Abstract
In the early 1970s, Manfred Eigen and colleagues developed the quasispecies model (qs) for the population-based origin of RNAs representing the early genetic code. The Eigen idea is basically that a halo of mutants is generated by error-prone replication around the master fittest type which will behave similarly as a biological population. But almost from the start, very interesting and unexpected observations were made regarding competition versus co-operation which suggested more complex interactions. It thus became increasingly clear that although viruses functioned similar to biological species, their behavior was much more complex than the original theory could explain, especially adaptation without changing the consensus involving minority populations. With respect to the origin of natural codes, meaning, and code-use in interactions (communication), it also became clear that individual fittest type-based mechanisms were likewise unable to explain the origin of natural codes such as the genetic code with their context- and consortia-dependence (pragmatic nature). This, instead, required the participation of groups of agents competent in the code and able to edit code because natural codes do not code themselves. Three lines of inquiry, experimental virology, quasispecies theory, and the study of natural codes converged to indicate that consortia of co-operative RNA agents such as viruses must be involved in the fitness of RNA and its involvement in communication, i.e., code–competent interactions. We called this co-operative form quasispecies consortia (qs-c). They are the essential agents that constitute the possibility of evolution of biological group identity. Finally, the basic interactional motifs for the emergence of group identity, communication, and co-operation—together with its opposing functions—are explained by the “Gangen” hypothesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the history of science and philosophy, there has been a long-standing debate between fundamentally different views of life, such as those that emphasize co-operation and symbiosis, and in contrast to this views that promote the central importance of ‘selfish’ individuals (Villarreal and Witzany 2013b; Witzany 2006, 2007). The adaptability of individual-based selection fundamentally stems from errors, a few of which might improve survival. But is it really possible to originate a natural language-like code based on errors? Do we know any empirical example in that a natural language or code used by living agents to organize and coordinate their behavior resulted out of selecting a variety of errors? In contrary, all empirical facts indicate natural language and code using agents that coordinate their behavior by information-exchange, -modulation, and -innovation, i.e., generating new sequence-based content. This means natural codes basically result out of social interactions of agents competent to use codes in the context (pragmatics) of real-life world experience and –history (Witzany 2014, 2015a). Applied to the virosphere, we outlined this in detail as a reply to outdated opinions concerning the role of virus within the tree of life (Villarreal and Witzany 2010). One crucial question therefore is Is really new sequence space the result of selection of an abundance of replication errors (Eigen 1971), or in contrary, is it the result of the practical competence of living agents to generate new sequences? Are, e.g., RNA viruses “error-prone” or is this a misleading term for a highly productive behavior?
Here we reconsider the importance to life of collective behaviors in RNA species from the perspective of infectious and transmissible agents as currently known as viruses and virus-like agents.
Therefore, we argue for the better explanatory power of highly productive RNA quasispecies consortia (qs-c) for the evolution, conservation, and plasticity of genetic identities. As will be presented below, the biological capability to generate a toxic (viral) code can clearly differentiate populations from one another and can force the coherence within one population. The code generation of a nucleic acid sequence that functions as preclusion of certain RNA consortia by toxic effect which is not counter-regulated by an antitoxin generates a consortia which are not excluded, and therefore can generate a “self,” which originates a biological identity. Such basic RNA behavior is completely absent in abiotic environments.
Falsified: Most Viruses are Disease-Causing Selfish Parasites
In contrast to former wide spread opinions that viruses are strictly selfish and disease-causing agents, two different and important assumptions are investigated here:
-
(1)
Empirical evidence now clearly supports the view that in most cases, viruses follow a persistent non-lytic lifestyle which has been documented in a variety of exemplars and are prevalent in every known cellular hosts. Although in last decades we heard of “exotic” viral lifestyles, such as temperate phages or lysogenic strains of viral genomes, the predominant opinion about virus life styles was that of epidemic character of viral diseases. Recent metagenomic studies about viruses have fundamentally falsified this perspective. Although the previous disease-causing designation remains true, it is now acknowledged that the large majority of virus infections on this planet do not display this disease causing behavior. Now we know most viruses are inapparent settlers of cellular host organisms, both within the nucleus in the genome or as non-genomic settlers such as plasmids. We have heard much about non-coding RNAs, mobile genetic elements, repeat sequences, and introns— all of them formerly being termed “junk,” but now we can imagine these as remnants of persistent viral infection events. Thus, the persistent viral lifestyle is the most dominant biological lifestyle on this planet. And from this perspective, cellular host organisms look like islands in an ocean of the global virosphere. Viruses and virus-derived parts represent the most abundant genetic information on the planet overrepresenting cellular genetic information ten times. If we ignore eukaryotes and only consider prokaryotic life, we have a number of prokaryote viruses of 1031, which means if we line up the length of their virions, we get 40 million light years (Rohwer 2014). The visible living world of organisms of all domains is literarily the tip of the iceberg surrounded by biology’s viral “dark matter” of which we are currently scratching the surface at best (Youle et al. 2012). Importantly, a key feature of this viral lifestyle is that only few need to remain as functional agents, such as mammalian endogenous retroviruses needed for in the syncytia, which regulates mammalian pregnancy (Perot et al. 2012). Here, a much larger number of retroviral-derived code (LTRs) are providing the regulatory network of the placenta. In most cases, parts of infectious agents remain as defectives some known as LTRs, non-LTRs, SINEs, LINEs, and Alu’s which are later on co-adapted for cellular needs such as regulation tools in all steps and fine-tuned substeps of cellular functions such as, e.g., transcription, translation, epigenetics, repair, and immunity (Villarreal 2005; Roossinck 2015; Chalopin et al. 2012; Slotkin and Martienssen 2007; Conley and Jordan 2012).
-
(2)
The second but not less important point is that viruses can co-operate, that is they interact to build groups that invade host genomes and even compete as a group for limited resources such as host genomes. This leads to an extraordinary effective result and a key behavioral motif that is able to integrate a persistent lifestyle into cellular host organisms, the “addiction” modules: former competing viral groups are counterbalancing each other together with the host immune system (Villarreal 2012). Although rather stable under certain circumstances, this addiction balance can also get out of balance which means the competing viral features may become virulent again. But when stable, we can find such counter-regulating paired genes of the addiction modules, as in the RM (restriction/modulation) systems as well as in TA (toxin/antitoxin) systems. Insertion/deletion functions represent similar modules as do RM systems. This ‘infectious’ colonization by new addiction modules is a main process in generating new sequence space without error-replication and therefore in the evolution, conservation and plasticity of genetic identities.
Because this view contradicts well-entrenched mainstream perspectives on virus–host relationships, we will remind the reader throughout this article on these key features.
Before Group Selection Might be Considered, Group Identity Must Emerge
The historical dilemma of co-operation (symbiosis) versus competition (selfish behaviors) in biology was thought to have been resolved in the 1960s by evolutionary biologist. With the introduction of kin selection and later game theory, it seemed apparent that individual-based selection could mostly account for co-operation (Eigen and Winkler 1983). Thus, the acceptance of the individual-selfish types became well established. This view, which requires unending competition and conflict, also results in the domination and use of the war-like metaphors.
However, absent from all of these early debates was the concept that biological groups must initially form some type of group identity before group selection might be considered or understood. Group identity seemed to be a concept that applied to things like human social behavior and not to the behavior of genes, ribonucleoproteins, viruses, or organisms. But with the development of metagenomic analysis, we can now clearly see, e.g., major viral influence on prokaryotic genetic diversity. Viruses are often thought to affect their host survival via ‘killing the winner’ (most numerous/successful host). Thus, lytic viruses would seem able to keep the dominant host type in check. Viruses can also transfer sequences and genes between virus and host. It has been proposed that viruses are a main driving force in microbial diversification and diversity (Weinbauer and Rassoulzadegan 2004). Some authors prefer a war-like metaphor to describe the virus–host relationship (Forterre and Prangishvili 2009, Brüssow 2009). Similar reports are arguing for a destructive/constructive dynamic between killer virus and surviving host (Koonin 2011, Koonin and Dolja 2014). Thus, this is the accepted neodarwinian perspective of individual fittest type selection as essentially put forward by Dawkins (Dawkins 2006) and his selfish gene narrative.
