Abstract
Circadian rhythms not only influence the overall daily routine of organisms but also directly affect life activities to varying degrees. Circadian locomotor output cycle kaput (Clock), the most critical gene in the circadian rhythm feedback system, plays an important role in the regulation of biological rhythms. Here, we aimed to elucidate the evolutionary history of the clock gene family in a taxonomically diverse set of vertebrates, providing novel insights into the evolution of the clock gene family based on 102 vertebrate genomes. Using genome-wide analysis, we extracted 264 clock sequences. In lobe-finned fishes and some basal non-teleost ray-finned fishes, only two clock isotypes were found (clock1 and clock2). However, the majority of teleosts possess three clock genes (two clock1 genes and one clock2 gene) owing to extra whole-genome duplication. The following syntenic analysis confirmed that clock1a, clock1b, and clock2 are conserved in teleost species. Interestingly, we discovered that osteoglossomorph fishes possess two clock2 genes. Moreover, protein sequence comparisons indicate that CLOCK protein changes among vertebrates were concentrated at the N-terminal and poly Q regions. We also performed a dN/dS analysis, and the results suggest that clock1 and clock2 may show distinct fates for duplicated genes between the lobe-finned and ray-finned fish clades. Collectively, these results provide a genome-wide insight into clock gene evolution in vertebrates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Circadian rhythms, biological rhythms with a period of approximately 24 h, are an adaptation of life to the light and dark cycles of the Earth (Pittendrigh 1993; Dunlap 1999; Bell-Pedersen et al. 2005). According to Hall, Rosbash, and Young, circadian rhythmicity is regulated, controlled, and maintained by a transcription/translation negative feedback loop composed of a group of genes and proteins that are highly conserved throughout evolution (Young 2000; Reppert and Weaver 2001; Panda et al. 2002a, b; Panda et al. 2002a, b; Hardin 2005; Buhr and Takahashi 2013). They were awarded the Nobel Prize in Physiology or Medicine in 2017 for their work on this mechanism. In general, the circadian rhythm machinery is composed of three portions: an input channel, core oscillator, and output channel (Reddy et al. 1984; Allada and White 1998; Brown et al. 2005; Yu and Paul 2006; Schmutz et al. 2010). For simplicity, circadian rhythms are driven by external environmental factors such as light and temperature through an input channel. In the core oscillator, a series of reactions translate these signals into physiological, behavioral, and hormonal rhythms through an output channel (Pando and Sassone-Corsi 2002). CLOCK, which is encoded by the clock gene, is a vital participant in the core oscillator with a highly conserved basic helix-loop-helix (bHLH)-period-arylhydrocarbon receptor nuclear translocator-single mind (PAS) domain (Vitaterna et al. 1994; Antoch et al. 1997; King et al. 1997a, b). The first vertebrate clock was identified in a mouse (Mus musculus) using forward genetic mutagenesis screens, molecular cloning, and biochemical, physiological, and behavioral characterization (Reddy et al. 1984; Konopka and Benzer 1971; Bargiello et al. 1984; Takahashi et al. 1994). The clock gene in mice is located in the middle of Chr5 and includes 24 exons of approximately 100 kb (King et al. 1997a, b). In 1998, the fruit fly clock gene was screened and named Jrk. Subsequently, the gene was rechristened dClock because it was highly similar to the mouse clock gene (Allada et al. 1998). Based on this previous study, the human clock gene was cloned in 1999. The hclock gene lies on the long arm of Chr4 with 20 exons and encodes a protein of 846 amino acids (aa) (Shearman et al. 2000).
Moreover, clock plays an important role in physiology. In many species, the circadian clock is expressed throughout the body, including peripheral tissues (Della Ragione F et al. 2005; Dibner-C et al. 2010; Markowska M et al. 2017). Therefore, tissues with circadian clock properties can be divided into central clocks and peripheral clocks according to their regulatory function levels. Periphery clocks are able to maintain their own clock, while simultaneously receiving and sending input and output to the suprachiasmatic nucleus (SCN). For vertebrates, the pacing point of the biological clock is the SCN and the pineal gland; the pineal gland in fish, amphibians, reptiles, and birds is sensitive to light, besides it also controls the elimination melatonin produces other rhythms in addition to the circadian rhythm, such as body temperature and eating (Menaker M et al. 1983; Saha S et al. 2019). In mammals, the pineal gland and the suprachiasmatic nucleus jointly control the rhythm, but there are many other evidences suggest that there are other pacing points, such as the retina (Lowrey et al. 2000; Mohawk et al. 2012; Touitou Y et al. 2020). Circadian rhythms can be regulated by changing the time phase and altering the expression of clock under various light, temperature, and feeding conditions. Knockout of clock causes biological circadian rhythm disorder or loss, increased appetite, excessive obesity, and a tendency for hyperlipidemia in mice (Turek et al. 2005). Moreover, the circadian rhythm in mice is accelerated when clock transcription is artificially increased. More than 100 metabolism-related genes show a circadian rhythm in liver tissue, and expression of all these genes is reduced to varying degrees after clock knockout (Okano et al. 2001; Oishi et al. 2003). In addition, other studies have shown that the expression of clock in the pineal gland of birds is similar to that in the SCN of mammals (Woller and Gonze 2013). In our bodies, dawn and dusk coordinate or entrain the circadian clock through neural pathways associating the retina to the SCN so that the master clock and its output rhythms do not drift from 24 h but keep pace with the solar day (Clayton et al. 2001; Panda et al. 2002a, b). Transient disruption of circadian timing such as that from a long-distance flight may lead to jet lag, and chronic alteration of the central clock mechanism in shift workers may contribute to poor health and sleep disorders. In brief, the circadian clock in organisms drives daily variations in many physiological and behavioral processes, including the sleep–wake cycle and body temperature, hormone levels, cognition, and memory (Yu and Paul 2006).
Wang first discussed clock evolution in a research article on six teleosts (Wang 2008). Toloza-Villalobos et al. described the diversification of six circadian clock gene families in eight teleost fishes (Toloza-Villalobos et al. 2015). Comparative studies of the clock gene family in vertebrates have mainly focused on the limited number of species included in previous studies; large-scale studies of the entire family of clock genes in vertebrates have not been reported. With the development of high-throughput sequencing and the availability of genome data in many databases, we can explore evolution of the clock gene family in more detail.
In our current study, we analyzed 102 genomes of vertebrate species. Among fishes, several species with particular living habitats, such as cavefishes (Astyanax mexicanus, Sinocyclocheilus anshuiensis, and Triplophysa rosa) and mudskippers (Periophthalmus magnuspinnatus and Boleophthalmus pectinirostris), those in a key phylogenetic position such as Sarcopterygii (Latimeria chalumnae), Chondrichthyes (Callorhinchus milii), Actinopterygii (Lepisosteus oculatus, Scleropages formosus, and Paramormyrops kingsleyae), and polyploid fishes (three Sinocyclocheilus species including S. grahami, S. rhinoceros, and S. anshuiensis), as well as two tetraploid salmonid species, Salmo salar and Oncorhynchus mykiss, were examined. Using a large number of species, this study attempted to address the following core questions: (1) what the general evolutionary pattern of clock genes is in vertebrates; (2) whether certain branches of vertebrates show evidence of evolution that is different from that in other branches; and (3) whether there are different evolutionary patterns for clock in Actinopterygii after fish-specific genome duplication (FSGD). This study contributes to a better understanding of the molecular evolution of clock in vertebrates.
Materials and Methods
Gene Acquisition and Identification
In total, we analyzed 102 vertebrate genomes with relatively high-quality assembled sequences. All of these were downloaded from the National Center for Biotechnology Information (NCBI) (Table S1), with the exception of the genome of T. rosa, which was assembled in our laboratory. Each genome sequence was initially aligned and subjected to a tBLASTn (version 2.6.0 + , NCBI, Bethesda, MD, USA) search with an E-value of 1e–5, employing the protein-nucleotide alignment strategy using the CLOCK protein of human, zebrafish, chicken, and anole lizard as queries (more query sequences in Table S2) (Matsuda et al. 2013). The alignment results were analyzed using a Perl script to obtain the best hit for each alignment. Following this, we extended 5–10 kbp upstream and downstream of the best hits to acquire candidate sequence segments that contained complete genes. Finally, the FGENESH + program was employed to predict the full-length of clock genes (Salamov and Solovyev 2000).
