresman JB, Fox DJ (2019) Gaussian 16, revision C.01, Wallingford CT: Gaussian Inc.
Froeyen M, Lescrinier E, Kerremans L, Rosemeyer H, Seela F, Verbeure B, Lagoja I, Rozenski J, Aerschot AV, Busson R, Herdewijn P (2001) α-Homo-DNA and RNA form a parallel oriented Non-A, Non-B-Type double helical structure. Chem: A Eur J 7:5183–5194
Fuller WD (1972) Studies in prebiotic synthesis VI. Synthesis of purine nucleosides. J Mol Biol 67:25–33
Fuller WD, Sanchez RA, Orgel LE (1972) Studies in prebiotic synthesis. VII solid -state synthesis of purine nucleosides. J Mol Evol 1:249–257
Gatlin LL (1972) Information theory and the living system. Columbia University Press, New York
Gesteland RF, Cech TR, Atkins JF (eds) (1999) The RNA World. Cold Spring Harbor Laboratory Press, New York
Gindulyte A, Bashan A, Agmon I, Massa L, Yonath A, Karle J (2007) The transition state for formation of the peptide bond in the ribosome. Proc Natl Acad Sci USA 108:13327–13332
Gonzalez MA, Jimenez Requejo JL, Palacios Albarran JC, Gabis Perez JA (1986) Facile preparation of C-glycosylbarbiturates and C-glycosylbarbituric acids. Carbohydr Res 158:53–66
Guesnet J-L, Vovelle F, Thuong NT, Lancelot G (1990) 2D NMR studies and 3D structure of the parallel-stranded duplex oligonucleotide Acrm5-alpha-d(TCTAAACTC)-beta-d(AGATTTGAG) via complete relaxation matrix analysis of the NOE effects and molecular mechanics calculations. Biochemistry 29:4982–4991
Gull M (2014) Prebiotic phosphorylation reactions on the early Earth. Challenges 5:193–212
Gull M, Cafferty BJ, Hud NV, Pasek MA (2017) Silicate-promoted phosphorylation of glycerol in non-aqueous solvents: A prebiotically plausible route to organophosphates. Life 7:7030029
Hehre WJ, Radom L, Pople JA, Schleyer P (1986) Ab initio molecular orbital theory. John Wiley and Sons Ltd, New York
Hertwig RH, Koch W (1997) On the parameterization of the local correlation functional. What is Becke-3-LYP? Chem Phys Lett 268:345–351
Hirakawa Y, Kakegawa T, Furukawa Y (2022) Borate-guided ribose phosphorylation for prebiotic nucleotide synthesis. Sci Rep 12:11828
Hooft G, Phillips WD, Zeilinger A, Allen R, Baggott J, Bouchet FR, Cantanhede SMG, Castanedo LAM, Cetto AM, Coley AA, Dalton BJD, Fahimi P, Franks S, Frano A, Fry ES, Goldfarb S, Langanke K, Matta CF, Nanopoulos D, Orzel C, Patrick S, Sanghai VAA, Schuller IK, Shpyrko O, Lidström S (2024) The sounds of science—a symphony for many instruments and voices: part II. Phys Script 99:052501
Holm NG (2012) The significance of Mg in prebiotic geochemistry. Geobiology 10:269–279
Hud NV, Anet FAL (2000) Intercalation-mediated synthesis and replication: a new approach to the origin of life. J Theor Biol 205:543–562
Hud NV, Cafferty BJ, Krishnamurthy R, Williams LD (2013) The origin of RNA and “my grandfather’s axe.” Chem Biol 20:466–474
HyperChem release 7.0 for Windows: Reference Manual (2002). Hypercube, Inc., Waterloo
Jeilani YA, Nguyen MT (2020) Autocatalysis in formose reaction and formation of RNA nucleosides. J Phys Chem B 124:11324–11336
Jissy AK, Ramana JHV, Datta A (2011) π-Stacking interactions between G-quartets and circulenes: a computational study. J Chem Sci 123:891–900
Johnson ER, Mackie ID, DiLabio GA (2009) Dispersion interactions in density-functional theory. J Phys Org Chem 22:1127–1135
Josa D, Rodríguez-Otero J, Cabaleiro-Lago EM, Rellán-Piñeiro M (2013) Analysis of the performance of DFT-D, M05–2X and M06–2X functionals for studying π π interactions. Chem Phys Lett 557:170–175
Joyce GF, Orgel LE (1999) Prospects for understanding the origin of the RNA world. In: The RNA world, 2nd edn. Cold Spring Harbor Press, New York, pp 49–78
Kaur S, Sharma P, Wetmore SD (2017) Structural and electronic properties of barbituric acid and melamine-containing ribonucleosides as plausible components of prebiotic RNA: implications for prebiotic self-assembly. Phys Chem Chem Phys (PCCP) 19:30762–30771
Kaur S, Sharma P, Wetmore SD (2019) Can cyanuric acid and 2, 4, 6-triaminopyrimidine containg ribonucleosides be components of prebiotic RNA? Insights from QM calculations and MD simulations. ChemBioChem 20:1415–1415
