Abstract
The evolutionary history of Class I aminoacyl-tRNA synthetases (aaRS) through the reconstruction of ancestral sequences is presented. From structural molecular modeling, we sought to understand its relationship with the acceptor arms and the tRNA anticodon loop, how this relationship was established, and the possible implications in determining the genetic code and the translation system. The results of the molecular docking showed that in 7 out 9 aaRS, the acceptor arm and the anticodon loop bond practically in the same region. Domain accretion process in aaRS and repositioning of interactions between tRNAs and aaRS are illustrated. Based on these results, we propose that the operational code and the anticodon code coexisted, competing for the aaRS catalytic region, while consequently contributed to the stabilization of these proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In biological systems, nucleic acids are responsible for storing genetic information that can later be translated into proteins. These two components, proteins and nucleic acids, are crucial for the proper functioning of living beings, and the correspondence between both is guaranteed by the genetic code. The translation system is one of the best-conserved processes throughout evolution, and its components are also well conserved. One of the central parts in the translation process, present in all domains of the tree of life, is aminoacyl-tRNA synthetases (aminoacyl-tRNA ligase, aaRS), enzymes responsible for the activation and binding of an amino acid to its cognate tRNA. The aminoacylation is catalyzed in two stages, where first the amino acid is activated by an ATP molecule and then immediately transferred to the 3′ end of the cognate tRNA (Crnković et al. 2019). For several decades, aaRSs have been the target of several hypotheses regarding its evolution, structural organization, and their relationship with the genetic code. aaRSs were divided into two structurally and evolutionary unrelated classes, each one containing 10 proteins (Eriani et al. 1990). This division was based on structural and functional correlations observed in the way the aaRS of each class aminoacylates the tRNA acceptor arm (Eriani et al. 1990). Posteriorly, phylogenetic studies supported this division and shed light on the factors that influenced the evolution and diversification of these two classes of proteins (Nagel and Doolittle 1991). It was proposed that in the beginning of the biological systems, the aminoacylation reaction could have been performed by RNA catalysts with low specificity. In fact, some studies support the idea that structural determinants for aminoacylation, the tRNA acceptor arm, and the anticodon loop were already present before the appearance of aaRS (Ribas De Pouplana and Schimmel 2001). During the evolutionary process, these catalytic RNAs would have been replaced by ancestral enzymes. After its emergence, the evolution of aaRS would be guided by subsequent events of gene duplication and diversification that improved specificity and catalytic capacity (Nagel and Doolittle 1995). Treangen and Rocha (2011) proposed that in addition to the processes of gene duplication, after the appearance of prokaryotes, the horizontal gene transfer (HGT) process played an important role in the expansion of these proteins. In addition to the HGT process, several other events, such as recombination, exchange, and fusion of domains, would also have been essential in structuring this family of proteins (Saha et al. 2009; Chaliotis et al. 2017). Regarding the evolutionary history and the selective pressures that may have guided aaRS, it can be inferred that the substrates must have had a strong influence, since, during the aminoacylation process, the protein acts independently of other cellular processes. In some scenarios, it is hypothesized that one of the determinants for the diversification of aaRS may have been its relationship with amino acids and that their chemical characteristics drove the evolution in the N-terminal region, responsible for the recognition and activation of amino acids in modern structures (Nagel and Doolittle 1995; de Farias and Guimarães 2008). In this context, some studies have proposed that the amino acids-binding domain would have appeared before the anticodon-binding domain; therefore, the primitive aaRS should be restricted to the catalytic portion, with the anticodon-binding domain being a later acquisition (Caetano-Anollés et al. 2013). In line with the proposals that relate the evolution of aaRS with their cognate amino acids, some models suggest that tRNA is the result of changes such as the addition and duplication of nucleotides, events that started from the CCA end of the acceptor arm, the region responsible for binding to the amino acid (Sun and Caetano-Anollés 2008). It was proposed that initially, there was an operational code (De Duve 1988), which would work only with the tRNA accepting arm portion. These suggestions are based on the observation that aminoacylation can occur without the need for the recognition of the anticodon (Hou and Schimmel 1988; Park and Schimmel 1988; Hamann and Hou 1995). However, some authors have proposed that the anticodon arm may have been the first part to emerge (Szathmáry 1999; de Farias 2013; de Farias et al. 2014, 2019). Farias et al. (2014) proposed a model in which tRNA and aaRS co-evolved from changes in the second base of the anticodon, a change that would alter the stereochemical properties of the anticodon, creating a selective pressure for the diversification of aaRS. In a conciliatory proposal, Farias et al. (2019) suggested that in its origin, the catalytic site of the aaRS could interact with the tRNA anticodon arm; thus, they presented a model for the origin of the genetic code in which the operational and the anticodon codes evolved at the same time.
