Abstract
Across the streptophyte lineage, which includes charophycean algae and embryophytic plants, there have been at least four independent transitions to the terrestrial habitat. One of these involved the evolution of embryophytes (bryophytes and tracheophytes) from a charophycean ancestor, while others involved the earliest branching lineages, containing the monotypic genera Mesostigma and Chlorokybus, and within the Klebsormidiales and Zygnematales lineages. To overcome heat, water stress, and increased exposure to ultraviolet radiation, which must have accompanied these transitions, adaptive mechanisms would have been required. During periods of dehydration and/or desiccation, proteomes struggle to maintain adequate cytoplasmic solute concentrations. The increased usage of charged amino acids (DEHKR) may be one way of maintaining protein hydration, while increased use of aromatic residues (FHWY) protects proteins and nucleic acids by absorbing damaging UV, with both groups of residues thought to be important for the stabilization of protein structures. To test these hypotheses we examined amino acid sequences of orthologous proteins representing both mitochondrion- and plastid-encoded proteomes across streptophytic lineages. We compared relative differences within categories of amino acid residues and found consistent patterns of amino acid compositional fluxuation in extra-membranous regions that correspond with episodes of terrestrialization: positive change in usage frequency for residues with charged side-chains, and aromatic residues of the light-capturing chloroplast proteomes. We also found a general decrease in the usage frequency of hydrophobic, aliphatic, and small residues. These results suggest that amino acid compositional shifts in extra-membrane regions of plastid and mitochondrial proteins may represent biochemical adaptations that allowed green plants to colonize the land.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
At least four independent transitions from water to terrestrial habitats occurred during the diversification of the streptophytes, which include charophycean green algae and embryophytic plants. One of these represents a major organismal radiation during the history of life on earth, with the evolution of embryophytes (bryophytes + tracheophytes) from a freshwater charophyceae ancestor (Kenrick and Crane 1997; Bateman et al. 1998; Qiu and Palmer 1999; Karol et al. 2001; Lewis and McCourt 2004; Qiu et al. 2006, 2007; Turmel et al. 2006; Lemieux et al. 2007), which is estimated to have occurred between 425 and 490 million years ago (Mya) (Sanderson 2004).
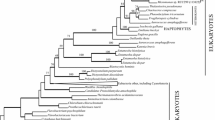
Another transition involves the unicellular monotypic genera Mesostigma and Chlorokybus that form the earliest branching lineage in the streptophyte tree (Fig. 1) (Qiu et al. 2006; Lemieux et al. 2007). Mesostigma occupies the water column in freshwater habitats, while Chlorokybus has adapted to occupy sub-aerial habitats (Lewis and McCourt 2004; Lemieux et al. 2007). Klebsormidiales is another early branching lineage that contains members that occupy terrestrial habitats, in addition to two genera from the Zygnematales that can form sub-aerial mats (i.e., Cylindrocystis, Mesotaenium) (Lewis and McCourt 2004).
Streptophyta phylogeny (Karol et al. 2001; Qiu et al. 2006, 2007) showing major evolutionary grades of green algae, ‘bryophytes’, and the tracheophyte lineage. Proteomic sequence data, derived from plastid (Pt) and mitochondrial (Mt) proteomes, were obtained for taxa presented in the tree as indicated within closed brackets. Branches with broken lines indicate an ‘aquatic’ → ‘terrestrial’ transition, where proteomes may have optimized amino acid composition to cope with higher temperatures, increased ultraviolet radiation (UV), and cellular desiccation/dehydration. Major morphological adaptations for terrestrialization are described in external text-boxes at respective nodes in the tree, and an estimated date of divergence (Sanderson 2004) is provided within an internal box
To overcome increased water stress, heat, and exposure to UV, which must have accompanied these transitions, adaptive mechanisms may have been required (Graham et al. 2000; Proctor and Pence 2002; Waters 2003; Oliver et al. 2005; Rensing et al. 2008; Wang et al. 2009b; Khandelwal et al. 2010; Richardt et al. 2010; Wolf et al. 2010).
Desiccation tolerance in plants involves mechanisms of cellular protection for vegetative structures, spores, seeds, and pollen; research on this topic is centered on the accumulation of osmoregulating free amino acids (e.g., proline) in the cellular solution (Blum and Ebercon 1976; Carmo-Silva et al. 2009; Khandelwal et al. 2010; Richardt et al. 2010), over-expression of abiotic stress related genes (Frank et al. 2005; Close 1997; Cuming 1999; Proctor and Smirnoff 2000; Proctor et al. 2007; Finkelstein et al. 2008; Hájek and Beckett 2008; Rensing et al. 2008; Khandelwal et al. 2010), and epigenetic modifications (Granot et al. 2009).
Based on phylogenetic mapping, Oliver et al. (2005) suggest that primitive land plants were vegetatively and reproductively tolerant of desiccation, with presence of active dehydrin/rehydrin proteins found in liverwort and moss lineages (Waters 2003; Close 1997; Cuming 1999; Proctor et al. 2007; Hájek and Beckett 2008; Rensing et al. 2008). With the subsequent divergence of hornworts (Anthocerotophyta), this protective mechanism seems to have been lost, but had reappeared independently in various tracheophyte lineages, presumably as they adapted to arid habitats (Oliver et al. 2005; Granot et al. 2009).
Cellular dehydration/desiccation affects the concentration of cellular solutes and is therefore likely to interrupt the balanced organization of water molecules required for protein folding and stabilization (Mrabet et al. 1992; Jaenicke and Böhm 1998; Borders et al. 1994; Strub et al. 2004). Both these processes are usually dependent on a protein’s surface–polar interface with water (Klotz 1958; Bryant 1996; Killian and Heijne 2000; Smith et al. 2004).
UV radiation is another problem terrestrial plants face; cells, cellular components and proteomes and DNA therein may all be subject to resulting oxidative damage (Tyrrell 1994; Brosche and Strid 2003; Rensing et al. 2008; Wolf et al. 2010). The penetration of UV is mainly limited to surfaces and near-surfaces of cells, and in chloroplasts, can alter thylakoid integrity thereby compromising photosynthesis (Tyrrell 1994; Kovács and Keresztes 2002; Rodrigues et al. 2006).