Kim H-J, Furukawa Y, Kakegawa T, Bita A, Scorei R, Benner SA (2016) Evaporite borate-containing mineral ensembles make phosphate available and regiospecifically phosphorylate ribonucleosides: borate as a multifaceted problem solver in prebiotic chemistry. Angew Chem Int Ed 55:15816–15820
Klamt A (2011) The COSMO and COSMO-RS solvation models. Adv Rev 1:699–709
Klamt A, Schüümann G (1993) COSMO: a new approach to dielectric screening in solvents with explicit expressions for the screening energy and its gradient. J Chem Soc Perkin Trans 2(2):799–805
Kolb VM, Dworkin JP, Miller SL (1994) Alternative bases in the RNA world: the prebiotic synthesis of urazole and its ribosides. J Mol Evol 38:549–557
Krishnamurthy R (2012) Role of pKa of nucleobases in the origins of chemical evolution. Acc Chem Res 45:2035–2044
Lancelot G, Guesnet J-L, Roig V, Thuong NT (1987) 2D-NMR studies of the unnatural duplex α-d(TCTAAAC)-β-d(AGATTTG). Nucleic Acids Res 15:7531–7547
Lancelot G, Guesnet J-L, Vovelle F (1989) Solution structure of the parallel-stranded duplex oligonucleotide α-d( TCTAAAC)-β-d(AGATTTG) via complete relaxation matrix analysis of the NOE effects and molecular mechanics calculations. Biochemistry 28:7871–7878
Lee EJ, Lim HK, Cho YS, Hah SS (2013) Peptide nucleic acids are an additional class of aptamers. RSC Adv 3:5828–5831
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation energy formula into a functional of the electron density. Phys Rev B 37:785–789
Lehman N (2015) The RNA World: 4,000,000,050 years old. Life 5:1583–1586
Li C, Cafferty BJ, Karunakaran SC, Schuster GB, Hud NV (2016) Formation of supramolecular assemblies and liquid crystals by purine nucleobases and cyanuric acid in water: implications for the possible origins of RNA. Physical Chemistry Chemical Physics (PCCP) 18:20091–20096
Lubczak J, Mendyk E (2008) Stable enol form of barbituric acid. Heterocycl Commun 14:149–154
Mamajanov I, Engelhart AE, Bean HD, Hud NV (2010) DNA and RNA in anhydrous media: duplex, triplex, and G-quadruplex secondary structures in a deep eutectic solvent. Angew Chem Int Ed 49:6310–6314
Massa L, Matta CF, Yonath A, Karle J (2010) Quantum transition state for peptide bond formation in the ribosome. In: Matta CF (ed) Quantum biochemistry: electronic structure and biological activity, vol 2. Wiley, Weinheim, pp 501–515
Mateo-Martí E, Pradie C-M (2010) A novel type of nucleic acid-based biosensors: the use of PNA probes, associated with surface science and electrochemical detection techniques. Intell Biosens. https://doi.org/10.5772/7160
McCollom TM (2013) Miller-Urey and beyond: what have we learned about prebiotic organic. Annu Rev Earth Planet Sci 41:207–229
Meggers E, Zhang L (2010) Synthesis and properties of the simplified nucleic acid glycol nucleic acid. Acc Chem Res 43:1092–1102
Menchise V, De Simone G, Tedeschi T, Corradini R, Sforza S, Marchelli R, Capasso D, Saviano M, Pedone C (2003) Insights into peptide nucleic acid (PNA) structural features: the crystal structure of a D-lysine-based chiral PNA-DNA duplex. Proc Natl Acad Sci USA 100:12021–12026
Miertuš S, Scrocco E, Tomasi J (1981) Electrostatic interaction of a solute with a continuum. a direct utilization of ab initio molecular potentials for the prevision of solvent effects. Chem Phys 55:117–129
Miertuš S, Tomasi J (1982) Approximate evaluations of the electrostatic free energy and internal energy changes in solution processes. Chem Phys 65:239–245
Miller SL (1953) Production of amino acids under possible primitive earth conditions. Science 117:528–529
Miller SL (1955) Production of some organic compounds under possible primitive earth. J Am Chem Soc 77:2351–2361
Montero LA (2019) GRANADA programme: distribución aleatoria de moléculas alrededor de un sistema poliatómico central (random distribution of molecules around a central polyatomic system). University of Havana, Faculty of Chemistry
Montero LA, Esteva AM, Molina J, Zapardiel A, Hernández L, Márquez H, Acosta A (1998) A theoretical approach to analytical properties of 2,4-diamino-5-phenylthiazole in water solution. tautomerism and dependence on pH. J Am Chem Soc 120:12023–12033