Herein, we analyze the evolutionary history of Class I aaRSs through the reconstruction of ancestral sequences. From structural molecular modeling, we seek to understand its relationship with the acceptor arms and the tRNA anticodon loop, how this relationship was established, and the possible implications in determining the genetic code and the translation system.
In this context, three scenarios are possible: (i) only the acceptor arm of the tRNA interacts with the ancestral portion of the aaRS, thus, being the most primitive operational code; (ii) only the tRNA anticodon loop interacts with the ancestral portion of aaRS, and in this scenario, the genetic code would be more primitive, and (iii) Both the acceptor arm and the anticodon loop interact with the ancestral portion of aaRS, in this scenario, the operational code, and the genetic code would be contemporaneous and would play complementary roles. We also analyzed, from the reconstruction of ancestral sequences, the possible most primitive portions of Class I aaRS, as well as the observed interactions that could serve as a substrate for the construction of hypotheses about the evolutionary forces that led to the stabilization of the interaction between tRNAs and aaRS.
Material and Methods
Data Sources
We retrieved 4533 sequences of 9 aaRSs from GenBank (https://www.ncbi.nlm.nih.gov/genbank/). For each type of aaRS, redundant and duplicate sequences were removed, and the quality of the sequences were checked manually. The number of sequences for each aaRS was proportional between the 3 domains of life (Archaea, Bacteria, and Eukarya) to avoid bias in the ancestral reconstruction. For each aminoacyl-tRNA-synthetase, the following numbers of sequences were used: ArgRS (600), CysRS (600), GluRS (216), IleRS (600), MetRS (417), TyrRS (600), TrpRS (600), and ValRS (300).
Reconstruction of Ancestral Sequences
For the reconstruction of ancestral sequences, the alignment of the sequences was initially performed by MAFFT software (Version 7.0) using the BLOSUM45 substitution matrix with a 1.53 penalty for gap openings (https://mafft.cbrc.jp/alignment/server/index.html) (Katoh et al. 2019). A study comparing multiple alignment tools identified MAFFT as the best software for this type of approach (Vialle et al. 2018). To determine the best evolutionary model, the alignment was submitted to the ModelTest-NG software (Version 0.1.5) (Darriba et al. 2020) hosted on the Cipres server (https://www.phylo.org/). For the reconstruction of ancestral sequences, Hanson et al. (2010) showed that the best method for this kind of approach is the maximum likelihood (ML). Thus, phylogenetic trees were inferred using the IQ-Tree software (http://www.iqtree.org/) (Version 1.6.12) (Nguyen et al. 2015). Ancestral sequences were obtained using the MEGAX software (Version 10.0) using the ML method (Kumar et al. 2018). The alignments and trees used for the reconstruction of the ancestral sequences are available in the Supplementary Material. Probabilities of ancestral sites are available in Supplementary Data; probability values range from 0 to 1.
