Abstract
We report the cloning and sequence analysis of Echis ocellatus cDNAs coding for dimeric disintegrin subunits and for the short disintegrin ocellatusin. All the dimeric disintegrin subunit messengers belong to the short-coding class, indicating that short messengers may be more widely distributed than previously thought. Mass spectrometric analysis of the HPLC-separated venom proteins was performed to characterize the dimeric disintegrins expressed in the venom proteome. In addition to previously reported EO4 and EO5 heterodimers, a novel dimeric disintegrin containing RGD- and KGD-bearing subunits was identified. However, a WGD-containing polypeptide encoded by clone Eo1-1 was not detected in the venom, suggesting the occurrence of larger genomic than proteomic diversity, which could represent part of a non-venom-secreted reservoir of disintegrin that may eventually acquire physiological relevance for the snake upon changes of ecological niches and prey habits. On the other hand, the realization of the existence of two distinct messengers coding for the short disintegrin ocellatusin reveals key events of the evolutionary emergence of the short disintegrin ocellatusin from a short-coding dimeric disintegrin precursor by two nucleotide mutations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Approximately 130 million years ago snakes diverged from lizards and since then they developed their own venom apparatus (Kochva 1987; Fry and Wüster 2004; Fry et al. 2005). Snake venoms are composed of proteins belonging to only a few major families, including enzymes (serine proteinases, Zn2+-metalloproteases, L-amino acid oxidases, phospolipases A2 [PLA2]) and proteins without enzymatic activity (disintegrins, C-type lectins, natriuretic [bradykinin-potentiating] peptides, cyteine-rich secretory protein [CRISP] toxins, nerve and endothelial growth factors, cystatin, and Kunitz-type protease inhibitors) (Juárez et al. 2004; Fry 2005). These toxins evolved from proteins with a normal physiological function and were recruited into the venom proteome before the diversification of the advanced snakes (Fry 2005) (superfamily Colubroidea), which make up over 80% of the about 2900 species of snake currently described (Vidal 2002), hence predating the evolution of the intricate front-fanged venom delivery mechanisms.
Venom proteins play a number of adaptative roles: immobilizing, paralyzing, killing, and digesting prey, and deterring competitors. Notably, most venom toxins are extensively cross-linked by disulfide bonds and have flourished into functionally diverse, toxin multigene families that exhibit interfamily, intergenus, interspecies, and intraspecific variability. The existence in the same venom of a functionally diverse isoform of the same protein family reflects an accelerated, by gene duplication, Darwinian evolution (Moura da Silva et al. 1996; Menez 2002; Tani et al. 2002). Rapid co-evolution between snakes and their prey in driving the evolution of venom proteins has been discussed (Daltry et al. 1996). Studies on the underlying mechanisms of this protein structural/functional variation are of considerable theoretical interest in the field of molecular evolution, in the development of new research tools and drugs of potential clinical use, and for antivenom production strategies.
Viper venom disintegrins are a family of small (40–100 amino acids), cysteine-rich polypeptides that selectively block the function of integrin receptors (Calvete et al. 2005; Calvete 2005) and result from proteolytic processing of larger mosaic PII and PIII metalloprotease precursors (Kini and Evans 1992) or are synthesized from short-coding mRNAs (Okuda et al. 2002). Currently, the disintegrin family can be conveniently divided into five groups according to their length and number of disulfide bonds (Calvete et al. 2003). The first group includes short disintegrins, composed of 41–51 residues and 4 disulfide bonds. The second group is formed by the medium-sized disintegrins, containing about 70 amino acids and 6 disulfide bonds. The third group includes long disintegrins, with an ∼84-residue polypeptide cross-linked by 7 disulfide bonds. The fourth subfamily, the PIII disintegrins, is modular proteins containing an N-terminal disintegrin-like domain of about 100 amino acids including 8 disulfide bonds and a C-terminal 110- to 120-residue cysteine-rich domain cross-linked by 6 disulfides (Calvete et al. 2002a). Unlike the PII (short, medium, and long) and PIII disintegrins, which are single-chain molecules, the fifth group is composed of homo- and heterodimers. Dimeric disintegrins contain subunits of about 67 residues with 10 cysteines involved in the formation of 4 intrachain disulfide bonds and 2 interchain cystine linkages (Calvete et al. 2000b; Bilgrami et al. 2004, 2005). Bilitoxin-1 represents a unique homodimeric disintegrin comprising disulfide-bonded polypeptides, each containing 15 cysteinyl residues (Nikai et al. 2000). The current view is that the structural diversity of disintegrins has been achieved during ophidian evolution through the selective loss of disulfide bonds (Calvete et al. 2003).