Increased concentrations of free aromatic compounds (e.g., tannins, flavonoids) (Cooper-Driver and Bhattacharya 1998, Waters 2003), and the release of isoprene hydrocarbon compounds from photosynthetic surfaces are thought to be adaptive mechanisms for protection from UV damage, and heat induced production of reactive oxygen species (Hanson et al. 1999; Sharkey et al. 2008).
As demonstrated above, many mechanisms have evolved that allow plants to deal with environmental stresses during their movement to terrestrial habitats. One additional mechanism that has not been considered so far in the studies of plant colonization of land is the alteration of the amino acid composition, i.e., increase or decrease in the use of certain residues such as those with either polar charge or UV absorbing aromatic rings.
Increased use of polar charged amino acids, particularly the EKR residues, may provide protection from desiccation through thermostability and hydrophilicity (Haney et al. 1999; Killian and Heijne 2000; Nishio et al. 2003; Brocchieri 2004), while increased use of aromatic amino acids (FHWY) on protein surfaces has been suggested to provide thermostability for thermophilic proteins, due primarily to the absorptive properties of the side-chain aromatic ring (Kannan and Vishveshwara 2000). In early land plant lineages such whole proteome adaptations might have been especially important in the light-capturing chloroplast proteome, with UV-B known to damage photosystem components (Tyrrell 1994; Kovács and Keresztes 2002; Rodrigues et al. 2006).
If proteomic adaptations occurred during terrestrialization events in streptophytes, we reasoned that a signature might be found in the amino acid composition of transitional lineages when compared to their closest freshwater aquatic relatives. To answer this question we used concatenated orthologous sets of genes containing 55 chloroplast (plastid), and 13 mitochondrial protein-coding genes, and compared amino acid compositional frequencies between (1) the ‘bryophyte’ and ‘charophycean’ green algal grades, (2) the tracheophyte lineage and ‘bryophytes’, and (3) Chlorokybus and Mesostigma. We also investigated whether proteins/protein regions exposed to the organellar fluid might be under stronger selection for protein hydrophilicity and aromaticity, by comparing transmembrane domains with those of extra-membrane regions.
Materials and Methods
Taxon Sampling and Protein Alignments
Organellar sequences for sampled taxa were obtained from the NCBI GenBank database (http://www.ncbi.nlm.nih.gov) (Fig. 1; refer to Supplementary Table S1 for GenBank accession numbers). All sampled genes are presented in Supplementary Table S2.
Amino acid sequence alignments for the 55 plastid and 13 mitochondrial gene datasets (Supplementary Table S2) were concatenated and analyzed as individual compartments (Figs. 2, 3, 4), and also as classes of structurally related proteins (Supplementary Tables S2, S3).
Percentage difference between mean usage frequencies for categories of amino acid residues. Comparisons are between Chlorkybus–Mesostigma (hatched bars), bryophytes–green algae (excluding Chlorkybus) (gray bars), tracheophytes–bryophytes (open bars), across concatenated sets of orthologous protein-coding genes (n = number of genes). Taxa sampled for a the plastid proteome (Pt) include six alga, four bryophytes, and 42 tracheophytes; b mitochondrial proteome (Mt) includes four alga, three bryophytes, and six tracheophytes. For plastid and mitochondrial data-sets, the Mann–Whitney test was used to evaluate statistical significance (see “Materials and Methods” section). The level of significance is indicated by *P < 0.05; **P < 0.01
Boxplots showing the frequency (%) of the EKR residue category in extra-membrane regions of a chloroplast (Pt), and b mitochondria (Mt), for algae (excluding Chlorokybus), bryophytes, and tracheophytes. Boxes show mean, SD, and values with extreme variance (black circles). Frequency values for Chlorokybus (C.a) are schematically shown in filled circles. Between-group comparisons that are significantly different (P ≤ 0.001) are indicated with brackets
Boxplots showing the frequency (%) of a polar, and b hydrophobic residue categories in extra-membrane regions of the mitochondrion, for algae (excluding Chlorokybus), bryophytes, and tracheophytes. Boxes show mean, SD, and values with extreme variance (black circles). Frequency values for Chlorokybus (C.a) are schematically shown in filled circles. Between-group comparisons that are significantly different (P ≤ 0.001) are indicated with brackets
Alignment of amino acid sequences was performed using Muscle (Edgar 2004). Amino acid alignment length was 13873 and 4276 for the plastid and mitochondrial matrices, respectively. We determine the amino acid composition frequencies for both gapped and ungapped alignments. For consistency, we have used the data obtained from ungapped alignments as similar results were obtained for both analyses.
The amino acid matrix length for transmembrane domains was 2133 (plastid), and 1509 (mitochondrion), while for extra-membrane regions the length was 11740 (plastid) and 2767 (mitochondrion). Both the concatenated plastid and mitochondrial extra-membrane datasets also contained non-membrane-bound genes (refer to Supplementary Table S2).
A manual inspection was performed to identify non-orthologous regions, facilitating an unbiased and accurate estimation of amino acid frequency (alignments available upon request).
Regarding taxon selection, we had to consider the presence of RNA editing sites in the genomic sequences, as these sites can significantly alter the amino acid composition (Giegé and Brennicke 1999; Kugita et al. 2003; Wolf et al. 2004; Jobson and Qiu 2008), and for this reason we only used sequences in which cDNA data were available in GenBank. For the hornwort Megaceros aenigmaticus (Li et al. 2009) we sequenced the cDNA of all mitochondrial genes represented in our dataset (Supplementary Table S2). We did not use available programs to approximate RNA editing sites due to the sensitivity of our analyses: i.e., even just a few undetermined sites can alter the results significantly, as determined by mitochondrial protein alignments for the bryophytes Pleurozia purpurea (GenBank accession: NC_013444) (Wang et al. 2009a) and Phaeoceros laevis (GenBank accession: NC_013765) (Xue et al. 2010), and tracheophyte Isoetes engelmannii (Grewe et al. 2009).
