Abstract
The genetic code has a high level of error robustness. Using values of hydrophobicity scales as a proxy for amino acid character, and the mean square measure as a function quantifying error robustness, a value can be obtained for a genetic code which reflects the error robustness of that code. By comparing this value with a distribution of values belonging to codes generated by random permutations of amino acid assignments, the level of error robustness of a genetic code can be quantified. We present a calculation in which the standard genetic code is shown to be optimal. We obtain this result by (1) using recently updated values of polar requirement as input; (2) fixing seven assignments (Ile, Trp, His, Phe, Tyr, Arg, and Leu) based on aptamer considerations; and (3) using known biosynthetic relations of the 20 amino acids. This last point is reflected in an approach of subdivision (restricting the random reallocation of assignments to amino acid subgroups, the set of 20 being divided in four such subgroups). The three approaches to explain robustness of the code (specific selection for robustness, amino acid–RNA interactions leading to assignments, or a slow growth process of assignment patterns) are reexamined in light of our findings. We offer a comprehensive hypothesis, stressing the importance of biosynthetic relations, with the code evolving from an early stage with just glycine and alanine, via intermediate stages, towards 64 codons carrying todays meaning.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
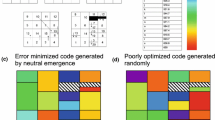
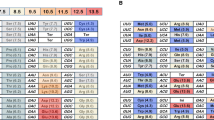
The genetic code is a basic feature of molecular biology. It sets the rules according to which nucleic-acid sequences are translated into amino acid sequences. The genetic code probably evolved by a process of gradual evolution from a proto-biological stage, via many intermediary stages, to its present form (see e.g. Crick 1968; Lehman and Jukes 1988; Vetsigian et al. 2006). During this process, error robustness was built into the code (see e.g. Ardell 1998; Caporaso et al. 2005; Crick 1968; Di Giulio 2008; Freeland et al. 2003; Higgs 2009; Ikehara et al. 2002; Massey 2008; Vetsigian et al. 2006; Wolf and Koonin 2007; Wong 2005). Two different kinds of error robustness can be observed (Vetsigian et al. 2006) by even the most superficial inspection of the standard genetic code (SGC). On one hand, codons assigned to the same amino acid are almost always similar, see Table 1. As an example, all codons ending with a pyrimidine (U or C) in a codon box (the four codons sharing first and second nucleotides) are without exception assigned to the same amino acid (e.g. UAU and UAC both code for Tyr). On the other hand, similar codons are mostly assigned to similar amino acids, e.g. codons with U in the second position are all assigned to hydrophobic amino acids (Woese 1965; Woese et al. 1966a, b). This is illustrated in Table 1, when looking at the values of polar requirement: overall, low values of polar requirement correspond to hydrophobic amino acids.
Three main approaches exist to explain the emergence of this robustness of the code: specific selection for robustness (see e.g. Freeland and Hurst 1998a; Haig and Hurst 1991; Vetsigian et al. 2006), amino acid-RNA interactions leading to assignments (see e.g. Woese 1965; Yarus et al. 2009), and a slow growth process of assignment patterns reflecting the history of amino acid repertoire growth (see e.g. Crick 1968; Di Giulio 2008; Massey 2006; Wong 1975). The concept that all three competing hypotheses are important has also been brought forward (Knight et al. 1999). In the present study we make adjustments to earlier mathematical work in this field (see e.g. Buhrman et al. 2011; Freeland and Hurst 1998a; Haig and Hurst 1991), which integrate the three concepts into a single mathematical model. We will now, one by one, introduce these three adjustments.
Polar Requirement
The polar requirement (Woese et al. 1966a) is not just a measure related to hydrophobicity. Several different measures of hydrophobicity exist, each focusing on different aspects of it. Polar requirement specifically focuses on the nature of the interaction between amino acids and nucleic acids. Stacking interactions between e.g. the planar guanidinium group of arginine and the planar purine ring systems and pyrimidine ring systems of RNA is an example of that. Woese chose to chemically model the nucleotide rings by using pyridine as the solvent system in the measurements leading to the polar requirement scale (Woese 1965, 1967, 1973; Woese et al. 1966a, b). This interaction between amino acids and nucleic acids has been stressed as an especially important aspect of early protein chemistry because one possibility for the very first function of coded peptides was suggested (Noller 2004) to be the enlargement of the number of conformations accessible for RNA (realized by the binding of small, oligopeptide cofactors). Thus, polar requirement could have been among the most important aspects of an amino acid during early stages of genetic code evolution.
The remarkable character of polar requirement as a measure of amino acids in connection to the genetic code was found again and again throughout the years. Firstly, Woese found that distinct amino acids coded by codons differing only in the third position are very close in polar requirement, despite differences in general character (Woese et al. 1966b). The pair cysteine and tryptophan nicely exemplifies this. Secondly, Haig and Hurst (1991) discovered that polar requirement showed the SGC to be special to a much larger degree than another scale of hydrophobicity [the hydropathy scale of Kyte and Doolittle (1982)]. Thirdly, when Mathew and Luthey-Schulten updated the values of polar requirement (Mathew and Luthey-Schulten 2008) by in silico methods (the most important change was believed to be due to a cellulose–tyrosine interaction artefact in the original experiments), the SGC showed a further factor 10 increase (Butler et al. 2009) in error robustness calculations. In all these developments the expectation that polar requirement would behave in a special way, as interaction between nucleotides and amino acids is biochemically important, was more than borne out by the results. One of the adjustments we introduce in our work compared to our earlier calculations (Buhrman et al. 2011) is that in the present work we use the new, updated values of polar requirement (see Table 1).
Aptamers
Oligonucleic-acid molecules that bind to a specific target molecule (e.g. a specific amino acid) are called aptamers (Ellington and Szostak 1990). Over the last two decades, many results have been obtained regarding specific binding of amino acids by RNA aptamers, mainly by Yarus and co-workers (Illangasekare and Yarus 2002; Majerfeld and Yarus 1994; Yarus et al. 2009). For several amino acids, codons and anticodons were found in binding sites, in quantities higher than would be expected to occur by chance (Yarus et al. 2009). In Table 2, a list of occurrences of anticodons in binding sites of RNA sequences is given, together with the articles in which these sequences were reported. Please note that the definition of anticodons used in these articles is: triplets complementary to codons. These anticodons are therefore not necessarily identical to the triplets found in tRNA molecules which are normally meant with the word ‘anticodon’. As an example: the triplet AUG is considered as an His anticodon because it is complementary to the His codon CAU. In tRNAs, however, the anticodon recognizing CAU is GUG (see Grosjean et al. 2010; Johansson et al. 2008) for reviews on codon–anticodon interaction). We summarize published details on the aptamers for seven amino acids, and subsequently formulate a conclusion regarding the implications of the existence of these molecules for genetic-code error-robustness calculations. This conclusion is based on reasoning presented by the Yarus group concerning the existence of specific relationships between certain triplets and certain amino acids. These relationships could have led to evolutionary conserved assignments of these amino acids to these triplets, e.g. by a mechanism as presented in (Yarus et al. 2009).