Opposing Forces Linked to Survival: Cumulative Evolution
However, the above scenario of killer virus and war-like metaphor is only half the story and not the innovative half.
The amazing hidden half of the story: Although viruses can clearly kill their host, they can also colonize their host without disease and thus sometimes protect the host from the same or similar virus. Together with lysis and protection, we see a virus-colonized host that is both symbiotic and innovative (acquiring new competent code). Virus together with their non-infectious defective variants can sometimes persist in their host for long—even permanent—times without causing harm. Most importantly, the acquired virus information indeed can then provide the basis of antiviral defense (Villarreal 2005, 2009a, 2011a). This new virus–host combination can thus provide an ‘antiviral’ survival advantage, especially in a virus-rich habitat. Since it is transmissible, such survival can also apply to groups that were virus colonized (not necessarily direct kin). This virus/host relationship is not one that adheres to a war-like metaphor. But it represents a pervasive yet ancient and ongoing force in the origin and evolution of life.
The opposing forces of virus killing and virus protection together function to define and provide an acquired group identity linked to survival. Both the destructive and the immune function to destruction promote the synthesis of new identity. This represents three steps: evolution, conservation, and plasticity of genetic identities.
As noted, viruses are the most abundant and diverse genetic entities on earth (Ryan 2009, Moelling 2013, Rohwer 2014). As they seek to either colonize or lytically replicate in their host, they also provide a dynamic, consortial, and history-dependent behavior for cumulative evolution.
Cumulative evolution at the level of nucleic acid codes (genetic code based on DNA or RNA) is not solely what remains out of natural selection processes of mutations (error replication). Cumulative evolution is more like a “ratchet effect”, which as originally used was metaphor-like means for the cultural transmission of learned experiences to the next generations. This learning must accumulate so that every new generation must not need to repeat all innovative thoughts and techniques (Tomasello 2014). Interestingly, such “ratchet” (cumulative) effects are basically evident in the virus–host relationship of persistent colonizations, although this is a genotypic level in contrast the phenotypic level where this effect is originally described. Organism immunity, especially adaptive immunity (CRIPRs/Cas, VDJ-systems), represents excellent and highly efficient examples for this “ratchet” effect as they integrate history-dependent experiences with infectious elements into their newly ‘adapted’ genetic identity.
Additionally, the “marking” of DNAs via methylation and histone modification links the genome to environmental experiences in a memory-like learning function and is used in context-dependent “interpretation”, i.e., in transcriptional modifications (alternative splicing, RNA editing) and translation. We can consider how epigenetics might contribute to a cumulative evolution. Since it marks regions of DNA for non-expression, compared to unmarked regions, the expression patterns in subsequent generations can be affected thus show adaptation without genetic change. Interestingly, the origin of most all of these marking epigenetic systems appears to relate directly to the silencing and persistence of virus-like parasitic code. This marking allows the epigenetic inheritance of the differentiation in the variety of tissues of several organs throughout all known species during the timely regulated steps in developmental stages. Therefore, epigenetics is a rather intriguing example of how evolution, conservation, and plasticity of genetic identities are interlinked without the error-replication narrative, but related to genetic parasite colonization.
Therefore, we should not think in terms of destruction and error, but think of the great and innovative power of the ever-present virosphere (Villarreal 2009a), linking together former competing agents into groups that are competent to generate new genetic code sequences, with new genetic identities. Thus, the persistence of virus derived information matters in a big way that affects group identity. This perspective represents a major adjustment to our thinking.
Techniques, Tools, and Strategies of Co-operative Agents of the Virosphere
As mentioned before, a main strategy by which viruses affect host group survival is via ‘addiction modules’ (Villarreal 2012). Viruses can persist in the host by ‘addicting’ them. In this, they are providing protection (antitoxin) against the ever-present killing (toxin) by virus. This module provides a core ‘social force,’ linking two lineages (viral and host) into one group. Thus, it is via addiction modules that viruses provide a path to co-operation and symbiosis. Such a relationship was initially discovered via viruses of bacteria that persist but do not integrate into host DNA (episomal phage). And although this lesson in addiction was acquired from persisting DNA viruses of bacteria, it has led to generalized idea of virus-mediated group identity (Supplementary File 1). We can now also apply this idea to populations of infectious RNA agents, which will also apply to the very origins of life.
The study of RNA virus in the 1970s led to quasispecies theory, which is based on the idea that ‘error prone’ replication of master fittest type results in a population of mostly less fit variants around the master type (Eigen 1993; Eigen and Schuster 1977; Witzany 1995). As mentioned in the introduction, “error” is not the appropriate term to describe agents that are competent to use the natural genetic code and that act co-operatively also. Experimental virology since 1990 has redefined quasispecies as being co-operative. However, besides being co-operative, quasispecies are also exclusive of other quasispecies (Domingo et al. 2012). They have group recognition, which means they are able to differentiate self from non-self. These two features allow quasispecies to promote the emergence of group identity. The concept of group identity has not historically been crucial or important in biological theories. However, as we will assert below, group identity provides a fundamental and innovative force in biology that affects both the very earliest and most recent events in evolution (Villarreal 2005, 2014). When such concepts are applied to the behaviors and evolution of pre-cellular RNA populations of relatively simple structures (stem loops), we can define a basal and still active force in the emergence of life (Villarreal and Witzany 2013a).
Evolution of Early (RNA Based) Life was Communal
At least with the emergence of DNA-based life forms, we see clonal and individual types can exist. However, with the viruses, we still see the communal character that must have prevailed during early life. As noted, the ‘killing vs. survival’ character of virus and host has led too many to consider war-like metaphors. But the necessity of group identity (a general issue in the biology of cells, tissues, organs, and organisms) also requires that individual agents be able to participate in a coherent group, i.e., to co-operate (Villarreal 2009b).
The crucial difference to former concepts is that with group identity via counter-regulated addiction modules, we see that two opposing components must be present and work coherently to define the group as a whole. This means biological identity inherently is constituted by dynamic interactions of co-operative groups. Science has long approached complex systems from a reductionist (individual) perspective. But when the many make up the one (such as a co-operative group), coherent non-reductionistic approaches become necessary to understand the system (Woese 2004). Let us now consider how the virus, the most selfish of all genetic agents, can also inform us about co-operation and group behaviors.
Overall Conceptual Objectives: Group Identity as Essential Feature of Consortia
Thus, a core assertion is that a ‘virus addiction module’ is a general and essential strategy for existence of life in the virosphere. But because viruses are transmissible and can persist in specific host populations, this leads to a form of group immunity/identity since identical but uncolonized host populations remain susceptible to the killing actions of the lytic viruses. In this way, we see that the viruses are providing the necessary opposing functions for addiction (persistence/protection and lytic/killing).
The colonized host must retain persisting viruses (or defectives) in order to survive in the omnipresent virosphere and also to be able to kill competing but uncolonized host. It is important to note that this is a main underlying mechanism of host cell group identity (Villarreal 2012).
This concept would seem to suggest a deep importance as to why prokaryotes, for example, must retain cryptic virus information (Supplementary File 2) and might better explain the highly dynamic interactions of bacteria and their viruses. But the existence of virus mediated group identity can have very deep implications for all life. Indeed it can lead us to propose a role for this in the origin of life itself.
How Might a Virus Lifestyle Predate the Origin of Life?