The sensitivity of the analyses also limited us to using only those taxa that contained genomes with the full complement of expressed target genes (Supplementary Table S2).
Several lineages of green algae and early land plants formed two individual grades before and after the colonization of land by plants (Karol et al. 2001, Qiu et al. 2006, 2007), and they showed similar frequencies of amino acid composition. Thus, we lumped the various lineages in these two paraphyletic groups into two groups for the sake of convenient comparison. If the strict clade-to-clade comparison approach was taken, the trend revealed here would just be as distinct or even more so since there would be several lineages on both sides of the terrestrialization event for comparison.
Amino Acid Compositional Analyses and Statistics
The amino acid frequency was determined for all residues across the sampled taxa in data-sets for plastid and mitochondrial proteomes (Supplementary Table S1). Previous studies have calculated an ‘expected’ frequency from codon usage data, and correlated this with the observed usage frequency (King and Jukes 1969). As we wanted to find broad patterns across amino acid categories and taxonomic groups, we took a different approach by calculating the average frequency across taxonomic groupings of streptophytes: freshwater algae (Mesostigmatales, Chlorokybales, Zygnematales, Coleochaetales, Charales), and Embryophytes; bryophytes (Marchantiophyta, Bryophyta, Anthocerotophyta), and tracheophytes (Lycopodiophyta, Moniliformopses Spermatophyta) (sampled taxa are shown in Fig. 1, and Supplementary Table S1).
We categorized residues based on major biochemical side-chain properties using the EMBL-EBI ‘Amino Acid Properties and Substitutions’ website (http://www.ebi.ac.uk), and the Amino Acid Properties’ website (http://www.mcb.ucdavis.edu/courses/bis102/AAProp.html), and formed sub-categories of related residues based on those deemed important in studies of adaptive thermostability (Deckert et al. 1998; Kumar and Nussinov 2001). The analyzed categories are as follows; polar side-chains ‘charged’ (DEHKR), charged polar side-chains ‘negative’ (DE), charged polar side-chains ‘positive’ (HKR), charged polar side-chains deemed important for protein thermostabilization (EKR), combined polar side-chains (DEHKNQRST), uncharged polar side-chains ‘polar-neutral’ (NQST), side-chains ‘aromatic’ (FHWY), side-chains ‘aliphatic’ (ILV), residues ‘hydrophobic’ (AGFILMPVW), and residues ‘small’ (ACDGNSTV).
Data presented in Fig. 2 are the relative difference for the average compositional frequency between taxonomic groups [{AA freq. Group 1 − AA freq. Group 2}/{MAX AA freq.}]. This procedure was performed to account for differences in residue number per amino acid category.
We also partitioned the data into the following functional gene classes: ATP = ATP synthases (plastid, mitochondrion); NDH = nicotinamide dehydrogenases (plastid, mitochondrion); PET = cytochrome-bf (plastid); PSI-II = components of photosystem I and II (plastid); and COX = cytochrome-c-oxidases (mitochondrion). For these classes we present relative difference for the average compositional frequency results for charged EKR, polar, and aromatic groups (Supplementary Table S3).
Transmembrane domain boundaries were estimated for the plastid and mitochondrial protein alignments (see above for length), across all sampled taxa, using the Sequence Annotation feature in the Protein Knowledge Database (UniProtKB) (http://www.uniprot.org).
Amino acid frequency results (relative difference for the average compositional frequency) were determined separately for alignments containing either transmembrane domains or extra-membrane regions, across all residue categories (Supplementary Table S4).
To examine the effect of nucleotide shifts between taxa, we did not look at the nucleotide compositional changes themselves; instead we reasoned that such changes should be more accurately reflected by relative difference between combined AT rich (FYMINK) and GC rich (GARP) amino acid residues (Supplementary Table S4). Our main reason for using this approach was the presence of RNA editing in organelle genomes, which results in an observed non-correspondence of genomic DNA → protein sequence (i.e., genomic GC and AT do not fully reflect the amino acid composition) (see above). In any case, the nucleotide frequency for taxa used in plastid and mitochondrial data-sets were presented in Jobson and Qiu (2008).
We compared taxonomic group means (average compositional frequency per taxonomic grouping), and assessed significance (P ≤ 0.05) using the Mann–Whitney test (Fig. 2; Supplementary Tables S3, S4). We also visualized the compositional data as boxplots (Figs. 3, 4, 5), and compared the significance of frequency difference (P ≤ 0.05) between taxonomic groups using one-way ANOVA in R version 2.9.2 software (http://www.R-project.org) (R Development Core Team 2009). For the Mesostigma/Chlorokybus comparison we did not perform statistical analyses on the relative frequency differences.
Boxplots showing the frequency (%) of the aromatic residue category in extra-membrane regions of a plastid (Pt), and b mitochondria (Mt), for algae (excluding Chlorokybus), bryophytes, and tracheophytes. Boxes show mean, SD, and values with extreme variance (black circles). Frequency values for Chlorokybus (C.a) are schematically shown in filled circles. The value for Mesostigma (M.v) is shown with open circle, and the value for Chara (C.v) is also indicated
Results and Discussion
Most research linking protein level amino acid compositional shifts with adaptation to new habitats has focused on extreme examples comparing optimal growth temperatures across mesophilic versus thermophilic prokaryotes. In particular, the latter group shows an observed pattern of increased use of thermostabilizing polar charged amino acids, particularly the hydrophilic EKR category (Haney et al. 1999; Killian and Heijne 2000; McDonald 2001; Nishio et al. 2003; Singer and Hickey 2003; Brocchieri 2004), at the expense of uncharged polar residues (Haney et al. 1999; Kumar and Nussinov 2001; McDonald 2001). Differing interpretations of the above observations have lead to two sides; those that attribute this pattern to mutational biases driven by nucleotide compositional shifts (Foster et al. 1997; Gu et al. 1998; Jabbari et al. 2003; Lobry and Chessel 2003; Wang et al. 2004), and those tending toward the involvement of adaptive processes (Deckert et al. 1998; Haney et al. 1999; Killian and Heijne 2000; McDonald 2001; Nishio et al. 2003; Brocchieri 2004; Tekaia and Yeramian 2006; Boussau et al. 2008; Saelensminde et al. 2009).