For Ile, Trp, and His, three binding motifs were described, respectively named the ‘UAUU-motif’ (Lozupone et al. 2003), the ‘CYA-motif’ (Majerfeld et al. 2010, Majerfeld and Yarus 2005), and the ‘histidine-motif’ (Majerfeld et al. 2005). As can be seen from the names, the anticodons UAU for Ile, and CCA for Trp, are characteristic for the motifs (‘CYA’ stands for ‘CUA or CCA’). In the case of His, both GUG and AUG (the anticodons for the two His codons CAC and CAU) are found in quantities higher than would be expected by chance (Majerfeld et al. 2005).
Although binding sites for Phe and Tyr have so far not been studied as extensively as those for Ile, Trp, and His, the analysis of Yarus et al. (2009) shows that the anticodons (GAA and AAA for Phe, and GUA and AUA for Tyr) are present in the binding sites more often than would be expected on a random basis.
Both the CCU anticodon (Janas et al. 2010) and the UCG anticodon (Yarus et al. 2009) are present in Arg binding sites more often than would be expected on a random basis. Thus, a physico-chemical background was observed, compatible with: (1) Arg having more than four codons, and (2) all six Arg codons sharing the same middle nucleotide.
A similar observation can be made for Leu, the other amino acid which is encoded by six codons all having the same middle nucleotide. For this amino acid, however, only a single RNA sequence was found binding the amino acid with specificity (Yarus et al. 2009). Inspection of this sequence shows anticodons UAG, GAG, and CAA to be present in its binding parts.
Taking the combined results of Yarus and co-workers into consideration, we propose to fix assignments of Ile, Trp, His, Phe, Tyr, Arg, and Leu for calculations using random variants of the SGC.
Gradual Growth
In ‘Methods’ section we present our approach in detail. We use Haig and Hurst’s ‘mean square’ measure, [as first proposed in Haig and Hurst (1991)] to quantify the error robustness of a given code. With this measure, a relatively error-robust code gets a low value when compared to the average value of a large set of codes produced by random allocation of amino acid assignments [see Buhrman et al. (2011) for a more in-depth treatment of the approach]. The space of codes allowed to exist by the allocation procedure can be large [in the original work of Haig and Hurst (1991) the space has a size of exactly 20! codes, which is \(\approx 2.433 \times 10^{18}\) codes]. We call a code optimal if it reaches the minimum in error robustness calculations among all possible codes in a particular setting.
In 1975, Wong proposed the coevolution theory of the genetic code (Wong 1975). According to this proposal, SGC codons assigned to an amino acid biosynthetically derived from another amino acid, were originally assigned to that ‘precursor’ amino acid. As an example: Pro is biosynthetically derived from Glu. According to coevolution theory, the four Pro codons (CCN) would have originally encoded Glu. Without embracing all details of the original coevolution theory, or modern refinements of the theory (Di Giulio 2008; Wong 2007), something remarkable can be noted as a result of this way of looking at the SGC. Shikimate-derived amino acids (Phe, Tyr, and Trp) all have U in the first position of the codon (Phe: UUY; Tyr: UAY; and Trp: UGG). Glu-derived amino acids (Pro, Gln, and Arg) almost always have C in the first position of the codon (Pro: CCN; Gln: CAR, which stands for ‘CAA or CAG’; and Arg: AGR and CGN, where N stands for all four nucleotides). Asp-derived amino acids (Ile, Met, Thr, Asn, and Lys) all have A in the first position of the codon (Ile: AUY and AUA; Met: AUG; Thr: ACN; Asn: AAY; and Lys: AAR). Codons with G in the first position all code for amino acids produced in Urey–Miller experiments Footnote 1 (Val: GUN; Ala: GCN; Asp: GAY; Glu: GAR; and Gly: GGN). This ‘layered structure’ of the SGC was first pointed out explicitly by Taylor and Coates (1989). It may indeed suggest a sequential development of the repertoire of amino acids specified in the developing code, and a possibly sequential introduction of use of G, A, C, and U as first nucleotide in codons. The ‘layered structure’ of the SGC is a regularity different from the well-known error-robust distribution of polar requirement (Haig and Hurst 1991), which is pronounced in the first and the third, but not in the second position of the codon (please note: having, as a group, all the same nucleotide in the first position, gives error robustness for the group character to changes in the second and third position). As is shown in ‘Appendix: Molecular Structure Matrix’, it is possible to prove the presence of the ‘layered structure’ quantitatively, when the appropriate set of values is developed and used as input.
Freeland and Hurst (1998b) followed the concept of Taylor and Coates, and formally divided the 20 amino acids in four groups of five amino acids each: Gly, Ala, Asp, Glu, and Val in a first group which could be called ‘the prebiotic group’; a second group of amino acids with codons starting with A (Ile, Met, Thr, Asn, and Lys); a third group with codons mainly starting with C (Leu, Pro, His, Gln, and Arg); and, finally, a group with codons mainly starting with U (Phe, Ser, Tyr, Cys, and Trp). Division of the set of twenty in these four subsets was subsequently incorporated in the calculations on code error robustness (Freeland and Hurst 1998b). This approach reduced the size of the space from which codes could be sampled randomly in a drastic way: from a size of about \(2 \times 10^{18}\) codes (see above) to a size of (5!)4 codes (which is exactly \(2.0736 \times 10^{8}\) codes). This space was called the ‘historically reasonable’ set of possible codes (Freeland and Hurst 1998). By sampling from the historically reasonable set of possible codes, we incorporate in the current study the notion of a chronologically-determined, layered structure of the SGC.
Integration of assumptions
We have found that if: (1) the updated values for polar requirement are used as amino acid attributes; (2) the assignments of seven amino acids to codons are fixed following the rationale given above; and (3 the subdivision leading to the historically reasonable set of possible codes is used to define the space of code variations [which is also reduced in size by (2)], then the SGC is optimal. It is important to note that the constraints applied drastically reduce the size of the space: with applying both (2) and (3), the ‘realistic space’ has a size of 11,520 codes.
Methods
We use the mean-square method developed by Alff-Steinberger (1969), Wong (1980), Di Giulio (1989), and Haig and Hurst (1991). For the mathematical formulation, we follow the approach of Buhrman et al. (2011) and consider the undirected graph G = (V, E) that has the 61 codons Footnote 2 as its vertices and an edge between any two codons if they differ in only one position, yielding 263 edges. A code F maps each codon c to exactly one amino acid F(c). We denote by rF(c) the polar requirement of the amino acid that codon c encodes in the code F and by r the full vector of 20 values. The mean square error function of code F is then given by
where the αc,c' are the weights of the different mutations that can occur (corresponding to edges of the graph) and \(N=\sum_{\{c,c'\} \in E} \alpha_{c,c'}\) is the total weight. Following Haig and Hurst (1991), we use a subscript 0 to indicate the overall measure. If we set all 263 weights αc,c' to 1, we get the original function described by Haig and Hurst (1991), which we simply denote by MS0(F). We also consider the following set of weights introduced by Freeland and Hurst (1998a), which differentiates between transition errors (i.e. U to C, C to U, A to G, G to A) and transversion errors and the position where they occur in the codon:
-
αc,c' = 0.5 if (c, c′) is a transversion in the first position or a transition in the second position,
-
αc,c' = 0.1 if (c, c′) is a transversion in the second position,
-
αc,c' = 1 otherwise.