Is not a virus a parasite of a cell therefore only able to emerge after the emergence of cells (and ribosomes)? But a virus at its core is a molecular genetic parasite that can parasitize every nucleic acid sequence even other virus systems. This means any proto-biotic replicator system can and likely will be susceptible to virus emergence and colonization.
However, the virus-mediated addiction module leads us to also think how opposing functions might emerge and support consortia functions or group identity in early life. This can now provide us with a major insight. If virus can function as a consortia (an essential interacting group), then it might provide mechanisms from which consortial functions themselves could emerge in the origin of proto-biotic life (Villarreal and Witzany 2013b). Genetic parasites can act as a group (qs-c). But for the groups to be coherent, they must attain group identity and this is typically via an addiction strategy (Villarreal 2009a). In general, antiviral (and antiproviral) systems (such as CRISPRs in archaea and bacteria) will themselves emerge in host from virus-derived information. Most importantly, it is the viruses that are providing the crucial functions needed for antiviral defense.
Such thinking, however, does not seem to adhere to the tenants of natural selection in which the variants from an individual fittest type must undergo natural selection. Instead, it fundamentally derives from an external consortia representing diverse information and is not generated by variation from a fittest individual. There need never have been an individual fittest type to get the consortia started. To initate the functional activity of a consortia, there need never have been an individual fittest type. The consortia itself provides the needed function. In addition, the transmissive (infectious-horizontal) nature of the consortia agents means that they need not decend from a common ancestor.
Instead, group identity becomes crucial and the participating agents become ‘one’ and this must express coupled opposing protective/replicative and destructive functions in order to define the group identity.
The opposing functions are the basis of addiction modules. Thus, the emergence of group identity becomes an essential and very early event in the emergence of life. This is coherent to the basically group behavior of RNA-based agents that are competent to use the natural genetic code as outlined in the introduction. This group selection and group identity are needed to create information coherence and network formation and to establish a system of communication, i.e., code–competent interactions: The identity serves as information also for the ones that do not share this identity (Villarreal and Witzany 2010, 2013b). This is the beginning of self/non-self differentiation capability.
When viral consortia successfully colonize a host, new virus–host combined identity (immunity) results. This colonization also creates new regulatory networks which will typically involve new addiction modules. With this new identity, there will also emerge an enhanced (cumulative) host complexity along with new virus/host evolutionary ecology (Villarreal 2005, 2011b). Survival in the virosphere will have been significantly modified. And a new host–host interaction and group survival mediated by virus will also result. Again we look at the core of evolution, conservation, and plasticity of genetic identities.
A Network is not a Linear Order of Nodes
However, in contrast to traditional and linear thinking, all these features (group identity, addiction modules, regulatory complexity, network emergence, host–virus ecology, and host–host competition) are fundamentally interlinked and consortial (Villarreal and Witzany 2013b). They are inherently network phenomena. But such networks will have a major historic and stochastic dependence. Thus, they will inherently resist any formal predictive analysis calculation as is done in systems biology. These identity networks cannot be teased apart to define a single and linear logic as is currently accepted for individual fittest type selection. Nor can one ‘node’ be set to one specific function as the participating agents will be context dependent and multifunctional. For example, one cannot understand the origin and function of the viral ‘toxin’ (including lysis) without also considering the viral ‘antitoxin’ (the opposition of “self” by persistence or defectives). And these two functions will respond rather differently to self and non-self agents (such as another virus). They must be considered to have emerged together in a particular context.
We will now apply this network membership idea to the origin of RNA-based life below. However, such opposing and multifunctional requirements for network identity will almost certainly confuse us. Our very language compels us to generate a coherent line up according grammar rules and as a result to think in linear terms and syntactic sequence order (like this text). And we have become imprinted to think first of individual fittest type mechanisms in order to explain co-operative and complex systems. We do not, for example, consider the possibility of gang-like agent action, as being important for innovative and multifunctional solutions to problems (as presented in the “Gangen” hypothesis below).
The Basic Motif: Virus-Derived Addiction Modules Promote Group Identity
The discovery of addiction modules and their relationship to persisting virus has mostly been in the context of bacterial dsDNA viruses. And from this perspective, we can also infer that collective action of dispersed seemly defective (cryptic) viruses is able to provide specific adaptive functions (such as mobilization, network control). But as asserted in our introduction, a host cell population that is persistently colonized by such a ‘controlled or cryptic’ virus set will also be able to provide information that resist the action of the equivalent lytic virus(es). Thus, a competing identical population of host cells that are not persistently colonized with the same cryptic virus will be susceptible to viral lysis when it becomes exposed to populations of cells that are persistently (non-lytically) infected (Fig. 1).
Schematic of virus affects on population-based host survival. The five diffuse circles (left) represent a host population free of the infectious virus in question. When exposed, many members will succumb to the toxic (acute) affects of virus infection (crossed lines). Some, however, may be stably colonized (shown with dark center). This host population has acquired a new virus derived instruction set that also provided immunity to the same (and often other) viruses (shown by broken lines between cells). If this population retains some capacity to produce infectious virus, or if the virus remains prevalent, when it encounters another naive population, the uncolonized population will crash due to virus toxicity. The virus colonized population will be favored. Reproduced with permission from Villarreal (2011b)
This is essentially why a ‘lysogenic’ strain of bacteria will lyse an identical bacterial strain that is not lysogenic when the two populations are mixed. The lysogenic strain can ‘reach out’ and kill its otherwise identical neighbor via transmissible virus (Villarreal 2011a, 2012). Since this can happen with episomally persisting agents, it need not directly involve the host DNA genome content (it can be epigenomic). The history of virus exposure and colonization will therefore determine whether a specific host population will be lysed or resist a particular virus. As mentioned before, this is historically derived and stochastic, however, and cannot be predicted. But to continue to favor survival of the virus persisting population, these cells must maintain both the capacity to resist virus as well as the capacity (or a habitat with the capacity) for the production of lytic virus.
Hence, virus ‘junk’ must remain as it is crucial for this. This is thus a ‘virus addiction module’ with both protective and destructive functions which are required to favor the survival of persistently infected populations, especially in a diverse and omnipresent virosphere.
In this way, viruses are promoting the emergence of a group identity in its host. The bacterial identity will be very much determined by its colonizing set of genetic parasites (Villarreal 2011b, 2012). Although such assertions seem broadly important, in our judgment what is even more broadly significant is that this situation defines a strategy by which a collective set of ‘sub-functional’ and opposing agents can participate in the genesis of a new collective function and group identity. This then importantly requires a coherent network that is inclusive of opposing functions (various TAs), but favors persistence of the parasite derived new information.
Importantly, this new information is not the result of error-replication but a result of module-like linked genetic contents. This fundamental difference to error-replication narratives proposes new nucleic acid sequence constructions by integration of larger content arrangements into a coherent syntax without destroying the already existing sequence content. Cryptic prophages are indeed the main source of new TA sets in prokaryotes, but such new sets must counter or interact with prior TA sets to persist. They must become coherent with their host. The big implication of this is that such a strategy should also apply to various RNA agents thought to have participated in the origin of life.
What addiction modules and group identity can now allow us to explore is how a collective of sub-functional RNA agents might have been able to become a coherent group that has both function and a TA system needed for group identity. RNA is the crucial population of agents that needs to be understood as a defined population. For it is this defined consortia that underlies the origin of life and the regulations of much of the complexity of higher organisms. Can virus addiction and group identity help explain innovative RNA functions?
Origin and Function of RNA and Virus is Linked to Addiction, Persistence, and Group Behavior
RNA is active directly (enzymatically, genetically) and not only as a messenger. Thus, RNA is more fundamental molecule of life than is the DNA. Accordingly, and in keeping with population-based functions of RNA, we have previously proposed that DNA can be considered as a habitat for a consortia RNA (Villarreal and Witzany 2013a; Witzany 2015b). Thus, ribosomes can be considered as functional RNA consortia that have inhabited DNA (discussed further below). In a sense, a focus on DNA genomes (including viral) give us a skewed view that DNA initiates major creative changes (via errors), with little or no role of RNA populations and diversity and genetic innovation.