Here we take a first look at this question across episodes of terrestrialization in the streptophyte lineage (Fig. 1). We examined the amino acid frequency of extant taxa (i.e., tree terminals), under the assumption that particular “fixed” amino acid usage frequency patterns had changed in response to shifts in the ecological/physiological environment; in our case, freshwater aquatic versus terrestrial habitats. It should be noted that our analyses are constrained by the assumption of within-taxon amino acid compositional stability over a period of >400 million years (Sanderson 2004).
Residues for Protecting Proteins Against Cellular Dehydration and Desiccation, and Maintaining Their Thermal Stability
Our comparison of amino acid frequencies in extra-membrane regions for bryophytes and algae revealed a positive difference in frequency of charged residues, including negative, positive, and EKR categories. This pattern is consistent with that of the Chorokybus–Mesostigma comparison, with the strongest consistent increase observed for positively charged residues (HKR) in both plastid and mitochondria (Fig. 2a, b), while the greatest difference was for EKR (bryophyte–algae; P ≤ 0.05) in the plastid genome (Fig. 2a).
We then examined the within-group distribution for EKR (Fig. 3) and found an unbiased increase in the plastid proteome across all bryophytes versus algae (P ≤ 0.05), and a clear increase for Chlorokybus relative to the five other algal species (Fig. 3a). This trend was also observed in the mitochondrion (Fig. 3b). When we looked at the plastid EKR frequency in major gene classes, we found that all contribute positively to this pattern, with ATP, NDH, PET, and ribosomal/polymerase contributing a significant effect (Supplementary Table S3). For the mitochondrion, the NDH gene class contributes all of the positive EKR effect observed in Fig. 2b, while ATP and COX classes contribute a slightly negative trend (Supplementary Table S3).
When we combined all polar residues (DEHKNQRST) for the bryophyte–algae comparison we found that the mitochondrial frequency increases significantly (P ≤ 0.001) (Figs. 2b, 4a), with all gene classes contributing to this positive pattern (Supplementary Table S3), while for the same comparison in plastids, the effect is dampened, due mainly to the corresponding decrease of polar-neutral residues (NQST) (Fig. 2a). A reverse trend was found for aliphatic and hydrophobic residues (Fig. 2b), with significant losses of the latter (P ≤ 0.001) found in the mitochondrion (Fig. 4b), and a non-significant loss for both categories in plastids (Fig. 2a). For the plastid proteome we instead found that the small residue category decreases significantly (P ≤ 0.05), while remaining constant for the mitochondrion (Fig. 2b).
For the Chlorokybus mitochondrial proteome we find that the compositional frequencies of the combined polar (Fig. 4a), and hydrophobic categories (Fig. 4b) had strongly diverged from Mesostigma, reaching levels comparable to the mean of the bryophyte grade. This pattern of positive increase is observed in all mitochondrial gene classes, but only ATP and PET classes for the plastid proteome (Supplementary Table S3).
The above analyses of usage frequency within categories of amino acid residues revealed patterns within extra-membranous regions, under the hypothesis that natural selection for hydrophilicity would be stronger on exposed regions of proteins. To further test this idea we also performed the same analyses on transmembrane domains and found that in the plastid proteome both the bryophyte–algae and Chorokybus–Mesostigma comparisons were generally consistent with this hypothesis: extra-membrane regions show a stronger increase in the charged polar residue categories than that of the transmembrane domains (Supplementary Table S4A).
We found that the pattern for the mitochondrion was less clear (Supplementary Table S4B). This may be due to the limited transmembrane data (see “Materials and Methods” section), and the low percentage of charged polar residues normally located in transmembrane domains (i.e., mean frequency of EKR in mitochondria: transmembrane = 2.2%; extra-membrane = 14.3%), with a similar contrast found in the plastid proteome (data not shown). However, a more robust comparison can be made using the combined polar category, with average transmembrane mitochondrial and plastid frequencies of 29 and 33%, versus 54 and 57% for extra-membrane regions, respectively. From this comparison it seems generally clear that as streptophytes transitioned to terrestrial habits, polar residues were gained less frequently in transmembrane domains than that in extra-membrane regions (Supplementary Table S4).
We have uncovered a consistent pattern of a positive difference for charged residues as early land plants (represented by the bryophyte grade) colonized land, and as Chlorokybus adapted to aerial environments. Further, the clear algae to bryophyte EKR increase in plastids is made more striking when one considers that the ‘fixed’ frequencies for algae are represented by terminal taxa across four major nodes of streptophyte evolution (Fig. 1), with the sub-aerial Chlorokybus being the only taxon to deviate from the algal trend.
Aromatic Residues for Protecting Proteins and Nucleic Acids Against UV Damage
Increased usage of aromatic amino acids (HFWY) is observed in extra-membrane regions in the plastid proteome across all bryophytes versus all but one algal taxon (Chara) (Fig. 5a), with this comparison approaching significance (P = 0.06) (Fig. 2a). For the bryophyte–algae comparison of the plastid proteome, all gene classes, except PET, contribute to the positive HFWY usage pattern, with that for PSI-II and NDH found to be significantly higher (P ≤ 0.05) (Supplementary Table S3). The lowest levels were found for Mesostigma, with a relatively small increase (0.1%) in Chlorokybus versus Mesostigma (Fig. 5a).
Of the obligate aquatic algae, Chara shows the highest levels for HFWY in plastids, deviating from the algal mean by 0.6%, followed in usage frequency by the first branching bryophyte Marchantia (Figs. 1, 3c). Charales (represented by our sampled Chara) are thought to be the sister lineage to embryophytes (Kenrick and Crane 1997; Bateman et al. 1998; Qiu and Palmer 1999; Karol et al. 2001; Lewis and McCourt 2004; Qiu et al. 2006, 2007; Lemieux et al. 2007; Qiu 2008) (Fig. 1), and perhaps under this phylogenetic scenario (refer to Turmel et al. 2006 for an alternate phylogenetic placement of Chara), the observed build-up of aromatic residues was a pre-adaptation for life on land (Fig. 1). To examine this hypothesis, we trialed the inclusion of Chara within the bryophyte group and found HFWY is significantly elevated (P ≤ 0.001) when compared to the other aquatic algae (Fig. 5a).