Using weights for different codon positions implies the existence of a tRNA with a triplet anticodon during the process of code evolution. As we consider a process of gradual expansion of the repertoire of amino acids during the evolution of the SGC (see e.g. Ardell 1998, Crick 1968, Lehman and Jukes 1988) as the most likely mechanism—with duplication of tRNA genes, and subsequent divergence [cf. (Ohno 1970)] of their sequences and functions—we think this assumption is acceptable. This assumption does not necessarily imply the existence of protein aminoacyl-tRNA synthetases during all or part of the process of code evolution, as there could originally have been ribozymes which fulfilled their function. The value of error-robustness of a code F using the set of weights introduced above will be denoted by MS FH0 (F).
In principle, there are at least three ways in which one can improve the model of Haig and Hurst (1991) to reflect biological reality more accurately. The first possibility is to change how the level of error robustness is measured, e.g. by introducing weighting factors as described above. Variations of the weighting factors used in the calculation show an even higher error robustness of the SGC, as noticed by e.g. Butler et al. (2009), Freeland and Hurst (1998a), Gilis et al. (2001). The rationale behind changing weighting factors is improved reflection of natural selection pressures. It is, however, difficult to decide which weighting factors adequately reflect the natural selection pressures operating during the early evolution of the genetic code [see comment 4 of Ardell in Novozhilov et al. (2007) and the exchange of thoughts with respect to ‘column 4’ in Higgs (2009)].
The second way to improve the model is to change the set of values representing amino acid properties used as input in the error-robustness calculation. For instance, one can use the values of hydropathy from Kyte and Doolittle (1982), or the matrix of Gilis et al. (2001) instead of the polar requirement scale. In our paper, we use the values of the 2008 update of polar requirement by in silico methods (Mathew and Luthey-Schulten 2008) given in Table 1. Work concerning the issue what an ‘ideal’ set of 20 values would look like, and work considering different known sets of amino acid properties is presented in ‘Appendices: Inverse Parametric Optimization and Scan of Other Amino Acid Properties’.
The third way to improve the model is to change the size of the space from which random codes are sampled (Buhrman et al. 2011). The incentive to enlarge that space [as was done in Buhrman et al. (2011)] is the wish to work from a space that encompasses all possible codes, or at least, all known codes. As indicated in Buhrman et al. (2011), larger spaces are increasingly difficult to work with. The frequency distributions obtained by sampling from the larger spaces in Buhrman et al. (2011) highly coincide with the frequency distribution obtained from the original space [as presented in Haig and Hurst (1991)]. From this viewpoint, working in the original space is acceptable as a simplification. In the current study, we shrink the size of the space, based on considerations of fixed assignments of certain codons, and combining this with the constraint of the historically reasonable set of possible codes of Freeland and Hurst (1998b), as outlined in ‘Introduction’ section.
MATLAB-programs were used for the error-robustness calculations and visualizations. All software can be found as supplemental information, or downloaded from https://github.com/cschaffner/gcode.
Results
Among all genetic codes (in this particular setting of the problem), the SGC is optimal in terms of error-robustness if:
-
1.
We use the updated values of polar requirement (Mathew and Luthey-Schulten 2008).
-
2.
We use fixation for Phe, Tyr, Trp, His, Leu, Ile, and Arg, based on aptamer experiments (Janas et al. 2010; Yarus et al. 2009).
-
3.
We use the historically reasonable set of possible codes (Freeland and Hurst 1998b).
Figure 1 shows a histogram of MS FH0 (F)-values resulting from this procedure. When, the original error function MS0(F) from Haig and Hurst (1991) is used, the result is essentially the same: the SGC is the optimal code. We wondered if by fixation of just one or two more assignments, the SGC would be optimal in the space resulting from the combination of these fixations with the random permutations of amino acid assignments according to the method used by Haig and Hurst (1991), without the constraint of the historically reasonable set of possible codes (Freeland and Hurst 1998b). This was not the case (as is reported in ‘Appendix: Minimal Number of Fixed Assignments’).
Discussion
What is the biological relevance of the mathematical result presented, if any? Can we indeed conclude that natural selection steered the translation system toward better and better variants of the assignments (in terms of error-robustness) within realistic boundaries? Stated differently, when making a model, should one respect that seven assignments are fixed, and that the system evolved gradually (as reflected by using the historically reasonable set of possible codes), until the optimal code (within these boundaries) was reached? Or is it rash to arrive at such a conclusion, and could one imagine positive selection for error-robustness to be an illusion?
The space of codes resulting from the constraints imposed on the calculations is a space of very limited size: only 11,520 codes (\(2!\times 2!\times 4!\times 5!\)). The fact that the SGC is optimal in this space is impressive, but of a different order of magnitude than the near-optimalities in significantly larger spaces presented in earlier studies (e.g. Buhrman et al. 2011; Butler et al. 2009; Freeland and Hurst 1998a; Freeland et al. 2000; Gilis et al. 2001). The impact of the different fixed assignments varies: for the MS0-values, it would theoretically suffice to fix the three assignments of Phe, Trp, and Arg (or any set containing them) in order to find the SGC to be optimal in the resulting space. Footnote 3 In this way, the SGC can be thought of as the global optimum in a space of \(3! \times 4!\times 5!\times 5!=2073600\) codes. We further refrain from presenting it thus, because in doing so we would abandon the physico-chemical facts which were the starting point for our calculations with fixed assignments.
It is also possible to increase the number of fixed assignments (and in this way decrease the size of the space of random code variants) even further. A recent article (Johnson and Wang 2010) suggests that more than the seven assignments (listed in Table 2) are fixed.
The logical extreme of fixing assignments is that all assignments of the SGC are fixed, as argued recently by Erives (2011). In his theory, a kind of RNA cage (pacRNA: proto-anti-codon RNA) is presented, in which different amino acids are bound by different kinds of ‘walls’, which are exposing anticodons to the different amino acids. Although this model combines elegant explanations for several aspects of present-day tRNA functioning, it is very hard to get an objective measure for the specificity of amino acid-anticodon interactions in this model. In particular, the different possibilities allowed by ‘breathing’ of the cage cast doubt on interaction specificity. Some objections can also be raised regarding the tRNA activation mechanism. Yarus and co-workers recently reported a very small ribozyme (only five nucleotides in length), which was experimentally shown to aminoacylate certain small RNAs using aminoacyl-NMPs as activated precursors (Turk et al. 2010; Yarus 2011). Such an early activation mechanism, using NTPs as source of energy, is different from the one in Erives’ model, where the \(5^\prime\) end of the pacRNA is performing this role.