Sequence Analysis and Structural Characterization
All high-confidence CLOCK sequences were submitted to the MEME program to identify conserved motifs in CLOCK (version 4.11.2, http://alternate.meme-suite.org/tools/meme) with the following parameters: any number of repetitions, maximum of 10 misfits, and an optimum motif width of 6–50 aa residues and threshold is lower than the e-0 range (Bailey 2009). The exon–intron structural information for clock was extracted using a Perl script and identified in the Gene Structure Display Server (GSDS, http://gsds.cbi.pku.edu.cn/) (Hu et al. 2015). Finally, we analyzed the phylogenetic relationships, gene structures, and conserved motifs of clock genes in TBtools (Chen et al. 2018).
Phylogenetic Analysis and Structural Differences Between CLOCK Proteins
Phylogenetic analysis was performed using the predicted protein sequences encoded by clock genes. A multiple codon-base alignment was assembled using MAFFT (version 7.149b) with—auto strategy, and the alignment was adjusted in Gblocks (version 0.9b, Jose Castresana) (Talavera and Castresana 2007; Katoh and Standley 2013). Subsequently, we predicted the best nucleotide substitution model for the data using IQtree (version 1.6.1) ModelFinder under Bayesian information criterion (Nguyen et al. 2015). The parameters in the best nucleotide substitution model, TVMe + R6, were applied to IQtree to construct phylogenetic trees using the maximum likelihood (ML) method and 1000 replicates to calculate branch supports.
Detection of Differences Between CLOCK Protein Sequences
We first selected human CLOCK PAS-A (6QPJ) and CLOCK:BMAL1 transcriptional activator complex (4F3L) as templates from the Protein Data Bank (Huang et al. 2012; Wang et al. 2013). Because the addition of more species left the display of results mostly unchanged, we selected the following five species to analyze the CLOCK protein sequence. Representative sequences from zebrafish (Danio rerio), anole lizard (Anolis carolinensis), chicken (Gallus gallus), house mouse (Mus musculus), and tropical clawed frog (Xenopus tropicalis) were aligned with the human CLOCK protein. Previous studies have reported that teleosts such as zebrafish possess two clock1 genes (clock1a and clock1b) (Wang 2008); thus, we chose five relative well-annotated species from the teleosts to identify the differences between CLOCK1a and CLOCK1b. Finally, to obtain the evolutionary profile of BMAL1 in a CLOCK:BMAL1 heterodimer, the BMAL1 sequences from the species mentioned above were analyzed as well. CLOCK and BMAL1 interface analysis were computed on Discovery studio 3.1 (Accelrys, San Diego, CA, USA). Jalview (version 2, http://www.jalview.org/) and PyMOL (version 2.4, https://pymol.org/2/) were applied to visualize the alignment results and predict the secondary structure using Jpred Secondary Structure Prediction module.
Synteny Analysis and Conservation Strategy Identification
To estimate the conservation and homology of clock genes, we investigated several genes located upstream and downstream of clock within lobe-finned fish and ray-finned fish genomes in Ensembl, and verified these synteny analyses in Genomicus mapper; human clock1 and clock2 were used as anchor sites (https://www.genomicus.biologie.ens.fr/genomicus-98.01/cgi-bin/search.pl). Orthologous genes in the regions of human clock1 and clock2 loci (approximately 5–10 Mb) were searched in most lobe-finned fish clades. Finally, we selected four genes (kit, kdr, cep135, and aasd) and another four genes (chst10, lonrf2, rpl31, and cont11) upstream and downstream of each clock paralog in the lobe-finned fish clade using Ensembl datasets. In ray-finned fish, nine gene (cep135, clock1a, teme165, tyrp1b, clock1b, kitb, zc3h12b, clock2, and brwd3) locations were conserved. The eight genes in humans and chickens mentioned above were used as reference sequences for mammals and birds, respectively. Sequences of these eight genes in anole lizard and tropical clawed frog were downloaded as queries for reptiles and amphibians, respectively. For ray-finned fish, nine genes (cep135, clock1a, teme165, tyrp1b, clock1b, kitb, zc3h12b, clock2, and brwd3) compared with orthologous genes in zebrafish (D. rerio) were used as the query sequences to search for syntenic locations in genomes. Then, the strategy of alignment to nucleotides was employed to examine these extracted syntenic genes in lobe-finned fish, cartilaginous fish, and ray-finned fish. The tBLASTn results were further analyzed using a Perl script to obtain the best hits.
dN/dS Analysis
To understand clock substitution changes in vertebrates, we selected 29 species, including X. tropicalis, Homo sapiens, Mus musculus, and D. rerio model species and representative species from the other classes represented, to estimate the substitution rates for clock CDS sequences. We constructed a ML tree (in supplementary materials) based on these 29 species and estimated the dN/dS [ratio of nonsynonymous substitutions (dN) and synonymous substitutions (dS)] of each clock gene by the codeml modules in PAML package (version 4.9) (Yang 2007). The topology of the ML tree is consistent with Fig. 3, and thus we used the same clade mark. To compare the selection between the two principal clades (clock1 and clock2), a “two-ratio model” was used. We labeled the two principal clades (with different # labels in the root of each clade) and run a branch-specific model. Following this, we compared the likelihood of this model with the one-ratio model assuming a global dN/dS along the whole tree (model = 0 in control file) through a Likelihood Ratio Test (LRT).
To compare the selection intensity between lobe-finned and ray-finned fishes in clock genes, we performed the following analysis. We implemented the two-ratio model described above on a phylogenetic tree of clock2 sequences. Then we labeled lobe-finned (clade 1) and ray-finned (clade 2) with different # labels, and compared the likelihood of this model with one-ratio model to assess whether dN/dS is significantly different between species groups. For clock1 gene, we implemented a three-ratio model on a phylogenetic tree of clock1 sequences. We labeled: (a) lobe-finned clock1 (clade 3), (b) ray-finned clock1a (clade 4) and (c) ray-finned clock1b (clade 5), and compared the likelihood of this model with one-ratio model to assess whether dN/dS is significantly different between clock1 clades.
To test whether the functional and structural divergence of clock genes after gene duplication was driven by positive selection (particularly the residue H84), branch and branch-site models were used to detect accelerated evolution and positive selection on specific (labeled) branches (Bielawski and Yang 2003). On clock2 phylogeny, we labeled ancestral branches of lobe-fined and ray-fined fishes to identify adaptive changes after split of these two groups and compared the likelihood of this model with that of one-ratio model. The same for clock1, labeled the ancestral branches of clade 3 and superclade 4–5.
To test for positive selection after clock1 gene duplication in ray-finned fishes, a phylogeny with clades 4 and 5 was used, and labeled the ancestral branches of the two clades. L. oculatus and S. formosus were used as outgroup here.
Results
Identification and Phylogenetic Analysis of Vertebrate Clock Genes
We identified a total of 264 clock genes from 102 genomes of vertebrates, including 10 mammals, 23 birds, 10 reptiles, 2 amphibians, and 57 fish species (see more details in Table S3). The accession numbers of these genomes are provided in Table S1. The corresponding CLOCK protein sequences contained approximately 850 aa each, with different species showing only a few differences.
For fish in the lobe-finned clade, we acquired only two clock genes. This suggests that the clock gene family is highly conserved in the lobe-finned fish clade. Previous studies have identified three clock members (defined as clock1a, clock1b, and clock2) in teleosts such as zebrafish, medaka, and Fugu (Wang 2008). Results of our current study support the finding that most teleosts with FSGD possess three clock isotypes (clock1a, clock1b, and clock2), whereas Osteoglossiformes fishes such as P. kingsleyae and S. formosus possess two clock2 genes (clock2ba and clock2bb; clock2a was lost in the fish common ancestor). In addition, certain tetraploid fishes that have undergone more than one whole-genome duplication (WGD) event, such as S. salar and O. mykiss, possess four clock genes, and three Sinocyclocheilus fishes, namely S. grahami (Sg), S. rhinoceros (Sr), and S. anshuiensis (Sa), contain six clock genes. Nonetheless, a few teleosts such as Gadus morhua and Gasterosteus aculeatus were found to have two clock sequences each (Fig. 1). Furthermore, the lobe-finned fish Latimeria chalumnae, basal non-teleost ray-finned fish Lepisosteus oculatus, and cartilaginous fish Callorhinchus milii possess two clock genes as expected. To visualize the gene structure, we used local MEME prediction and TBtools for modification (Fig. 2). The results show that CLOCK1 of most vertebrate species contain around 14–18 motifs (Figure S1). Interestingly, both mudskippers possessed an extra motif 9 (red box shown in Fig. 2 & Figure S1).
