Abstract
Snakes in the family Elapidae largely produce venoms rich in three-finger toxins (3FTx) that bind to the \(\upalpha _1\) subunit of nicotinic acetylcholine receptors (nAChRs), impeding ion channel activity. These neurotoxins immobilize the prey by disrupting muscle contraction. Coral snakes of the genus Micrurus are specialist predators who produce many 3FTx, making them an interesting system for examining the coevolution of these toxins and their targets in prey animals. We used a bio-layer interferometry technique to measure the binding interaction between 15 Micrurus venoms and 12 taxon-specific mimotopes designed to resemble the orthosteric binding region of the muscular nAChR subunit. We found that Micrurus venoms vary greatly in their potency on this assay and that this variation follows phylogenetic patterns rather than previously reported patterns of venom composition. The long-tailed Micrurus tend to have greater binding to nAChR orthosteric sites than their short-tailed relatives and we conclude this is the likely ancestral state. The repeated loss of this activity may be due to the evolution of 3FTx that bind to other regions of the nAChR. We also observed variations in the potency of the venoms depending on the taxon of the target mimotope. Rather than a pattern of prey-specificity, we found that mimotopes modeled after snake nAChRs are less susceptible to Micrurus venoms and that this resistance is partly due to a characteristic tryptophan\(\rightarrow\)serine mutation within the orthosteric site in all snake mimotopes. This resistance may be part of a Red Queen arms race between coral snakes and their prey.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Micrurus is part of the group of elapids known as the ‘true coral snakes’ along with Micruroides and Sinomicrurus. Micrurus is the most speciose genus of elapids and can be found from Argentina to the southern United States of America (Campbell and Lamar 2004). Genetic and morphological evidence clearly distinguishes between two main clades within Micrurus: short-tailed and long-tailed (Campbell and Lamar 2004; Jowers et al. 2019, 2023). In general, the short-tailed species are found in South America and possess a triadal color pattern (e.g., red-black-white-black-white-black-red) and tricolored tails, but include some bicolored species with bicolored tails as well. The long-tailed species are somewhat more variable; these mostly Central and North American species include bicolored, monadal (e.g., red-yellow-black-yellow-red), and triadal patterns with mostly bicolored tails (Campbell and Lamar 2004; Jowers et al. 2019, 2023). These clades are often referred to in the literature as ‘triadal’ and ‘monadal’ (Jowers et al. 2019, 2023; Gómez et al. 2021), but exceptions to these rules of color patterns abound. For the purposes of this paper, we will use the ‘short-tailed’ and ‘long-tailed.’ The venom composition of the two groups also differs somewhat in that the venoms from short-tailed species are usually (though not exclusively) dominated by three-finger toxins (3FTx). In contrast, long-tailed species produce venoms that range from being almost entirely composed of 3FTx to mostly phospholipase A\(_2\)s (PLA\(_2\)s) (Lomonte et al. 2016; Sanz et al. 2019).
One of the most widespread toxic activities of 3FTx is antagonistic binding to the orthosteric site (the acetylcholine binding region) of nicotinic acetylcholine receptor (nAChR) \(\upalpha _1\) subunits (two of which along with a \(\upbeta\), \(\updelta\), and \(\upepsilon\), form the heteropentameric muscular nAChR subtype (Tae and Adams 2023)), which are located at the neuromuscular junction (Galzi et al. 1991; Nirthanan and Gwee 2004). This disrupts muscle contraction which can cause flaccid paralysis; this mode of action is often referred to as \(\upalpha\)-neurotoxicity (Barber et al. 2013). The plesiotypic form of 3FTx—which can be found in venoms from many caenophidian snakes at varying abundances—contain 10 cysteine residues, which form 5 disulfide bonds to stabilize the structure of the toxin (Utkin et al. 2015). Ancestors of the family Elapidae evolved a derived form with only 8 cysteines; these ‘short-chain 3FTx’ lost the second and third cysteines—which together form a disulfide bridge in Loop I of the plesiotypic toxins—and are much more potent as a result (Fry et al. 2003). A further derived form known as ‘long-chain 3FTx’—also found only in elapid venoms—has evolved two new cysteines which form a disulfide bridge and stabilize Loop II (Utkin et al. 2015; Nirthanan and Gwee 2004). Although these long-chain toxins also contain 10 cysteines, the placement of cysteine residues is distinct from the plesiotypic form and protein sequence phylogenetic analyses confirm that they are derived from short-chain toxins (Fry et al. 2003; Utkin et al. 2015; Koludarov et al. 2023). Long-chain and short-chain neurotoxins are known to compete with each other when binding to the nAChR \(\upalpha _1\) subunit, but they exhibit different specificities and kinetics (short-chain toxins bind to and dissociate from the nAChR much quicker) indicating slightly different biochemical mechanisms of action and target sites (Ackermann and Taylor 1997; Silva et al. 2018). Recent studies using cryogenic electron microscopy to investigate the binding of 3FTx to nAChRs confirm that plesiotypic, short-chain, and long-chain toxins all bind to the target receptors in noticeably different fashions (Rahman et al. 2020; Nys et al. 2022; Shenkarev et al. 2022).
Bio-layer interferometry (BLI) can be used as a method to assess the binding of \(\upalpha\)-neurotoxic snake venoms to mimotopes (small synthetic molecules which mimic an epitope) that are designed to resemble the nAChR orthosteric site from a wide range of taxa (Kamat and Rafique 2017; Zdenek et al. 2019; Harris et al. 2020a, b, c). Past studies have shown that similar mimotopes interact strongly with \(\upalpha\)-bungarotoxin (the best-studied long-chain neurotoxin), but this has not been explicitly demonstrated for short-chain neurotoxins (Bracci et al. 2001, 2002; Kasher et al. 2001; Katchalski-Katzir et al. 2003). Other studies investigated mimotopes of the orthosteric site as potential treatments to inhibit the \(\upalpha\)-neurotoxic activity of snake venoms by acting as decoys that keep the toxins from binding to the genuine receptors and found that the results were quite consistent for long-chain toxins, but unreliable for short-chain toxins (Albulescu et al. 2019; Kudryavtsev et al. 2020; Lynagh et al. 2020). In this study, we use BLI to investigate the patterns of \(\upalpha\)-neurotoxicity between species of Micrurus and across different targets.
The long-chain neurotoxins seem to have originally evolved shortly after the common ancestor of Micrurus and the more derived elapids diverged (Dashevsky and Fry 2018; Koludarov et al. 2023); 3FTx with the additional cysteines characteristic of long-chain neurotoxins have been published from all lineages of elapids besides Calliophis and the true coral snakes despite extensive transcriptomic investigations of these basal genera (Corrêa-Netto et al. 2011; Margres et al. 2013; Guerrero-Garzón et al. 2018; Tan et al. 2019; Bénard-Valle et al. 2020; Dashevsky et al. 2021; The UniProt Consortium 2023). Therefore, results obtained by studying Micrurus venoms more clearly demonstrate the effects of short-chain neurotoxins without the confounding presence of long-chain toxins. This makes these venoms an ideal test of whether and how short-chain neurotoxins bind to mimotopes of the nAChR orthosteric site. No other research has examined the binding of short-chain neurotoxins to \(\upalpha _1\) subunit mimotopes specifically, and previous studies employing this method have only demonstrated significant binding from venoms that contain both short- and long-chain neurotoxins (Zdenek et al. 2019; Harris et al. 2020a, c).
Due to the high sensitivity of the BLI method, it can distinguish differences in how well a venom can bind to the different mimotopes, which can help assess the prey-specificity (greater toxicity against preferred prey species than other taxa) of that venom (Zdenek et al. 2019; Harris et al. 2020b, c). Prey specificity is a widespread phenomenon in snake venoms (da Silva and Aird 2001; Mackessy et al. 2006; Pawlak et al. 2006, 2008; Barlow et al. 2009; Heyborne and Mackessy 2013; Modahl et al. 2018; Sousa et al. 2018) and has been documented in Micrurus venoms using in vitro tests of whole venom (da Silva and Aird 2001). By using this BLI assay our results will clarify if this specificity is achieved through variation in the ability of these venoms to bind to the orthosteric site of the nAChR in their prey.