Taking all considerations sketched above into account, it is possible to draw a tentative picture of genetic code evolution which is compatible with the indications concerning which aspects of code evolution are important. Code evolution probably followed classical mechanisms of gene duplication and subsequent diversification (here of ‘tRNA’ genes and genes involved in aminoacylation). Evolution would be mainly by stop-to-sense reassignments (Lehman and Jukes 1988), with occasional reassignments in only slightly different new or developing uses of codons [cf. Ardell 1998; Vetsigian et al. 2006], not yet massively present in protein-coding sequences [cf. the frozen accident concept (Crick 1968)]. In a proto-biological stage, RNA would be absent while very small peptides could have been synthesized, e.g. by the salt-induced peptide formation (SIPF) reaction (Rode et al. 1999; Schwendinger and Rode 1989). Under prebiotic conditions especially Ala and Gly would be expected to be present in relatively large amounts (see e.g. Higgs and Pudritz 2009; Philip and Freeland 2011). Asp-containing peptides could possibly play a role in the origin of RNA, as they could position Mg2+ ions in the correct orientation to help polymerize nucleotides, and, concomitantly, keep these ions from stimulating RNA hydrolysis (Szostak 2012). Asp content of peptides could be enriched in the presence of carboxyl-group binding montmorillonite surfaces (Rode et al. 1999).
In the first stages of coded peptide synthesis, GCC and GGC probably were the only codons in mRNAs (Eigen and Schuster 1978), and coded peptides would consist of Ala and Gly. The remaining codons effectively would be stop codons (Lehman and Jukes 1988), although functioning without release factors: water would break bonds between tRNA and peptide whenever codons stayed unoccupied for too long. The ‘single-step biosynthetic distance’ between Ala and pyruvate suggests a carbon storage role for these peptides; Gly allowing folding of such molecules. A mRNA/tRNA system functioning without a ribosome has been proposed by several authors (Crick et al. 1961; Lehman and Jukes 1988; Woese 1973). The first rRNA could then have been functioning in improved termination (see above). At this stage the proposal that coded peptides enlarge the possible range of RNA conformations should be taken into account (Noller 2004).
In the next stage of coded peptide synthesis, Asp and Val could have been added to the repertoire (see e.g. Ardell 1998; Eigen and Schuster 1978; van der Gulik et al. 2009; Higgs 2009; Ikehara 2002). This would have been a crucial step: enabling directed production of the important Asp-containing peptides (van der Gulik et al. 2009; Szostak 2012) as well as formation of something resembling protein structure, characterized by hydrophobic cores (Val) and hydrophilic exteriors (Asp). The emerging polypeptides could have functioned in carbon storage, as mentioned above. Having started with trinucleotide codons, this aspect was retained, not because four nucleotide codons are in principle impossible, but this system allowed a further robust development (cf. Vetsigian et al. 2006). Depletion of prebiotic pools of either Ala, Gly, Asp, or Val (e.g. by excessive storage in coded peptides) could have led to the biosynthetic routes involving Gly, Ser, Val, Asp, Ala, and pyruvate. In this way the lack of an amino acid could in principle be resolved by use of the other three (cf. the hypothesized carbon storage function of coded peptides).
In a further stage, Ser, and Asp-derived amino acids like Asn and Thr would be added to the repertoire. Asn would be the first amino acid with an entirely biosynthetic origin (it is relatively unstable, and does not accumulate prebiotically). The production of Asn is known to be originally linked to enzymatic conversion of Asp to Asn on a tRNA (see e.g. Wong 2007). When instead of two molecules of pyruvate, one molecule of pyruvate and one molecule of alpha-keto-butyrate are fed into the Val biosynthesis pathway, Ile is produced instead. Therefore, when both Thr and Val biosynthesis are present, the evolution of just one enzyme (making alpha-keto-butyrate from Thr) suffices for the emergence of Ile. Aptamers can handle this amino acid, and these two factors (easy development from existing biochemistry and easy manipulation by RNA) could be responsible for the ‘choice’ of Ile (cf. Philip and Freeland 2011).
Larger amino acids like His and Gln would have appeared in a later stage of code development than Asp-derived amino acids like Asn and Thr. The reactions catalyzed by the few enzymes in the Leu biosynthesis, which are not enzymes involved in Val biosynthesis (apart from leucine aminotransferase) are reminiscent of the first three reactions of the citric acid cycle (Voet and Voet 1995). Jensen (1976) hypothesized that originally enzymes would have had much broader substrate specificity. With the citric acid cycle being ‘old’, as well as important for bio-energetic reasons, and Val biosynthesis being present, the system could have produced an excess of Leu. Again, aptamers would be able to ‘handle’ Leu. Existing biochemistry and aptamer potential would thus answer the question why Ile and Leu are part of the Set of Twenty, and e.g. norleucine and alpha-amino-butyric acid are not (cf. Philip and Freeland 2011). Linked to the citric acid cycle and important in nitrogen management are Glu and Gln. A further expansion of the repertoire with a Glu-derived amino acid is the expansion with Arg. Two of the enzymes of the urea (nitrogen management) cycle are related to pyrimidine synthesis enzymes, two others to purine synthesis enzymes (Berg et al. 2007). The last enzyme in the cycle is arginase. This suggests an ancient accumulation of Arg as a side effect of RNA synthesis, upon Glu becoming a major cell component. Arginase could function in bringing the Arg concentration down to acceptable levels. Aptamers could also have evolved to manipulate Arg levels, allowing Arg to become part of the Set of Twenty. Again Jensen’s concept of primordial broad substrate specificity (Jensen 1976) is essential to get a possible answer to the ‘Why these 20?’ question: Arg could be part of the set, rather than ornithine and citrulline, because Arg accumulates, and Arg can be manipulated by aptamers.
In an advanced stage of code development aromatic amino acids would be added to the repertoire, and release factors would evolve. Van der Gulik and Hoff (2011) have argued that codons UUA, AUA, UAA, CAA, AAA, GAA, UGA, and AGA could not function unambiguously until the anticodon modification machinery was developed, which is seen by them as the last development leading to the full genetic code. Because archaea and bacteria have different solutions for the ‘AUA problem’ [agmatidinylation vs. lysidinylation (van der Gulik and Hoff 2011)], unambiguous sense assignment of AUA must have been late indeed.
The SGC has probably evolved in a genetic environment characterized by rampant horizontal gene-flow (Vetsigian et al. 2006). The interaction between genetic systems with slightly different, still-evolving codes, is thought to have caused both universality and optimality of the SGC (Vetsigian et al. 2006). Universality, because the genetic code functioned as an innovation sharing protocol (Vetsigian et al. 2006). Optimality, because competition allowed selection for the ability to translate the genetic information accurately (Vetsigian et al. 2006). The work presented in our paper illuminates constraints within which this process of genetic code development took place. Both the step-by-step increasing complexity of biochemistry, and the stereochemical relationship between at least some amino acids and triplets, are factors which have to be taken into account.
In summary, although there are at least two different lines of research suggesting a greater number of fixed assignments than the seven given in Table 2 [based on the work of Yarus and co-workers (Janas et al. 2010; Yarus et al. 2009)], for now it is not clear that more [or even all (Erives 2011)] assignments are fixed. Thus, the observed error-robustness still needs explanation. It is possible that the optimality of the SGC we found results from positive selection for error-robustness, though starting within a more restricted set of possibilities than previously thought.