A basal role of small RNA in regulation has not been as obvious in prokaryotes compared to eukaryotes. Yet, we have long also known that RNA must indeed have been more basal if there ever indeed did exist an RNA world that created life. And this world must also have created DNA. Several investigators have noted the striking strategic similarities between the RNA-based defense systems of prokaryotes and eukaryotes (Karginov and Hannon 2010; Cooper and Overstreet 2014; Koonin and Krupovic 2014). Yet these systems share no homology with each other and must have derived from distinct ancestors. However, in both these domains of life, as well as in the interferon system of vertebrates and all RNA viruses, small RNAs with double-strand (stem-loop) regions are crucial for cellular defense recognition and response. Above, we noted that small RNAs can also be components of TA systems. Historically, little attention was given to the role of small RNA in basic regulatory functions of cells. More recently, we have come to learn that here too small RNA’s (such as CRISPRS or even tRNAs) are much more involved in basic regulatory functions (Nicolas et al. 2013. And in eukaryotes, there has been a big change in our thinking regarding regulation by RNA (Cech and Steitz 2014). Thus, we propose to place RNA-based regulation in a more fundamental role (Supplementary File 3).
As with transition from prokaryotes to eukaryotes, we see some striking differences regarding the activity and amount of parasitic RNA agents. Prokaryotic genetic changes are mostly driven by dsDNA parasites (virus and plasmids) as noted above. In eukaryotes, RNA agents (retroviruses and retroposons) are much more diverse, numerous and dynamic and are providing multiple levels of regulatory complexity. We have recently come to realize that transcription of such retroposon sequence (previously considered junk) is abundant and often produces non-coding RNAs, all being part of stem-loop regions (Villarreal and Witzany 2013b).
It is such RNA that is involved in complex multicellular identity. However, as we will now present, retroviruses are the major initiators of retroposon mediated changes in eukaryotes and the fitness of retroviral RNA (like all RNA) is fundamentally consortial. But this is not the quasispecies as most have come to understand it is based on error-replication and master fittest type concept of Manfred Eigen. It is fundamentally a co-operative and counteractive version of quasispecies (Domingo et al. 2012) that also supports group identity. Life will only emerge from consortial systems with group identity competent to use and edit the natural genetic code. Some RNA-based life forms, like ‘RNA only’ viruses, cannot persist as DNA and persist as RNA. Similar to the ancestral RNA, early life must have also persisted either as a dynamic RNA population or as a sequestered (static) RNA population. One present day example of dynamic RNA only persistence is Hepatitis C Virus HCV. The creative and consortial action of RNA populations remains a potent and ongoing force in the evolution, conservation, and plasticity of the genetic identities of the most complex life forms that continue to inhabit DNA.
Retroviral Networks Regulate Evolution and Development
Retroviruses also clearly generate and operate via quasispecies (Villarreal 2009c). But in contrast to the RNA-only viruses, retroviruses persist as and are copied from DNA and have also provided a large amount of genomic DNA sequence especially long terminal repeats (LTRs) as found in most eukaryotes (Shapiro 2005). If such genomic endogenous retrovirus (ERV) sequences are also produced by QS-mediated evolution, then their involvement in the formation of new or edited networks regulating host functions might be understood as resulting from a consortial RNA-based process with inherent coherence. Indeed, understanding the origin of transposable RNA-based networks (and network security) has always been challenging as networks do not fit into tree-based analogies (Bapteste et al. 2013; Daly et al. 2011; Feschotte 2008).
It also appears that various small non-coding RNAs participate in ‘multi-task’ networks and such RNAs tend to be transcribed from ‘junk’ retroposons (Mattick 2011; Mattick and Gagen 2001; Mattick and Makunin 2006; Pheasant and Mattick 2007). In terms of active editors of the human genome, there are about 330,000 solo LTRs (Oliver and Greene 2011, 2012) each of which must have initially corresponded to an intact ERV (~10 kb) subsequently lost by deletion. This means that 3.3 gB of human DNA (current size of our genome) was once retrovirus during our evolution. But such LTRs are highly involved in the emergence of new regulatory networks, such as the origin of the placenta (Bièche et al. 2003; Chuong et al. 2013; Emera and Wagner 2012; Harris 1998; Nakagawa et al. 2013) (re-regulating 1500 genes) Lynch et al. 2011, 2012) or in the African primates where alteration of 320,000 LTR p53 binding sites occurred onto the p53 cell cycle control network (Wang et al. 2007). These primate p53 network changes also relate to (co-operate with) changes in brain specific microRNAs (Le et al. 2009), alterations to DNA methylation involved in controlling SINE-derived RNA transcription (Leonova et al. 2013), as well as Alu-derived transcription (Zemojtel et al. 2009), an interconnected situation as seen in other networks. (Supplementary File 4)
Group Identity and Co-operativity of an RNA Collective: Essential Roles of Defective Minorities
In proposing the qs-c concept, it was argued that agent diversity (not errors) was essential for the capacity of a collective of RNA agents to function co-operatively (Villarreal and Witzany 2013b). Thus, an identity group of sub-functional RNA agents would be the predecessors of RNA-based life (not a functioning individual). And the type of Darwinian (individual fittest type) selection we are now so familiar with would not emerge until DNA emerged to provide individual genomes. DNA essentially functions as a habitat for the living RNA collective (Villarreal and Witzany 2013b). But with early life (prior to DNA), such an RNA collective must have been able to operationally hold itself together in order to function as a selectable population. And this would most likely be via both a commonly shared syntax of a natural nucleic acid code and a dynamic state (with ongoing replication), given the unstable highly productive (former ‘error-prone’) nature of RNA. Thus, in order to behave as a population or group, a qs-c must have some process that compels its coherence. Fundamentally, this is inherent in a qs-c behavior. But robust coherence of a population would also require a process that prevents the occurrence of both overly potent individual defectives as well as overly active individual replicators.
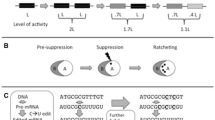
Significantly, self-parasitizing defectives can provide this control. Thus, there is an essential requirement for defective minorities providing functional (inhibitory) diversity. Minorities in the population will also retain memory (pre-requisite for learning) from past group selection events. And group process must also oppose non-members of other qs-c’s, just as observed with the RNA viruses. These negative (toxic) functions themselves are also likely to emerge from co-operative action of sub-functional RNA agents. Hence, to attain coherent group behavior, the group must also attain coherent group identity. And the occurrence of any participating TA function would need to be coherent with the rest of the TAs found in the population. In terms of the emergence of an RNA ribozyme-based living collective, it will need to have both opposing ribozyme activities; replication (ligation) and endonuclease to provide a coherent TA set (see Fig. 2).
Schematic for a ‘gangen’ of RNA that promotes the emergence of group identity, communication, and co-operativity from an RNA collective that also requires opposing functions. Reproduced with permission from Villarreal (2014)
Thus, the collective must initially emerge as a collective, with group identity mediated by co-operative sub-functional agents that together provide both the ligase (positive) and endonuclease (negative) functional features of an addiction module. And this means there was no ancestral individual fittest type, the collective was always a dynamic network with clearly defined membership (security, immunity), that depends on internal competition, co-operation, and opposing functions (antisense), while retaining a history of RNA agent colonization and their corresponding TA sets. All these aspects must attain coherence and provide coherent communication, genetic code-use, and group identity. In the origin of life, this was mediated mostly via a collective of stem-loop RNAs. Villarreal calls this hypothesis for the emergence of collective RNA-based life the ‘Gangen’ hypothesis as diagramed in Fig. 2. All the features noted above are included in the diagram (Villarreal 2014).