However, if coevolution can drive the evolution of prey-selective venom, then it must also be considered that there is an opposing evolutionary pressure for the prey of Micrurus to become less susceptible to the toxins of their predators. Following the life-dinner principle, the selection pressure on prey populations may be even greater because the consequence of a predation attempt for them is death rather than a missed meal (Dawkins and Krebs 1979). Resistance and reduced susceptibility to snake \(\upalpha\)-neurotoxins has been reported from a number of taxa which are either predators or prey of venomous snakes (Holding et al. 2016; Khan et al. 2020). Mongooses, pigs, and hedgehogs—which are known to feed on venomous snakes—are less susceptible to elapid \(\upalpha\)-neurotoxins due to convergent amino acid substitutions in the orthosteric binding site of the muscular nAChR subunit and similar mutations can be found in elapids themselves (Barchan et al. 1992, 1995; Takacs et al. 2001, 2004; Drabeck et al. 2015). Primates have also been shown to possess nAChR mutations which reduce their susceptibility to \(\upalpha\)-neurotoxins (Harris et al. 2021). Whether the immunity in elapids evolved as auto-resistance, in response to intraguild predation, or both, is unclear. Some lizards and other snakes also show reduced susceptibility to the effects of both long-chain and short-chain neurotoxins (Burden et al. 2006). Further, a study addressing the effects of M. nigrocinctus venom on different taxa found that cows are less susceptible than horses (Bolaños et al. 1975). Another demonstrated a marked difference between two genera of dipsadine snakes that are known prey of M. nigrocinctus: Geophis spp. were susceptible while Ninia spp. were more resistant (Urdaneta et al. 2004). The latter study incubated M. nigrocinctus venom with N. maculata blood serum which had a protective effect when injected into mice; this suggests that the mechanism of resistance is that the serum of N. maculata is able to directly inhibit the toxins rather than the presence of mutations to their muscular nAChR subunit allowing them to resist the toxic effects (Urdaneta et al. 2004). A more recent experiment using the venom of M. tener found that the snake species Conopsis lineata could survive higher venom doses than mice, but that both groups were paralyzed at the same dosage (Bénard-Valle et al. 2014).
Based on the evidence that some Micrurus venoms exhibit prey-selectivity (da Silva and Aird 2001) and some natural prey might have a reduced susceptibility to their venom (Urdaneta et al. 2004), we wanted to test the effects of both short-tailed (8 venoms) and long-tailed (7 venoms) Micrurus venoms using BLI. We analyzed these venoms using 12 different mimotopes designed to resemble the muscular nAChR orthosteric site from a range of taxa, including some that are natural prey of Micrurus and others that are not. We also wanted to assess if the serine (S) at position 187 of the muscular nAChR (the orthosteric site runs from 187–200 in the overall sequence of the receptor subunit) found in most snake lineages conferred any resistance/ reduced susceptibility since this is a significant biochemical change from the ancestral tryptophan (W) (Khan et al. 2020). To further test this, we designed a mutant based on the blind snake mimotope, substituting the natural 187 S for the ancestral W. Taken together, these data provide insights into the evolution of venom function within Micrurus and their coevolution with prey taxa.
Results and Discussion
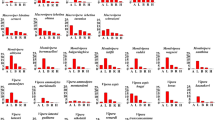
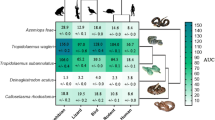
The binding of Micrurus toxins to muscular nAChR mimotopes varies greatly depending on the individual venom as well as the target: A Example results of Micrurus corallinus A venom binding to Dipsadine mimotope. Shaded regions indicate the area under the curve for each replicate that is then averaged. B Cells show the area under the curve (Mean ± Standard Deviation, \(N = 3\)) for the association step of the binding of each venom to each mimotope. Phylogeny on the left displays relationships between the venoms tested (short-tailed and long-tailed clades indicated by arrows, 3FTx-heavy venoms in purple and PLA2-heavy venoms in teal), while the one on top displays the relationship between organisms on which mimotopes were modeled (clade of mimotopes based on snake sequences indicated with arrow). Crotalus horridus venom was used as a negative control as it contains many related protein families, but none which target the nAChR (Rokyta et al. 2013, 2015). Phylogenetic topology for Micrurus species was primarily adapted from the results of two recent phylogenies (Jowers et al. 2023; Reyes-Velasco et al. 2020) and checked for consistency with previous findings (Slowinski 1995; Gutberlet Jr and Harvey 2004; Pyron et al. 2013; Figueroa et al. 2016; Lee et al. 2016; Lomonte et al. 2016; Zheng and Wiens 2016; Zaher et al. 2019; Jowers et al. 2019; Gómez et al. 2021)
Consistent with previous studies on neurotoxic snake venoms, the BLI method was able to detect orthosteric binding by Micrurus venoms, and interesting patterns emerged both in terms of the differences within Micrurus and between the targets (Fig. 1). Our results clearly show that the strength of binding measured by our assay depended on the species of origin for both the venom and mimotope. While short-tailed Micrurus venoms elicited very weak or no binding at all compared to the control of Crotalus horridus (a species chosen for its lack of nAChR-binding neurotoxins (Rokyta et al. 2013, 2015; Harris et al. 2020b), the venoms of many long-tailed species showed considerable binding to the targets (Fig. 1). Since Micrurus venoms do not contain long-chain neurotoxins, this confirms that the assay can be used to analyze the activity of short-chain neurotoxins that bind to the orthosteric site of the nAChR. A previous study that investigated the interaction of short-chain neurotoxins and nAChR mimotopes had reported very little binding, but those mimotopes were designed to resemble the orthosteric binding region of the \(\upalpha _7\) subunit (Albulescu et al. 2019), which is not a physiologically relevant target since it is located within the central nervous system, rather than at the neuromuscular junction (Gotti and Clementi 2004). 3FTx circulating in the bloodstream or lymphatic system are too large to reach the central nervous system but can readily affect the neuromuscular junction, which is why \(\upalpha\)-neurotoxins have evolved to immobilize prey by inhibiting this more accessible physiological target. Short-chain neurotoxins are known to bind almost exclusively to the \(\upalpha _1\) rather than the \(\upalpha _7\) subunit (Ackermann and Taylor 1997; Servent et al. 1997). Because the mimotopes in this study were designed to resemble the \(\upalpha _1\) subunit, this easily explains why our assay could measure the effects of these toxins where previous research had not. While this might be taken as evidence that decoy peptides that resemble the \(\upalpha _1\) subunit could have even greater potential as a therapeutic tool to help mitigate the neurotoxic symptoms of bites from coral snakes than the \(\upalpha _7\)-derived ones (Albulescu et al. 2019), other patterns in our results cut against this conclusion.
One of the striking trends in our data was the variability within and between species of Micrurus. The venom with the largest effect on average (and for every individual mimotope) was a sample of M. corallinus, while the other individual of this species that we tested had only the 6th highest average effect. This discrepancy between the two samples could result from regional variation within this species or could be an indicator of potential cryptic speciation in this lineage. While there is no direct evidence for this latter hypothesis, species complexes are so widespread in this incredibly speciose genus (Roze 1996; Campbell and Lamar 2004; Terribile et al. 2018; Nascimento et al. 2019) and new species which bear strong external similarity to their congeners are described with enough frequency (Di-Bernardo et al. 2007; Pires et al. 2014; da Silva et al. 2015; Bernarde et al. 2018) that the possibility bears mentioning. There is also a phylogenetic trend: the long-tailed Micrurus elicited a much greater response than their short-tailed relatives. Several short-tailed Micrurus venoms did not show appreciably stronger binding than our negative control (Crotalus horridus) venom. This is especially interesting in light of previous findings that the venoms produced by some long-tailed species (such as M. diastema, M. browni, M. tener, and M. fulvius) are dominated by PLA2s—which are not known to bind to nAChRs—rather than 3FTx (Lomonte et al. 2016). Due to the variability of each venom’s binding across the different mimotopes, we could not simply compare the means for each venom. Instead, we created a factor denoting whether the predominant toxin family in a given venoms was 3FTxs or PLA2s and used two-way unbalanced ANOVA with type III sum of squares to simultaneously test the effects of this factor, the mimotope, and their interaction. This analysis did not find a significant effect for the dominant toxin family (\(\textit{p} =0.12\)) or the interaction between toxin family and sensor (\(\textit{p} =0.99\)), but the sensor alone did have a significant effect (\(\textit{p} =0.004\)). We then repeated this analysis using a factor capturing the short- and long-tailed clades rather than the dominant toxin family. We found that the average difference between short-tailed and long-tailed species was greater than twofold for each mimotope, that this was a statistically significant trend (\(\textit{p} <0.001\)), that the variation across mimotopes was also significant (\(\textit{p} =0.02\)), but the interaction between the two was not (\(\textit{p} =0.71\)).