Notes
For a recent update on prebiotic synthesis see (Parker et al. 2011) and references therein.
In the original calculation, Haig and Hurst ignored the three ‘stop codons’ encoding chain termination.
When using the Freeland and Hurst weights (and hence the MS FH0 -values), it is possible to fix another set of three amino acids Phe, His, Trp in order to make the SGC optimal.
References
Aboderin AA (1971) An empirical hydrophobicity scale for α-amino-acids and some of its applications. Int J Biochem 2(11):537–544
Alff-Steinberger C (1969) The genetic code and error transmission. Proc Natl Acad Sci USA 64(2):584–591
Ardell DH (1998) On error minimization in a sequential origin of the standard genetic code. J Mol Evol 47(1):1–13
Berg JM, Tymoszko JL, Stryer L (2007) Biochemistry, 6th edn. W.H. Freeman and Company, New York, p 664
Biou V, Gibrat JF, Levin JM, Robson B, Garnier J (1988) Secondary structure prediction: combination of three different methods. Protein Eng 2(3):185–191
Buhrman H, van der Gulik PTS, Kelk SM, Koolen WM, Stougie L (2011) Some mathematical refinements concerning error minimization in the genetic code. IEEE/ACM Trans Comput Biol Bioinf 8(5):1358–1372
Burkard R, Derigs U (1980) Assignment and matching problems: solution methods with FORTRAN-programs. Lecture notes in economics and mathematical systems. Springer-Verlag, Berlin. http://books.google.nl/books?id=0jwZAQAAIAAJ
Burkard RE, Rendl F (1984) A thermodynamically motivated simulation procedure for combinatorial optimization problems. Eur J Oper Res 17(2):169–174
Butler T, Goldenfeld N, Mathew D, Luthey-Schulten Z (2009) Extreme genetic code optimality from a molecular dynamics calculation of amino acid polar requirement. Phys Rev E 79(6):060,901(R)
Caporaso JG, Yarus M, Knight R (2005) Error minimization and coding triplet/binding site associations are independent features of the canonical genetic code. J Mol Evol 61(5):597–607
Cornette JL, Cease KB, Margalit H, Spouge JL, Berzofsky JA, DeLisi C (1987) Hydrophobicity scales and computational techniques for detecting amphipathic structures in proteins. J Mol Biol 195(3):659–685
Crick FHC (1968) The origin of the genetic code. J Mol Biol 38(3):367–379
Crick FHC, Barnett L, Brenner S, Watts-Tobin RJ (1961) General nature of the genetic code for proteins. Nature 192(4809):1227–1232
Di Giulio M (1989) The extension reached by the minimization of the polarity distances during the evolution of the genetic code. J Mol Evol 29(4):288–293
Di Giulio M (2008) An extension of the coevolution theory of the origin of the genetic code. Biol Direct 3:37
Eigen M, Schuster P (1978) A principle of natural self organization. Part C: the realistic hypercycle. Naturwissenschaften 65(7):341–369
Eisenberg D, McLachlan AD (1986) Solvation energy in protein folding and binding. Nature 319(6050):199–203
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346:818–822
Eppstein D (2003) Setting parameters by example. SIAM J Comput 32(3):643–653
Erives A (2011) A model of proto-anti-codon RNA enzymes requiring L-amino acid homochirality. J Mol Evol 73:10–22. doi:10.1007/s00239-011-9453-4
Freeland SJ, Hurst LD (1998a) The genetic code is one in a million. J Mol Evol 47(3):238–248
Freeland SJ, Hurst LD (1998b) Load minimization of the genetic code: history does not explain the pattern. Proc R Soc B Biol Sci 265(1410):2111–2119
Freeland SJ, Knight RD, Landweber LF, Hurst LD (2000) Early fixation of an optimal genetic code. Mol Biol Evol 17(4):511–518
Freeland SJ, Wu T, Keulmann N (2003) The case for an error minimizing standard genetic code. Orig Life Evol Biosp 33(4-5):457–477
Gilis D, Massar S, Cerf NJ, Rooman M (2001) Optimality of the genetic code with respect to protein stability and amino-acid frequencies. Genome Biol 2(11):R49
Grantham R (1974) Amino acid difference formula to help explain protein evolution. Science 185(4154):862–864
Grosjean H, de Crecy-Lagard V, Marck C (2010) Deciphering synonymous codons in the three domains of life: co-evolution with specific tRNA modification enzymes. FEBS Lett 584(2):252–264
Haig D, Hurst LD (1991) A quantitative measure of error minimization in the genetic code. J Mol Evol 33(5):412–417
Higgs PG (2009) A four- column theory for the origin of the genetic code: tracing the evolutionary pathways that gave rise to an optimized code. Biol Direct 4:16
Higgs PG, Pudritz RE (2009) A thermodynamic basis for prebiotic amino acid synthesis and the nature of the first genetic code. Astrobiology 9(5):483–490
Ikehara K (2002) Origins of gene, genetic code, protein and life: comprehensive view of life systems from a GNC-SNS primitive genetic code hypothesis. J Biosci 27(2):165–186
Ikehara K, Omori Y, Arai R, Hirose A (2002) A novel theory on the origin of the genetic code: a GNC-SNS hypothesis. J Mol Evol 54(4):530–538
Illangasekare M, Yarus M (2002) Phenylalanine-binding RNAs and genetic code evolution. J Mol Evol 54(3):298–311
Janas T, Widmann JJ, Knight R, Yarus M (2010) Simple, recurring RNA binding sites for l-arginine. RNA 16(4):805–816
Jensen RA (1976) Enzyme recruitment in evolution of new function. Annu Rev Microbiol 30:409–425
Johansson MJO, Esberg A, Huang B, Bjork GR, Bystrom AS (2008) Eukaryotic wobble uridine modifications promote a functionally redundant decoding system. Mol Cell Biol 28(10):3301–3312
Johnson DBF, Wang L (2010) Imprints of the genetic code in the ribosome. Proc Natl Acad Sci USA 107(18):8298–8303
Kawashima S, Ogata H, Kanehisa M (1999) AAindex: amino acid index database. Nucleic Acids Res 27(1):368–369
Knight RD, Freeland SJ, Landweber LF (1999) Selection, history and chemistry: the three faces of the genetic code. Trends Biochem Sci 24(6):241–247
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105–132
Lehman N, Jukes TH (1988) Genetic code development by stop codon takeover. J Theor Biol 135(2):203–214
Li Y, Pardalos P, Resende M (1994) A greedy randomized adaptive search procedure for the quadratic assignment problem. Quadratic Assign Relat Probl 16:237–261
Lozupone C, Changayil S, Majerfeld I, Yarus M (2003) Selection of the simplest RNA that binds isoleucine. RNA 9(11):1315–1322
Majerfeld I, Chocholousova J, Malaiya V, Widmann J, McDonald D, Reeder J, Iyer M, Illangasekare M, Yarus M, Knight R (2010) Nucleotides that are essential but not conserved; a sufficient l-tryptophan site in RNA. RNA 16(10):1915–1924
Majerfeld I, Puthenvedu D, Yarus M (2005) RNA affinity for molecular l-histidine; genetic code origins. J Mol Evol 61:226–235
Majerfeld I, Yarus M (1994) An RNA pocket for an aliphatic hydrophobe. Nat Struct Biol 1(5):287–292
Majerfeld I, Yarus M (2005) A diminutive and specific RNA binding site for l-tryptophan. Nucleic Acids Res 33(17):5482–5493. doi:10.1093/nar/gki861
Massey SE (2006) A sequential "2-1-3" model of genetic code evolution that explains codon constraints. J Mol Evol 62(6):809–810