The ‘Gangen’ Hypothesis
The term comes from archaic Nordic. Gangen was an early Nordic term applied to pathways (gangway) but also led to descriptions of collectives (gangs) with clear collective functional abilities and group identities although participating members may highly vary according to dynamic changes in the real-life world environment. A Gangen (unlike a collective) must attain group identity (Villarreal 2014). Thus, it also describes the emergence of commonly shared code-use, group membership, and the collective living functions of the RNA agents. Membership is not a byproduct of individual selection, but enforced by the required toxic capacity within the collective. Note also that this collective, because it is dynamic and depends on diversity, will also retain memory of its history such as remaining minorities. Also the remaining minorities share this competence to use the natural genetic code. Emergence of a Gangen is therefore not a simple, chemically predetermined event. It depended on historic and stochastic agents that were able to join the collective and add and edit genetic code and its meaning (use).
This hypothesis provides a distinction between the principles of chemistry and biology (a living restricted collective with history and communication). Clearly, there is more to understanding the emergence of life than this hypothesis alone can account for. For example, the physical containment of the qs population (such as via membranes) or the source of metabolic energy, substrates, etcetera, and the role of amino acids as catalytic RNA primers or markers of replicator identity is not addressed and will not be considered here.
But there is an additional feature that should be emphasized, that is communication (code dependent interactions). Transmission of infectious code defines the origin of the virosphere. This issue reduces to the idea that a collective of agents (RNA) with inherent toxic and antitoxic features should be able to transmit (communicate) these agents and their features to nearby competing populations (via simple diffusion). Such transmission is essentially infectious and very much like a virus (or viroid). But in communicating RNA-based TAs, it strongly favors the survival of the RNA population with the compatible addiction modules that will inhibit agent toxicity (prevent lysis via ‘defective’ code) and allow persistence of the new agents. This is thus the survival of the persistently colonized (infected) set, which is an inherently symbiotic and consortial process. It also promotes increasing complexity (and identity/immunity) of the host collective via new agent colonization and stable addition. Thus, the transmission of RNA agents attains both communication (competent nucleic acid code-use) and recognition of group membership. In this way, the emergence of a ‘virosphere’ must also have been an early event in the origin of life, one that will shape communication of natural code and create group identity (Villarreal 2014) and therefore clearly represents evolution, conservation, and plasticity of genetic identities.
This concept differs fundamentally from current (and highly successful) view based on individual type selection of DNA-based organisms. Below, we assemble some evidence from study or RNA, which clearly supports the existence of collective phenomena in the origin of life.
Quasispecies Consortia (qs-c): Origin of Ribozymes and Co-operating Stem-loop RNAs
As noted, all investigations into the role of RNA in the origin of life assume that some form of ‘master fittest type’ of RNA existed, which was able to function as a ribozyme and inefficiently copies itself with a high error rate as essentially outlined initially by Eigen (Eigen 1971, 2013; Eigen and Schuster 1977). Thus, the original RNA replicator must have functioned as an individual ribozyme molecule. However, it has been noted that group selection of early replicators along with compartmentalization might be required to integrate information in the origin of life (Szathmary and Demeter 1987). Previously, however, the qs-c version of RNA selection has not been considered, and as it asserts that a sub-functional collective of RNA agents would be ancestral to effective ribozyme-based replication, it makes a very different prediction with respect to the unnecessary master fittest type (Villarreal and Witzany 2013b).
An assembly of sub-functioning RNAs could be produced by pure chemical mechanisms. But for this assembly to form a functional collective, it must attain a “Gangen” state (see above): A collective state of group identity based on commonly shared genetic code-use and co-operative functionality. Thus, bridging the split from proto-biotic assemblies to biotic groups requires agents competent for genetic code-use and co-operation. This can then provide both the combined positive and negative chemical activities of ribozymes.
Recently, there has been an accumulation of experimental evidence that RNA ribozymes do act and emerge from collectives that can also form networks. Very small hairpin ribozymes are known to have catalytic activity (Yarus 2011; Muller et al. 2012). Populations of evolving ligase ribozymes have been maintained by in vitro serial diluted passage (McGinness et al. 2002). More recently, the participation of two RNAs that participate in each other’s synthesis from four substrates (via co-operation) has been observed (Lincoln and Joyce 2009; Ferretti and Joyce 2013). Others have also used multiple (up to 4) stem-loop ribozymes together to select for combined ribozyme activity (Gwiazda et al. 2012). Similarly, 4 subfunctional fragments of group I intron ribozyme can self-assemble too into an autocatalytic ribozyme (Hayden and Lehman 2006). It has been established that group I ribozymes must undergo co-operative interactions that depend on native helix orientation to attain their functional 3D folds (Behrouzi et al. 2012). Co-operative fragments of RNA replicators have also been observed to spontaneously self-assemble and generate a network with co-operative catalytic activity (Vaidya et al. 2012). In such a network, a single RNA molecule can be multifunctional in an RNA pathway (Vaidya 2012).
Together, these results provide strong experimental evidence of the co-operative potential of sub-functional ribozymes. However, in none of these discussions has the issue of network membership (or group identity) been considered. According to the “Gangen” hypothesis, network membership along with their various addiction modules would also be essential for life to emerge from the RNA world. Accordingly, the ligation and endonuclease activity of ribozymes would need to emerge together to provide a TA set of functions.
It is now possible to consider many ancient and recent issues in evolutionary biology from the perspective of qs-c and the co-operative interaction of stem-loop RNAs. For example, ribosomes are essentially very complex ribozymes which are composed of a complicated set of covalently linked stem-loop RNAs which interact in complex ways to provide it with its core function, catalytic synthesis of peptide bonds (Bokov and Steinberg 2009). Given that their individual stem loops appear to have distinct evolutionary histories, the ribosome seems to represent consortia of stem loops that were built up historically over time during evolution (Harish and Caetano-Anollés 2012). Thus, when ribosomes became a resident of DNA in the first cells, the stem-loop RNA consortia were made stable. This again represents evolution, conservation, and plasticity of a genetic identity.
We can also evaluate very recent events in evolution from the qs-c perspective. Consider the extremely complex neuronal cell identity and communication issues that must apply to the nervous system of hominids. There is an emerging view for a basic role of non-coding retroposon-derived stem-loop RNAs in cell identity and neuronal network formation (Oldham et al. 2006; Qureshi and Mehler 2009; Barry and Mattick 2012; Qureshi and Mehler 2012). Indeed, as we reevaluate the human genome and consider the presence of several hundred thousand solo LTRs (many human specific), we might reconsider how these ‘many’ infectious agents co-operated to become the ‘one’ RNA collective we call human.
Conclusion
Viruses and virus-like infectious genetic parasites are the most abundant living entities on earth that outdate cellular life more than ten times. All living cellular organisms have always operated in a virosphere. And a virosphere is essentially a network of infectious genetic agents. The real survival of all organisms must always be considered in the context of its virosphere. This realization is thus a very recent but major shift in our thinking. Most experiments that evaluate the fitness (survival) of an organism ignore the virosphere and thus provide both artificial and unrealistic situations and outcomes for survival which have fundamentally misled us. For example, when we establish a sterile mouse colony free of all the usual persistent mouse viruses, we create an artificial laboratory habitat for survival. When we clone E. coli free of temperate and lytic phage, we similarly create an artificial laboratory artifact for survival. To get a coherent view on in vivo habitats in the context of real-life circumstances, we always must assume a virosphere perspective.