Phylogenetic relationships, gene structures, and conserved motifs in clock genes. a Phylogenetic tree of 42 clock genes. The unrooted maximum likelihood phylogenetic tree was constructed in IQtree using full-length nucleotide sequences of 42 clock genes, with 1000 bootstrap test replicates. b Distribution of conserved motifs in clock genes. Ten putative motifs are indicated by different colored boxes (see more details in supplementary materials). c Exon/intron organization of clock genes. Yellow boxes represent exons, and black lines with the same lengths represent introns. The regions upstream and downstream of clock genes (untranslated regions, UTR) are indicated by blue boxes. The lengths of exons can be inferred by the scale at the bottom. For additional information on species motifs, see Figure S1 (Color figure online)
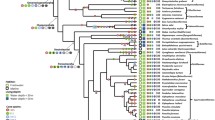
To understand the relationships among clock genes in vertebrates, we constructed a robust phylogenetic tree using the ML method from the 264 predicted clock sequences, with the sea lamprey (Petromyzon marinus) clock gene as the outgroup (Fig. 3). The sea lamprey is a relatively primitive species, a member of the Cyclostomata. It is the closest group in evolutionary relationship with other jawed vertebrates and shares the closest common ancestor. According to the well-supported phylogenetic topology, the 264 genes were classified into two distinct subfamilies (clock1 and clock2), which were further split into five sub-lineages of two lobe-finned fish clades (clade 1 and clade 3) and three ray-finned fish clades (clade 2, clade 4, and clade 5) because of FSGD. All clock genes from the lobe-finned fish clade formed a sister group with those of the ray-finned fish species, indicating consistency of the topology. Based on these results, combined with the domain structures predicted in MEME and gene structures predicted by annotation files (gff3), we determined that the clock gene family has two lineages: clock1 and npas2, which we renamed clock2, consistent with other reports (Wang 2008; Bailey et al. 2009).
Maximum likelihood phylogram depicting relationships among clock sequences from 102 representative vertebrates Phylogenetic reconstructions were based on the coding sequences of clock genes. A lamprey sequence was used as the outgroup. Different colors represent different clades: the blue circle represents clock1, and the yellow circle represents clock2. The same clade is shown in the background color. Clade 1: lobe-finned fish clock2; Clade 2: ray-finned fish clock2; Clade 3: lobe-finned fish clock1; Clade 4: ray-finned fish clock1b; Clade 5: ray-finned fish clock1a (Color figure online)
Protein Structure Comparison and Sequence Analysis
To further characterize clock1 and clock2, we chose one or two representative species from each group and aligned sequences of proteins encoded by these genes (Fig. 4). The comparison showed that CLOCK1 and CLOCK2 are highly conserved across the species selected, with each preserving the four conserving domains, bHLH, PAS-A, PAS-B, and poly Q (not shown) at the C-termini of the sequences (Yoshitane et al. 2009).
Alignment of CLOCK1 and CLOCK2 protein sequences from representative vertebrate species. The human CLOCK sequence was employed as the reference for comparison and numbering. Sequence alignments were realized in MAFFT and colored using Jalview. The secondary structures (α helix, red bar; β fold, green bar) are shown under the sequences. The functional domains also are indicated (black bar). Essential functional sites are indicated with light blue box for R39, dark blue box for E43, green box for R47, yellow box for 84th site amino acid and red box for others. Color codes for conservation vary according to clade. a CLOCK2 alignment for representative vertebrate species. b CLOCK1a and CLOCK1b alignment for five teleost fishes. c CLOCK1 alignment for representative vertebrate species. The complete sequences see Figure S2 (a-c) in supplementary materials (Color figure online)
CLOCK2 sequences from 10 representative vertebrate species were selected for alignment using human CLOCK2 as the template. We found that differences in CLOCK2 were mainly present in the N terminus and C-terminal poly Q regions. The important residue sites of CLOCK2 in vertebrates are marked with a red box in Fig. 4a. Three phosphorylation sites (Ser38, Ser42, and Ser427 not shown) in CLOCK1, numbered according to the human template, are conserved in CLOCK2 (Huang et al. 2012). The other vital residues are components of the CLOCK-BMAL1 heterodimer, interacting with each site of the E-box, with Arg39, Glu43, and Arg47 (Huang et al. 2012). In particular, we found that H84 in tetrapods is Q84 in ray-finned fish, and that this site was conserved in CLOCK1. This site is related to domains that facilitate CLOCK and BMAL1 recognition. In general, CLOCK2 sequences are relatively conserved at vital sites, with the secondary structures of CLOCK2 consisting of 25 α-helices and 23 β-strands.
As mentioned above, most ray-finned fish that had undergone FSGD had two isotypes of clock1 named clock1a and clock1b. To further determine the differences between CLOCK1a and CLOCK1b, we selected 10 representative sequences from the ray-finned fish for alignment with human CLOCK1 (protein ID: 6QPJ). CLOCK1a contains approximately 890 aa, whereas CLOCK1b contains 820 aa. Many variable sites in CLOCK1a and CLOCK1b were found downstream of the PAS-B domain. However, the phosphorylation sites, core hydrophobic residues, bHLH with PAS recognition sites, and residues that react with E-box (marked by red boxes) are almost identical in CLOCK1a and CLOCK1b, as shown in Fig. 4b. The unique notable difference was the variation of the critical site H84 to N84 in Fugu. In general, functional domains were highly conserved between CLOCK1a and CLOCK1b.
Similarly, an alignment of CLOCK1 protein sequences from 10 vertebrate species is shown in Fig. 4c. Compared with that in amphibians and fish, CLOCK1 in mammals, birds, and reptiles contained an extra 9–10 aa residues at the N terminus, except for the anole lizard. The pivotal sites (Ser38, Arg39, Ser42, Glu43, Arg47, His84, Leu57, Leu74, Phe104, Leu105, Leu113, and Trp362 in CLOCK1) are marked with red boxes in Fig. 4c (Huang et al. 2012). The results for Fugu and zebrafish CLOCK1b at these sites are also consistent with zebrafish having variant N84 and Fugu possessing a hypervariable region in the bHLH domain. Generally, CLOCK1 was more conserved than CLOCK2, with secondary structures of CLOCK1 consisting of 32 α-helices and 19 β-strands. In the 3D CLOCK:BMAL1 complex structure, each domain interacts primarily with the corresponding domain of its partner subunit, so that CLOCK bHLH interacts with BMAL1 bHLH, and CLOCK PAS-A (or PAS-B) interacts with BMAL1 PAS-A (or PAS-B) (Huang et al. 2012). The CLOCK:BMAL1 heterodimer interface analysis indicates that the CLOCK H84 interface contacts with BMAL1 D86 (Fig. 5; Table S4). In addition, the BMAL1 alignment results suggest that, in ray-finned fish, D86 is replaced by K86 (Figure S2), and that the amino acid residues that mediate the CLOCK:BMAL1 hydrophobic interactions are conserved (L95 and L115).