Massey SE (2008) A neutral origin for error minimization in the genetic code. J Mol Evol 67(5):510–516
Mathew DC, Luthey-Schulten Z (2008) On the physical basis of the amino acid polar requirement. J Mol Evol 66(5):519–528
MATLAB: version 7.12.0 (R2011a) The MathWorks Inc., Natick, Massachusetts (2011)
Meirovitch H, Rackovsky S, Scheraga HA (1980) Empirical studies of hydrophobicity. 1. Effect of protein size on the hydrophobic behavior of amino acids. Macromolecules 13(6):1398–1405
Miyazawa S, Jernigan RL (1985) Estimation of effective interresidue contact energies from protein crystal structures: quasi-chemical approximation. Macromolecules 18(3):534–552
Miyazawa S, Jernigan RL (1999) Self-consistent estimation of inter-residue protein contact energies based on an equilibrium mixture approximation of residues. Proteins 34(1):49–68
Noller HF (2004) The driving force for molecular evolution of translation. RNA 10(12):1833–1837
Novozhilov AS, Wolf YI, Koonin EV (2007) Evolution of the genetic code: partial optimization of a random code for robustness to translation error in a rugged fitness landscape. Biol Direct 2:24
Ohno S (1970) Evolution by gene duplication. Springer, Berlin
Oobatake M, Ooi T (1977) An analysis of non-bonded energy of proteins. J Theor Biol 67(3):567–584
Parker ET, Cleaves HJ, Dworkin JP, Glavin DP, Callahan M, Aubrey A, Lazcano A, Bada JL (2011) Primordial synthesis of amines and amino acids in a 1958 Miller H2S-rich spark discharge experiment. Proc Natl Acad Sci USA 108(14):5526–5531
Philip GK, Freeland SJ (2011) Did evolution select a nonrandom "alphabet" of amino acids? Astrobiology 11(3):235–240
Ponnuswamy PK, Prabhakaran M, Manavalan P (1980) Hydrophobic packing and spatial arrangement of amino acid residues in globular proteins. Biochim Biophys Acta 623(2):301–316
Rahman S, Bashton M, Holliday G, Schrader R, Thornton J (2009) Small molecule subgraph detector (SMSD) toolkit. J Cheminform 1(1):12. doi:10.1186/1758-2946-1-12 http://www.jcheminf.com/content/1/1/12
Rode BM, Son HL, Suwannachot Y, Bujdak J (1999) The combination of salt induced peptide formation reaction and clay catalysis: a way to higher peptides under primitive earth conditions. Orig Life Evol Biosph 29(3):273–286
Schwendinger MG, Rode BM (1989) Possible role of copper and sodium in prebiotic evolution of peptides. Anal Sci 5:411–414
Sweet RM, Eisenberg D (1983) Correlation of sequence hydrophobicities measures similarity in three-dimensional protein structure. J Mol Biol 171(4):479–488
Szostak JW (2012) The eightfold path to non-enzymatic rna replication. J Syst Chem 3:2
Taylor FJR, Coates D (1989) The code within the codons. BioSystems 22(3):177–187
Turk RM, Chumachenko NV, Yarus M (2010) Multiple translational products from a five-nucleotide ribozyme. Proc Natl Acad Sci USA 107(10):4585–4589
van der Gulik P, Massar S, Gilis D, Buhrman H, Rooman M (2009) The first peptides: the evolutionary transition between prebiotic amino acids and early proteins. J Theor Biol 261(4):531–539
van der Gulik PTS, Hoff WD (2011) Unassigned codons, nonsense suppression, and anticodon modifications in the evolution of the genetic code. J Mol Evol 73(3-4):59–69
Vetsigian K, Woese C, Goldenfeld N (2006) Collective evolution and the genetic code. Proc Natl Acad Sci USA 103(28):10,696–10,701
Voet D, Voet JG (1995) Biochemistry, 2nd edn, Wiley, New York, p 773
Woese CR (1965) Order in the genetic code. Proc Natl Acad Sci USA 54(1):71–75
Woese CR (1967) The genetic code. Harper and Row, New York
Woese CR (1973) Evolution of the genetic code. Naturwissenschaften 60(10):447–459
Woese CR, Dugre DH, Dugre SA, Kondo M, Saxinger WC (1966a) On the fundamental nature and evolution of the genetic code. Cold Spring Harb Symp Quant Biol 31:723–736
Woese CR, Dugre DH, Saxinger WC, Dugre SA (1966b) The molecular basis for the genetic code. Proc Natl Acad Sci USA 55(4):966–974
Wolf YI, Koonin EV (2007) On the origin of the translation system and the genetic code in the RNA world by means of natural selection, exaptation, and subfunctionalization. Biol Direct 2:14
Wong JT (1975) A co-evolution theory of the genetic code. Proc Natl Acad Sci USA 72(5):1909–1912
Wong JT (1980) Role of minimization of chemical distances between amino acids in the evolution of the genetic code. Proc Natl Acad Sci USA 77(2 II):1083–1086
Wong JT (2007) Question 6: coevolution theory of the genetic code: a proven theory. Orig Life Evol Biosph 37(4-5):403–408
Wong JTF (2005) Coevolution theory of genetic code at age thirty. BioEssays 27(4):416–425
Yarus M (2011) The meaning of a minuscule ribozyme. Philos Trans R Soc B Biol Sci 366(1580):2902–2909
Yarus M, Widmann JJ, Knight R (2009) RNA-amino acid binding: a stereochemical era for the genetic code. J Mol Evol 69(5):406–429
Acknowledgements
We thank the EiC and two anonymous reviewers for suggestions which improved the manuscript. Part of this research has been funded by NWO-VICI Grant 639-023-302, by the NWO-CLS MEMESA Grant, by the Tinbergen Institute, and by a NWO-VENI Grant.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendices
Appendices
Four further observations are reported here. Firstly, as explained in ‘Introduction’ section , consideration of the biosynthetic pathways leading to the different amino acids suggests an aspect of organization of the SGC, in which GNN codons tend to be assigned to ‘prebiotic amino acids’, ANN codons to comparatively small, aspartate-derived amino acids, CNN codons to larger amino acids, and UNN codons to the largest, or (in the case of cysteine) the most instable and reactive amino acid. In other words: the first position of the codon might have a link with the complexity of biochemistry, e.g. the UNN codons being the only ones encoding aromatic amino acids and the instable cysteine, and reflecting the most advanced stage of biochemistry during the evolution of the genetic code (when the biochemistry was sufficiently complex to handle cysteine, and to build tryptophan). In ‘Appendix :Molecular Structure Matrix’, we study this link with the biosynthetic development of amino acids by measuring how many one-atom changes are required to transform one amino acid into another. With respect to this distance measure, amino acids derived from the same precursor (like e.g. Ile and Thr) are comparatively close, because they share structure parts. Changing the second position of the codon (in the case of Ile and Thr: changing AUU to ACU) would then replace an amino acid by one with a comparatively similar structure, reflecting their membership of the same biosynthetic family. If the error-robustness calculation is performed with these molecular-structure distances, the SGC is found to have error protection in substitution mutations in the second position (and therefore grouping e.g. ANU codons together). The results are given in ‘Appendix :Molecular Structure Matrix’.