Montero LA, Molina J, Fabian J (2000) Water clusters for calculations of association energy. Int J Quantum Chem 79:8–16
Nelson KE, Levy M, Miller SL (2000) Peptide nucleic acids rather than RNA may have been the first genetic molecule. Proc Natl Acad Sci USA 97:3868–3871
Neveu M, Kim HJ, Benner SA (2013) The ‘“strong”’ RNA world hypothesis: fifty years old. Astrobiology 13:391–403
Ni G, Yuqi D, Tang F, Liu J, Zhao H, Chen Q (2019) Review of α-nucleosides: from discovery, synthesis to properties and potential applications. RSC Adv 9:14302–14320
Nielsen PE (1993) Peptide nucleic acid (PNA): a model structure for the primordial genetic material? Orig Life Evol Biosph 23:323–327
Nielsen PE (1996) Peptide nucleic acid (PNA). Implications for the origin of the genetic material and the homochirality of life. AIP Conf Proc 379:55–61
Nielsen PE (2007) Peptide nucleic acids and the origin of life. Chem Biodivers 4:1996–2002
Nielsen PE, Buchardt O, Egholm M, Norden B (1994) DNA-like double helix formed by peptide nucleic acid. Nature 368:561–563
Nielsen PE, Egholm M, Berg RH, Buchardt O (1991) Sequence-selective recognition of DNA by strand displacement with a thymine-substituted polyamide. Science 254:1497–1500
Orgel LE (2004) Prebiotic chemistry and the origin of the RNA world. Crit Rev Biochem Mol Biol 39:99–123
Orio M, Pantazis DA, Neese F (2009) Density functional theory. Photosynth Res 102:443–453
Paoletti J, Bazile D, Morvan F, Imbach J-L, Paoletti C (1989) α-DNA VIII:thermodynamic parameters of complexes formed between the oligo-alphadeoxynucleotides: α-d(GGAAGG) and α-d(CCTTCC) and their complementary oligo-betadeoxynucleotides: β-d(CCTTCC) and β-d(GGAAGG) are different. Nucleic Acid Res 17:2693–2704
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Pérez-Fernández C, Vega J, Rayo-Pizarro P, Mateo-Marti E, Ruiz-Bermejo M (2022) Prebiotic synthesis of noncanonical nucleobases under plausible alkaline hydrothermal conditions. Sci Rep 12:15140
Rao L, Ke H, Fu G, Xu X, Yan Y (2009) Performance of several density functional theory methods on describing hydrogen-bond interactions. J Chem Theory Comput 5:86–96
Ricardo A, Carrigan MA, Olcott AN, Benner A (2004) Borate minerals stabilize ribose. Science 303:196
Saladino R, Crestini C, Pino S, Constanzo G, Di Mauro E (2012) Formamide and the origin of life. Phys Life Rev 9:84–104
Salván CM, Bouza M, Fialho DM, Burcar BT, Fernández FM, Hud NV (2020) Prebiotic origin of pre-RNA building blocks in a urea “warm little pond” scenario. ChemBioChem 21:3504–3510
Sefah K, Yang Z, Bradley KM, Hoshika S, Jiménez E, Zhang L, Zhu G, Shanker S, Yu F, Turek D, Tan W, Benner SA (2014) In vitro selection with artificial expanded genetic information systems. Proc Natl Acad Sci USA 111:1449–1454
Sheng Y, Bean HD, Mamajanov I, Hud NV (2009) Comprehensive investigation of the energetics of pyrimidine nucleoside formation in a model prebiotic reaction. J Am Chem Soc 131:16088–16095
Simonović M, Steitz TA (2009) A structural view on the mechanism of the ribosome-catalyzed peptide bond formation. Biochem Biophys Acta 1789:612–623
Singh P, Singh A, Kaur J, Holzer W (2014) H-bond activated glycosylation of nucleobases: implications for prebiotic nucleoside synthesis. RSC Adv 4:3158–3161
Smith JE, Mowles AK, Mehta AK, Lynn DG (2014) Looked at life from both sides now. Life 4:887–902
Šponer JE, Šponer J, Fuentes-Cabrera M (2011) Prebiotic routes to nucleosides: a quantum chemical Insight into the energetics of the multistep reaction pathways. Chem A Eur J 17:847–854. https://doi.org/10.1002/chem.201002057
Šponer JE, Szabla R, Góra RW, Saitta AM, Pietrucci F, Saija F, Di Mauro E, Saladino R, Ferus M, Civišh S, Šponer J (2016) Prebiotic synthesis of nucleic acids and their building blocks at the atomic level—merging models and mechanisms from advanced computations and experiments. Phys Chem Chem Phys 18:20047–20066
Stewart JJP (2023) "MOPAC," Stewart Computational Chemistry, http://openmopac.net/.