To understand the genomic basis of the accelerated evolution of disintegrins, and the molecular mechanism underlying their structural diversification, we have started the genomic analysis of cDNAs encoding disintegrins from a venom gland library of Echis ocellatus.
Materials and Methods
cDNA Library Synthesis and Sequencing
Total RNA was extracted (using TriReagent, Life Technologies, UK) from pooled venom glands of 10 wild-caught specimens of E. ocellatus (Kaltungo, Nigeria), of different ages and of both sexes, and maintained at the herpetarium of the Liverpool School of Tropical Medicine, 3 days after venom extraction when toxin gene transcription rates are at a peak (Paine et al. 1992). Messenger RNA was purified using a single-step oligo(DT) cellulose purification (Amersham, UK) according to the manufacturer’s protocols. A Gateway CloneMiner cDNA library construction kit (Invitrogen, UK) was used to construct a directional plasmid cDNA library according to the manufacturer’s protocol. Fractions corresponding to the first 480 ng of eluted cDNA were pooled and used for recombination and transformation into the final library. Plasmid DNA from 1000 randomly selected colonies was sequenced (Lark technologies, UK) using M13 forward primers. To identify potential toxins, Blastn or Blastx was performed on sequenced clones against databases of UniProt, translations from EMBL, and all Serpentes nucleotide sequence or protein sequences including ESTs.
cDNA Cloning and Sequencing
A forward primer, 5′-CCAAATCCAGC/TCTCCAAAATG-3′, and a reverse primer, 5′-TTCCAG/TCTCCATTGTTGG/TTTA, complementary to the highly conserved 5′- and 3′-noncoding regions of cDNA encoding for elegantin-2a from Trimeresurus elegans (GenBank accession number AB059572), elegantin-1a from T. elegans (GenBank accession number AB059571), and HR2a from Trimeresurus flavoviridis (accession code AY037808) were synthesized. The specific sense primer 5′-GAAAAGGAAGACG ACTGTGAATC-3′ was synthesized based on the middle portion of the clone obtained from our E. ocellatus EST data. To obtain the full-length cDNAs encoding these precursors, we designed an antisense primer based on the poly(A) signal: 5′-TTTTT TTTTTTTTTTTTTTTTTA/C/G-3′. DNAs were amplified by PCR using total RT products and cDNA as templates. AmpliTaq Gold (Roche), a highly processive 5′–3′ DNA polymerase that lacks 3′–5′ exonuclease activity, was used. The PCR protocol included an initial denaturation step at 95°C for 6 min followed by 35 cycles of denaturation (1 min at 94°C), annealing (1 min at 55°C), and extension (1 min at 74°C) and a final extension for 7 min at 72°C. The amplified fragments were purified using the Wizard SV Gel and PCR Clean-Up System (Promega), ligated into a TA cloning vector (pCR-2.1-TOPO; Invitrogen, Groning, Netherlands), and used to transform chemically competent Escherichia coli (TOP 10F; Invitrogen). Positive clones, selected by growing the transformed cells in LB medium containing 10 μg/ml ampicillin, were confirmed by PCR amplification using the above primers.