The anchored thalli of Characeae, which often extends to the surface of the water column (de Bakker et al. 2005; Becker and Marin 2009), may be occasionally exposed to stronger UV when water levels decrease in its littoral habitat (de Bakker et al. 2005). In comparison to the relatively large and branched filamentous whorls found in the Characeae, the single-celled Mesostigma, branched filaments of Coleochaetales, and conjugating cells and unbranched filaments of the Zygnematales are less complex in morphology and possibly less exposed to UV (de Bakker et al. 2005).
In the mitochondrion, we find very little change between algae and bryophytes in the mitochondrion (Fig. 2b, 5b), which suggests that any adaptive increase in HFWY is restricted to the plastid proteome. In contrast to the pattern observed in plastids, for the mitochondrion Mesostigma strongly exceeded the frequency of Chlorokybus, with a 0.6% decrease in HFWY for Chlorokybus relative to the algal mean (Fig. 5a).
The above findings, restricted mainly to the light-absorbing plastid proteome, and interestingly, to the protein classes involved in photon capture (PSI-II), and cyclic electron transport (NDH), are suggestive of adaptation for dealing with increased UV damage to thylakoid membranes encountered during the colonization of land by plants (Tyrrell 1994; Wolf et al. 2010).
Evolution of Vascularized Tissues
Regarding niche occupation, it is conceivable that early land plants would have been highly restricted due to their shallowly placed rhizoids, with extant taxa generally limited to moist substrates. In contrast, the tracheophyte body plan enabled the occupation of dryer habitats through evolution of highly specialized regulation of cellular water relations; efficient water uptake through branched root systems, and control of loss via specialized leaf surface stomata (Kramer and Boyer 1995; Graham et al. 2000; Chaves et al. 2003; Chen et al. 2005; Song et al. 2005; Carmo-Silva et al. 2009).
Our bryophyte–algae comparison uncovered a pattern of increased usage for both polar and aromatic residues, and we now ask whether this pattern of usage frequency continued throughout the evolution of the tracheophyte lineage (Fig. 1). During terrestrialization we observe a significant build-up of polar charged amino acids in the plastid proteome (Figs. 2a, 3a). However, after the evolution of vascularized tissues we only detect a weak increase, or decrease in the case of EKR (Figs. 2a, 3a), with only Adiantum and all four grass taxa (Supplementary Table S1) reaching levels comparable to the bryophyte mean (~18.8%) (Fig. 3a). Wang et al. (2004) suggest that this increase in grasses is more related to mutational bias, and we also observe increased levels for the GC rich GARP residues after vascularization for both proteomes (Supplementary Table S4), which suggests that nucleotide compositional shifts may have, in part, contributed to the pattern. Banerjee et al. (2006) compared amino acid frequency across nuclear genomes of A. thaliana and O. sativa and found evidence for natural selection as a possible driver of increased hydrophobicity; however, in contrast to our organellar results, their study found that hydrophobic residues were favored by the grass taxon O. sativa.
The mitochondrial proteome does not conform to the above plastid pattern for charged side-chain categories, with generally stronger positive change occurring during/after vascularization, with corresponding significant losses of both aromatic and hydrophobic residues (Fig. 2b). For the hydrophobic category, the pattern is very weak, while for the non-charged and small categories there was a significant positive frequency difference (P < 0.05 and >0.001, respectively) (Fig. 2a).
Effects of Mutational Biases
Although our observed amino acid compositional shifts are suggestive of adaptive evolution for life on land, we are aware that random forces may have been involved in shaping these patterns (Foster et al. 1997; Gu et al. 1998; Jabbari et al. 2003; Lobry and Chessel 2003; Wang et al. 2004). Natural selection acts on individual point mutations, and therefore, it is difficult to envision the concerted fixation of adaptive substitutions across whole genomes, even over long periods of time (Jobson et al. 2010). We also note that the habitat transitions examined in this study likely involved major changes in population structure, with this factor previously shown to drive higher levels of protein evolution in large mammals with smaller population sizes, possibly due to a corresponding weakness of purifying selection (Popadin et al. 2007). In this study we were not able to differentiate potential mutational effects from those driven by purifying selection. However, we tested the effect of GC bias on our observed EKR and HFWY patterns by comparing the relative difference for GC rich (GARP) and AT rich (FYMINK) codons and found no significant between-group bias for the bryophytes–algae comparison.
From these results we find very little observed change in either plastid (P = 0.24) or mitochondrial proteomes (P = 0.35) (Supplementary Table S4). However, we did find significant biases of GARP for the bryophyte–tracheophyte comparisons, with a significant difference for tracheophytes relative to bryophytes in both plastid (P ≤ 0.001) and mitochondrial proteomes (P ≤ 0.001) (Supplementary Table S4).
Perhaps the most striking result is the similarity in magnitude of GARP-shift between transmembrane domains and extra-membrane regions (Supplementary Table S4), a pattern that does not correspond with the negative usage frequency of EKR in plastid transmembrane domains for the bryophytes–algae comparison (Supplementary Table S4).
It is also worth mentioning that the genetic code presumably evolved to alternate in response to changing environments by assigning the thermostable nucleotides GC to codons that encode thermostabilizing amino acid residues (Xia and Li 1998; Ikehara 2002). In support of this idea, shifts in nucleotide composition have been linked to environmental change (Cruveiller et al. 1999; Naya et al. 2002; Knight et al. 2004; Musto et al. 2004; Romero et al. 2009), and it is becoming evident that compositional shifts may be shaped by environmental conditions (Foerstner et al. 2005; Romero et al. 2009).