Outliers to this general pattern include the idiosyncratic species M. laticollaris at the base of the short-tailed clade, which was comparable to most long-tailed venoms, and the two North American species (M. tener and M. fulvius) which had much less effect than other long-tailed species. The similarity of M. laticollaris to long-tailed species is especially surprising because the sequence of its primary \(\upalpha\)-neurotoxin—MlatA1, UniProt K9MCH1 (Carbajal-Saucedo et al. 2013; Guerrero-Garzón et al. 2018)—is most similar to those of other short-tailed species: the top 10 most similar sequences in UniProt from other species all belong to short-tailed rather than long-tailed Micrurus (The UniProt Consortium 2023). This activity shift could potentially be an evolutionary innovation related to the large clades of short-chain 3FTx that are unique to long-tailed Micrurus and display strong signals of positive selection (Dashevsky and Fry 2018). However, the relatively similar results between M. laticollaris and most of the long-tailed species suggest an alternate hypothesis: that high binding may instead be an ancestral trait and that the main branch of the short-tailed phylogeny evolved away from toxins that bind to the orthosteric site. This scenario also accords with the overall evolutionary history of the 3FTx since the ancestral state of the family—represented by the plesiotypic toxins—is to bind to the orthosteric site, and this activity has been widely demonstrated for more derived toxins as well (Barber et al. 2013; Utkin et al. 2015; Rahman et al. 2020; Nys et al. 2022; Shenkarev et al. 2022).
This does not imply that species which produced small effects in this assay are necessarily less dangerous. Indeed, several of these species have been responsible for fatal bites in the past (Norris et al. 2009; Otero-Patiño 2014; Bucaretchi et al. 2016). This does not even necessarily mean that venoms which produce small effects do not inhibit the nAChR: post-synaptic neurotoxicity has been demonstrated in the venoms from a range of short-tailed (M. dissoleucus (Renjifo et al. 2012), M. laticollaris (Carbajal-Saucedo et al. 2013, 2014), M. lemniscatus (Cecchini et al. 2005; Floriano et al. 2019), M. mipartitus (Renjifo et al. 2012), M. obscurus (Yang et al. 2017), M. pyrrhocryptus (Yang et al. 2017), and M. surinamensis (Harris et al. 2020a)) and long-tailed (M. dumerilii (Serafim et al. 2002), M. fulvius (Snyder et al. 1973; Yang et al. 2017; Foo et al. 2019), and M. tener (Yang et al. 2017)) species. Earlier work has suggested that short-chain toxins might act as allosteric inhibitors of the nAChR: toxins which bind to a site other than the orthosteric site but still disrupt the receptor’s normal function (Nirthanan 2020; Harris et al. 2020a). This site may be located at other regions of the \(\upalpha _1\) subunit or other subunits of the muscular nAChR (which also include \(\upbeta\), \(\updelta\), and \(\upepsilon\) subunits). This previous research was focused on aquatic elapids including M. surinamensis, which is a short-tailed species, and our results suggest that allosteric 3FTx may be a feature in most of the rest of the clade as well. It has been put forward that allosteric 3FTx may be able to incapacitate prey faster than the more common orthosteric toxins, but would likely not bind as strongly (Barber et al. 2013; Harris 2022).
The possibility of wide spread allosteric toxins in Micrurus venoms casts doubts on the potential efficacy of decoy peptides as a potential therapeutic to combat bites from the genus. Some more recent studies into this area have used decoy peptides that do resemble the orthosteric site of the muscular nAChR and showed very potent binding with long-chain toxins (Kudryavtsev et al. 2020; Lynagh et al. 2020), but if there are short chain toxins that are able to cause severe neurotoxicity without significant affinity for the orthosteric site, then such peptides are unlikely to be of much use in combatting their effects. Even beyond the possibility of allosteric 3FTx, Micrurus venoms have been shown to contain potent neurotoxins from other toxin families that act through different mechanisms, including targeting the presynaptic side of the neuromuscular junction (Dal Belo et al. 2005; Oliveira et al. 2008; Bohlen et al. 2011; Terra et al. 2015; Floriano et al. 2019; Santos et al. 2020). This is particularly relevant for species that are known to produce primarily PLA2s—which are often presynaptically neurotoxic—in their venom rather than 3FTx, such as some populations of M. laticollaris, M. lemniscatus, M. ibiboboca, M. diastema, M. browni, M. tener, and M. fulvius (Lomonte et al. 2016; Sanz et al. 2019). Since the BLI method can only assay binding to the orthosteric site, toxins that act on other sites, be they elsewhere on the receptor or on the other side of the neuromuscular synapse, will not display any binding in our results.
Another interesting facet of our results is the fact that mimotopes derived from taxa other than snakes tend to be more sensitive to Micrurus venoms (non-snake mimotopes were almost twice as susceptible as snake mimotopes on average, Fig. 2). To analyze these data, we used a similar approach as when we compared short-tailed and long-tailed venoms, but used a new factor based on whether the mimotope was derived from the nAChR sequence of a non-snake or a snake species and then analyzed this factor, the species of origin for the venom, and their interaction. This once again confirmed that there is significant variation between the potency of the venom from different species (\(\textit{p} <0.001\)), but also that non-snake mimotopes are more easily bound by Micrurus venoms than those derived from snakes (\(\textit{p} <0.001\)), and these variables interact significantly as well (\(\textit{p} <0.001\)). Given the high proportion of snakes in the diets of most Micrurus species (Jackson and Franz 1981; Roze 1996; Marques and Sazima 1997; Marques 2002; Ávila et al. 2010; da Silva Banci 2017), our hypothesis of prey-specificity driving venom evolution would generally predict increased potency against mimotopes derived from snake sequences. As discussed earlier, our assay is a window into a specific fraction of the total activity of a snake’s venom. It is possible that the holistic potency of these venoms would be greater against snake rather than non-snake targets, but within the limitation of our results we find evidence for prey resistance.
To further investigate the resistance of snake mimotopes, we created a pseudo-ancestral mimotope by modifying the sequence of the most basal snake mimotope (blind snake) to revert the derived snake serine (S) at position 187 back to the tryptophan (W) that most other taxa possess at that position. Two-way ANOVA shows that Micrurus venoms more easily bind to the pseudo-ancestral blind snake mimotope than the normal blind snake mimotope (\(\textit{p} <0.001\)). Variation between venoms from different species of Micrurus remained significant (\(\textit{p} < 2 \times 10^{-16}\)) and the interaction between the two variables was marginally significant (\(\textit{p} =0.04\)). While these results clearly indicate that the W187S mutation reduces susceptibility to the binding of Micrurus toxins, Welch’s two-sample t-test shows that the ratio of the average response to a given venom of non-snake mimotopes to that of snake mimotopes (\(2.01\ \pm \ 0.65\)) is significantly greater than that the ratio between the pseudo-ancestral blind snake mimotope and the normal one (\(1.43\ \pm \ 0.40\), \(p =0.007\), Fig. 2). The source of this increased resistance is somewhat enigmatic since no other derived mutations are found in all and only snake sequences. Interestingly, the resistance conferred by this mutation seems to operate through biochemical mechanisms that do not interfere with the greater potency of the toxins from long-tailed species. The long-tailed species have similar response ratios between non-snake and snake mimotopes as the short-tailed species.