Isolation and Characterization of Venom Proteins
Pooled venom extracted from 150 wild-caught Echis ocellatus (Kaltungo, Nigeria) of different ages and of both sexes (maintained at the herpetarium of the Liverpool School of Tropical Medicine) was lyophilized and stored at 4°C in a dark bottle. For reverse-phase HPLC separation, 2 mg of the venom was dissolved in 100 μl of 5% acetonitrile and 0.1% trifluoroacetic acid (TFA). Insoluble material was removed by centrifugation in an Eppendorf centrifuge (Hamburg, Germany) at 13,000g for 10 min at room temperature. Soluble proteins were separated with an ETTAN LC HPLC system (Amersham Biosciences) using a Lichrospher RP100 C18 column (250 × 4 mm, 5-mm particle size; Merck, Darmstadt, Germany) eluted at 1 ml/min with a linear gradient of 0.1% TFA in water (solution A) and in acetonitrile (solution B), first isocratically (5% B) for 5 min, followed by linear gradients of 5–45% B for 120 min and 45–70% B for 20 min. Protein detection was at 215 nm and peaks were collected manually. The isolated protein fractions were analyzed by SDS-PAGE, N-terminal sequencing (using an Applied Biosystem´s Procise 492 sequencer), and matrix-assisted laser-desorption ionization time-of-flight mass spectrometry (MALDI-TOF-MS) using a Voyager-DE Pro instrument (Applied Biosystems) and sinapinic acid (3,5–dimethoxy-4-hydroxycinnamic acid; Sigma) saturated in 70% acetonitrile and 0.1% TFA as matrix, with and without prior reduction with DTT (10 mM for 15 min at 65°C) and pyridylethylation with 4-vinylpyridine (50 mM for 60 min at room temperature). The mass calibration standard consisted of a mixture of the following proteins, whose isotope-averaged molecular mass in daltons is given in parentheses: bovine insulin (5734.5), E. coli thioredoxin (11,674.5), and horse apomyoglobin (16,952.6). Molecular masses were also determined by electrospray ionization mass spectrometry with a triple quadrupole-ion trap hybrid instrument (QTrap from Applied Biosystems) equipped with a nanospray source (Protana, Denmark).
Phylogenetic Analysis
The program MEGA (Molecular Evolutionary Genetic Analysis; http://www.megasoftware.net (Kumar et al. 2001)) was employed for inferring phylogenies (evolutionary trees) from a multiple alignment of disintegrin sequences.
Database Accession Codes
The cDNA sequences have been deposited with the EMBL Nucleotide Sequence Data Bank (http://www.ebi.ac.uk/ ) and have the following accession codes: AM117387 (Dim-3 SP6), AM117388 (Eo-10), AM117389 (Eo-12), AM117390 (Eo-10c1), AM117391 (Eo1-1), and AM117392 (Eo10-c10).
Results and Discussion
Full-length disintegrin messengers were cloned from a cDNA library constructed from Echis ocellatus venom gland polyadenylated RNA using a forward primer for the highly conserved 5′ and either a reverse primer for the 3′ noncoding region of known disintegrins or an antisense poly(dT) primer based on the poly(A) signal. The deduced amino acid sequences of the seven cDNAs are displayed in Figs. 1 and 2.
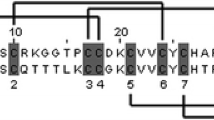
DNA and deduced amino acid sequences of Echis ocellatus dimeric disintegrin subunits. The nucleotide sequences are numbered in the 5′–3′ direction from the initial codon ATG to the stop codon TGA or TAA (asterisks). The signal sequence and the short- ened prepeptide regions are underlined and in italics and double underlined, respectively. The predicted mature protein sequences are in boldface. The integrin-binding tripeptide sequences are shaded. A, clone Dim-3 SP6; B, clone Eo-10; C, clone Eo-12; D, clone Eo-10c1; and E, clone Eo1-1.
cDNA and deduced amino acid sequence of ocellatusin long (clone Eo-00006; A) and short (clone Eo10c-10; B) precursors. The nucleotide sequences are numbered in the 5′–3′ direction, from the initial codon ATG to the stop codon TGA (asterisk). The N-terminal regions of the signal sequence, the pro-domain, the metalloproteinase, and the disintegrin domains are labeled. The signal sequences are underlined. The positions of the Cys-switch site (KMCGV) within the pro-domain, the Zn+2-binding motif (HEMGHNLGMNH) of the metalloprotease domain, and the RGD sequence of the disintegrin domain are shaded. The shortened propeptide region of the short-coding ocellatusin precursor and its homologous region in the long precursor are in italics and double underlined. The short-disintegrin ocellatusin is indicated in boldface. The tyrosine residue of Eo-00006 and Eo-10c1, which in dimeric disintegrin subunits corresponds to a conserved cysteine involved in an intersubunit disulfide, is in boldface, underlined, and boxed. The short disintegrin-specific cysteine residues at positions 468 and 102 of the long and the short precursors, respectively, are double underlined.