Synteny Analysis
Conserved syntenic regions, defined by two or more closely linked orthologous genes on a single chromosome or chromosomal fragment in each of two or more different species, provide crucial information regarding how genes and genomes evolve. Our synteny analysis assumed that genes in the chromosomal neighborhood of orthologous genes are likely orthologous, despite the fact that chromosomal rearrangements such as fusions, fissions, translocations, and inversions often disorganize the conserved gene order (Wood et al. 2005; Kasahara et al. 2007; Postlethwait 2007; Guyomard et al. 2012). Our synteny analysis showed that clock1 and clock2 share a conserved suite of upstream and downstream genes across species, although some species showed gene loss (Fig. 6). We observed that four conserved genes, KIT proto-oncogene receptor tyrosine kinase (kit), kinase insert domain receptor (kdr), centrosomal protein 135 (cep135), and aminoadipate-semialdehyde dehydrogenase (aasd), neighbor clock1 in lobe-finned fish. We collected transmembrane protein 165 (tmem165) and tyrosinase-related protein 1b (tyrp1b) for a ray-finned fish clock1 synteny analysis because of the lower synteny in this clade.
Comparison of the order of genes surrounding clock1 and clock2 in chromosomes of several vertebrates. Orthologous genes flanking clock1 and clock2 show highly conserved synteny among the vertebrates examined. Orthologous genes are shown in the same color. For additional information on synteny of species, see Figure S3 (NOTE: npas2 here is equivalent to clock2)
Subsequently, we performed a clock2 synteny analysis between lobe-finned fish and ray-finned fish (Fig. 6). In the lobe-finned fish clade, all clock2 genes shared a conserved suite of carbohydrate sulfotransferase 10 (chst10), LON peptidase N-terminal domain and ring finger 2 (lonrf2), ribosomal protein L31 (rpl31), and CCR4-NOT transcription complex subunit 11 (cnot11) around them, whereas none of these synteny genes could be identified in ray-finned fish. Hence, we selected zinc finger CCCH-type containing 12B (zc3h12b) and bromodomain and WD repeat domain containing 3 (brwd3) as candidate genes, localized near clock2 in ray-finned fish. Notably, all four of these genes near clock2 in ray-finned fish were absent in Osteoglossiformes such as P. kingsleyae and S. formosus. Therefore, we selected another four genes, melatonin receptor 1C (mtnr1c), VMA21 vacuolar H + -ATPase homolog (vma21), ephrin-B1 (efnb1), and ectodysplasin A (eda), to conduct a synteny analysis in P. kingsleyae and S. formosus. By performing a comparative synteny analysis between lobe-finned fish and ray-finned fish, we found that synteny in the lobe-finned fish clade was more conserved than that in ray-finned fish. Our synteny results supported the rationale to classify the two lineages as clock1 and clock2.
Positional Selection Analysis
To determine the selective modes that might have acted on these ancient clock duplicates, their ratio of nonsynonymous substitutions per nonsynonymous site (dN) to the numbers of synonymous substitutions per synonymous site (dS) were computed. Generally, dN/dS > 1 indicates positive selection, dN/dS = 1 indicates neutral selection, and dN/dS < 1 suggests purifying selection (Yang and Bielawski 2000; Hurst 2002). Among the calculations for 73 sequences from 29 vertebrates (Fig. 7), all dN/dS values were considerably less than 1, indicating that purifying selection acted on these genes in the vertebrate species during clock evolution (Table 1 & Table S5). However, the clock2 dN/dS values were higher than those for clock1 in the lobe-finned fish clade (P < 0.05), suggesting that the clock2 gene in lobe-finned fish may have experienced more intense selection for overlapping roles in a suprachiasmatic circadian clock (DeBruyne et al. 2007). The clock1a, clock1b, and clock2 dN/dS values in ray-finned fish were not significantly different (P > 0.05).
Ratios of nonsynonymous and synonymous substitutions (dN/dS) estimated with the Codeml module in PAML Significant differences among the five clades are marked with asterisks (* < 0.05, ** < 0.01). Clade 1: lobe-finned fish clock2; Clade 2: ray-finned fish clock2; Clade 3: lobe-finned fish clock1; Clade 4: ray-finned fish clock1b; Clade 5: ray-finned fish clade clock1a
Discussion
Possible Reasons for the Presence of Different Copies Among Vertebrates
During evolution, genes are often subject to duplication events such as WGD, large-scale segmental duplication, and small-scale gene duplication (Bridges 1936; Glasauer and Neuhauss 2014). WGD has been proposed to play a predominant role in providing additional genetic material for the occurrence of new genes, allowing organisms to acquire novel characteristics to survive natural challenges (Stephens 1951; Kaessmann 2010). Ohno proposed two rounds of WGD in early vertebrate evolution, with one round of WGD occurring before the Agnatha–Gnathostomata split and two rounds happening before the Chondrichthyes–Osteichthyes split, both of which provided raw materials for the evolutionary diversification of vertebrates (Taylor et al. 2001; Zhang 2003; Christoffels et al. 2004). Furthermore, after the second round of WGD in a common ancestor of vertebrates around 320 Mya, a third round of genome duplication occurred in the stem lineage of teleost fishes after the Actinopterygian–Sarcopterygian split (Amores et al. 1998; Taylor et al. 2003; Meyer and Van de Peer 2005). Acipenseridae, Catostomidae, Cobitidae, Cyprinidae, and Salmonidae fish families have even undergone a fourth round of WGD (Taylor et al. 2003; Vandepoele et al. 2004).
Results of our study are consistent with previous results showing that all vertebrates have undergone at least two rounds of WGD. Almost all species of lobe-finned fishes and some basal non-teleost ray-finned fishes included in the present study had two copies of both clock1 and clock2 genes, whereas most teleost fishes had three copies of clock such as clock1a, clock1b, and clock2 or clock1a, clock2ba, and clock2bb. Tetraploid teleosts such as S. salar and O. mykiss possessed four clock copies, and the three Sinocyclocheilus species each possessed six copies. In general, our results indicate that copy number variations in clock in vertebrates were mainly caused by a combination of WGD and gene loss, which is consistent with results from a recent genome-wide analysis of duplicate genes (Li et al. 2018). Because of selective loss of genes, the number of duplicates in diploids and tetraploids does not always correspond to a two-four-eight model. For diploid teleosts, most species examined in our present study have three members, clock1a, clock1b, and clock2. The clock2a gene was lost from the last common ancestor of teleosts, because no clock2a was found in the clade (Fig. 3) (Wang 2008; Toloza-Villalobos et al. 2015). This phenomenon is more common in tetraploid teleosts such as S. salar and O. mykiss, which possess two clock1a (clock1aa and clock1ab; lost clock1b) and two clock2b (clock2ba and clock2bb; lost clock2a), and the three Sinocyclocheilus fishes, which have six clock gene members, including two clock1a, two clock1b, and two clock2b (lost clock2a). In these five tetraploid species, distinct evolutionary models were apparent. Based on previous research, salmonid gene fractionation may still be occurring because of an additional and relatively recent WGD event that has been dated to 100–25 Mya (Berthelot et al. 2014). Compared with the third common teleost WGD, the rediploidization process in salmonids is ongoing, and only half of the protein-coding genes have been retained as duplicate copies. The ancient rediploidization event revealed here might explain why both of the salmonids lost clock1b. The detailed picture of clock1b loss will remain unclear until more salmonid species have been examined. Results for the three Sinocyclocheilus species revealed that intact remnant clock genes were conserved in this clade. Compared to that of the salmonid species mentioned above, diversification of Sinocyclocheilus species occurred recently, perhaps at the beginning of the rediploidization process when the common carp lineage appeared at the forth genome duplication event, which occurred approximately 8 Mya (Xu et al. 2014, 2019a, b). Thus, all three Sinocyclocheilus species possessed more members of the remaining clock than salmonids.
Combining results of previous studies with our own, we concluded that the evolutionary path of clock was as follows: the common ancestor of vertebrates had one clock gene, which is still retained in lamprey. During the second round of genome duplication in Chondrichthyes and Osteichthyes, clock gave rise to clock1 and clock2. In the subsequent third round of genome duplication in teleost fishes, clock1 gave rise to clock1a and clock1b, both of which have been preserved in almost all teleost fishes. In addition, clock2 turned into clock2a and clock2b, one of which (clock2b) has been maintained in the majority of teleosts (Wang 2008). During evolution, osteoglossomorph fishes (P. kingsleyae and S. formosus) maintained the two clock2 members but lost clock1b. The remnant clock1a of the two osteoglossiform fishes form a clade with spotted gar clock1 placed at the base of the phylogenetic tree, suggesting that Osteoglossiformes are relatively primitive among teleosts. Our estimated phylogenies of members of the clock gene family confirmed that spotted gar diverged from teleosts before the FSGD, whereas osteoglossomorphs experienced FSGD (Braasch et al. 2016; Bian et al. 2016). Among teleosts, Osteoglossomorpha has been considered an ancient group, with fossil records dating back to the late Jurassic (Bian et al. 2016). On the basis of the homology analysis, combined with fossil evidence, the two clock2 genes of osteoglossomorph fishes might represent an ancient branch of a special duplication.