Although Eigen’s quasispecies concept predominated evolution concepts nearly half a century, it could not coherently explain empirical data of RNA groups and viruses that co-operate. In this review, we have demonstrated and exemplified that evolution, conservation, and plasticity of genetic identities are the result of co-operative consortia of RNA stem loops that are competent to communicate, i.e., build groups that use natural genetic code and edit this code, even by the generation of new sequences without error replication. The highly productive (not “error-prone”) capability to generate new sequences allows such groups to constantly infect other nucleic sequence-based agents, whether they have virus-like or cellular genomes. The generation of such new sequences by co-operating RNA stem-loop groups leads to identity groups of viruses that can function as toxic and antitoxic codes. Infected host organisms are the habitats in which such formerly competing agent groups (“Gangen”) now unify in addiction modules, which provide group identity such as TA, RM, and ID modules. Thus, all of the former competing groups become unified into stable/unstable modules that are counter-regulated, to provide immunity and memory systems, such as VDJ and CRISPRS/Cas, against related genetic parasites for the host. In this way what historically seemed to be competing and selfish viruses can instead provide a unifying collective of viruses (and their defective participants) to better explain evolution, conservation as well as the plasticity and often cumulative genetic identities of complex organisms more coherently than the previous quasispecies concept.
References
Anand SK, Griffiths MW (2003) Quorum sensing and expression of virulence in Escherichia coli O157:H7. Int J Food Microbiol 85:1–9
Archer EJ, Simpson MA, Watts NJ, O’Kane R, Wang B, Erie DA et al (2013) Long-range architecture in a viral RNA genome. Biochemistry 52:3182–3190
Asadulghani M, Ogura Y, Ooka T, Itoh T, Sawaguchi A, Iguchi A et al (2009) The defective prophage pool of Escherichia coli O157: prophage-prophage interactions potentiate horizontal transfer of virulence determinants. PLoS Pathog 5:e1000408
Bapteste E, van Iersel L, Janke A, Kelchner S, Kelk S, McInerney JO et al (2013) Networks: expanding evolutionary thinking. Trends Genet 29:439–441
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Barry G, Mattick JS (2012) The role of regulatory RNA in cognitive evolution. Trends Cogn Sci 16:497–503
Behrouzi R, Roh JH, Kilburn D, Briber RM, Woodson SA (2012) Cooperative tertiary interaction network guides RNA folding. Cell 149:348–357
Berkhout B, Van Wamel JLB (2000) The leader of the HIV-1 RNA genome forms a compactly folded tertiary structure. RNA 6:282–295
Bièche I, Laurent A, Laurendeau I, Duret L, Giovangrandi Y, Frendo J-L et al (2003) Placenta-specific INSL4 expression is mediated by a human endogenous retrovirus element. Biol Reprod 68:1422–1429
Bokov K, Steinberg SV (2009) A hierarchical model for evolution of 23S ribosomal RNA. Nature 457:977–980
Bondy-Denomy J, Davidson AR (2014) When a virus is not a parasite: the beneficial effects of prophages on bacterial fitness. J Microbiol 52:235–242
Briones C, Stich M, Manrubia SC (2009) The dawn of the RNA world: toward functional complexity through ligation of random RNA oligomers. RNA 15:743–749
Brüssow H (2009) The not so universal tree of life or the place of viruses in the living world. Philos Trans R Soc Lond B 364:2263–2274
Canchaya C, Fournous G, Chibani-Chennoufi S, Dillmann ML, Brüssow H (2003) Phage as agents of lateral gene transfer. Curr Opin Microbiol 6:417–424
Canchaya C, Fournous G, Brüssow H (2004) The impact of prophages on bacterial chromosomes. Mol Microbiol 53:9–18
Carbonell A, Flores R, Gago S (2012) Hammerhead ribozymes against virus and viroid RNAs. In: Erdmann VA, Barciszewski J (eds) From nucleic acids sequences to molecular medicine. Springer, Berlin, Heidelberg, pp 411–427
Cech TR, Steitz JA (2014) The noncoding RNA revolution—trashing old rules to forge new ones. Cell 157:77–94
Chalopin D, Tomaszkiewicz M, Galians D, Volff JN (2012) LTR retroelement-derived protein-coding genes and vertebrate evolution. In: Witzany G (ed) Viruses: essential agents of life. Springer, Dordrecht, pp 269–282
Chikova AK, Schaaper RM (2005) The bacteriophage P1 hot gene product can substitute for the Escherichia coli DNA polymerase III theta subunit. J Bacteriol 187:5528–5536
Chuong EB, Rumi MAK, Soares MJ, Baker JC (2013) Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat Genet 45:325–329
Conley AB, Jordan IK (2012) Endogenouis retroviruses and the epigenome. In: Witzany G (ed) Viruses: essential agents of life. Springer, Dordrecht, pp 309–323
Cooper EL, Overstreet (2014) Diversity, evolution, and therapeutic applications of small RNAs in prokaryotic and eukaryotic immune systems. Phys Life Rev 11:113–134
Daly T, Chen XS, Penny D (2011) How old are RNA networks? In: Collins LJ (ed) RNA infrastructure and networks. Springer, New York, pp 255–273
Dawkins R (2006) The selfish gene: 30th anniversary edition. Oxford University Press, Oxford
Domingo E, Sheldon J, Perales C (2012) Viral quasispecies evolution. Microbiol Mol Biol Rev 76:159–216
Dy RL, Przybilski R, Semeijn K, Salmond GPC, Fineran PC (2014) A widespread bacteriophage abortive infection system functions through a Type IV toxin–antitoxin mechanism. Nucleic Acids Res 42:4590–4605
Edgar R, Qimron U (2010) The Escherichia coli CRISPR system protects from λ lysogenization, lysogens, and prophage induction. J Bacteriol 192:6291–6294
Eigen M (1971) Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften 58:465–523
Eigen M (1993) Viral quasispecies. Sci Am 269:42–49
Eigen M (2013) From strange simplicity to complex familiarity: a treatise on matter, information, life and thought. Oxford University Press, Oxford
Eigen M, Schuster P (1977) The hypercyde. A principle of natural self-organization. Naturwissenschaften 64:541–565
Eigen M, Winkler R (1983) The Laws of the game: how the principles of nature govern chance. Princeton University Press, Princeton
Emera D, Wagner GP (2012) Transposable element recruitments in the mammalian placenta: impacts and mechanisms. Brief Funct Genomics 11:267–276
Ferretti AC, Joyce G (2013) Kinetic properties of an RNA enzyme that undergoes self-sustained exponential amplification. Biochemistry 52:1227–1235
Feschotte C (2008) Transposable elements and the evolution of regulatory networks. Nat Rev Genet 9:397–405
Fineran PC, Blower TR, Foulds IJ, Humphreys DP, Lilley KS, Salmond GP (2009) The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair. Proc Natl Acad Sci USA 106:894–899
Flores R, Gago-Zachert S, Serra P, Sanjuan R, Elena SF (2014) Viroids: survivors from the RNA world? Annu Rev Microbiol 68:395–414
Forterre P, Prangishvili D (2009) The great billion-year war between ribosome- and capsid-encoding organisms (cells and viruses) as the major source of evolutionary novelties. Ann N Y Acad Sci 1178:65–77
Garcia-Contreras R, Zhang XS, Kim Y, Wood TK (2008) Protein translation and cell death: the role of rare tRNAs in biofilm formation and in activating dormant phage killer genes. PLoS ONE 3(6):e2394