cDNA Sequences of Dimeric Disintegrin Subunits
The deduced amino acid sequences of five E. ocellatus cDNAs (Fig. 1) show extensive sequence similarity to known sequences of subunits of dimeric disintegrins, including the number and position of the 10 cysteine residues (Calvete et al. 2003) (Fig. 3). Clones Dim-3 SP6 and Eo-10 (Figs. 1A and 1B, respectively) encode isoforms differing in their primary structures only at residue 54 (Fig. 3). These RGD-containing disintegrin chains are identical to EO4A (Calvete et al. 2003) except for (i) position 48, which is asparagine in EO4A and histidine in the E. ocellatus disintegrin cDNA, and (ii) the last four C-terminal residues, which could not be unambiguously determined by protein chemical methods and are shown here to correspond to GKYD. Except for a K/Q substitution at position 14, clone Eo-12 (Fig. 1C) corresponds to the previously reported MLD-containing A-subunit of dimeric disintegrin EO5 in E. ocellatus venom (Calvete et al. 2003). The deduced sequences of clones Eo-10c1 and Eo1-1 represent novel disintegrin chains. Eo-10c1 (Fig. 1D) is another isoform of Dim-3 SP6 and Eo-10 expresses a KGD integrin-inhibitory tripeptide instead of RGD.
A Multiple amino acid sequence alignment of the Echis ocellatus cDNA-deduced disintegrin sequences with those of dimeric disintegrin subunits. B Alignment of the cDNA-deduced amino acid sequences of the long (Eo-00006) and short (Eo10c-10) ocellatusin precursors with that of the protein isolated from the venom of Echis ocellatus (Smith et al. 2002). Cysteine residues are shaded in pale gray. The integrin-binding tripeptides are shown in boldface. The tyrosine residue of Eo-00006 and Eo-10c1, which in dimeric disintegrin subunits corresponds to a conserved cysteine involved in an intersubunit disulfide, is highlighted in boldface, double underlined, and marked with an asterisk. The short disintegrin-specific cysteine residue is labeled by a filled diamond.When available, the Swiss-Prot/TrEMBL ( http://www.us.expasy.org/sprot/ ) databank accession codes are given in brackets.
The WGD-containing polypeptide encoded by cDNA Eo1-1 has not been reported in Echis ocellatus venom (Calvete et al. 2003) and may therefore correspond to a non-venom-secreted or to a novel disintegrin messenger. To check these possibilities, we reexamined the disintegrin content of the pooled venoms used for cDNA library synthesis and sequencing: venom proteins were separated by reverse-phase HPLC (Fig. 4) and each chromatographic fraction was analyzed by MALDI-TOF mass spectrometry. Dimeric disintegrin candidates were protein fractions of 13- to 15-kDa molecular mass, which generated 7- to 8-kDa subunits following reduction and pyridylethylation. The only Echis ocellatus venom proteins fulfilling these criteria were the previously reported EO4 and EO5 heterodimers (Calvete et al. 2003) (Fig. 4).
Reverse-phase HPLC separation of Echis ocellatus venom proteome. Protein fractions whose identities were characterized by combination of N-terminal sequencing and mass spectrometry are labeled. Peaks containing the short disintegrin ocellatusin and the dimeric disintegrins EO4 and EO5 are highlighted in boldface. DC-fragments, disintegrin/cysteine-rich domains of PIII metalloproteases; PLA2, phospholipase A2; SVMPs, snake venom metalloproteases; LAO, L-amino acid oxidase.
The molecular masses of these dimeric disintegrins were determined by nanoelectrospray ionization mass spectrometry. The mass spectrum of EO4 displayed a major (14,488 ± 2 Da) and two minor (14,583 ± 2 Da and 14,684 ± 2 Da) ions, and that of Eo5 showed a single ion of 14,520 ± 1 Da. MALDI-TOF mass analysis of the reduced and pyridylethylated (PE) proteins showed that the 14,488-Da EO4 species was composed of subunits of molecular masses of 8418 and 8196 Da. The 8196-Da subunit corresponds to the calculated molecular mass of PE-EO4B (=PE-EO5B), and the 8418-Da polypeptide has the isotope-averaged mass calculated for the cDNA-deduced PE-amino acid sequences of clone Dim-3 SP6 (8417.7 Da; =PE-EO4A). On the other hand, minor PE-EO4 ions at m/z 8293 and 8390 indicated that the 14,583-Da and the 14,684-Da EO4 species correspond to (EO4A)-S-S-(EO4B + C-terminal proline) (Mcalc: 14,583.2 Da) and to (EO4A)-S-S-(Eo-10c1) (Mcalc: 14,684.6 Da), respectively. The latter protein, which we call EO4-KGD to distinguish it from the previously characterized EO4-VGD species, represents a novel dimeric disintegrin. Its integrin-binding motif (KGD) has been reported to inhibit the platelet αIIbβ3 integrin with a high degree of selectivity (Scarborough et al. 2001).