Secondly, we tried to find numerical values for the 20 amino acids which make the SGC optimal in terms of error robustness among all possible genetic codes. Using a numerical optimization approach developed by Eppstein (2003), we were able to find 20 such values. In fact, many different sets of 20 values have this property. Details about these SGC-optimality calculations can be found in ‘Appendix: Inverse Parametric Optimization’.
Thirdly, we screened a large list of physico-chemical amino acid characteristics on their performance in our error-robustness calculations. Polar requirement was one of the best performing measures. This strongly supports the remark by Haig and Hurst [‘The natural code is very conservative with respect to polar requirement. The striking correspondence between codon assignments and such a simple measure deserves further study’ (Haig and Hurst 1991)]. The observation of Vetsigian, Woese, and Goldenfeld (‘Although we do not know what defines amino acid ‘similarity’ in the case of the code, we do know one particular amino acid measure that seems to express it quite remarkably in the coding context. That measure is amino acid polar requirement […]’ (Vetsigian et al. 2006) should also be mentioned. More details are given in ‘Appendix: Scan of Other Amino Acid Properties’.
Finally, we wondered if, by fixing just one or two more assignments, the SGC would be optimal without using the subdivision leading to the historically reasonable set of possible codes (as explained in ‘Introduction’ section) This was not the case. When working with Haig–Hurst weights (i.e. equal weighting), there exist 34 sets of 9 fixed assignments which do have this characteristic. However, none of these 34 sets consists of the seven fixed assignments based on aptamer considerations plus two more amino acids. The smallest set containing the seven has size 10. When working with Freeland–Hurst weights (see ‘Methods’ section), sets of 8 or 9 fixed assignments with the required characteristic, do not exist. This work is presented in ‘Appendix: Minimal Number of Fixed Assignments’.
Molecular Structure Matrix
Polar requirement is just one physico-chemical aspect of amino acids. The discovery that only 1 in 10000 random codes has a lower error-robustness value than the SGC when polar requirement is used as an amino acid characteristic (Haig and Hurst 1991) is compelling evidence that error robustness is present in the SGC. When a conservative attitude is taken, and a phenomenon is considered noteworthy only when the probability to encounter it as a random effect is <0.1 %, the SGC is clearly noteworthy. If one considers the error-robustness values for the three positions separately [please refer to Buhrman et al. (2011) for details] the results in the left column of Fig. 2 are obtained. The third position is in the <0.1 % category, the first position is in the <1 % category, while the second position, with about 22 %, is not even in the <5 % category, and can thus not be considered special.
Histograms of the MS-values of 10 million random samples using updated polar requirement (Mathew and Luthey-Schulten 2008) (4 histograms on the left) and molecular-structure distances from Table 3 squared (4 histograms on the right). The top row shows the MS0 value, the second row is the component from the first codon position (MScore1), third and forth row the components from the middle (MScore2) and last (MScore3) codon position. In contrast to the original definition (Haig and Hurst 1991) of MSi for i ≥ 1, we have chosen to normalize MScorei with the same constant as MS0 so that MS0 = ∑ 3 i=1 MScorei. The dashed red line indicates the value of the SGC
This result is not entirely satisfactory, because the codons of several pairs of similar amino acids are related by second position changes. For instance, a change from phenylalanine (Phe) to tyrosine (Tyr) is clearly a conservative change from a biological viewpoint. To develop a measure for this kind of amino acid relatedness, we introduce a new way of measuring amino acid similarity by one-atom changes which yields a measure of similarity in terms of molecular structure. We should stress that this measure does not reflect actual chemical reactions/steps. As an example, we compute the distance between Phe and Tyr to be 3 as follows: the hydrogen atom at the end of the side chain of Phe is taken off as a first step. An oxygen atom is placed on the position, which the hydrogen atom had before as a second step. The Tyr molecule is completed by addition of an hydrogen atom on top of this oxygen atom, producing the hydroxyl group at the end of the side chain of Tyr, and this is the third and final step. Generally, the distance between two molecules is defined to be the minimal number of ‘allowed one-atom changes’ to transform one molecule into the other, where the allowed one-atom changes are the following:
-
taking off or attaching an arbitrary single atom,
-
creating or destroying a single bond (thereby possibly opening or closing a ring structure),
-
changing a single bond to a double bond or vice versa.
It is not hard to see that an algorithmic way of computing the distance between two molecules m1 and m2 is to find the maximal common sub-graph mc of their molecular structure, and to sum up how many steps are required to go from m1 to mc and from m2 to mc. The distance matrix between the 20 amino acids in Table 3 has been obtained in this way, using the Small Molecule Subgraph Detector (SMSD) toolkit (Rahman et al. 2009) to find the maximal common subgraph and post-processing this information with a python script. The software code can be found in the supplemental information.
In order to perform the error-robustness calculations, we followed the procedure by Haig and Hurst (1991) and considered the squared distances. In this way, the zeroes in the diagonal remain zero. The values for small changes become slightly larger (so the edge from Phe to Tyr gets a value 9), while the values for large changes (like going from Gly to Tyr) become considerably larger (in the case of Gly to Tyr 20 becomes 400). Large changes thus get stronger emphasis (Di Giulio 1989). Whether squaring is the right way to make these kind of calculations has been discussed elsewhere (Ardell 1998; Freeland et al. 2000); we just want to compare molecular structure as an input to characteristics like polar requirement, hydropathy, volume and isoelectric point, as studied by Haig and Hurst (1991). The histograms of the error-robustness in terms of molecular structure are shown in the right column of Fig. 2.
Although not producing (unlike polar requirement) a result in the <0.1 % category, it is still remarkable that the SGC is, with 0.151 %, in the <1 % category when molecular structure is used as input. This means that this matrix is performing better than volume or the hydropathy scale of hydrophobicity in the work of Haig and Hurst (1991). Even more remarkable, the error robustness comes mainly from the second position, using this measure (Fig. 2).
Inverse Parametric Optimization
Instead of asking the question ‘What is the most error-robust genetic code in terms of e.g. polar requirement?’, one could also ask the question ‘Is there a set of numerical values for the 20 amino acids such that the SGC is the optimal code in terms of error robustness?’ If one particular set of 20 values turns out to have that property, one can compare this set with different sets of amino acid characteristics, and suggest which characteristic resembles the ‘ideal values’ best. This then might be the factor playing a selective role during evolution of the SGC.