Conclusions
In this study we uncovered consistent patterns of amino acid compositional fluxuation that correlate with episodes of terrestrialization; positive change in usage frequency for residues with charged side-chains, and a general decrease in the usage frequency of aliphatic, hydrophobic, and small residues, and a positive change in the usage of aromatic residues in the plastid proteome. These results agree with a hypothesis that amino acid residue changes may protect proteins and nucleic acids from UV, heat, and water stress. Since the same patterns were observed in two independent terrestrialization events, charophycean algae → bryophytes, and Mesostigma → Chlorokybus, these results suggest that the amino acid compositional changes reflect adaptive evolution at the biochemical level when these photosynthetic eukaryotes moved from water to land. Once more extensive genomic data are available for comparison, taxa representing the Klebsormidiales, and Cylindrocystis and Mesotaenium of the Zygnematales should also be assessed.
References
Banerjee T, Gupta SK, Ghosh TC (2006) Compositional transitions between Oryza sativa and Arabidopsis thaliana genes are linked to the functional change of encoded proteins. Plant Sci 170:267–273
Bateman RM, Crane PR, DiMichele WA, Kenrick PR, Rowe NP, Speck T, Stein WE (1998) Early evolution of land plants: phylogeny, physiology, and ecology of the primary terrestrial radiation. Annu Rev Ecol Syst 29:263–292
Becker B, Marin B (2009) Streptophyte algae and the origin of embryophytes. Ann Bot 103:999–1004
Blum A, Ebercon A (1976) Genotypic responses in sorghum to drought stress. III. Free proline accumulation and drought resistance. Crop Sci 16:428–431
Borders CL Jr, Broadwater JA, Bekeny PA, Salmon JE, Lee AS, Eldridge AM, Pett VB (1994) A structural role for arginine in proteins: multiple hydrogen bonds to backbone carbonyl oxygens. Protein Sci 3:541–548
Boussau B, Blanquart S, Necsulea A, Lartillot N, Gouy M (2008) Parallel adaptations to high temperatures in the Archaean eon. Nature 456:942–945
Brocchieri L (2004) Environmental signatures in proteome properties. Proc Natl Acad Sci USA 101:8257–8258
Brosche M, Strid A (2003) Molecular events following perception of ultraviolet- B radiation by plants. Physiol Plant 117:1–10
Bryant RG (1996) The dynamics of water-protein interactions. Annu Rev Biophys Biomol Struct 25:29–53
Carmo-Silva AE, Francisco A, Powers SJ, Keys AJ, Ascensão L, Parry MAJ, Arrabaça MC (2009) Grasses of different C4 subtypes reveal leaf traits related to drought tolerance in their natural habitats: changes in structure, water potential, and amino acid content. Am J Bot 96:1222–1235
Chaves MM, Maroco JP, Pereira JS (2003) Understanding plant responses to drought—from genes to the whole plant. Funct Plant Biol 30:239–264
Chen Z, Hong X, Zhang H, Wang Y, Li X, Zhu JK, Gong Z (2005) Disruption of the cellulose synthase gene, AtCesA8/IRX1, enhances drought and osmotic stress tolerance in Arabidopsis. Plant J 43:273–283
Close T (1997) A commonality in the response of plants to dehydration and low temperature. Physiol Plantarum 100:291–296
Cooper-Driver GA, Bhattacharya M (1998) Role of phenolics in plant evolution. Phytochemistry 49:1165–1174
Cruveiller S, Jabbari K, D’Onofrio G, Bernardi G (1999) Different hydrophobicities of orthologous proteins from Xenopus and human. Gene 238:15–21
Cuming AC (1999) LEA proteins. In: Shewry PR, Casey R (eds) Seed proteins. Kluwer, Netherlands, pp 753–780
de Bakker NV, van Bodegom PM, van de Poll WH, Boelen P, Nat E, Rozema J, Aerts R (2005) Is UV-B radiation affecting charophycean algae in shallow freshwater systems? New Phytol 166:957–966
Deckert G, Warren PV, Gaasterland T, Young WG, Lenox AL, Graham DE, Overbeek R, Snead MA, Keller M, Aujay M, Huber R, Feldman RA, Short JM, Olsen GJ, Swanson RV (1998) The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature 392:353–358
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
Foerstner KU, von Mering C, Hooper SD, Bork P (2005) Environments shape the nucleotide composition of genomes. EMBO Rep 6:1208–1213
Foster PG, Jermiin LS, Hickey DA (1997) Nucleotide composition bias affects amino acid content in proteins coded by animal mitochondria. J Mol Evol 44:282–288
Frank W, Ratnadewi D, Reski R (2005) Physcomitrella patens is highly tolerant against drought, salt and osmotic stress. Planta 220:384–394
Giegé P, Brennicke A (1999) RNA editing in Arabidopsis mitochondria effects 441 C to U changes in ORFs. Proc Natl Acad Sci USA 96:15324–15329
Graham LE, Cook ME, Busse JS (2000) The origin of plants: body plan changes contributing to a major evolutionary radiation. Proc Natl Acad Sci USA 97:4535–4540
Granot G, Sikron-Persi N, Gaspan O, Florentin A, Talwara S, Paul LK, Morgenstern Y, Granot Y, Grafi G (2009) Histone modifications associated with drought tolerance in the desert plant Zygophyllum dumosum Boiss. Planta 231:27–34
Grewe F, Viehoever P, Weisshaar B, Knoop V (2009) A trans-splicing group I intron and tRNA-hyperediting in the mitochondrial genome of the lycophyte Isoetes engelmannii. Nucleic Acids Res 37:5093–5104
Gu X, Hewett-Emmett D, Li WH (1998) Directional mutational pressure affects the amino acid composition and hydrophobicity of proteins in bacteria. Genetica 102(103):383–391
Hájek T, Beckett RP (2008) Effect of water content components on desiccation and recovery in sphagnum mosses. Ann Bot 101:165–173