Snake nAChR are consistently less vulnerable to \(\upalpha\)-neurotoxins, partially due to the W187S mutation: response of non-snake mimotopes to each Micrurus venom are on average higher than the response of snake mimotopes to that same venom (indicated by the ratios greater than 1), similarly the pseudo-ancestral blind snake mimotope without the W187S mutation are more vulnerable than the normal blind snake mimotope
While 3FTx are an ancient family of snake toxins and have even been reported from very basal snake lineages including Pythonidae and Cylindrophiidae (Fry et al. 2013; Hargreaves et al. 2014), there is no published evidence they evolved before the divergence of the thread snakes and blind snakes from the more derived Alethinophidian snakes (Dashevsky et al. 2018; Koludarov et al. 2023). If indeed \(\upalpha\)-neurotoxic 3FTx had not evolved before the most recent common ancestor of all extant snakes, selection to resist the binding of these toxins cannot explain the widespread occurrence of the W187S mutation. This tryptophan residue is largely conserved in most other chordate taxa and negative selection has been shown to act on this site in most lineages (Khan et al. 2020). A similar scenario has been observed in lineages that can resist toad toxins, but related taxa in toad-free environments tend to lose the adaptation (Ujvari et al. 2015), suggesting the adaptation comes at a physiological cost and is lost when it is no longer offset by the benefit of resistance. It is unclear whether the W187S mutation arose as a deleterious trait and persisted through chance, if it was a neutral mutation because something about the biology of snakes negated the penalty that other taxa would face, or if the common ancestor of snakes was exposed to some selective pressure that made the mutation adaptive at the time. Since then, however, snakes whose venoms contain 3FTx—many of which are ophiophagus—have become widespread. Whatever its origins, the W187S mutation certainly seems to have been exapted to help protect snakes armed with these neurotoxins from the risk of self-envenomation and, perhaps more importantly, to protect snakes broadly from predation by their neurotoxic relatives.
Another factor that is not captured by our results is the possibility of prey taxa producing venom inhibitors rather than altering the targets, as has been well characterized for other families of snake toxins (Holding et al. 2016). Indeed, preliminary evidence suggests this might be the case for some snakes (Bolaños et al. 1975; Rodrigues et al. 2020, 2021). Coupled with the W187S mutation at the muscular nAChR orthosteric site, such venom inhibitors could have a synergistic effect and grant high levels of resistance to \(\upalpha\)-neurotoxicity. Along with other factors, including relaxed selective constraint and pressure from predators, multi-faceted resistance has been suggested as one potential reason why so many neurotoxic snake venoms seem to be toxic to the point of overkill (Broad et al. 1979; Mebs 2001; Kundu et al. 2015; Aird et al. 2017; Gangur et al. 2018; Healy et al. 2019). If their prey (other snakes) are resistant then venomous snakes might need to evolve toward larger doses and/or more toxic venoms due to Red Queen dynamics (Van Valen 1973). Since the model organisms (usually mice) are not participants in this specific arms race, the tests carried out on them may wrongfully suggest the venoms in question are much more toxic than they are in practice in their proper ecological context. With this in mind, the reduced susceptibility to \(\upalpha\)-neurotoxins we observed among snakes relative to non-snake taxa may be just part of an ongoing arms race between predator and prey. Two studies have been carried out comparing the effects of Micrurus venoms on dipsadine snake species and lab mice and both suggest that these snakes are much less susceptible to the lethal effects of the venoms, with the least resistant snake species exhibiting LD50s threefold greater than the mice (Urdaneta et al. 2004; Bénard-Valle et al. 2014). However, there is substantial variation even within this subfamily of snakes and the more recent study found that there was no difference in the dosage required to paralyze the mice and snakes, which is a more ecologically relevant measure when considering the predatory purpose of the venom (Bénard-Valle et al. 2014).
Conclusion
This study examines a very particular mode of action among Micrurus venoms—binding to the orthosteric site of the nAChR—and reveals two particularly interesting patterns. The first is that there is substantial variation between and within Micrurus species and that the phylogenetic pattern suggests that the ancestral state of the genus was likely to produce toxins which bind strongly to the orthosteric site. Several lineages—including most of the short-tailed clade as well as the two North American species in the long-tailed clade—have shifted away from this specific target. The second major trend was that targets which mimicked snake sequences were less susceptible to Micrurus venoms than those which mimicked other taxa. This is due, in part, to a W\(\rightarrow\)S mutation in the orthosteric region, which is shared by snakes, but rare elsewhere. However, there is much more to even a simple venom than its potency at one specific site of one physiological target and these results may not be representative of the broader picture of the arms race between \(\upalpha\)-neurotoxic elapids and their snake prey.
Methods
Venom Samples
All venoms were stored as lyophilized powder as part the long-term cryogenic collections in the Venom Evolution Lab. Venoms for Micrurus fulvius (FL, USA), M. obscurus (Brazil), and M. tener (TX, USA) were supplied by Miami Serpentarium. Nathaniel Frank of MToxins Venom Lab provided M. corallinus A venom. Venom from M. pyrrhocryptus was from an individual in the captive collection of Iwan Hendrikx that originally came from Suriname. José A. Portes-Junior, Anita M. Tanaka-Azevedo, Kathleen F. Grego, and Sávio S. Sant’Anna of Instituto Butantan provided the majority of the Brazilian venoms: M. altirostris (pooled from 3 individuals), M. corallinus B (pooled from 3 individuals), M. frontalis, M. hemprichii, M. ibiboboca (pooled from 2 individuals), and M. lemniscatus. Venoms of Mexican Micrurus from Alejandro Alagón—M. browni, M. diastema, M. distans, and M. laticollaris—were manually extracted from Micrurus specimens kept at the “Herpetario Cantil” from Instituto de Biotecnología–Universidad Nacional Autónoma de México. After extraction, venoms were diluted in MilliQ water, centrifuged 3 min at 10,000 RCF to remove insoluble material, lyophilized and stored at 4 °C until shipment.
As no live animals were used for this study, and all venoms were from previously collected stocks, no animal ethics approvals were required for this work. These lyophilized venoms were resuspended in water, centrifuged (4 °C, 5 min at 14,000 RCF), and diluted into a solution of 1 \(\frac {\textrm{mg}}{\textrm{ml}}\) of venom in a 1:1 mixture of water and glycerol. Protein concentrations were measured using a NanoDrop 2000 UV–Vis Spectrophotometer (Thermofisher, Sydney, NSW, Australia).
Mimotope Production and Preparation
The sequences for the muscular nAChR were downloaded from the following UniProt entries: Tetronarce californica (fish, P02710), Xenopus tropicalis (amphibian, F6RLA9), Sarcophilus harrisii (marsupial, G3W0J0), Rattus norvegicus (rodent, P25108), Gallus gallus (bird, E1BT92), Gekko japonicus (gecko, GenBank: XM015426640), and Anolis carolinensis (anole, H9GA55), Barisia imbricata (alligator lizard, A0A859JD35), Anilios bituberculatus (blind snake, UniProt: A0A7L5PIU6), Boa constrictor (boa, A0A7L5PLV8), Oxyrhopus rhombifer (Dipsadine, A0A7L5PLX4), and Pantherophis spiloides (Colubrine, A0A7L5PL14). These sequences were then aligned using AliView 1.18 (Larsson 2014) and were trimmed down to the 14 amino acids of the orthosteric binding site. The alignment of these sites can be found in the Supplementary Data S1.
Subsequently, following previous protocols (Chiappinelli et al. 1996; Bracci et al. 2001; Kasher et al. 2001; Bracci et al. 2002) mimotopes of these sequences were developed by GenicBio Ltd (Shanghai, China). As per previous studies (Bracci et al. 2001), the cysteine doublet in the orthosteric binding site sequence was replaced in peptide synthesis steps with serine doublet to avoid uncontrolled postsynthetic thiol oxidation. Research has shown this has no effect on the analyte-ligand complex formation (Tzartos and Remoundos 1990; McLane et al. 1991, 1994). The mimotope was further synthesized to a biotin linker bound to two aminohexanoic acid (Ahx) spacers to form a 30 Å linker between biotin and the peptide. This provides conformational freedom for the analyte-ligand complex. The purpose of adding the biotin-Ahx complex is to allow the biotin molecule to bind non-covalently to the avidin pocket of the streptavidin-coated disposable biosensors used in the bio-layer interferometry assay. Dried stocks of synthesized mimotopes were solubilized in 100% dimethyl sulfoxide (DMSO) and then diluted 1:10 in deionized water to make a final working stock concentration of 50 \(\frac{{\upmu }{\textrm{g}}}{\textrm{ml}}\) and stored at −80 \({}^{\circ }\)C until use.