Moreover, the results from the synteny analysis revealed that clock1 and clock2 genes localize on different chromosomes or chromosomal fragments in the same species. Interestingly, we found that the syntenic genes were not conserved between lobe-finned fish and ray-finned fish. Only two genes, kit and cep135, both near the clock1 gene, were shared in ray-finned fish and lobe-finned fish. Furthermore, syntenic genes located around clock1a and clock1b in the ray-finned fish were not completely identical. There was no homologous gene shared by lobe-finned fish and ray-finned fish that localized in regions flanking clock2 in our study. It seemed that clock genes experienced rearrangement after the split between the lobe-finned and ray-finned fishes. In particular, no collinear sequences were found in osteoglossomorphs (P. kingsleyae and S. formosus), so another four candidate genes were selected to conduct the synteny analysis. Interestingly, we found that syntenic genes in lobe-finned fish are markedly more conserved than those in ray-finned fish. This phenomenon may reflect the fact that ray-finned fish lineages are more inclined to inter-chromosomal rearrangements than lobe-finned fish lineages, leading to shorter conserved syntenic blocks in ray-finned fish compared with those in lobe-finned fish (Braasch et al. 2016; Ravi and Venkatesh 2018). Moreover, FSGD events also led to shorter syntenic blocks through differential gene loss without rearrangements (Ravi and Venkatesh 2018). The disrupted syntenic blocks are very widespread in ray-finned fish genomes. For example, Xu et al. (2019a, b) reported that the syntenic region of the tph locus contained seven genes, and this block was entirely conserved in tetrapods. However, in zebrafish, tph was duplicated to tph1a and tph1b, owing to the FSGD, and the regions of tph1a and tph1b retained only one and two genes, respectively (Xu et al. 2019a, b).
Adaptive Evolution of Clock Genes in Vertebrates
The Earth turns on its axis every 24 h, and almost all life on the planet shows circadian rhythmicity that follows daily changes caused by this autogiration (Abhilash et al. 2017). The molecular CLOCK that controls circadian rhythms was revealed to be an important regulator of physiology and disease (Yi et al. 2010). In humans, CLOCK and BMAL1 form a CLOCK:BMAL1 heterodimer through their PAS domains. By binding the E-boxes in other CCGs, the CLOCK:BMAL1 heterodimer drives transcription of these genes (Allada et al. 1998; Bielawski and Yang 2003; Yoshitane et al. 2009). Thus, elucidating critical sites of protein functional diversity and understanding how gene families evolve are core evolutionary biology interests. In our protein sequence analysis, critical sites associated with the E-box were largely conserved, except for Fugu CLOCK1b. Previous research (Wang 2008) has shown that Fugu clock1b exhibits a higher dN/dS value than other clock genes, and our results are consistent with these results. In addition, CLOCK2 H84 in ray-finned fish was turned into Q84/N84, indicating the diversity of CLOCK2 between lobe-finned and ray-finned fishes. Changes in this site suggest diverse modes of CLOCK1:BMAL1 and CLOCK2:BMAL1 dimer recognition in teleosts. Because these two amino acids carry different charges, changes at this site suggest diverse modes of CLOCK:BMAL1 dimer recognition and transactivation activity in vertebrates. Huang et al. (2012) showed that the CLOCK:BMAL1 bHLH dimer interface is largely mediated by conserved hydrophobic interactions. This transformation seems to enhance the formation of a stable heterodimeric complex and increases the transactivation activity of CLOCK:BMAL1.
Moreover, previous studies have shown both structural and functional evolutionary divergence in the clock gene family (DeBruyne et al. 2007; Wang 2008). Thus, we analyzed selected aspects of structural and functional divergence in the clock family and attempted to elucidate patterns of selection pressure in the five clades. Results showed that clade 3 (Fig. 3, lobe-finned fish clock1) shows a more intense purifying selection compared with clade 1 (lobe-finned fish clock2, P < 0.01, Fig. 7). During evolution, the transcription factor CLOCK2 was able to serve as a functional substitute for CLOCK1 in the master brain clock to regulate circadian rhythmicity in mice (Glasauer and Neuhauss 2014). In homozygous CLOCK2-mutant mice, which do not express functional CLOCK2 (Garcia et al. 2000), robust circadian rhythms control locomotor behavior suggests that clock1 and clock2 have overlapping functions (Dudley et al. 2003). The results of our protein sequence analysis and structural characterization show that CLOCK2 is less conserved than CLOCK1, but preserves at pivotal sites, and this suggests that lobe-finned fish CLOCK2 plays overlapping roles in the circadian rhythm compared with ray-finned fish (DeBruyne et al. 2007; Dibner et al. 2010; Partch et al. 2014). In other words, CLOCK1 has a more prominent role than CLOCK2 in controlling circadian gene expression (DeBruyne et al. 2007). However, CLOCK2 was reported to have other functions, including roles in memory, mood regulation, and ingestion (Garcia et al. 2000; Dudley et al. 2003; Ozburn et al. 2017). Thus, the sequence variability owing to relaxed constraints might have allowed the acquisition of new function by CLOCK2 (neofunctionalization) (Antoch et al. 1997; Lowrey et al. 2000; DeBruyne et al. 2007). In most ray-finned fish, FSGD resulted in an extra clock gene; hence, three clades remained (clade 2, clade 4, and clade 5). These clades showed relatively low dN/dS values with no significant difference between them. Although these three clades showed dN/dS values less than 1, it is possible that some genes are undergoing neofunctionalization and have relaxed functional constraints (He et al. 2005). From a molecular viewpoint, the CLOCK2 pivotal site related to CLOCK:BMAL1 interactions in fish species (H84, which became Q 84) suggest that clock2 may be undergoing neofunctionalization and have played multiple roles during evolution (Kovanen et al. 2010). In clock1a and clock1b, the relatively high dN/dS value compared to that of lobe-finned fish clock1 (clade 3, P < 0.05) suggests that the duplicated members have relaxed functional constraints and play a sub-functional role in circadian rhythms (Wang 2008).
In vertebrate circadian clock gene networks, the CLOCK:BMAL1 complex binds to regulatory elements containing E-boxes in a set of rhythmic genes that encode the repressor proteins period (encoded by PER1, PER2, and PER3) and cryptochrome (encoded by CRY1 and CRY2) (Gekakis et al. 1998; Kume, et al. 1999; Shearman et al. 2000). Previous studies have shown that both bmal and clock genes were duplicated and lost after FGSD (Wang 2008, 2009; Toloza-Villalobos et al. 2015). Although clock plays a key role in the regulation of biological rhythms (DeBruyne et al. 2007; Ray et al. 2020), the number of bmal and clock genes is the same in different species. For example, the human genome contains two clock and bmal genes, whereas zebrafish contains three clock and bmal genes (Wang 2008, 2009). At the same time, based on dN/dS analyses, the evolution rate of clock and bmal genes is less than one, and these two genes have asymmetric evolutionary rates between duplicates (Wang 2008, 2009). In addition, our sequence analysis results showed (not listed in the text) that the 86th amino acid of the BMAL1 protein sequence (interface contact with the 84th position of CLOCK) in ray-finned fish, with the substitution of H84 by Q84, also leads the D86–K86 change. Therefore, we conclude that the CLOCK:BMAL1 dimer is co-evolving.