The isotope-averaged masses of the PE-subunits of the 14,520-Da dimeric disintegrin, 8452 and 8196 Da, confirmed the identity of this protein as EO5 (see Table 2 in Ref. 14). This protein is constituted by an 8196-Da VGD-containing B-subunit (=EO4-VGD B-subunit) and an MLD-containing A-subunit coded for by clone Eo-12 (Fig. 1C).
It is worthwhile to note that E. ocellatus venom glands appear to synthesize non-venom-expressed (or very low-abundance) transcripts (Eo-10, Eo1-1) (Fig. 1) encoding putative dimeric disintegrin subunits. The WGD integrin-binding motif of Eo1-1 has been reported in studies on other viper disintegrins to enhance the inhibitory effect on the RGD-dependent integrins αIIbβ3, αVβ3, and α5β1 (Calvete et al. 2002). The reason for the apparently larger transcriptomic than proteomic diversity remains unclear, although it suggests an economy of protein production. Alternatively, these messengers may exhibit an individual or a temporal expression pattern over the lifetime of the snake. We hypothesize that the non-venom-secreted or very low-abundance (<0.05% of the total venom proteins) disintegrins may play a hitherto unrecognized physiological function or may simply represent orphan molecules which may eventually become functional for the adaptation of snakes to changing ecological niches and prey habits. Alternatively, as pointed out in the accompanying article (Juárez et al. 2006) on the Bitis arietans BA-5A disintegrin, it is also possible that the undetected Echis ocellatus disintegrins may be translated but not secreted into the venom. Clearly, further work is needed to clarify this point.
Our results also show that short messengers coding for dimeric disintegrin subunits may be more widely distributed than previously thought, perhaps representing the canonical structure of dimeric disintegrin precursors. In support of this view, the two dimeric disintegrin messengers found in the transcriptome of Bitis gabonica, gabonin-1 and gabonin-2 (Francischetti et al. 2004), and all the full-length disintegrin cDNAs characterized from venom gland cDNA libraries of Macrovipera lebetina and Cerastes vipera (Sanz et al. 2006), all belong to the short-coding region class. Our working hypothesis is that removal of the metalloprotease domain may represent a step in the evolutionary pathway yielding dimeric disintegrins from a PII-metalloprotease-coding precursor (Fig. 5).
Illustration depicting the proposed common ancestry of the messenger precursors coding for the short disintegrin ocellatusin and dimeric disintegrins EO4 and EO5. The proposed evolutionary pathway of EO4 and EO5 includes the removal of the metalloproteinase domain from a PII metalloproteinase precursor gene. A key event in the emergence of ocellatusin appears to be the substitution of the second N-terminal cysteine residue (Cys7 in the dimeric disintegrin subunit precursor) by tyrosine, thereby impairing dimerization through homologous CysA7-CysB12 and CysA12-CysB7 linkages, along with the appearance of a novel cysteine residue at position 102 (short-coding precursor numbering) between the 9th and the 10th cysteine of the precursor (C), enabling the short disintegrin-specific disulfide bond depicted by the dashed line, and the proteolytic processing of the N- and the C-terminal regions (scissors). The two conserved disulfide bonds in the structures of dimeric disintegrin subunits and the short disintegrins are represented by thick lines.