Bio-Layer Interferometry
Full details of the assay, including a full methodology and data analysis, can be found in the validation study (Zdenek et al. 2019) and further publications using this protocol (Harris et al. 2020a, b, c, 2021; Chowdhury et al. 2022). In summary, the BLI assay was performed on the Octet Red 96 system (ForteBio, Fremont, CA, USA). Venom (analyte) samples were diluted at 1:20 (a final experimental concentration of 50 \(\frac{{\upmu }{\textrm{g}}}{\textrm{ml}}\) per well). Mimotope aliquots were diluted at 1:50 (a final concentration of 1 \(\frac{{\upmu }{\textrm{g}}}{\textrm{ml}}\) per well). The assay running buffer was 1X DPBS with 0.1% BSA and 0.05% Tween-20. Before experimentation, streptavidin biosensors were hydrated in the running buffer for 30–60 min. Elimination of bound venom toxins (regeneration) was performed using a standard acidic solution glycine buffer (10 mM glycine (pH 1.5\(-\)1.7) in ddH2O). Raw data are provided in Supplementary Data S2 and line charts similar to Fig. 1A can be found in Supplementary Figs. S1–S12. All data obtained from BLI on Octet Red 96 system (ForteBio) were processed following the protocol in the validation study for this assay (Zdenek et al. 2019). In brief, the raw data were exported to a.csv file. Area under the curve for the association step of each samples was estimated using trapezoidal approximation in Microsoft Excel 15.28. Because venoms are heterologous mixtures of multiple different toxins of unknown molarity in the assay we were unable to estimate the on and off rates of the binding reactions necessary to calculate dissociation constants. The presence of multiple toxins, some of which were able to weakly associate with the mimotopes also complicated the dissociation phase of the reaction which is why we focused on the association phase. Further statistical analyses were carried out in the RStudio 2023.06.1+524 implementation of R 4.3.1 (RStudio Team 2015; R Core Team 2015). These data can be found in the Supplementary Material.
References
Ackermann EJ, Taylor P (1997) Nonidentity of the α-neurotoxin binding sites on the nicotinic acetylcholine receptor revealed by modification in α-neurotoxin and receptor structures. Biochemistry 36(42):12836–12844
Aird SD, Arora J, Barua A et al (2017) Population genomic analysis of a pitviper reveals microevolutionary forces underlying venom chemistry. Genome Biol Evol 9(10):2640–2649. https://doi.org/10.1093/gbe/evx199
Albulescu LO, Kazandjian T, Slagboom J et al (2019) A decoy-receptor approach using nicotinic acetylcholine receptor mimics reveals their potential as novel therapeutics against neurotoxic snakebite. Front Pharmacol. https://doi.org/10.3389/fphar.2019.00848
Ávila RW, Kawashita-Ribeiro RA, Ferreira VL et al (2010) Natural history of the coral snake Micrurus pyrrhocryptus cope 1862 (Elapidae) from semideciduous forests of Western Brazil. South Am J Herpetol 5(2):97–101. https://doi.org/10.2994/057.005.0204
Barber CM, Isbister GK, Hodgson WC (2013) Alpha neurotoxins. Toxicon 66:47–58. https://doi.org/10.1016/j.toxicon.2013.01.019
Barchan D, Kachalsky S, Neumann D et al (1992) How the mongoose can fight the snake: the binding site of the mongoose acetylcholine receptor. Proc Natl Acad Sci 89(16):7717–7721
Barchan D, Ovadia M, Kochva E et al (1995) The binding site of the nicotinic acetylcholine receptor in animal species resistant to α-bungarotoxin. Biochemistry 34(28):9172–9176
Barlow A, Pook CE, Harrison RA et al (2009) Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc Royal Soc B: Biol Sci 276(1666):2443–2449. https://doi.org/10.1098/rspb.2009.0048
Bénard-Valle M, Carbajal-Saucedo A, de Roodt A et al (2014) Biochemical characterization of the venom of the coral snake Micrurus tener and comparative biological activities in the mouse and a reptile model. Toxicon 77:6–15. https://doi.org/10.1016/j.toxicon.2013.10.005
Bénard-Valle M, Neri-Castro E, Yañez-Mendoza MF et al (2020) Functional, proteomic and transcriptomic characterization of the venom from Micrurus browni: identification of the first lethal multimeric neurotoxin in coral snake venom. J Proteomics 225:103863. https://doi.org/10.1016/j.jprot.2020.103863
Bernarde PS, Turci LCB, Abegg AD et al (2018) A remarkable new species of coralsnake of the Micrurus hemprichii species group from the Brazilian Amazon. Salamandra 54(4):249–258. https://doi.org/10.5281/zenodo.2586853
Bohlen CJ, Chesler AT, Sharif-Naeini R et al (2011) A heteromeric Texas coral snake toxin targets acid-sensing ion channels to produce pain. Nature 479(7373):410–414. https://doi.org/10.1038/nature10607
Bolaños R, Piva A, Taylor R et al (1975) Natural resistance of bovine animals to Micrurus nigrocinctus venom. Toxicon 13(5):369–370
Bracci L, Lozzi L, Lelli B et al (2001) Mimotopes of the nicotinic receptor binding site selected by a combinatorial peptide library. Biochemistry 40:6611–6619. https://doi.org/10.1021/bi0023201
Bracci L, Lozzi L, Pini A et al (2002) A branched peptide mimotope of the nicotinic receptor binding site is a potent synthetic antidote against the snake neurotoxin alpha-bungarotoxin. Biochemistry 41:10194–10199. https://doi.org/10.1021/bi0256025
Broad AJ, Sutherland SK, Coulter AR (1979) The lethality in mice of dangerous Australian and other snake venom. Toxicon 17(6):661–664
Bucaretchi F, Capitani EMD, Vieira RJ et al (2016) Coral snake bites (Micrurus spp.) in Brazil: a review of literature reports. Clin Toxicol 54(3):222–234. https://doi.org/10.3109/15563650.2015.1135337
Burden SJ, Hartzell HC, Yoshikami D (2006) Acetylcholine receptors at neuromuscular synapses: phylogenetic differences detected by snake α-neurotoxins. Proc Natl Acad Sci 72(8):3245–3249. https://doi.org/10.1073/pnas.72.8.3245
Campbell JA, Lamar WW (2004) The venomous reptiles of the Western Hemisphere. Cornell University Press, Ithaca
Carbajal-Saucedo A, López-Vera E, Bénard-Valle M et al (2013) Isolation, characterization, cloning and expression of an alpha-neurotoxin from the venom of the Mexican coral snake Micrurus laticollaris (Squamata: Elapidae). Toxicon 66:64–74. https://doi.org/10.1016/j.toxicon.2013.02.006
Carbajal-Saucedo A, Floriano RS, Belo CAD et al (2014) Neuromuscular activity of Micrurus laticollaris (Squamata: Elapidae) venom in vitro. Toxins 6(1):359–370. https://doi.org/10.3390/toxins6010359
Cecchini AL, Marcussi S, Silveira LB et al (2005) Biological and enzymatic activities of Micrurus sp. (Coral) snake venoms. Comp Biochem Physiol A Mol Integr Physiol 140(1):125–134. https://doi.org/10.1016/j.cbpb.2004.11.012
Chiappinelli VA, Weaver WR, McLane KE et al (1996) Binding of native κ-neurotoxins and site-directed mutants to nicotinic acetylcholine receptors. Toxicon 34(11/12):1243–1256
Chowdhury A, Zdenek CN, Fry BG (2022) Diverse and dynamic alpha-neurotoxicity within venoms from the palearctic viperid snake clade of Daboia, Macrovipera, Montivipera, and Vipera. Neurotox Res. https://doi.org/10.1007/s12640-022-00572-w
Corrêa-Netto C, Junqueira-de Azevedo IdLM, Silva DA et al (2011) Snake venomics and venom gland transcriptomic analysis of Brazilian coral snakes, Micrurus altirostris and M. corallinus. J Proteomics 74(9):1795–1809. https://doi.org/10.1016/j.jprot.2011.04.003