Garrett RA, Vestergaard G, Shah SA (2011) Archaeal CRISPR-based immune systems: exchangeable functional modules. Trends Microbiol 19:549–556
Gazit E, Sauer RT (1999) Stability and DNA binding of the phd protein of the phage P1 plasmid addiction system. J Biol Chem 274:2652–2657
Gomaa AA, Klumpe HE, Luo ML, Selle K, Barrangou R, Beisel CL (2014) Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems. mBio 5(1):e00928–13
Guo P, Cheng Q, Xie P, Fan Y, Jiang W, Qin Z (2011) Characterization of the multiple CRISPR loci on Streptomyces linear plasmid pSHK1. Acta Biochim Biophys Sin 43:630–639
Gwiazda S, Salomon K, Appel B, Muller S (2012) RNA self-ligation: From oligonucleotides to full length ribozymes. Biochimie 94:1457–1463
Harish A, Caetano-Anollés G (2012) Ribosomal history reveals origins of modern protein synthesis. PLoS ONE 7:e32776
Harris JR (1998) Placental endogenous retrovirus (ERV): structural, functional, and evolutionary significance. BioEssays 20:307–316
Haurwitz RE, Jinek M, Wiedenheft B, Zhou K, Doudna JA (2010) Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329:1355–1358
Hayden EJ, Lehman N (2006) Self-assembly of a group i intron from inactive oligonucleotide fragments. Chem Biol 13:909–918
Hazan R, Engelberg-Kulka H (2004) Escherichia coli mazEF-mediated cell death as a defense mechanism that inhibits the spread of phage P1. Mol Genet Genomics 272:227–234
Hazan R, Sat B, Reches M, Engelberg-Kulka H (2001) Postsegregational killing mediated by the P1 phage “addiction module” phd-doc requires the Escherichia coli programmed cell death system mazEF. J Bacteriol 183:2046–2050
Karginov FV, Hannon GJ (2010) The CRISPR system: small RNA-guided defense in bacteria and archaea. Mol Cell 37:7–19
Kim Y, Wang X, Ma Q, Zhang XS, Wood TK (2009) Toxin-antitoxin systems in Escherichia coli influence biofilm formation through YjgK (TabA) and fimbriae. J Bacteriol 191:1258–1267
Koonin EV (2011) The logic of chance: the nature and origin of biological evolution. FT Press, Upper Saddle River
Koonin EV, Dolja VV (2014) Virus world as an evolutionary network of viruses and capsidless selfish elements. Microbiol Mol Biol Rev 78:278–303
Koonin EV, Krupovic M (2014) Evolution of adaptive immunity from transposable elements combined with innate immune systems. Nat Rev Genet. doi:10.1038/nrg3859
Lawrence CM, Menon S, Eilers BJ, Bothner B, Khayat R, Douglas T, Young MJ (2009) Structural and functional studies of archaeal viruses. J Biol Chem 284:12599–12603
Le MT, Teh C, Shyh-Chang N, Xie H, Zhou B, Korzh V et al (2009) MicroRNA-125b is a novel negative regulator of p53. Genes Dev 23:862–876
Lehnherr H, Yarmolinsky MB (1995) Addiction protein Phd of plasmid prophage P1 is a substrate of the ClpXP serine protease of Escherichia coli. Proc Natl Acad Sci USA 92:3274–3277
Lehnherr H, Maguin E, Jafri S, Yarmolinsky MB (1993) Plasmid addiction genes of bacteriophage P1: doc, which causes cell death on curing of prophage, and phd, which prevents host death when prophage is retained. J Mol Biol 233:414–428
Leonova KI, Brodsky L, Lipchick B, Pal M, Novototskaya L, Chenchik AA et al (2013) p53 cooperates with DNA methylation and a suicidal interferon response to maintain epigenetic silencing of repeats and noncoding RNAs. Proc Natl Acad Sci 110:E89–E98
Lillestøl RK, Redder P, Garrett RA, Brügger K (2006) A putative viral defence mechanism in archaeal cells. Archaea 2:59–72
Lincoln TA, Joyce GF (2009) Self-sustained replication of an RNA enzyme. Science 323:1229–1232
Lynch VJ, Leclerc RD, May G, Wagner GP (2011) Transposon-mediated rewiring of gene regulatory networks contributed to the evolution of pregnancy in mammals. Nat Genet 43:1154–1159
Lynch VJ, Nnamani M, Brayer KJ, Emera D, Wertheim JO, Pond SLK, et al. (2012) Lineage-specific transposons drove massive gene expression recruitments during the evolution of pregnancy in mammals. arXiv: 1208.4639. http://arxiv.org/abs/1208.4639. Accessed 18 Dec 2012
Magnuson R, Lehnherr H, Mukhopadhyay G, Yarmolinsky MB (1996) Autoregulation of the plasmid addiction operon of bacteriophage P1. J Biol Chem 271:18705–18710
Marraffini LA, Sontheimer EJ (2010) Self versus non-self discrimination during CRISPR RNA-directed immunity. Nature 463:568–571
Mattick JS (2011) The central role of RNA in human development and cognition. FEBS Lett 585:1600–1616
Mattick JS, Gagen MJ (2001) The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms. Mol Biol Evol 18:1611–1630
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 1:R17–29
McGinness KE, Wright MC, Joyce GF (2002) Continuous in vitro evolution of a ribozyme that catalyzes three successive nucleotidyl addition reactions. Chem Biol 9:585–596
Miller WA, White KA (2006) Long-distance RNA-RNA interactions in plant virus gene expression and replication. Ann Rev Phytopathol 44:447–467
Mochizuki T, Sako Y, Prangishvili D (2011) Provirus induction in hyperthermophilic archaea: characterization of aeropyrum pernix spindle-shaped virus 1 and aeropyrum pernix ovoid virus 1. J Bacteriol 193:5412–5419
Moelling K (2013) What contemporary viruses tell us about evolution: a personal view. Arch Virol 158:1833–1848
Mojica FJM, Díez-Villaseñor C, García-Martínez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182
Moons P, Werckx W, Van Houdt R, Aertsen A, Michiels CW (2006) Resistance development of bacterial biofilms against bacteriophage attack. Commun Agric Appl Biol Sci 71:297–300
Muller S, Appel B, Krellenberg T, Petkovic S (2012) The many faces of the hairpin ribozyme: Structural and functional variants of a small catalytic RNA. IUBMB Life 64:36–47
Nakagawa S, Bai H, Sakurai T, Nakaya Y, Konno T, Miyazawa T et al (2013) Dynamic evolution of endogenous retrovirus-derived genes expressed in bovine conceptuses during the period of placentation. Genome Biol Evol 5:296–306
Nicolas FE, Torres-Martinez S, Ruiz-Vazquez RM (2013) Loss and retention of RNA interference in fungi and parasites. PLoS Pathog 9(1):e1003089
Nozawa T, Furukawa N, Aikawa C, Watanabe T, Haobam B, Kurokawa K et al (2011) CRISPR inhibition of prophage acquisition in streptococcus pyogenes. PLoS ONE 6(5):e19543
Ogura Y, Ooka T, Asadulghani M, Terajima J, Nougayrede JP, Kurokawa K et al (2007) Extensive genomic diversity and selective conservation of virulence-determinants in enterohemorrhagic Escherichia coli strains of O157 and non-O157 serotypes. Genome Biol 8:R138
Oldham MC, Horvath S, Geschwind DH (2006) Conservation and evolution of gene coexpression networks in human and chimpanzee brains. Proc Natl Acad Sci USA 103:17973–17978
Oliver KR, Greene WK (2011) Mobile DNA and the TE-Thrust hypothesis: supporting evidence from the primates. Mob DNA 2:8–25
Oliver KR, Greene WK (2012) Transposable elements and viruses as factors in adaptation and evolution: an expansion and strengthening of the TE-Thrust hypothesis. Ecol Evol 2:2912–2933
Ooka T, Ogura Y, Asadulghani M, Ohnishi M, Nakayama K, Terajima J et al (2009) Inference of the impact of insertion sequence (IS) elements on bacterial genome diversification through analysis of small-size structural polymorphisms in Escherichia coli O157 genomes. Genome Res 19:1809–1816