Diversification of Clock Genes in Cavefishes
According to results of our phylogenetic analysis and protein structural comparison, the clock gene family in some cavefishes such as Astyanax mexicanus and T. rosa have three and S. anshuiensis possesses six members. Moreover, the sequences of cavefish clock family genes are similar (Table S1). In the laboratory, the adult surface Mexican tetra shows robust circadian rhythms. These genes have been retained in cave populations but with substantial alterations (Beale et al. 2013). In addition, different groups from various caves display subtle differences. In other words, Mexican tetra cave populations have rhythms that differ among diverse populations (Beale et al. 2013). These differences may result from increased levels of light-inducible gene expression in cavefish, including expression of members of the circadian rhythm repressor per family (Beale et al. 2013, 2016). From a molecular standpoint, cavefish appear as if they have experienced constant light rather than perpetual darkness. The other cavefish, S. anshuiensis, was reported to have weaker circadian rhythms because both copies of Skp1 proteins had deletions at their N-termini, and expression levels of the rhythm pathway genes were decreased (Yang et al. 2016). The cavefish T. rosa also had three complete clock genes. Further examination of the expression of cavefish clock and its regulatory mechanism should add to our understanding of cave animal adaptations. In addition to biological rhythms, the clock gene is reported to have other physiological functions in vertebrates, such as roles in fertility, seasonality, and cancer regulation (Turek et al. 2005; Kovanen et al. 2016; Abhilash and Sharma 2016; Abhilash et al. 2017; Jankowski and Dmitrzak-Weglarz 2017). Thus, we speculated that the difference between cave and non-cave fish clock genes more likely results from expression and regulation, and it is reasonable that the clock gene is preserved in cavefish. In general, the results of our current study provide a genome-wide view of the evolution of clock gene family in vertebrates.
Conclusions
Through a genomic survey, this study provides genome-wide insights into clock family genes in vertebrates. Combining the results of a phylogenetic analysis with synteny identification, we found that copy number variations in vertebrate clock genes were mainly associated with WGDs and gene losses. By comparing CLOCK1 and CLOCK2 protein sequences, we also revealed many similarities and differences between clock1 and clock2. Furthermore, dN/dS results suggested that clock genes had different fates following duplication between ray-finned and lobe-finned fishes. In addition, in the special and primitive clade of teleosts, the clock genes of osteoglossomorph fishes show an opposite pattern of duplication. Therefore, teleosts have adopted various strategies to adapt to diverse environments after FSGD. Moreover, although the cavefish possessed clock genes like other species, different levels of rhythm showed that further expression experiments should be performed to illuminate the role that clock plays in these cave species.
References
Abhilash L, Sharma VK (2016) On the relevance of using laboratory selection to study the adaptive value of circadian clocks. Physiol Entomol 41:293–306. https://doi.org/10.1111/phen.12158
Abhilash L, Shindey R, Sharma VK (2017) To be or not to be rhythmic? A review of studies on organisms inhabiting constant environments. Biol Rhythm Res 48:677–691. https://doi.org/10.1080/09291016.2017.1345426
Allada R, White NE, So WV, Hall JC, Rosbash M (1998) A mutant drosophila homolog of mammalian clock disrupts circadian rhythms and transcription of period and timeless. Cell 93:791–804. https://doi.org/10.1016/S0092-8674(00)81440-3
Amores A, Force A, Yan YL, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish hox clusters and vertebrate genome evolution. Science 282:1711–1714. https://doi.org/10.1126/science.282.5394.1711
Antoch MP, Song EJ, Chang AM, Vitaterna MH, Zhao YL, Wilsbacher LD, Sangoram AM, King DP, Pinto LH, Takahashi JS (1997) Functional identification of the mouse circadian clock gene by transgenic bac rescue. Cell 89:655–667. https://doi.org/10.1016/S0092-8674(00)80246-9
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren JY, Li WW, Noble WS (2009) Meme suite: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. https://doi.org/10.1093/nar/gkp335
Bargiello TA, Jackson FR, Young MW (1984) Restoration of circadian behavioural rhythms by gene transfer in drosophila. Nature 312:752–754. https://doi.org/10.1038/312752a0
Beale A, Guibal C, Tamai TK, Klotz L, Cowen S, Peyric E, Reynoso VH, Yamamoto Y, Whitmore D (2013) Circadian rhythms in mexican blind cavefish astyanax mexicanus in the lab and in the field. Nat Commun 4:2769. https://doi.org/10.1038/ncomms3769
Beale AD, Whitmore D, Moran D (2016) Life in a dark biosphere: a review of circadian physiology in “arrhythmic” environments. J Comp Physiol B 186:947–968. https://doi.org/10.1007/s00360-016-1000-6
Bell-Pedersen D, Cassone V, Earnest D et al (2005) Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat Rev Genet 6:544–556. https://doi.org/10.1038/nrg1633
Berthelot C, Brunet F, Chalopin D, Juanchich A, Bernard M, Noel B, Bento P, Da Silva C, Labadie K, Alberti A et al (2014) The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates. Nat Commun 5:3657. https://doi.org/10.1038/ncomms4657
Bian C, Hu Y, Ravi V, Kuznetsova IS, Shen X, Mu X, Sun Y, You X, Li J, Li X et al (2016) The Asian arowana (Scleropages formosus) genome provides new insights into the evolution of an early lineage of teleosts. Sci Rep 6:24501. https://doi.org/10.1038/srep24501
Bielawski JP, Yang Z (2003) Maximum likelihood methods for detecting adaptive evolution after gene duplication. J Struct Funct Genom 3:201–202. https://doi.org/10.1023/A:1022642807731
Braasch I, Gehrke AR, Smith JJ, Kawasaki K, Manousaki T, Pasquier J, Amores A, Desvignes T, Batzel P, Catchen J et al (2016) The spotted gar genome illuminates vertebrate evolution and facilitates human-teleost comparisons. Nat Genet 48:427–437. https://doi.org/10.1038/ng0616-700c
Bridges C (1936) The bar “gene” a duplication. Science 83:210–211. https://doi.org/10.1126/science.83.2148.210
Brown SA, Ripperger J, Kadener S, Fleury-Olela F, Vilbois F, Rosbash M, Schibler U (2005) Period1-associated proteins modulate the negative limb of the mammalian circadian oscillator. Science 308:693–696. https://doi.org/10.1126/science.1107373
Buhr ED, Takahashi JS (2013) Molecular components of the mammalian circadian clock. Handb Exp Pharmacol 217:3–27. https://doi.org/10.1007/978-3-642-25950-0_1
Chen C, Xia R, Chen H, He Y (2018) Tbtools, a toolkit for biologists integrating various hts-data handling tools with a user-friendly interface. Biorxiv.