Molecular Modeling
The three-dimensional models of ancestral proteins were generated using the I-Tasser server (https://zhanglab.ccmb.med.umich.edu/I-TASSER/) (Roy et al. 2010). The structures were refined using the GalaxyWeb server (http://galaxy.seoklab.org/cgi-bin/submit.cgi?type=REFINE) (Ko et al. 2012). The structural alignment of three-dimensional models was performed with the help of the TM-Align server (https://zhanglab.ccmb.med.umich.edu/TM-align/) (Zhang 2005). For structural alignments, modern aaRS structures were chosen, obtained from the PDB database (rcsb.org) (Berman 2000), with the best resolution and associated with their cognate tRNA. The validation of the three-dimensional structure of the modeled proteins was done through the PROCHECK server (https://servicesn.mbi.ucla.edu/PROCHECK/) (Laskowski et al. 1993).
Molecular Docking
For molecular docking analyzes, three-dimensional structures of cognate tRNAs including the anticodon loop and the acceptor arm region were used. Molecular docking was performed between the acceptor arm and the anticodon stem with their respective ancestral protein. For molecular docking, the Hex 8.0.0 program (Ritchie and Venkatraman 2010) was used according to the following parameters: correlation type—shape + electro + DARS, FFT mode—5 dimensions, sampling method—range angles, post-processing—DARS minimization. The other parameters were used in the standard configuration.
Results and Discussion
Structural Evolution of Class I aaRS
The ancestral sequences were generated from modern sequences obtained for each of the 9 aaRSs. Two cutoff points were used for the reconstruction of ancestral sequences. The first cutoff point with complete deletion returns an ancestral sequence with the positions aligned in all modern sequences (100%), while the cutoff point of partial deletion corresponds to the positions aligned in 99% of modern sequences. These two cutoff points are used to observe two distinct moments in the aaRS evolution process, with the sequences obtained with 100% of the aligned positions hypothesized as older and those with 99% as having already gone through a stage of incorporation of new parts in the protein. For each cutoff point, two ancestral sequences were generated, resulting in 4 ancestral sequences for each aaRS. The Class I aminoacyl-tRNA synthetases are highly conserved proteins, with small variations throughout the domains of life, having a catalytic region that binds to the tRNA acceptor arm and an anticodon-binding region, also having two structurally conserved motifs, HIGH and KMSKS, in addition to a Rossmann-fold domain present in the catalytic site of the enzymes (Cusack et al. 1991). The Class I aaRS couples the aminoacyl to the 2′-hydroxyl of the tRNA acceptor stem, while, in Class II, the aminoacyl is attached to the 3′-hydroxyl of the tRNA.
For Class I, it was possible to analyze 9 aaRS according to data availability and biological limitations. In the case of LysRS, despite being found in both Classes I and II, in this work, we considered it as a member of class II because it is present in most prokaryotes and eukaryotes, unlike LysRS class I, which is only found in archaea and some bacteria (Ambrogelly et al. 2002). In the case of GlnRS, this is absent in most prokaryotes, as it involves a gene resulting from a process of duplication and subsequent horizontal gene transfer (Lamour et al., 1994; Brown and Doolittle, 1999). In this circumstance, the Gln-tRNAGln formation process is conducted with the aid of GluRS in a pathway involving the amidotransferase enzyme (Wu et al., 2009). Among the structures of 9 aaRS obtained from the ancestral sequences from complete deletion (ArgRS, CysRS, GluRS, IleRS, LeuRS, MetRS, TyrRS, TrpRS, ValRS), 8 of them had the catalytic region conserved, and 6 out of 9 aaRSs analyzed, also had the region of connection to the anticodon loop conserved. The only exception was TrpRS, which had only the anticodon loop conserved.
These results corroborate previous studies that suggest the catalytic site as the first domain of aaRS to emerge, with the other domains having subsequently emerged from that catalytic region (Caetano-Anollés et al. 2013). Previous studies also have shown that aaRS when the anticodon-binding sites were deleted, maintained the ability to catalyze the aminoacylation process, even though this affected aaRS's ability to discriminate non-cognate tRNAs (Schwob and Söll 1993). The structures generated from partial deletion sequences (99%) show an increase in the complexity of aaRS, with the appearance of new structures. The increase in the structural complexity of the aaRS may have been due to domain recombination processes, as well as, through the extension by incorporation of new sequences in the original information. The data on the most primordial structure of aaRS, suggest that during the evolutionary process, there was the addition of structural portions that led to greater protection of the catalytic site, which may have led to an increase of efficiency in the aminoacylation process.