Let A be the set of amino acids and let \(\mathcal{F}\) be the set of all codes. We aim at solving the following problem: find a non-trivial vector \({\bf x} \in {\rm I R}_{\geq 0}^{A}\) of amino acid property values such that \(MS^{\alpha, {\bf x}}_0({\rm SGC}) = \mathop{\rm arg min}\limits_{F \in \mathcal{F}}MS^{\alpha, {\bf x}}_0(F)\).
To solve this problem, we used a modification of the method of Eppstein (2003). We define variables \({\bf x} \in {\rm I R}^{20}\) and consider the following constraint satisfaction problem: Find x such that
Note that the number of inequalities (3) equals the size of the code space, which can be quite large. To deal with the potentially large number of constraints we follow a cutting plane approach. We work with intermediate solutions \(\overline{{\bf x}}_i\), start with i = 0, and set \(\overline{{\bf x}}_0\) to some random values that satisfy constraints (1) and (2). We then solve the separation problem for the class of constraints (3). That is, we have to find a code F such that \(MS^{\alpha,\overline{{\bf x}_i}}_0(F) < MS^{\alpha,\overline{{\bf x}}_i}_0({\rm SGC})\) or prove that no such code exists. We can answer this question by finding
using the quadratic assignment approach described in Buhrman et al. (2011). In fact, for the actual procedure it suffices to use much faster QAP heuristics, e.g. based on simulated annealing (Burkard and Rendl 1984) or the GRASP heuristic (Li et al. 1994), instead of full QAP solvers. If we find an F with \(MS^{\alpha,\overline{{\bf x}_i}}_0(F) <\; MS^{\alpha,\overline{{\bf x}}_i}_0({\rm SGC})\), we have found a violated inequality
which we add to the constraint satisfaction problem. We solve this set of quadratic constraints using the non-linear constraint solver fmincon from MATLAB’s optimization toolbox (MATLAB 2011), obtain a new set of values \(\overline{{\bf x}}_{i+1}\) and iterate the process until no more violated inequalities can be separated. A final solution x* can be verified by a QAP solver such as Burkard and Derigs (1980). All software used is provided as supplemental information.
Using this procedure, we found many different sets of 20 values under which the SGC is optimal with respect to error-robustness. We steered the values towards the polar requirement values r by using the distance to r as the objective function in our approach. See Fig. 3 for an illustration of some of the solutions we found.
Eight examples of sets of values for the 20 amino acids that make the SGC the most error-robust genetic code. The (artificial) values are found by using inverse parametric optimization, as described in Appendix: Inverse Parametric Optimization. All sets have been normalized to have mean 0 and standard deviation 1. For comparison, we also show the original polar requirements on top (1), and the updated polar-requirement values on the second row (2). Value sets 3–6 make the SGC optimal with respect to MS0. Value sets 7–10 make the SGC optimal with respect to MS FH0
An analysis of the correlation coefficients of these ‘ideal’ values with a database of 744 known amino acid properties from the literature (AAindex: Kawashima et al. 1999) shows no correlation above 0.82 except with polar requirement. In other words, we do not know of any sets of straightforward physico-chemical amino acid properties which resemble one of these ‘ideal’ sets. This might suggest that a combination of several aspects of code evolution and amino acid properties [as suggested by e.g. Higgs (2009)] resulted in the configuration of the SGC.
Scan of Other Amino Acid Properties
We performed error-robustness calculations for all (complete) amino acid properties of the AAindex-database (Kawashima et al. 1999). For the purpose of comparison, we extended the database to include the original polar requirements (Woese et al. 1966a), and the updated polar requirements (Mathew and Luthey-Schulten 2008), as well as two sets of numerical values found by the procedure described in ‘Appendix: Inverse Parametric Optimization’.
In a first scan, 50000 random codes were sampled from
-
1.
all codes,
-
2.
codes with the seven assignments of Phe, Tyr, Trp, His, Leu, Ile, and Arg fixed,
-
3.
codes with seven fixed assignments and respecting the structure enforced by the constraint of the historically reasonable set of possible codes (all 11,520 codes were computed in this case).
For all of the three settings above, error-robustness values were computed using Haig–Hurst and Freeland–Hurst weights (the same random samples were used for the two weight sets, the results are thus statistically correlated).
Out of the 55 best-performing codes, the same calculations as above were performed with 106 samples. The 20 best performing properties are presented in Table 4. Not surprisingly, our two sets of (artificial) numerical values found by inverse parametric optimization (described in ‘Appendix Inverse Parametric Optimization’) end up on the top.
Furthermore, we observe that the SGC is error-robust in terms of several measures of polar requirement [as noted, e.g. in Vetsigian et al. (2006)]. One of these (for which this is not immediately obvious) is Grantham’s polarity scale (1974), which is a combination of Aboderin’s scale (1971) and polar requirement. It is especially noteworthy that the updated polar requirement (Mathew and Luthey-Schulten 2008) is consistently showing up within the best four sets of numerical values. When the sets found by inverse parametric optimization are left out, the updated values of polar requirement are in all three settings (no blocks fixed, 7 blocks fixed, and the set of 11,520 codes resulting from 7 fixed blocks plus the constraint of the historically reasonable set of possible codes) the best set of values when Freeland–Hurst weights are used.
Minimal Number of Fixed Assignments
In this appendix, we investigate how many amino acid assignments need to be fixed such that the SGC is the most error-robust genetic code with respect to the updated polar requirements (Mathew and Luthey-Schulten 2008), when we do not use the constraint of the historically reasonable set of possible codes.
For the case of the Haig–Hurst weights, there are 67 different minimal subsets \(S_1, S_2, \ldots, S_{67} \subseteq \{\hbox{Phe}, \hbox{Leu}, \hbox{Ile}, \ldots, \hbox{Ser}, \hbox{Gly}\)} such that for any \(i \in \{1,2,\ldots, 67\)}, fixing the assignments of all amino acids in Si makes the SGC the most error-robust genetic code. Any super-set of these 67 minimal subsets will also have this property, because fixing more assignments only limits the number of possible genetic codes. Out of the 67 minimal subsets, 34 of them are of size 9, 15 of size 10, 15 of size 11, and 3 of size 12.
When fixing the seven assignments of Phe, Tyr, Trp, His, Leu, Ile, and Arg (based on aptamer experiments) the minimal sets of assignments that need to be fixed in addition are: {Ser, Gln, Cys} or {Met, Ser, Gln}.
For the case of the Freeland–Hurst weights, there are 186 different minimal subsets: 2 subsets of size 10, 4 of size 11, 13 of size 12, 44 of size 13, 52 of size 14, 45 of size 15, 21 of size 16, and 5 of size 17. When fixing the seven assignments of Phe, Tyr, Trp, His, Leu, Ile, and Arg (based on aptamer experiments), there are 6 different minimal sets (of size 6) each of which can be fixed in addition in order to make the SGC the most error-robust genetic code.
Rights and permissions
About this article
Cite this article
Buhrman, H., van der Gulik, P.T.S., Klau, G.W. et al. A Realistic Model Under Which the Genetic Code is Optimal. J Mol Evol 77, 170–184 (2013). https://doi.org/10.1007/s00239-013-9571-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-013-9571-2