Christoffels A, Koh EG, Chia JM, Brenner S, Aparicio S, Venkatesh B (2004) Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Mol Biol Evol 21:1146–1151. https://doi.org/10.1093/molbev/msh114
Clayton JD, Kyriacou CP, Reppert SM (2001) Keeping time with the human genome. Nature 409:829–831. https://doi.org/10.1038/35057006
DeBruyne JP, Weaver DR, Reppert SM (2007) Clock and npas2 have overlapping roles in the suprachiasmatic circadian clock. Nat Neurosci 10:543–545. https://doi.org/10.1038/nn1884
Della Ragione F, Comitato R, Angelini F, D’Esposito M, Cardone A (2005) Molecular cloning and characterization of the clock gene period2 in the testis of lizard Podarcis sicula and its expression during seasonal reproductive cycle. Gene 363:105–12. https://doi.org/10.1016/j.gene.2005.08.018
Dibner C, Schibler U, Albrecht U (2010) The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu Rev Physiol 72:517–549. https://doi.org/10.1146/annurev-physiol-021909-135821
Dudley CA, Erbel-Sieler C, Estill SJ, Reick M, Franken P, Pitts S (2003) Altered patterns of sleep and behavioral adaptability in npas2-defficient mice. Science 301:379–383. https://doi.org/10.1126/science.1082795
Dunlap JC (1999) Molecular bases for circadian clocks. Cell 96:271–290. https://doi.org/10.1016/S0092-8674(00)80566-8
Garcia JA, Zhang D, Estill SJ, Michnoff C, Rutter J, Reick M, Scott K, Diaz-Arrastia R, McKnight SL (2000) Impaired cued and contextual memory in npas2-deficient mice. Science 288:2226–2230. https://doi.org/10.1126/science.288.5474.2226
Gekakis N, Staknis D, Nguyen HB, Davis FC, Wilsbacher LD, King DP, Takahashi JS, Weitz CJ (1998) Role of the CLOCK protein in the mammalian circadian mechanism. Science 280:1564–1569. https://doi.org/10.1126/science.280.5369.1564
Glasauer SM, Neuhauss SC (2014) Whole-genome duplication in teleost fishes and its evolutionary consequences. Mol Genet Genom 289:1045–1060. https://doi.org/10.1007/s00438-014-0889-2
Guyomard R, Boussaha M, Krieg F, Hervet C, Quillet E (2012) A synthetic rainbow trout linkage map provides new insights into the salmonid whole genome duplication and the conservation of synteny among teleosts. BMC Genet. https://doi.org/10.1186/1471-2156-13-15
Hardin PE (2005) The circadian timekeeping system of drosophila. Curr Biol 15:R714–R722. https://doi.org/10.1016/j.cub.2005.08.019
He X, Zhang J (2005) Rapid subfunctionalization accompanied by prolonged and substantial neofunctionalization in duplicate gene evolution. Genetics 169:1157–1164. https://doi.org/10.1534/genetics.104.037051
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G (2015) Gsds 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Huang NA, Chelliah Y, Shan YL, TaylorYoo CASH, Partch C, Green CB, Zhang H, Takahashi JS (2012) Crystal structure of the heterodimeric Clock:Bmal1 transcriptional activator complex. Science 337:189–194. https://doi.org/10.1126/science.1222804
Hurst L (2002) The ka/ks ratio: diagnosing the form of sequence evolution. Trends Genet 18:486–487. https://doi.org/10.1016/S0168-9525(02)02722-1
Jankowski KS, Dmitrzak-Weglarz M (2017) Arntl, clock and per3 polymorphisms—links with chronotype and affective dimensions. Chronobiol Int 34:1105–1113. https://doi.org/10.1080/07420528.2017.1343341
Kaessmann H (2010) Origins, evolution, and phenotypic impact of new genes. Genome Res 20:1313–1326. https://doi.org/10.1101/gr.101386.109
Kasahara M, Naruse K, Sasaki S, Nakatani Y, Qu W, Ahsan B, Yamada T, Nagayasu Y, Doi K, Kasai Y et al (2007) The medaka draft genome and insights into vertebrate genome evolution. Nature 447:714–719. https://doi.org/10.1038/nature05846
Katoh K, Standley DM (2013) Mafft multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
King DP, Vitaterna MH, Chang AM, Dove WF, Pinto LH, Turek FW, Takahashi JS (1997) The mouse clock mutation behaves as an antimorph and maps within the w-19h deletion, distal of kit. Genetics 146:1049–1060. http://orcid.org/0000-0003-0384-8878
King DP, Zhao YL, Sangoram AM, Wilsbacher LD, Tanaka M, Antoch MP, Steeves TDL, Vitaterna MH, Kornhauser JM, Lowrey PL, Turek FW, Takahashi JS (1997) Positional cloning of the mouse circadian clock gene. Cell 89:641–653. https://doi.org/10.1016/S0092-8674(00)80245-7
Konopka RJ, Benzer S (1971) Clock mutants of drosophila melanogaster. Proc Natl Acad Sci 68:2112–2116. https://doi.org/10.1073/pnas.68.9.2112
Kovanen L, Donner K, Kaunisto M, Partonen T (2016) CRY1 and CRY2 genetic variants in seasonality: A longitudinal and cross-sectional study. Psychiatry Res 242:101–110. https://doi.org/10.1016/j.psychres.2016.05.044
Kovanen L, Saarikoski ST, Aromaa A, Lonnqvist J, Partonen T (2010) Arntl (bmal1) and npas2 gene variants contribute to fertility and seasonality. PLoS ONE 5:e10007. https://doi.org/10.1371/journal.pone.0010007
Kume K, Zylka MJ, Sriram S, Shearman LP, Reppert SM (1999) mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98:193–205. https://doi.org/10.1016/S0092-8674(00)81014-4
Li Y, Lv Y, Bian C, You X, Deng L, Shi Q (2018) A comparative genomic survey provides novel insights into molecular evolution of l-aromatic amino acid decarboxylase in vertebrates. Molecules 23:917. https://doi.org/10.3390/molecules23040917
Lowrey PL, Shimomura K, Antoch MP, Yamazaki S, Zemenides PD, Ralph MR, Menaker M, Takahashi JS (2000) Positional syntenic cloning and functional characterization of the mammalian circadian mutation tau. Science 288:483–491. https://doi.org/10.1126/science.288.5465.483
Markowska M, Majewski PM, Skwarło-Sońta K (2017) Avian biological clock—immune system relationship. Dev Comp Immunol 66:130–138. https://doi.org/10.1016/j.dci.2016.05.017
Matsuda F, Tsugawa H, Fukusaki E (2013) Method for assessing the statistical significance of mass spectral similarities using basic local alignment search tool statistics. Anal Chem 85:8291–8297. https://doi.org/10.1021/ac401564v
Menaker M, Wisner S (1983) Temperature-compensated circadian clock in the pineal of Anolis. Proc Natl Acad Sci 80(19):6119–21. https://doi.org/10.1073/pnas.80.19.6119
Meyer A, Van de Peer Y (2005) From 2r to 3r: evidence for a fish-specific genome duplication (fsgd). Bioessays 27:937–945. https://doi.org/10.1002/bies.20293
Mohawk JA, Green CB, Takahashi JS (2012) Central and peripheral circadian clocks in mammals. Annu Rev Neurosci 35:45–462. https://doi.org/10.1146/annurev-neuro-060909-153128
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) Iq-tree: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Oishi K, Miyazaki K, Kadota K, Kikuno R, Nagase T, Atsumi G, Ohkura N, Azama T, Mesaki M, Yukimasa S et al (2003) Genome-wide expression analysis of mouse liver reveals clock-regulated circadian output genes. J Biol Chem 278:41519–41527. https://doi.org/10.1074/jbc.M304564200
Okano T, Yamamoto K, Okano K, Hirota T, Kasahara T, Sasaki M, Takanaka Y, Fukada Y (2001) Chicken pineal clock genes: Implication of bmal2 as a bidirectional regulator in circadian clock oscillation. Genes Cells 6:825–836. https://doi.org/10.1046/j.1365-2443.2001.00462.x
Ozburn AR, Kern J, Parekh PK, Logan RW, Liu Z, Falcon E, Becker-Krail D, Purohit K, Edgar NM, Huang Y et al (2017) Npas2 regulation of anxiety-like behavior and gabaa receptors. Front Molec Neurosci 10:360. https://doi.org/10.3389/fnmol.2017.00360
Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109:307–320. https://doi.org/10.1016/S0092-8674(02)00722-5
Panda S, Hogenesch J, Kay S (2002) Circadian rhythms from flies to human. Nature 417:329–335. https://doi.org/10.1038/417329a
Pando MP, Sassone-Corsi P (2002) Unraveling the mechanisms of the vertebrate circadian clock: zebrafish may light the way. BioEssays 24:419–426. https://doi.org/10.1002/bies.10091
Partch CL, Green CB, Takahashi JS (2014) Molecular architecture of the mammalian circadian clock. Trends Cell Biol 24:90–99. https://doi.org/10.1016/j.tcb.2013.07.002