Two Distinct Messengers Coding for the Short Disintegrin Ocellatusin
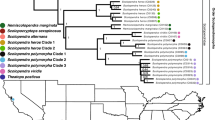
Figure 2 shows the DNA and deduced amino acid sequences of a long and a short full-length E. ocellatus cDNA, both encoding for the short disintegrin ocellatusin (Smith et al. 2002) (DCESGP...GEHDP, isotope-averaged M+H+ calc, 5594.3; M+H+ measured, 5594), the major secreted disintegrin in E. ocellatus venom (Fig. 4). The amino acid sequences located N-terminal of mature ocellatusin within the long and the short precursors, ELLQNSVNPCYDPVTCQPKEKE and NSAHPCYDPVTCQPKEKE, respectively (Figs. 2 and 3B), exhibit large similarity between themselves and with the N-terminal regions of the subunits of dimeric disintegrins (Fig. 3A). This fact strongly argues for a common ancestry of both ocellatusin messengers and the precursors of dimeric disintegrin chains. This hypothesis is supported by a phylogenetic analysis of the dimeric and the short disintegrins from Echis species (Fig. 6A), which shows that the two ocellatusin precursors, EO10c-10 and EO-00006, cluster into an intermediate group between the venom-secreted dimeric disintegrin subunits and the short disintegrins. Phylogeny reconstruction of the known disintegrins of Echis ocellatus using a maximum parsimony method (Fig. 6B) also supports the emergence of ocellatusin from EO10c-10 and EO-00006 transcripts.
A Cladogram for the multiple sequence analysis of the dimeric disintegrin subunits and the short disintegrins from Echis snake species. The ocellatusin precursors, EO10c-10 and EO-00006, are denoted by filled circles. The tree represents the minimum evolutionary distance estimated through neighbor joining using maximum likelihood distances. The length of the horizontal scale bar represents 20% divergence. For primary references of the analyzed disintegrins, consult Calvete et al. (2003). Maximum parsimony analysis produced a similar topology (B). C Scheme of the domain organization, disulfide bond patterns, and proposed evolutionary pathway from the PIII disintegrin/cysteine-rich proteins to short disintegrins. Structural features (the cysteine-rich domain of PIII disintegrin-like molecules, and class-specific disulfides) lost along the disintegrin diversification pathway are highlighted with thick lines. The proposed mechanism for the emergence of the short disintegrin ocellatusin from the long (EO-00006) and the short (EO10c-10) precursors is shown schematically in Fig. 5.
Snake venom PIII disintegrins evolved from the extracellular domains of cell membrane-anchored ADAM (a disintegrin and metalloprotease) molecules after mammals and reptiles diverged (Moura da Silva et al. 1996). Phylogenetic analysis, in conjunction with biochemical and genetic data, supports the model depicted in Fig. 6C, by which the structural diversification of the disintegrin family occurred through the successive loss of disulfide bonds (Calvete et al. 2003).
The most noticeable difference between ocellatusin and the dimeric disintegrin precursors is the presence at position 54 of the ocellatusin short-coding precursor (position 420 of the long precursor) of a tyrosine residue, which in all dimeric disintegrin subunits harbors a conserved cysteine residue (Figs. 2 and 3) involved in dimer formation (Calvete et al. 2000b; Bilgrami et al. 2004, 2005). In crystallographic studies, this Cys7 residue of each subunit of two E. carinatus sochureki dimeric disintegrins forms the two interchain cystine linkages (Cys7A-Cys12B and Cys12A-Cys7B) with cysteine-12 from the other subunit of the dimers (Bilgrami et al. 2004, 2005) (Fig. 5). Theoretically, the single amino acid substitution Cys7→Tyr, generated through a single nucleotide mutation, TGT→TAT or TGC→TAC, is the simplest molecular solution to hinder dimerization. Though experimental validation is needed, we hypothesize that the Cys7→Xaa mutation may also represent a key step along the evolutionary pathway of short disintegrins from dimeric disintegrin precursors. Substitution, by cysteine, of the residue located two positions N-terminal of the last cysteine residue of the disintegrin domain (residues 468 and 102 of the long and the short precursors, respectively) (Fig. 2) and subsequent proteolytic processing of the N- and the C-terminal regions are further molecular events needed for generating mature short disintegrins (Fig. 5). In this sense, it is noteworthy that in all cDNAs coding for short dimeric disintegrin messengers, residue 102 is either a serine invariably coded for by a TCT codon, including acostatin α from Agkistrodon contortrix contortrix (Okuda et al. 2002), clones Dim-3 SP6, Eo-10, Eo-12, Eo-10c1, and Eo1-1 from Echis ocellatus (Fig. 1), clones CV3 and CV11 from Cerastes vipera, and ML2/8/15 and ML3 from Macrovipera lebetina (Sanz et al. 2006), or a threonine (ACT codon) in gabonin-1 and gabonin-2 from Bitis gabonica (Francischetti et al. 2004). Substitution of serine for cysteine can be accomplished by a single C→G mutation, whereas changing threonine for cysteine needs a minimum of two mutations (i.e., ACT → TCT → TGT). This may in part explain the observation that short disintegrins are commonly found in the venoms of species from genera like Echis, which also express dimeric disintegrins containing the sequence “SXDC” but have not been reported in the venom gland transcriptome of Bitis gabonica (dimeric disintegrins with “TPDC” sequence).