da Silva Banci KR (2017) Feeding on elongate prey: additonal data for coral snake Micrurus corallinus (Elapidae) and comments on aposematism. Herpetol Notes 10:335–338
da Silva NJ, Aird SD (2001) Prey specificity, comparative lethality and compositional differences of coral snake venoms. Comp Biochem Physiol C: Toxicol Pharmacol 128(3):425–456. https://doi.org/10.1016/S1532-0456(00)00215-5
da Silva NJ, Pires MG, Zaher H et al (2015) A new species of monadal coral snake of the genus Micrurus (Serpentes, Elapidae) from western Amazon. Zootaxa 3974(4):538–554
Dal Belo CA, Leite GB, Toyama MH et al (2005) Pharmacological and structural characterization of a novel phospholipase A2 from Micrurus dumerilii carinicauda venom. Toxicon 46(7):736–750. https://doi.org/10.1016/j.toxicon.2005.07.016
Dashevsky D, Fry BG (2018) Ancient diversification of three-finger toxins in Micrurus coral snakes. J Mol Evol 86(1):58–67. https://doi.org/10.1007/s00239-017-9825-5
Dashevsky D, Debono J, Rokyta D et al (2018) Three-finger toxin diversification in the venoms of cat-eye snakes (Colubridae: Boiga). J Mol Evol 86(8):531–545. https://doi.org/10.1007/s00239-018-9864-6
Dashevsky D, Rokyta D, Frank N et al (2021) Electric blue: molecular evolution of three-finger toxins in the long-glanded coral snake species Calliophis bivirgatus. Toxins 13(2):124. https://doi.org/10.3390/toxins13020124
Dawkins R, Krebs JR (1979) Arms races between and within species. Proc Royal Soc London Ser B Biol Sci 205(1161):489–511
Di-Bernardo M, Borges-Martins M, Silva Da, (Jr.) NJ, (2007) A new species of coralsnake (Micrurus: Elapidae) from southern Brazil. Zootaxa. https://doi.org/10.11646/zootaxa.1447.1.1
Drabeck DH, Dean AM, Jansa SA (2015) Why the honey badger don’t care: convergent evolution of venom-targeted nicotinic acetylcholine receptors in mammals that survive venomous snake bites. Toxicon 99:68–72. https://doi.org/10.1016/j.toxicon.2015.03.007
Figueroa A, McKelvy AD, Grismer LL et al (2016) A species-level phylogeny of extant snakes with description of a new colubrid subfamily and genus. PLoS ONE 11(9):e0161070. https://doi.org/10.1371/journal.pone.0161070
Floriano RS, Schezaro-Ramos R, Silva NJ et al (2019) Neurotoxicity of Micrurus lemniscatus (South American coralsnake) venom in vertebrate neuromuscular preparations in vitro and neutralization by antivenom. Archiv Toxicol. https://doi.org/10.1007/s00204-019-02476-9
Foo CS, Jobichen C, Hassan-Puttaswamy V et al (2019) Fulditoxin, representing a new class of dimeric snake toxins, defines novel pharmacology at nicotinic acetylcholine receptors. Br J Pharmacol. https://doi.org/10.1111/bph.14954
Fry BG, Wüster W, Kini RM et al (2003) Molecular evolution and phylogeny of elapid snake venom three-finger toxins. J Mol Evol 57(1):110–129. https://doi.org/10.1007/s00239-003-2461-2
Fry BG, Undheim EAB, Ali SA et al (2013) Squeezers and leaf-cutters: differential diversification and degeneration of the venom system in toxicoferan reptiles. Mol Cell Proteomics 12(7):1881–1899. https://doi.org/10.1074/mcp.M112.023143
Galzi JL, Revah F, Bessis A et al (1991) Functional architecture of the nicotinic acetylcholine receptor: from electric organ to brain. Ann Rev Pharmacol Toxicol 31(1):37–72. https://doi.org/10.1146/annurev.pa.31.040191.000345
Gangur AN, Seymour JE, Liddell MJ et al (2018) When is overkill optimal? Tritrophic interactions reveal new insights into venom evolution. Theor Ecol 11(2):141–149. https://doi.org/10.1007/s12080-017-0354-z
Gómez JPH, Ramirez MV, Gómez FJR et al (2021) Multilocus phylogeny clarifies relationships and diversity within the Micrurus lemniscatus complex (Serpentes: Elapidae). Salamandra 57(2):10
Gotti C, Clementi F (2004) Neuronal nicotinic receptors: from structure to pathology. Prog Neurobiol 74(6):363–96. https://doi.org/10.1016/j.pneurobio.2004.09.006
Guerrero-Garzón JF, Bénard-Valle M, Restano-Cassulini R et al (2018) Cloning and sequencing of three-finger toxins from the venom glands of four Micrurus species from Mexico and heterologous expression of an alpha-neurotoxin from Micrurus diastema. Biochimie 147:114–121. https://doi.org/10.1016/j.biochi.2018.01.006
Gutberlet RL Jr, Harvey MB (2004) The evolution of New World venomous snakes. The venomous reptiles of the Western Hemisphere, vol 2. Cornell University Press, Ithaca, pp 634–682
Hargreaves AD, Swain MT, Logan DW et al (2014) Testing the Toxicofera: comparative transcriptomics casts doubt on the single, early evolution of the reptile venom system. Toxicon 92:140–156
Harris R (2022) The co-evolutionary dynamics of α-neurotoxic snake venoms and their nicotinic acetylcholine receptor targets. PhD Thesis, The University of Queensland, School of Biological Sciences
Harris RJ, Youngman NJ, Zdenek CN et al (2020a) Assessing the binding of venoms from aquatic elapids to the nicotinic acetylcholine receptor orthosteric site of different prey models. Int J Mol Sci 21(19):7377
Harris RJ, Zdenek CN, Debono J et al (2020b) Evolutionary interpretations of nicotinic acetylcholine receptor targeting venom effects by a clade of Asian Viperidae Snakes. Neurotox Res 38(2):312–318. https://doi.org/10.1007/s12640-020-00211-2
Harris RJ, Zdenek CN, Harrich D et al (2020c) An appetite for destruction: detecting prey-selective binding of α-neurotoxins in the venom of Afro-Asian elapids. Toxins 12(3):205
Harris RJ, Nekaris KAI, Fry BG (2021) Monkeying around with venom: an increased resistance to α-neurotoxins supports an evolutionary arms race between Afro-Asian primates and sympatric cobras. BMC Biol 19(1):253. https://doi.org/10.1186/s12915-021-01195-x
Healy K, Carbone C, Jackson AL (2019) Snake venom potency and yield are associated with prey-evolution, predator metabolism and habitat structure. Ecol Lett 22(3):527–537. https://doi.org/10.1111/ele.13216
Heyborne WH, Mackessy SP (2013) Identification and characterization of a taxon-specific three-finger toxin from the venom of the Green Vinesnake (Oxybelis fulgidus; family Colubridae). Biochimie 95(10):1923–1932
Holding ML, Drabeck DH, Jansa SA et al (2016) Venom resistance as a model for understanding the molecular basis of complex coevolutionary adaptations. Integr Comp Biol 56(5):1032–1043. https://doi.org/10.1093/icb/icw082
Jackson DR, Franz R (1981) Ecology of the eastern coral snake (Micrurus fulvius) in northern peninsular Florida. Herpetologica 1981:213–228
Jowers MJ, Garcia Mudarra JL, Charles SP et al (2019) Phylogeography of West Indies Coral snakes (Micrurus): Island colonisation and banding patterns. Zool Script 48(3):263–276. https://doi.org/10.1111/zsc.12346
Jowers MJ, Smart U, Sánchez-Ramírez S et al (2023) Unveiling underestimated species diversity within the Central American Coralsnake, a medically important complex of venomous taxa. Sci Rep 13(1):11674. https://doi.org/10.1038/s41598-023-37734-5
Kamat V, Rafique A (2017) Designing binding kinetic assay on the bio-layer interferometry (BLI) biosensor to characterize antibody-antigen interactions. Anal Biochem 536:16–31. https://doi.org/10.1016/j.ab.2017.08.002
Kasher R, Balass M, Scherf T et al (2001) Design and synthesis of peptides that bind α-bungarotoxin with high affinity. Chem Biol 8(2):147–155. https://doi.org/10.1016/S1074-5521(00)90063-2