Ortmann AC, Wiedenheft B, Douglas T, Young M (2006) Hot crenarchaeal viruses reveal deep evolutionary connections. Nat Rev Microbiol 4:520–528
Palmer KL, Whiteley M (2011) DMS3-42: The secret to CRISPR-dependent biofilm inhibition in Pseudomonas aeruginosa. J Bacteriol 193:3431–3432
Perot P, Bolze PA, Mallet F (2012) From viruses to genes: syncytins. In: Witzany G (ed) Viruses: essential agents of life. Springer, Dordrecht, pp 325–361
Pheasant M, Mattick JS (2007) Raising the estimate of functional human sequences. Genome Res 17:1245–1253
Prangishvili D (2013) The wonderful world of archaeal viruses. Ann Rev Microbiol 67:565–585
Qureshi IA, Mehler MF (2009) Regulation of non-coding RNA networks in the nervous system–what’s the REST of the story? Neurosci Lett 466:73–80
Qureshi IA, Mehler MF (2012) Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat Rev Neurosci 13:528–541
Redfield RJ, Campbell AM (1984) Origin of cryptic lambda prophages in Escherichia coli K-12. Cold Spring Harb Symp Quant Biol 49:199–206
Rohwer F (2014) Life in our phage world. San Diego, Wholon
Roossinck MJ (2015) Metagenomics of plant and fungal viruses reveals an abundance of persistent lifestyles. Front Microbiol 5:1–3
Ryan F (2009) Virolution. Collins, London
Shapiro JA (2005) Retrotransposons and regulatory suites. BioEssays 27:122–125
Slotkin RK, Martienssen R (2007) Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet 8:272–285
Snyder JC, Brumfield SK, Kerchner KM, Quax TEF, Prangishvili D, Young MJ (2013) Insights into a viral lytic pathway from an archaeal virus-host system. J Virol 87:2186–2192
Szathmary E, Demeter L (1987) Group selection of early replicators and the origin of life. J Theor Biol 128:463–486
Takagi H, Kakuta Y, Okada T, Yao M, Tanaka I, Kimura M (2005) Crystal structure of archaeal toxin-antitoxin RelE–RelB complex with implications for toxin activity and antitoxin effects. Nat Struct Mol Biol 12:327–331
Tomasello M (2014) A natural history of human thinking. Havard University Press, Cambridge
Touchon M, Charpentier S, Clermont O, Rocha EP, Denamur E, Branger C (2011) CRISPR distribution within the Escherichia coli species is not suggestive of immunity-associated diversifying selection. J Bacteriol 193:2460–2467
Tyndall C, Lehnherr H, Sandmeier U, Kulik E, Bickle TA (1997) The type IC hsd loci of the enterobacteria are flanked by DNA with high homology to the phage P1 genome: implications for the evolution and spread of DNA restriction systems. Mol Microbiol 23:729–736
Vaidya N (2012) Spontaneous Cooperative Assembly of Replicative Catalytic RNA Systems. Dissertations and Theses. http://pdxscholar.library.pdx.edu/open_access_etds/934
Vaidya N, Manapat ML, Chen IA, Xulvi-Brunet R, Hayden EJ, Lehman N (2012) Spontaneous network formation among cooperative RNA replicators. Nature 491:72–77
Villarreal LP (2005) Viruses and the evolution of life. ASM Press, Washington
Villarreal LP (2009a) Origin of group identity. Viruses, addiction and co-operation. Springer, New York
Villarreal LP (2009b) Persistence pays: how viruses promote host group survival. Curr Opin Microbiol 12:467–472
Villarreal LP (2009c) The source of self: genetic parasites and the origin of adaptive immunity. Ann N Y Acad Sci 1178:194–232
Villarreal LP (2011a) Viral ancestors of antiviral systems. Viruses 3:1933–1958
Villarreal L (2011b) Viruses and host evolution: virus-mediated self identity. In: Lopez-Larrea C (ed) Self and Non-Self. Landes Bioscience and Springer Science+Business Media, Austin, pp 185–217
Villarreal LP (2012) The addiction module as a social force. In: Witzany G (ed) Viruses: essential agents of life. Springer Science+Business Media, Dordrecht, pp 107–145
Villarreal LP (2014) Force for ancient and recent life: viral and stem-loop RNA consortia promote life. Ann N Y Acad Sci. doi:10.1111/nyas.12565
Villarreal LP, Witzany G (2010) Viruses are essential agents within the roots and stem of the tree of life. J Theor Biol 262:698–710
Villarreal LP, Witzany G (2013a) The DNA habitat and its RNA inhabitants: at the dawn of RNA sociology. Genomics Insights 6:1–12
Villarreal LP, Witzany G (2013b) Rethinking quasispecies theory: from fittest type to cooperative consortia. World J Biol Chem 4:71–82
Wang T, Zeng J, Lowe CB, Sellers RG, Salama SR, Yang M et al (2007) Species-specific endogenous retroviruses shape the transcriptional network of the human tumor suppressor protein p53. Proc Natl Acad Sci USA 104:18613–18618
Wang X, Kim Y, Ma Q, Hong SH, Pokusaeva K, Sturino JM, Wood TK (2010) Cryptic prophages help bacteria cope with adverse environments. Nat Commun 1:147
Weinbauer RG, Rassoulzadegan F (2004) Are viruses driving microbial diversification and diversity? Environ Microbiol 6:1–11
Witzany G (1995) From the „logic of the molecular syntax“to molecular pragmatism. Explanatory deficits in Manfred Eigen’ s concept of language and communication. Evol Cog 1:148–168
Witzany G (2006) Serial endosymbiotic theory (set): the biosemiotic update. Acta Biotheor 54:103–117
Witzany G (2007) From biosphere to semiosphere to social lifeworlds. Biology as an understanding social science. In: Barbieri M (ed) Biosemiotic research trends. Nova Science Publishers, New York, pp 185–213
Witzany G (2014) Pragmatic turn in biology: from biological molecules to genetic content operators. World J Biol Chem 5:279–285
Witzany G (2015a) Life is physics and chemistry and communication. Ann N Y Acad Sci 1341:1–9
Witzany G (2015) (ed) The DNA habitats and their RNA inhabitants, vol 1341. New York Academy of Sciences, New York
Woese C (2004) A new biology for a new century. Microbiol Mol Biol Rev 68:173–186
Wood TK (2009) Insights on Escherichia coli biofilm formation and inhibition from whole-transcriptome profiling. Environ Microbiol 11:1–15
Yang Z, Kim J, Zhang C, Zhang M, Nietfeldt J, Southward CM et al (2009) Genomic instability in regions adjacent to highly conserved pch prophage in E. coli O157:H7 generates diversity in expression patterns of the LEE pathogenicity island. J Bacteriol 191:3553–3568
Yarmolinsky MB (2004) Bacteriophage P1 in retrospect and in prospect. J Bacteriol 186:7025–7028
Yarus M (2011) The meaning of a minuscule ribozyme. Philos Trans R Soc Lond B 366:2902–2909
Youle M, Haynes M, Rohwer F (2012) Scratching the surface of biology’s dark matter. In: Witzany G (ed) Viruses: essential agents of life. Springer, Dordrecht, pp 61–81
Zegans ME, Wagner JC, Cady KC, Murphy DM, Hammond JH, O’Toole GA (2009) Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J Bacteriol 191:210–219
Zemojtel T, Kielbasa SM, Arndt PF, ChungHR Vingron M (2009) Methylation and deamination of CpGs generate p53-binding sites on a genomic scale. Trends Genet 25:63–66
Zhang Z, Liu Y, Wang S, Yang D, Cheng Y, Hu J et al (2012) Temperate membrane-containing halophilic archaeal virus SNJ1 has a circular dsDNA genome identical to that of plasmid pHH205. Virology 434:233–241
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Villarreal, L.P., Witzany, G. When Competing Viruses Unify: Evolution, Conservation, and Plasticity of Genetic Identities. J Mol Evol 80, 305–318 (2015). https://doi.org/10.1007/s00239-015-9683-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-015-9683-y