Coevolution Between aaRS Class I and tRNA
Molecular docking was performed between the three-dimensional structures obtained for ancestral sequences generated by complete deletion and partial deletion with the anticodon loop and the cognate tRNA acceptor arm of each aaRS. For docking, the entire surface of the proteins was considered, so that the acceptor arm and the anticodon loop would find the connection region with the lower energy. The binding energies are compiled in Table 1. The results show that both the anticodon loop and the tRNA acceptor arm, with only two exceptions (LeuRS and ValRS with the tRNA acceptor arm), linked with lower free energy in the structures of the complete deletion, while in the structures of partial deletion, there was an increase in the free energy (Table 1).
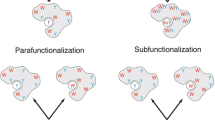
The lower free energy between aaRS with the tRNA acceptor arm and the anticodon loop lead us to hypothesize that, initially, tRNA and aaRS were more strongly linked. In a study evaluating the structural evolution of GlyRS, Farias et al. (2019) suggested that the anticodon loop worked primarily as a conformational stabilizer for the GlyRS catalytic site (Farias et al. 2019). Our results suggest a similar behavior, with a lower binding-free energy between the ancestral structure obtained by complete deletion and the tRNA, indicating that in this evolutionary stage, this interaction could have acted only to increase the stability of both molecules (Ribas de Pouplana 2020). As the protein became more complex, due to an exaptation process, the tRNA/aaRS complex began to work on the aminoacylation process, in a process of coevolution with the assembly of the primitive translation system. Another characteristic observed that reinforces the idea that tRNAs and aaRS may have initially acted to increase the stabilization of these molecules, is the preference for binding between tRNAs and aaRS in parts of the structure with low stability or unstructured properties (Fig. 1).
With the addition of structural layers in the aaRS, leading to greater protection of the catalytic site, the tRNAs modify the interaction sites, and at this point, the aminoacylation process must have started (Fig. 2). This scenario is in line with hypotheses that suggest that the primitive ribosome worked by producing peptides at random, without an established genetic code.
Domain accretion process in aaRS (CysRS) and repositioning of interactions between tRNAs and aaRS. In red, the structure obtained for the ancestral sequence determined by complete deletion. In blue, the structure obtained for the sequence determined by partial deletion, and in green, the structure of the modern protein. In black, the anticodon loop; in yellow, the acceptor arm; and in gray, the modern tRNA. With the increase in the complexity of aaRS, the acceptor arm and the anticodon loop reposition their interaction sites. The final complexification of aaRS was accompanied by the fusion of the acceptor arm with the anticodon loop, giving rise to the modern tRNA (Color figure online)
Here, we also evaluated the binding of the tRNA acceptor arm and the anticodon loop with ancestors aaRS. Several works deal with the origin of both regions and discuss which region first emerged in the tRNA. Currently, the most accepted hypothesis suggests an operational code, where the acceptor's arm is envisaged as the first region to emerge in the tRNA. This hypothesis is supported by numerous studies (Schwob and Söll 1993; Hou and Schimmel 1988; Park and Schimmel 1988), while other studies suggest the initial appearance of the anticodon loop (Szathmáry 1999). The result of the molecular docking showed that in 7 out 9 aaRS, the acceptor arm and the anticodon loop bond practically in the same region (Fig. 1). Based on these results, we propose that the operational code and the anticodon code coexisted, competing for the aaRS