Strecker A (1854) Ueber einen neuen aus Aldehyd—Ammoniak und Blausäure entstehenden Körper. Annalen der Chemie und Pharmacie 91:349–351
Sutherland J (2010) Ribonucleotides. Cold Spring Harbor Perspect Biol 2:a005439
Suárez-Marina I, Abul-Haija YM, Turk-MacLeod R, Gromski PS, Cooper GJT, Olivé AO, Colón-Santos S, Cronin L (2019) Integrated synthesis of nucleotide and nucleosides influenced by amino acids. Commun Chem 2:28
Thibaudeau C, Acharya P, Chattopadhyaya J (2005) Stereoelectronic Effects in Nucleosides and Nucleotides and their Structural Implications. Uppsala University Press, Uppsala
Thibaudeau C, Foldesi A, Chattopadhyaya J (1997) The first experimental evidence for a larger medium-dependent flexibility of natural β-D-nucleosides compared to the α-D-nucleosides. Tetrahedron 53:14043–14072
Thibaudeau C, Plavec J, Chattopadhyaya J (1994) Quantitation of the anomeric effect in adenosine and guanosine by comparison of the thermodynamics of the pseudorotational equilibrium of the pentofuranose moiety in N- and C-nucleosides. J Am Chem Soc 116:8033–8031
Tomasi J (2011) Selected features of the polarizable continuum model for the representation of solvation. Wires Computational Mol Sci 1:855–867
Tomasi J, Cappelli C, Mennucci B, Cammi R (2010) From molecular electrostatic potentials to solvation models and ending with biomolecular photophysical processes. In: Matta CF (ed) Quantum biochemistry: electronic structure and biological activity, vol 1. Wiley, Weinheim, pp 131–170. https://doi.org/10.1002/9783527629213.ch4
Trinks P (1987) Zur chemie der aminopyrimidine, PhD Thesis, ETH Zürich, Zürich
Volkenstein MV (1979) Mutations and the value of information. J Theor Biol 80:155–169
Volkenstein MV (2009) Entropy and information. Birkhäser Verlag AG, Basel
Wellinton KW, Benner SA (2006) A review: synthesis of aryl C-glycosides via the Heck coupling reaction. Nucleosides Nucleotides Nucleic Acids 25:1309–1333
Wu J, Meng Q, Ren H, Wang H, Wu J, Wang Q (2017) Recent advances in peptide nucleic acid for cancer bionanotechnology. Acta Pharmacol Sin 38:798–805
Wächtershäuser G (2014) The place of RNA in the origin and early evolution of the genetic machinery. Life 4:1050–1091
Zhang L, Peritz A, Meggers E (2005) A simple glycol nucleic acid. J Am Chem Soc 127:4174–4175
Zhao Y, Truhlar DG (2007) Density functionals for noncovalent interaction energies of biological importance. J Chem Theory Comput 3:289–300
Zhao Y, Truhlar DG (2008a) Density functionals with broad applicability in chemistry. Acc Chem Res 41:157–167
Zhao Y, Truhlar DG (2008b) The M06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four M06-class functionals and 12 other function. Theoret Chem Acc 120:215–241
Acknowledgements
The authors are grateful to The Governor General of Canada, Her Excellency the Right Honourable Mary Simon, for The Governor General Gold Medal awarded to L.A.M.C. for his PhD Thesis from Saint Mary’s University [1] from which some of the results covered in this paper derive. The authors thank Prof. Stacey D. Wetmore, Prof. Cory Pye, Prof. Genlou Sun, Prof. Fernando Cortéz Guzmán, and Prof. Lou Massa for suggestions and useful discussions. The Natural Sciences and Engineering Research Council of Canada (NSERC), Canada Foundation for Innovation (CFI), Research Nova Scotia (RNS, formerly the Nova Scotia Research Innovation Trust (NSRIT)), the Digital Research Alliance of Canada (formerly Compute Canada), Saint Mary’s University, and Mount Saint Vincent University are acknowledged for supporting this research. L.A.M.C. has also received funding for this work in the form of a Nova Scotia Graduate Scholarship (NSGS), a Scotia Scholars Award, and an Abe Leventhal Student Research Award from the Alzheimer Society of Nova Scotia. The authors are grateful to the Chemical Computing Group (CCG) for its support to L.A.M.C.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Yitzhak Tor.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Castanedo, L.A.M., Matta, C.F. Prebiotic N-(2-Aminoethyl)-Glycine (AEG)-Assisted Synthesis of Proto-RNA?. J Mol Evol 92, 449–466 (2024). https://doi.org/10.1007/s00239-024-10185-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-024-10185-w