Katchalski-Katzir E, Kasher R, Balass M et al (2003) Design and synthesis of peptides that bind α-bungarotoxin with high affinity and mimic the three-dimensional structure of the binding-site of acetylcholine receptor. Biophys Chem 100:293–305
Khan MA, Dashevsky D, Kerkkamp HM et al (2020) Widespread evolution of molecular resistance to snake venom α-neurotoxins in vertebrates. Toxins 2020:205
Koludarov I, Senoner T, Jackson TNW et al (2023) Domain loss enabled evolution of novel functions in the snake three-finger toxin gene superfamily. Nat Commun 14(1):4861. https://doi.org/10.1038/s41467-023-40550-0
Kudryavtsev DS, Tabakmakher VM, Budylin GS et al (2020) Complex approach for analysis of snake venom α-neurotoxins binding to HAP, the high-affinity peptide. Sci Rep 10(1):3861. https://doi.org/10.1038/s41598-020-60768-y
Kundu P, Venkitachalam S, Vidya TNC (2015) Why so toxic? Resonance 20(7):617–627. https://doi.org/10.1007/s12045-015-0220-5
Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30(22):3276–3278. https://doi.org/10.1093/bioinformatics/btu531
Lee MSY, Sanders KL, King B et al (2016) Diversification rates and phenotypic evolution in venomous snakes (Elapidae). Open Sci 3(1):150277. https://doi.org/10.1098/rsos.150277
Lomonte B, Rey-Suárez P, Fernández J et al (2016) Venoms of Micrurus coral snakes: evolutionary trends in compositional patterns emerging from proteomic analyses. Toxicon 122:7–25. https://doi.org/10.1016/j.toxicon.2016.09.008
Lynagh T, Kiontke S, Meyhoff-Madsen M et al (2020) Peptide inhibitors of the α-Cobratoxin-nicotinic acetylcholine receptor interaction. J Med Chem 63(22):13709–13718. https://doi.org/10.1021/acs.jmedchem.0c01202
Mackessy SP, Sixberry NM, Heyborne WH et al (2006) Venom of the Brown Treesnake, Boiga irregularis: ontogenetic shifts and taxa-specific toxicity. Toxicon 47(5):537–548. https://doi.org/10.1016/j.toxicon.2006.01.007
Margres MJ, Aronow K, Loyacano J et al (2013) The venom-gland transcriptome of the eastern coral snake (Micrurus fulvius) reveals high venom complexity in the intragenomic evolution of venoms. BMC Genom 14(1):1
Marques OA (2002) Natural history of the coral snake Micrurus decoratus (Elapidae) from the Atlantic Forest in southeast Brazil, with comments on possible mimicry. Amphibia-Reptilia 23(2):228–232
Marques OA, Sazima I (1997) Diet and feeding behavior of the coral snake, Micrurus corallinus, from the Atlantic forest of Brazil. Herpetol Natl Hist 5(1):88–93
McLane KE, Wu X, Diethelm B et al (1991) Structural determinants of α-bungarotoxin binding to the sequence segment 181–200 of the muscle nicotinic acetylcholine receptor subunit: effects of cysteine/cystine modification and species-specific amino acid substitutions. Biochemistry 30(20):4925–4934. https://doi.org/10.1021/bi00234a013
McLane KE, Wu X, Conti-Tronconi BM (1994) An α-bungarotoxin-binding sequence on the torpedo nicotinic acetylcholine receptor α-subunit: conservative amino acid substitutions reveal side-chain specific interactions. Biochemistry 33(9):2576–2585
Mebs D (2001) Toxicity in animals. Trends in evolution? Toxicon 39(1):87–96. https://doi.org/10.1016/S0041-0101(00)00155-0
Modahl CM, Mrinalini Frietze S et al (2018) Adaptive evolution of distinct prey-specific toxin genes in rear-fanged snake venom. Proc Royal Soc B Biol Sci 285(1884):20181003. https://doi.org/10.1098/rspb.2018.1003
Nascimento LRS, Silva NJJ, Feitosa DT et al (2019) Taxonomy of the Micrurus spixii species complex (Serpentes, Elapidae). Zootaxa. https://doi.org/10.11646/zootaxa.4668.3.4
Nirthanan S (2020) Snake three-finger α-neurotoxins and nicotinic acetylcholine receptors: molecules, mechanisms and medicine. Biochem Pharmacol 2020:114168. https://doi.org/10.1016/j.bcp.2020.114168
Nirthanan S, Gwee MCE (2004) Three-finger α-neurotoxins and the nicotinic acetylcholine receptor, forty years on. J Pharmacol Sci 94(1):1–17. https://doi.org/10.1254/jphs.94.1
Norris RL, Pfalzgraf RR, Laing G (2009) Death following coral snake bite in the United States: first documented case (with ELISA confirmation of envenomation) in over 40 years. Toxicon 53(6):693–697. https://doi.org/10.1016/j.toxicon.2009.01.032
Nys M, Zarkadas E, Brams M et al (2022) The molecular mechanism of snake short-chain α-neurotoxin binding to muscle-type nicotinic acetylcholine receptors. Nat Commun 13(1):4543. https://doi.org/10.1038/s41467-022-32174-7
Oliveira DA, Harasawa C, Seibert CS et al (2008) Phospholipases A2 isolated from Micrurus lemniscatus coral snake venom: behavioral, electroencephalographic, and neuropathological aspects. Brain Res Bull 75(5):629–639. https://doi.org/10.1016/j.brainresbull.2007.10.007
Otero-Patiño DR (2014) Snake bites in Colombia. In: Gopalakrishnakone P, Faiz SMA, Gnanathasan CA et al (eds) Clinical toxinology. Springer, Netherlands, pp 1–42
Pawlak J, Mackessy SP, Fry BG et al (2006) Denmotoxin, a three-finger toxin from the colubrid snake Boiga dendrophila (Mangrove Catsnake) with bird-specific activity. J Biol Chem 281(39):29030–29041. https://doi.org/10.1074/jbc.M605850200
Pawlak J, Mackessy SP, Sixberry NM et al (2008) Irditoxin, a novel covalently linked heterodimeric three-finger toxin with high taxon-specific neurotoxicity. FASEB J 23(2):534–545. https://doi.org/10.1096/fj.08-113555
Pires MG, Jr Da Silva, NJ, Feitosa DT, et al (2014) A new species of triadal coral snake of the genus Micrurus Wagler, 1824 (Serpentes: Elapidae) from northeastern Brazil. Zootaxa 3811(4):10. https://doi.org/10.11646/zootaxa.3811.4.8
Pyron RA, Burbrink FT, Wiens JJ (2013) A phylogeny and revised classification of Squamata, including 4161 species of lizards and snakes. BMC Evol Biol 13:93. https://doi.org/10.1186/1471-2148-13-93
R Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rahman MM, Teng J, Worrell BT et al (2020) Structure of the native muscle-type nicotinic receptor and inhibition by snake venom toxins. Neuron 106(6):952-962.e5. https://doi.org/10.1016/j.neuron.2020.03.012
Renjifo C, Smith EN, Hodgson WC et al (2012) Neuromuscular activity of the venoms of the Colombian coral snakes Micrurus dissoleucus and Micrurus mipartitus: an evolutionary perspective. Toxicon 59(1):132–142. https://doi.org/10.1016/j.toxicon.2011.10.017
Reyes-Velasco J, Adams RH, Boissinot S et al (2020) Genome-wide SNPs clarify lineage diversity confused by coloration in coralsnakes of the Micrurus diastema species complex (Serpentes: Elapidae). Mol Phylogenet Evol 147:106770. https://doi.org/10.1016/j.ympev.2020.106770
Rodrigues CFB, Serino-Silva C, Morais-Zani Kd et al (2020) BoaγPLI: structural and functional characterization of the gamma phospholipase A2 plasma inhibitor from the non-venomous Brazilian snake Boa constrictor. PLoS ONE 15(2):e0229657. https://doi.org/10.1371/journal.pone.0229657
Rodrigues CFB, Zdenek CN, Serino-Silva C et al (2021) BoaγPLI from Boa constrictor blood is a broad-spectrum inhibitor of venom PLA2 pathophysiological actions. J Chem Ecol. https://doi.org/10.1007/s10886-021-01289-4
Rokyta DR, Wray KP, Margres MJ (2013) The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venom-gland transcriptomics. BMC Genomics 14(1):394. https://doi.org/10.1186/1471-2164-14-394