Haney PJ, Badger JH, Buldak GL, Reich CI, Woese CR, Olsen GJ (1999) Thermal adaptation analyzed by comparison of protein sequences from mesophilic and extremely thermophilic Methanococcus species. Proc Natl Acad Sci USA 96:3578–3583
Hanson DT, Swanson S, Graham LE, Sharkey TD (1999) Evolutionary significance of isoprene emission from mosses. Am J Bot 86:634–639
Ikehara K (2002) Origins of gene, genetic code, protein and life: comprehensive view of life systems from a GNC-SNS primitive genetic code hypothesis. J Biosci 27:165–186
Jabbari K, Cruveiller S, Clay O, Bernardi G (2003) The correlation between GC3 and hydropathy in human genes. Gene 317:137–140
Jaenicke R, Böhm G (1998) The stability of proteins in extreme environments. Curr Opin Struct Biol 8:738–748
Jobson RW, Qiu YL (2008) Did RNA editing in plant organellar genomes originate under natural selection or through genetic drift? Biol Direct 3:43
Jobson RW, Dehne-Garcia A, Galtier N (2010) Apparent longevity-related adaptation of mitochondrial amino acid content is due to nucleotide compositional shifts. Mitochondrion 10:540–547
Kannan N, Vishveshwara S (2000) Aromatic clusters: a determinant of thermal stability of thermophilic proteins. Prot Eng 13:753–761
Karol KG, McCourt RM, Cimino MT, Delwiche CF (2001) The closest living relatives of land plants. Science 294:2351–2353
Kenrick P, Crane PR (1997) The origin and early evolution of plants on land. Nature 389:33–39
Khandelwal A, Cho SH, Marella H, Sakata Y, Perroud P-F, Pan A, Quatrano RS (2010) Role of ABA and ABI3 in desiccation tolerance. Science 327:546
Killian JA, Heijne G (2000) How proteins adapt to a membrane-water interface. Trends Biochem Sci 25:429–434
King JL, Jukes TH (1969) Non-Darwinian evolution. Science 164:788–798
Klotz IM (1958) Protein hydration and behavior. Science 128:815–822
Knight CG, Kassen R, Hebestreit H, Rainey PB (2004) Global analysis of predicted proteomes: functional adaptation of physical properties. Proc Natl Acad Sci USA 101:8390–8395
Kovács E, Keresztes A (2002) Effect of gamma and UV-B/C radiation on plant cells. Micron 33:199–203
Kramer PJ, Boyer JS (1995) Water relations of plants and soils. Academic Press, San Diego
Kugita M, Yamamoto Y, Fujikawa T, Matsumoto T, Yoshinaga K (2003) RNA editing in hornwort chloroplasts makes more than half the genes functional. Nucleic Acids Res 31:2417–2423
Kumar S, Nussinov R (2001) How do thermophilic proteins deal with heat? Cell Mol Life Sci 58:1216–1233
Lemieux C, Otis C, Turmel M (2007) A clade uniting the green algae Mesostigma viride and Chlorokybus atmophyticus represents the deepest branch of the Streptophyta in chloroplast genome-based phylogenies. BMC Biol 5:2
Lewis LA, McCourt RM (2004) Green algae and the origin of land plants. Am J Bot 91:1535–1556
Li L, Wang B, Liu Y, Qiu Y-L (2009) The complete mitochondrial genome sequence of the hornwort Megaceros aenigmaticus shows a mixed mode of conservative yet dynamic evolution in early land plant mitochondrial genomes. J Mol Evol 68:665–678
Lobry JR, Chessel D (2003) Internal correspondence analysis of codon and amino-acid usage in thermophilic bacteria. J Appl Genet 44:235–261
McDonald JH (2001) Patterns of temperature adaptation in proteins from bacteria Deinococcus radiodurans and Thermus thermophilus. Mol Biol Evol 18:741–749
Mrabet NT, Van den Broeck A, Van den brande I, Stanssens P, Laroche Y, Lambeir AM, Matthijssens G, Jenkins J, Chiadmi M, van Tilbeurgh H et al (1992) Arginine residues as stabilizing elements in proteins. Biochemistry 31:2239–2253
Musto H, Naya H, Zavala A, Romero H, Alvarez-Valín F, Bernardi G (2004) Correlations between genomic GC levels and optimal growth temperatures in prokaryotes. FEBS Lett 573:73–77
Naya H, Romero H, Zavala A, Alvarez B, Musto H (2002) Aerobiosis increases the genomic guanine plus cytosine content (GC%) in prokaryotes. J Mol Evol 55:260–264
Nishio Y, Nakamura Y, Kawarabayasi Y, Usuda Y, Kimura E, Sugimoto S, Matsui K, Yamagishi A, Kikuchi H, Ikeo K, Gojobori T (2003) Comparative complete genome sequence analysis of the amino acid replacements responsible for the thermostability of Corynebacterium efficiens. Genome Res 13:1572–1579
Oliver MJ, Velten J, Mishler BD (2005) Desiccation tolerance in bryophytes: a reflection of the primitive strategy for plant survival in dehydrating habitats? Integr Comp Biol 45:788–799
Popadin K, Polishchuk LV, Mamirova L, Knorre D, Gunbin K (2007) Accumulation of slightly deleterious mutations in mitochondrial protein-coding genes of large versus small mammals. Proc Natl Acad Sci USA 104:13390–13395
Proctor MCF, Pence VC (2002) In: Black M, Pritchard HW (eds) Desiccation and survival in plants: drying without dying. CABI, Wallingford, pp 207–237
Proctor MC, Smirnoff N (2000) Rapid recovery of photosystems on rewetting desiccation-tolerant mosses: chlorophyll fluorescence and inhibitor experiments. J Exp Bot 51:1695–1704
Proctor MC, Ligrone R, Duckett JG (2007) Desiccation tolerance in the moss Polytrichum formosum: physiological and fine-structural changes during desiccation and recovery. Ann Bot 99:75–93
Qiu Y-L (2008) Phylogeny and evolution of charophytic algae and land plants. J Syst Evol 46:287–306
Qiu Y-L, Palmer JD (1999) Phylogeny of early land plants: insights from genes and genomes. Trends Plant Sci 4:26–30