catalytic region, while consequently acting on the stabilization of these proteins. The aaRS catalytic region then served as a condition for fusion events between the acceptor arm and the anticodon loop to occur, events similar to those hypothesized by Root-Bernstein and collaborators (2016). The accommodation of the acceptor arm and the anticodon loop in the same region may also reflect a primitive structure of tRNAs, as suggested by Möller and Janssen (1992), which based on the statistical analysis of 1400 tRNA sequences, proposed that the acceptor arm and the anticodon loop arose from a single region derivate from positions 3–5 of primitive tRNAs, thus, having a similar chemical nature, which can explain the similarity between the anticodon loop and the acceptor arm of the tRNAs. This increase in the complexity of the tRNAs up to the current structures was accompanied by an increase in the free energy between the aaRS and the tRNAs (Table 1), guaranteeing certain plasticity in the structural relationship of both, and which could also reflect the co-optation of the tRNA so that it started to participate in the flow of information. This structural expansion of the tRNAs would also have acted as a selective force for the structural diversification of the aaRS, which after being established conformationally, would be under new selective pressure from changes in the second base of the anticodon, as proposed by Farias and collaborators (de Farias 2013; de Farias et al. 2014). Based on the results presented, we propose that the complexity of the tRNA and the aaRS emerged in a process of coevolution, with the tRNA initially acting as a conformational stabilizer of the aaRS, being subsequently co-opted to act in the flow of information and in the establishment of the process of translation and the genetic code.
Conclusion
The results presented were obtained by computational methods, and thus, we must keep in mind that analyses of this type present uncertainties and cannot be understood with the same degree of accuracy as the results obtained by experimentation; however, we believe that they can raise questions and hypotheses that can contribute to the development of the research field in the origin of life. The evolution of the aminoacyl-tRNA synthetases is an open field and many hypotheses have been postulated. The Rodin and Rodin (2008) hypothesis postulates that Class I and Class II aaRSs originated from a single ancestral gene. The two extant classes were coded from opposite strands of a double-stranded nucleic acid (Rodin et al. 2009). The latter is reinforced with the fact that Class I and Class II synthetases bind to the opposite sides of the tRNA, and hints with the idea that both classes evolved together. However, this hypothesis demands for an advance operational genetic code. This would imply that the evolution of the genetic code was uncoupled from the evolution of the aaRSs. Our docking experiments indicate the coupling of the synthetase evolution with that of the tRNA. Assuming the primordial tRNAs began as short RNAs, expanding to an acceptor stem helix and finally to a structure including the anticodon stem and loop; this, in turn, requires that the aaRSs shifted from reading an operational code in the acceptor arm to reading of the anticodon. Our structural analyses of the interactions of the tRNA with the contacting domains of the aminoacyl-tRNA synthetases class I led us to postulate a model of domain accretion process in aaRS and repositioning of interactions between tRNAs and aaRS, which allowed the interaction of the aaRS with both the acceptor stem and the anticodon.
References
Ambrogelly A, Korencic D, Ibba M (2002) Functional annotation of class I lysyl-tRNA synthetase phylogeny indicates a limited role for gene transfer. J Bacteriol 184(16):4594–4600. https://doi.org/10.1128/JB.184.16.4594-4600.2002