Pittendrigh C (1993) Temporal organization—reflections of a Darwinian clock-watcher. Annu Rev Physiol 55:16–54. https://doi.org/10.1146/annurev.ph.55.030193.000313
Postlethwait JH (2007) The zebrafish genome in context: ohnologs gone missing. J Exp Zool 308:563–577. https://doi.org/10.1002/jez.b.21137
Ravi V, Venkatesh B (2018) The divergent genomes of teleosts. Annu Rev Anim Biosci 6:47–68. https://doi.org/10.1146/annurev-animal-030117-014821
Ray S, Valekunja UK, Stangherlin A, Howell SA, Snijders AP, Damodaran G, Reddy AB (2020) Circadian rhythms in the absence of the clock gene Bmal1. Science 367:800–806. https://doi.org/10.1126/science.aaw7365
Reddy P, Zehring WA, Weeler DA, Pirrotta V, Hadfield C, Hall JC, Rosbash M (1984) Molecular analysis of the period locus in drosophila melanogaster and identification of a transcript involved in biological rhythms. Cell 38:701–710. https://doi.org/10.1016/0092-8674(84)90265-4
Reppert SM, Weaver DR (2001) Molecular analysis of mammalian circadian rhythms. Annu Rev Physiol 63:647–676. https://doi.org/10.1146/annurev.physiol.63.1.647
Saha S, Singh KM, Gupta BBP (2019) Melatonin synthesis and clock gene regulation in the pineal organ of teleost fish compared to mammals: Similarities and differences. Gen Comp Endocrinol 279:27–34. https://doi.org/10.1016/j.ygcen.2018.07.010
Salamov AA, Solovyev VV (2000) Ab initio gene finding in Drosophila genomic DNA. Genome Res 10:516–522. https://doi.org/10.1101/gr.10.4.516
Schmutz I, Ripperger JA, Baeriswyl-Aebischer S, Albrecht U (2010) The mammalian clock component peroid2 coordinates circadian output by interaction with nuclear receptors. Genes Dev 24:345–357. https://doi.org/10.1101/gad.564110
Shearman LP, Sriram S, Weaver DR, Maywood ES, Chaves I, Zheng BH, Kume K, Lee CC, van der Horst GTJ, Hastings MH, Reppert SM (2000) Interacting molecular loops in the mammalian circadian clock. Science 288:1013–1019. https://doi.org/10.1126/science.288.5468.1013
Stephens SG (1951) Possible significance of duplication in evolution. Adv Genet 4:247–265. https://doi.org/10.1016/S0065-2660(08)60237-0
Takahashi JS, Pinto LH, Vitaterna MH (1994) Forward and reverse genetic approaches to behavior in the mouse. Science 264:1724–1733. https://doi.org/10.1126/science.8209253
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577. https://doi.org/10.1080/10635150701472164
Taylor JS, Van de Peer Y, Braasch I, Meyer A (2001) Comparative genomics provides evidence for an ancient genome duplication event in fish. Philos Trans R Soc Lond Ser B 356:1661–1679. https://doi.org/10.1098/rstb.2001.0975
Taylor JS, Braasch I, Fricke T, Meyer A, Van de Peer Y (2003) Genome duplication, a trait shared by 22,000 species of ray-finned fish. Genome Res 13:382–390. https://doi.org/10.1101/gr.640303
Toloza-Villalobos J, Arroyo JI, Opazo JC (2015) The circadian clock of teleost fish: a comparative analysis reveals distinct fates for duplicated genes. J Mol Evol 80:57–64. https://doi.org/10.1007/s00239-014-9660-x
Touitou Y, Point S (2020) Effects and mechanisms of action of light-emitting diodes on the human retina and internal clock. Environ Res 190:109942. https://doi.org/10.1016/j.envres.2020.109942
Turek FW, Joshu C, Kohsaka A, Lin E, Ivanova G, McDearmon E, Laposky A, Losee-Olson S, Easton A, Jensen DR, Eckel RH, Takahashi JS, Bass J (2005) Obesity and metabolic syndrome in circadian clock mutant mice. Science 308:1043–1045. https://doi.org/10.1126/science.1108750
Vandepoele K, De Vos W, Taylor JS, Meyer A, Van de Peer Y (2004) Major events in the genome evolution of vertebrates: paranome age and size differ considerably between ray-finned fishes and land vertebrates. Proc Natl Acad Sci 101:1638–1643. https://doi.org/10.1073/pnas.0307968100
Vitaterna MH, King DP, Chang AM, Kornhauser JM, Lowrey PL, McDonald JD, Dove WF, Pinto LH, Turek FW, Takahashi JS (1994) Mutagenesis and mapping of a mouse gene, clock, essential for circadian behavior. Science 264:719–725. https://doi.org/10.1126/science.8171325
Wang H (2008) Comparative analysis of teleost fish genomes reveals preservation of different ancient clock duplicates in different fishes. Mar Genom 1:69–78. https://doi.org/10.1016/j.margen.2008.06.003
Wang H (2009) Comparative genomic analysis of teleost fish bmal genes. Genetica 136:149–161. https://doi.org/10.1007/s10709-008-9328-9
Wang Z, Wu Y, Li L, Su XD (2013) Intermolecular recognition revealed by the complex structure of human clock-bmal1 basic helix-loop-helix domains with e-box DNA. Cell Res 23:213–224. https://doi.org/10.1038/cr.2012.170
Woller A, Gonze D (2013) The bird circadian clock: insights from a computational model. J Biol Rhythms 28:390–402. https://doi.org/10.1177/0748730414531216
Woods IG, Wilson C, Friedlander B, Chang P, Reyes DK, Nix R, Kelly PD, Chu F, Postlethwait JH, Talbot WS (2005) The zebrafish gene map defines ancestral vertebrate chromosomes. Genome Res 15:1307–1314. https://doi.org/10.1101/gr.4134305
Xu P, Zhang X, Wang X, Li J, Liu G, Kuang Y, Xu J, Zheng X, Ren L, Wang G et al (2014) Genome sequence and genetic diversity of the common carp, cyprinus carpio. Nature Genet 46:1212–1219. https://doi.org/10.1038/ng.3098
Xu J, Li Y, Lv Y, Bian C, You X, Endoh D, Teraoka H, Shi Q (2019) Molecular evolution of tryptophan hydroxylases in vertebrates: a comparative genomic survey. Genes 10:203. https://doi.org/10.3390/genes10030203
Xu P, Xu J, Liu G, Chen L, Zhou Z, Peng W, Jiang Y, Zhao Z, Jia Z, Sun Y et al (2019) The allotetraploid origin and asymmetrical genome evolution of the common carp cyprinus carpio. Nat Commun 10:4625. https://doi.org/10.1038/s41467-019-12644-1
Yang ZH (2007) Paml 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591. https://doi.org/10.1093/molbev/msm088
Yang ZH, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15:496–503. https://doi.org/10.1016/S0169-5347(00)01994-7
Yang J, Chen X, Bai J, Fang D, Qiu Y, Jiang W, Yuan H, Bian C, Lu J, He S et al (2016) The sinocyclocheilus cavefish genome provides insights into cave adaptation. BMC Biol 14:1. https://doi.org/10.1186/s12915-015-0223-4
Yi C, Mu L, de la Longrais IA, Sochirca O, Arisio R, Yu H, Hoffman AE, Zhu Y, Katsaro D (2010) The circadian gene npas2 is a novel prognostic biomarker for breast cancer. Breast Cancer Res Treat 120:663–669. https://doi.org/10.1007/s10549-009-0484-0
Yoshitane H, Takao T, Satomi Y, Du NH, Okano T, Fukada Y (2009) Roles of clock phosphorylation in suppression of e-box-dependent transcription. Mol Cell Biol 29:3675–3686. https://doi.org/10.1128/MCB.01864-08
Young MW (2000) Marking time for a kingdom. Science 288:451–453. https://doi.org/10.1126/science.288.5465.451
Yu WH, Paul E (2006) Circadian oscillators of drosophila and mammals. J Cell Sci 119:4793–4795. https://doi.org/10.1242/jcs.03174
Zhang J (2003) Evolution by gene duplication: an update. Trends Ecol Evol 18:292–298. https://doi.org/10.1016/S0169-5347(03)00033-8
Acknowledgements
We thank members from Peng’s lab for constructive discussion. This work was funded by the grant from the National Natural Science Foundation of China (No. 31872204).
Funding
This work was funded by the grant from the National Natural Science Foundation of China (No. 31872204).
Author information
Authors and Affiliations
Contributions
ZP contributed to the study conception and design. YM, FS, and QZ contributed to material preparation, data collection and analysis. YM prepared the first draft of the manuscript and all authors commented on the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling Editor: David Alvarez-Ponce.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mao, Y., Shao, F., Zhao, Q. et al. Molecular Evolution of clock Genes in Vertebrates. J Mol Evol 89, 494–512 (2021). https://doi.org/10.1007/s00239-021-10020-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10020-6