Concluding Remarks
The specific goal of this article was to investigate the genomic basis of the accelerated evolution of disintegrins and the molecular mechanism underlying their structural diversification. To this end, we sought to analyze cDNAs encoding disintegrins from a venom gland library of Echis ocellatus. The combined transcriptomic and proteomic analysis of E. ocellatus disintegrins revealed a non-venom-secreted disintegrin, suggesting that the accepted view of an accelerated evolution within the disintegrin family might not be mirrored in the venom proteome. A “hidden reservoir” of non-venom-secreted toxins could represent a pool of orphan molecules which may eventually become of relevance, i.e., for the adaptation of snakes in response to a changing ecological niche and prey habits. The identification of two distinct messengers coding for the short disintegrin ocellatusin sheds light on key events of the evolutionary pathway leading to short-coding messengers for dimeric disintegrin subunits from a PII-metalloproteinase precursor and to the short disintegrin ocellatusin from a short-coding dimeric disintegrin precursor by a minimum of two nucleotide mutations (Cys→Tyr and Ser→Cys). It is notable that the native fold of short disintegrins adopts a disulfide bond pattern slightly different from that of the dimeric disintegrin chains (Calvete et al. 2003; Moreno-Murciano et al. 2003; Monleón et al. 2005) (Fig. 5), providing additional possibilities for the evolution of the structure and function of this family of integrin antagonists. This point is illustrated in Fig. 7, in which the three-dimensional structures of schistatin (homodimer; PDB code 1RMR) (Bilgrami et al. 2004) and echistatin (short disintegrin; PDB code 1RO3) (Monleón et al. 2005), both from E. carinatus sochureki, have been superimposed. The root mean square deviation (r.m.s.d.) for the topologically equivalent Cα atoms is 4–5 Å, and the loop that contains the RGD motif and the C-terminal tail, which exhibit concerted dynamics, are oriented differently in the two structures. However, the overall scaffolding of schistatin has been reported to display an r.m.s.d. of 1.6 Å for the Cα atoms with medium-sized trimestatin (Bilgrami et al. 2004) reflecting, at the structural level, the proposed divergent evolution of disintegrins by disulfide bond engineering (Calvete et al. 2003).
Superposition of Cα traces of echistatin (red) and schistatin (magenta and cyan). N-t, N-terminus; C-t, C-terminus. Intrachain disulfide bonds are shown in one subunit of schistatin as dashed green lines, and the two interchain cystine linkages of the homodimer are depicted in the space-filling model in green. Disulfide bonds of echistatin are shown as dashed orange lines, and the RGD motif of the short disintegrin is labeled. The arrow indicates the position of the N-terminal proteolytic cleavage (E65-D66 of the short-coding precursor shown in Fig. 2B) depicted as a scissor in Fig. 5.