Berman HM, Westbrook JD, Feng Z, Gilliland GL, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Brown JR, Doolittle WF (1999) Gene descent, duplication, and horizontal transfer in the evolution of glutamyl- and glutaminyl-tRNA synthetases. J Mol Evol 49(4):485–495. https://doi.org/10.1007/pl00006571
Caetano-Anollés G, Wang M, Caetano-Anollés D (2013) Structural phylogenomics retrodicts the origin of the genetic code and uncovers the evolutionary impact of protein flexibility. PLoS ONE. https://doi.org/10.1371/journal.pone.0072225
Chaliotis A, Vlastaridis P, Mossialos D, Ibba M, Becker HD, Stathopoulos C, Amoutzias GD (2017) The complex evolutionary history of aminoacyl-tRNA synthetases. Nucleic Acids Res 45(3):1059–1068. https://doi.org/10.1093/nar/gkw1182
Crnković A, Vargas-Rodriguez O, Söll D (2019) Plasticity and constraints of tRNA aminoacylation define directed evolution of aminoacyl-tRNA synthetases. Int J Mol Sci. https://doi.org/10.3390/ijms20092294
Cusack S, Härtlein M, Leberman R (1991) Sequence, structural and evolutionary relationships between class 2 aminoacyl-tRNA synthetases. Nucleic Acids Res 19:3489–3498. https://doi.org/10.1093/nar/19.13.3489
Darriba D, Posada D, Kozlov AM, Stamatakis A, Morel B, Flouri T (2020) Modeltest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37:291–294. https://doi.org/10.1093/molbev/msz189
de Duve C (1988) The second genetic code. Nature 333:117–118. https://doi.org/10.1038/333117a0
de Farias ST, Guimarães RC (2008) Aminoacyl-tRNA synthetase classes and groups in prokaryotes. J Theor Biol 250:221–229. https://doi.org/10.1016/j.jtbi.2007.09.025
de Farias ST, do Rêgo TG, José MV (2014) Evolution of tranfer RNA and the origin of the translation system. Front Genet 5:1–4. https://doi.org/10.3389/fgene.2014.00303
de Farias ST, Antonino D, Rêgo TG, José MV (2019) Structural evolution of Glycyl-tRNA synthetases alpha subunit and its implication in the initial organization of the decoding system. Prog Biophys Mol Biol 142:43–50. https://doi.org/10.1016/j.pbiomolbio.2018.08.007
De Farias ST (2013) Suggested phylogeny of tRNAs based on the construction of ancestral sequences. J Theor Biol 335:245–248. https://doi.org/10.1016/j.jtbi.2013.06.033
De Pouplana LR, Schimmel P (2001) Operational RNA code for amino acids in relation to genetic code in evolution. J Biol Chem 276:6881–6884. https://doi.org/10.1074/jbc.R000032200
Eriani G, Delarue M, Poch O, Gangloff J, Moras D (1990) Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347:203–206. https://doi.org/10.1038/347203a0
Hamann CS, Hou YM (1995) Enzymic aminoacylation of tRNA acceptor stem helices with cysteine is dependent on a single nucleotide. Biochemistry 34:6527–6532. https://doi.org/10.1021/bi00019a034
Hanson-Smith V, Kolaczkowski B, Thornton JW (2010) Robustness of ancestral sequence reconstruction to phylogenetic uncertainty. Mol Biol Evol 27:1988–1999. https://doi.org/10.1093/molbev/msq081
Hou Y-M, Schimmel P (1988) A simple structural feature is a major determinant of the identity of a transfer RNA. Nature 333:140–145. https://doi.org/10.1038/333140a0
Katoh K, Rozewicki J, Yamada KD (2018) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166. https://doi.org/10.1093/bib/bbx108
Ko J, Park H, Heo L, Seok C (2012) GalaxyWEB server for protein structure prediction and refinement. Nucleic Acids Res 40:294–297. https://doi.org/10.1093/nar/gks493
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lamour V, Quevillon S, Diriong S, N’Guyen VC, Lipinski M, Mirande M (1994) Evolution of the Glx-tRNA synthetase family: the glutaminyl enzyme as a case of horizontal gene transfer. Proc Natl Acad Sci USA 91(18):8670–8674. https://doi.org/10.1073/pnas.91.18.8670
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/s0021889892009944
Möller W, Janssen GMC (1992) Statistical evidence for remnants of the primordial code in the acceptor stem of prokaryotic transfer RNA. J Mol Evol 34:471–477. https://doi.org/10.1007/BF00160461