Qiu Y-L, Li L, Wang B, Chen Z, Knoop V, Groth-Malonek M, Dombrovska O, Lee J, Kent L, Rest J, Estabrook GF, Hendry TA, Taylor DW, Testa CM, Ambros M, Crandall-Stotler B, Duff RJ, Stech M, Frey W, Quandt D, Davis CC (2006) The deepest divergences in land plants inferred from phylogenomic evidence. Proc Natl Acad Sci USA 103:15511–15516
Qiu Y-L, Li L, Wang B, Chen Z, Dombrovska O, Lee J, Kent L, Li R, Jobson RW, Hendry TA, Taylor DW, Testa CM, Ambros M (2007) A non-flowering land plant phylogeny inferred from nucleotide sequences of seven chloroplast, mitochondrial and nuclear genes. Int J Plant Sci 168:691–708
R Development Core Team (2009) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, Shapiro H, Nishiyama T, Perroud PF, Lindquist EA, Kamisugi Y, Tanahashi T, Sakakibara K, Fujita T, Oishi K, Shin-I T, Kuroki Y, Toyoda A, Suzuki Y, Hashimoto S, Yamaguchi K, Sugano S, Kohara Y, Fujiyama A, Anterola A, Aoki S, Ashton N, Barbazuk WB, Barker E, Bennetzen JL, Blankenship R, Cho SH, Dutcher SK, Estelle M, Fawcett JA, Gundlach H, Hanada K, Heyl A, Hicks KA, Hughes J, Lohr M, Mayer K, Melkozernov A, Murata T, Nelson DR, Pils B, Prigge M, Reiss B, Renner T, Rombauts S, Rushton PJ, Sanderfoot A, Schween G, Shiu SH, Stueber K, Theodoulou FL, Tu H, Van de Peer Y, Verrier PJ, Waters E, Wood A, Yang L, Cove D, Cuming AC, Hasebe M, Lucas S, Mishler BD, Reski R, Grigoriev IV, Quatrano RS, Boore JL (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319:64–69
Richardt S, Timmerhaus G, Lang D, Qudeimat E, Corrêa LG, Reski R, Rensing SA, Frank W (2010) Microarray analysis of the moss Physcomitrella patens reveals evolutionarily conserved transcriptional regulation of salt stress and abscisic acid signalling. Plant Mol Biol 72:27–45
Rodrigues GC, Jansen MAK, van den Noort ME, van Rensen JJS (2006) Evidence for the semireduced primary quinone electron acceptor of photosystem II being a photosensitizer for UVB damage to the photosynthetic apparatus. Plant Sci 170:283–290
Romero H, Pereira E, Naya H, Musto H (2009) Oxygen and guanine-cytosine profiles in marine environments. J Mol Evol 69:203–206
Saelensminde G, Halskau Ø Jr, Jonassen I (2009) Amino acid contacts in proteins adapted to different temperatures: hydrophobic interactions and surface charges play a key role. Extremophiles 3:11–20
Sanderson MJ (2004) Molecular evidence on plant divergence times. Am J Bot 91:1656–1665
Sharkey TD, Wiberley AE, Donohue AR (2008) Isoprene emission from plants: why and how. Ann Bot 101:5–18
Singer GA, Hickey DA (2003) Thermophilic prokaryotes have characteristic patterns of codon usage, amino acid composition and nucleotide content. Gene 317:39–47
Smith JC, Merzel F, Bondar AN, Tournier A, Fischer S (2004) Structure, dynamics and reactions of protein hydration water. Philos Trans R Soc Lond B Biol Sci 359:181–189
Song CP, Agarwal M, Ohta M, Guo Y, Halfter U, Wang P, Zhu JK (2005) Role of an Arabidopsis AP2/EREBP-type transcriptional repressor in abscisic acid and drought stress responses. Plant Cell 17:2384–2396
Strub C, Alies C, Lougarre A, Ladurantie C, Czaplicki J, Fournier D (2004) Mutation of exposed hydrophobic amino acids to arginine to increase protein stability. BMC Biochem 5:9
Tekaia F, Yeramian E (2006) Evolution of proteomes: fundamental signatures and global trends in amino acid compositions. BMC Genomics 7:307
Turmel M, Otis C, Lemieux C (2006) The chloroplast genome sequence of Chara vulgaris sheds new light into the closest green algal relatives of land plants. Mol Biol Evol 23:1324–1338
Tyrrell RM (1994) The molecular and cellular pathology of solar ultraviolet radiation. Mol Asp Med 15:1–77
Wang HC, Singer GA, Hickey DA (2004) Mutational bias affects protein evolution in flowering plants. Mol Biol Evol 21:90–96
Wang B, Xue J-Y, Li L, Liu Y, Qiu Y-L (2009a) The complete mitochondrial genome sequence of the liverwort Pleurozia purpurea reveals extremely conservative mitochondrial genome evolution in liverworts. Curr Genet 55:601–609
Wang XQ, Yang PF, Liu Z, Liu WZ, Hu Y, Chen H, Kuang TY, Pei ZM, Shen SH, He YK (2009b) Exploring the mechanism of Physcomitrella patens desiccation tolerance through a proteomic strategy. Plant Physiol 149:1739–1750
Waters ER (2003) Molecular adaptation and the origin of land plants. Mol Phylogenet Evol 29:456–463
Wolf PG, Rowe CA, Hasebe M (2004) High levels of RNA editing in a vascular plant chloroplast genome: analysis of transcripts from the fern Adiantum capillus-veneris. Gene 339:89–97
Wolf L, Rizzini L, Stracke R, Ulm R, Rensing SA (2010) The molecular and physiological responses of Physcomitrella patens to ultraviolet-B radiation. Plant Physiol 153:1123–1134
Xia X, Li W-H (1998) What amino acid properties affect protein evolution? J Mol Evol 47:557–564
Xue J-Y, Liu Y, Li L, Wang B, Qiu YL (2010) The complete mitochondrial genome sequence of the hornwort Phaeoceros laevis: retention of many ancient pseudogenes and conservative evolution of mitochondrial genomes in hornworts. Curr Genet 56:53–61
Acknowledgments
We thank Jiayu Xue and Yang Liu for helpful contributions. This study was partly supported by NSF grant DEB 0531689 to Y.-L. Qiu.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jobson, R.W., Qiu, YL. Amino Acid Compositional Shifts During Streptophyte Transitions to Terrestrial Habitats. J Mol Evol 72, 204–214 (2011). https://doi.org/10.1007/s00239-010-9416-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-010-9416-1