Rokyta DR, Wray KP, McGivern JJ et al (2015) The transcriptomic and proteomic basis for the evolution of a novel venom phenotype within the Timber Rattlesnake (Crotalus horridus). Toxicon 98:34–48. https://doi.org/10.1016/j.toxicon.2015.02.015
Roze JA (1996) Coral snakes of the Americas: biology, identification, and venoms. Krieger Publishing Company
RStudio Team (2015) RStudio: integrated development environment for R. RStudio Inc, Boston
Santos RTFd, Silva MFP, Porto RM et al (2020) Effects of Mlx-8, a phospholipase A2 from Brazilian coralsnake Micrurus lemniscatus venom, on muscarinic acetylcholine receptors in rat hippocampus. J Venom Anim Toxins Incl Trop Dis 26:e20190041. https://doi.org/10.1590/1678-9199-jvatitd-2019-0041
Sanz L, Quesada-Bernat S, Ramos T et al (2019) New insights into the phylogeographic distribution of the 3FTx/PLA2 venom dichotomy across genus Micrurus in South America. J Proteomics 200:90–101. https://doi.org/10.1016/j.jprot.2019.03.014
Serafim FG, Reali M, Cruz-Höfling MA et al (2002) Action of Micrurus dumerilii carinicauda coral snake venom on the mammalian neuromuscular junction. Toxicon 40(2):167–174. https://doi.org/10.1016/S0041-0101(01)00217-3
Servent D, Winckler-Dietrich V, Hu HY et al (1997) Only snake curaremimetic toxins with a fifth disulfide bond have high affinity for the neuronal α7 nicotinic receptor. J Biol Chem 272(39):24279–24286
Shenkarev ZO, Chesnokov YM, Zaigraev MM et al (2022) Membrane-mediated interaction of non-conventional snake three-finger toxins with nicotinic acetylcholine receptors. Commun Biol 5(1):1–15. https://doi.org/10.1038/s42003-022-04308-6
Silva A, Cristofori-Armstrong B, Rash LD et al (2018) Defining the role of post-synaptic α-neurotoxins in paralysis due to snake envenoming in humans. Cell Mol Life Sci 75(23):4465–4478. https://doi.org/10.1007/s00018-018-2893-x
Slowinski JB (1995) A phylogenetic analysis of the new world coral snakes (Elapidae: Leptomicrurus, Micruroides, and Micrurus) based on allozymic and morphological characters. J Herpetol 29(3):325–338. https://doi.org/10.2307/1564981
Snyder GK, Ramsey HW, Taylor WJ et al (1973) Neuromuscular blockade of chick biventer cervicis nerve-muscle preparations by a fraction from coral snake venom. Toxicon 11(6):505–508
Sousa LF, Zdenek CN, Dobson JS (2018) Coagulotoxicity of Bothrops (Lancehead Pit-Vipers) venoms from Brazil: differential biochemistry and antivenom efficacy resulting from prey-driven venom variation. Toxins 10(10):411
Tae HS, Adams DJ (2023) Nicotinic acetylcholine receptor subtype expression, function, and pharmacology: therapeutic potential of α-conotoxins. Pharmacol Res 191:106747. https://doi.org/10.1016/j.phrs.2023.106747
Takacs Z, Wilhelmsen KC, Sorota S (2001) Snake α-neurotoxin binding site on the Egyptian Cobra (Naja haje) nicotinic acetylcholine receptor is conserved. Mol Biol Evol 18(9):1800–1809. https://doi.org/10.1093/oxfordjournals.molbev.a003967
Takacs Z, Wilhelmsen KC, Sorota S (2004) Cobra ( Naja spp. ) nicotinic acetylcholine receptor exhibits resistance to Erabu Sea Snake (Laticauda semifasciata) short-chain a-neurotoxin. J Mol Evol 58(5):516–526. https://doi.org/10.1007/s00239-003-2573-8
Tan KY, Liew JL, Tan NH et al (2019) Unlocking the secrets of banded coral snake (Calliophis intestinalis, Malaysia): a venom with proteome novelty, low toxicity and distinct antigenicity. J Proteomics 192:246–257. https://doi.org/10.1016/j.jprot.2018.09.006
Terra AL, Moreira-Dill LS, Simões Silva R et al (2015) Biological characterization of the Amazon coral Micrurus spixii snake venom: isolation of a new neurotoxic phospholipase A2. Toxicon 103:1–11. https://doi.org/10.1016/j.toxicon.2015.06.011
Terribile LC, Feitosa DT, Pires MG et al (2018) Reducing Wallacean shortfalls for the coralsnakes of the Micrurus lemniscatus species complex: present and future distributions under a changing climate. PLoS ONE 13(11):e0205164. https://doi.org/10.1371/journal.pone.0205164
The UniProt Consortium (2023) UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res 51(D1):D523–D531. https://doi.org/10.1093/nar/gkac1052
Tzartos SJ, Remoundos MS (1990) Fine localization of the major α-bungarotoxin binding site to residues α 189–195 of the Torpedo acetylcholine receptor: residues 189, 190, and 195 are indispensable for binding. J Biol Chem 265(35):21462–21467
Ujvari B, Casewell NR, Sunagar K et al (2015) Widespread convergence in toxin resistance by predictable molecular evolution. Proc Natl Acad Sci 112(38):11911–11916. https://doi.org/10.1073/pnas.1511706112
Urdaneta AH, Bolaños F, Gutiérrez JM (2004) Feeding behavior and venom toxicity of coral snake Micrurus nigrocinctus (Serpentes: Elapidae) on its natural prey in captivity. Comp Biochem Physiol C Toxicol Pharmacol 138(4):485–492. https://doi.org/10.1016/j.cca.2004.08.018
Utkin Y, Sunagar K, Jackson TN et al (2015) Three-finger toxins (3ftxs). In: Fry BG (ed) Venomous reptiles and their toxins: evolution, pathophysiology and biodiscovery. Oxford University Press, pp 215–227
Van Valen L (1973) A new evolutionary law. Evol Theor 1:1–30
Yang DC, Dobson J, Cochran C et al (2017) The bold and the beautiful: a neurotoxicity comparison of new world coral snakes in the Micruroides and Micrurus genera and relative neutralization by antivenom. Neurotox Res 32(3):487–495. https://doi.org/10.1007/s12640-017-9771-4
Zaher H, Murphy RW, Arredondo JC et al (2019) Large-scale molecular phylogeny, morphology, divergence-time estimation, and the fossil record of advanced caenophidian snakes (Squamata: Serpentes). PLoS ONE 14(5):e0216148. https://doi.org/10.1371/journal.pone.0216148
Zdenek CN, Harris RJ, Kuruppu S et al (2019) A taxon-specific and high-throughput method for measuring ligand binding to nicotinic acetylcholine receptors. Toxins 11(10):600. https://doi.org/10.3390/toxins11100600
Zheng Y, Wiens JJ (2016) Combining phylogenomic and supermatrix approaches, and a time-calibrated phylogeny for squamate reptiles (lizards and snakes) based on 52 genes and 4162 species. Mol Phylogenet Evol 94:537–547. https://doi.org/10.1016/j.ympev.2015.10.009
Funding
Open access funding provided by CSIRO Library Services. DD was funded by a UQ Centennial Scholarship from The University of Queensland, a Research Training Program scholarship from the Australian Government Department of Education and Training, and a CSIRO Early Research Career Postdoctoral Fellowship from the Commonwealth Science & Industry Research Organisation. AA was funded by Pronaii 303045 (CONAHCYT, Mexico). JAPJ was funded by the Fundação de Amparo á Pesquisa do Estado de São Paulo under the grant 2018/25749-8. BGF was funded by Australian Research Council Grant DP190100304.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Darin Rokyta.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dashevsky, D., Harris, R.J., Zdenek, C.N. et al. Red-on-Yellow Queen: Bio-Layer Interferometry Reveals Functional Diversity Within Micrurus Venoms and Toxin Resistance in Prey Species. J Mol Evol 92, 317–328 (2024). https://doi.org/10.1007/s00239-024-10176-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-024-10176-x