Nagel GM, Doolittle RF (1991) Evolution and relatedness in two aminoacyl-tRNA synthetase families. Proc Natl Acad Sci USA 88:8121–8125. https://doi.org/10.1073/pnas.88.18.8121
Nagel GM, Doolittle RF (1995) Phylogenetic analysis of the aminoacyl-tRNA synthetases. J Mol Evol 40:487–498. https://doi.org/10.1007/BF00166617
Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Park SJ, Schimmel P (1988) Evidence for interaction of an aminoacyl transfer RNA synthetase with a region important for the identity of its cognate transfer RNA. J Biol Chem 263:16527–16530. https://doi.org/10.1016/s0021-9258(18)37421-0
Ribas de Pouplana L (2020) The evolution of aminoacyl-tRNA synthetases: from dawn to LUCA. Enzymes 48:11–37. https://doi.org/10.1016/bs.enz.2020.08.001
Ritchie DW, Venkatraman V (2010) Ultra-fast FFT protein docking on graphics processors. Bioinformatics 26:2398–2405. https://doi.org/10.1093/bioinformatics/btq444
Rodin SN, Rodin AS (2008) On the origin of the genetic code: signatures of its primordial complementarity in tRNAs and aminoacyl-tRNA synthetases. Heredity 100:341–355. https://doi.org/10.1038/sj.hdy.6801086
Rodin AS, Rodin SN, Carter CW (2009) On primordial sense-antisense coding. J Mol Evol 69:555–567. https://doi.org/10.1007/s00239-009-9288-4
Root-Bernstein R, Kim Y, Sanjay A, Burton ZF (2016) tRNA evolution from the proto-tRNA minihelix world. Transcription 7:153–163. https://doi.org/10.1080/21541264.2016.1235527
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738. https://doi.org/10.1038/nprot.2010.5
Saha R, Dasgupta S, Basu G, Roy S (2009) A chimaeric glutamyl:glutaminyl-tRNA synthetase: implications for evolution. Biochem J 417:449–455. https://doi.org/10.1042/BJ20080747
Schwob E, Soll D (1993) Selection of a “minimal” glutaminyl-tRNA synthetase and the evolution of class I synthetases. EMBO J 12:5201–5208. https://doi.org/10.1002/j.1460-2075.1993.tb06215.x
Sun FJ, Caetano-Anollés G (2008) The origin and evolution of tRNA inferred from phylogenetic analysis of structure. J Mol Evol 66:21–35. https://doi.org/10.1007/s00239-007-9050-8
Szathmáry E (1999) The origin of the genetic code: Amino acids as cofactors in an RNA world. Trends Genet 15:223–229. https://doi.org/10.1016/S0168-9525(99)01730-8
Treangen TJ, Rocha EPC (2011) Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001284
Vialle RA, Tamuri AU, Goldman N (2018) Alignment modulates ancestral sequence reconstruction accuracy. Mol Biol Evol 35:1783–1797. https://doi.org/10.1093/molbev/msy055
Wu J, Bu W, Sheppard K, Kitabatake M, Kwon ST, Söll D, Smith JL (2009) Insights into tRNA-dependent amidotransferase evolution and catalysis from the structure of the Aquifex aeolicus enzyme. J Mol Biol 391(4):703–716. https://doi.org/10.1016/j.jmb.2009.06.014
Zhang Y, Skolnick J (2005) TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res 33:2302–2309. https://doi.org/10.1093/nar/gki524
Acknowledgements
MVJ was funded by Dirección General de Asuntos del Personal Académico (DGAPA) and Universidad Nacional Autónoma de México, UNAM (PAPIIT-IN201019).
Funding
MVJ was financially supported by PAPIIT-IN201019; UNAM; México.
Author information
Authors and Affiliations
Contributions
Conceptualization: STF, PHL, and MVJ; Methodology: STF and PHL; Formal Analysis: STF and PHL; Investigation: STF and PHL; Resources: MVJ; Writing: Original and draft Preparation: STF, PHL and MVJ.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Informed Consent
All authors consented.
Additional information
Handling editor: Ulrich Muller.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dantas, P.H.L.F., José, M.V. & de Farias, S.T. Structural Computational Analysis of the Natural History of Class I aminoacyl-tRNA Synthetases Suggests their Role in Establishing the Genetic Code. J Mol Evol 89, 611–617 (2021). https://doi.org/10.1007/s00239-021-10029-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10029-x