References
Bilgrami S, Tomar S, Yadav S, Kaur P, Kumar J, Jabeen T, Sharma S, Sinhg TP (2004) Crystal structure of schistatin, a disintegrin homodimer from saw-scaled viper (Echis carinatus) at 2.5 Å resolution. J Mol Biol 341:829–837
Bilgrami S, Yadav S, Sharma S, Perbandt M, Betzel C, Singh TP (2005) Crystal structure of the disintegrin heterodimer from saw-scaled viper (Echis carinatus) at 1.9 Å resolution. Biochemistry 44:11058–11066
Calvete JJ (2005) Structure-function correlations of snake venom disintegrins. Curr Pharm Des 11:829–835
Calvete JJ, Moreno-Murciano MP, Sanz L, Jürgens M, Schrader M, Raida M, Benjamin DC, Fox JW (2000a) The disulfide bond pattern of catrocollastatin C, a disintegrin/cysteine-rich protein isolated from Crotalus atrox venom. Protein Sci 9:1365–1373
Calvete JJ, Jürgens M, Marcinkiewicz C, Romero A, Schrader M, Niewiarowski S (2000b) Disulfide bond pattern and molecular modelling of the dimeric disintegrin EMF-10, a potent and selective integrin α5β1 antagonist from Eristocophis macmahoni venom. Biochem J 345:573–581
Calvete JJ, Fox JW, Agelan A, Niewiarowski S, Marcinkiewicz C (2002) The presence of the WGD motif in CC8 heterodimeric disintegrin increases its inhibitory effect on αIIbβ3, αVβ3, and α5β1 integrins. Biochemistry 41:2014–2021
Calvete JJ, Moreno-Murciano MP, Theakston RDG, Kisiel DG, Marcinkiewicz C (2003) Snake venom disintegrins:novel dimeric disintegrins and structural diversification by disulfide bond engineering. Biochem J 372:725–734
Calvete JJ, Marcinkiewicz C, Monleón D, Esteve V, Celda B, Juárez P, Sanz L (2005) Snake venom disintegrins: evolution of structure and function. Toxicon 45:1063–1074
Daltry JC, Wüster W, Thorpe RS (1996) Diet and snake venom evolution. Nature 379:537–540
Francischetti IMB, My-Pham V, Harrison J, Garfield MK, Ribeiro JMC (2004) Bitis gabonica (Gaboon viper) snake venom gland:toward a catalog for the full-length transcripts (cDNA) and proteins. Gene 337:55–69
Fry BG (2005) From genome to “venome”: molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res 15:403–420
Fry BG, Wüster W (2004) Assembling an arsenal: origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences. Mol Biol Evol 21:870–883
Fry BG, Vidal N, Norman JA, Vonk FJ, Scheib H, Ramjan SFR, Kuruppu S, Fung K, Hedges SB, Richardson MK, Hodgson WC, Ignjatovic V, Summerhayes R, Kochva E (2005) Early evolution of the venom system in lizards and snakes. Nature 439:584–588
Juárez P, Sanz L, Calvete JJ (2004) Snake venomics: characterization of protein families in Sistrurus barbouri venom by cysteine mapping, Nterminal sequencing, and tandem mass spectrometry analysis. Proteomics 4:327–338
Juárez P, Wagstaff SC, Oliver J, Sanz L, Harrison RA, Calvete JJ (2006) Molecular cloning of disintegrin-like transcript BA-5A from a Bitis arietans venom gland cDNA library: a putative intermediate in the evolution of the long-chain disintegrin bitistatin. J Mol Evol. Companion paper 10.1007/s00239-006-0268-z
Kini R, Evans HJ (1992) Structural domains in venom proteins: evidence that metalloproteinases and nonenzymatic platelet aggregation inhibitors (disintegrins) from snake venoms are derived by proteolysis from a common precursor. Toxicon 30:265–293
Kochva E (1987) The origin of snakes and evolution of the venom apparatus. Toxicon 25:65–106
Kumar S, Tamura K, Jakobsen IB, Nei M (2001) Mega2: Molecular Evolutionary Genetics Analysis software. Bioinformatics 17:1244–1245
Ménez A (2002) Perspectives in molecular toxinology. John Wiley & Sons, Chichester, UK
Monleón D, Esteve V, Kovacs H, Calvete JJ, Celda B (2005) Conformation and concerted dynamics of the integrin-binding site and the C-terminal region of echistatin revealed by homonuclear NMR. Biochem J 387:57–66
Moreno-Murciano MP, Monleón D, Marcinkiewicz C, Calvete JJ, Celda B (2003) NMR solution structure of the non-RGD disintegrin obtustatin. J Mol Biol 329:135–145
Moura-Da-Silva AM, Theakston RDG, Crampton JM (1996) Evolution of disintegrin cysteine-rich and mammalian matrix-degrading metalloproteinases:gene duplication and divergence of a common ancestor rather than convergent evolution. J Mol Evol 43:263–269
Nikai T, Taniguchi K, Komori Y, Masuda K, Fox JW, Sugihara H (2000) Primary structure and functional characterization of bilitoxin-1, a novel dimeric P-II snake venom metalloproteinase from Agkistrodon bilineatus venom. Arch Biochem Biophys 378:6–15
Okuda D, Koike H, Morita T (2002) A new gene structure of the disintegrin family: a subunit of dimeric disintegrin has a short coding region. Biochemistry 41:14248–14254
Paine MJ, Desmond HP, Theakston RD, Crampton JM (1992) Purification, cloning, and molecular characterization of a high molecular weight hemorrhagic metalloprotease, jararhagin, from Bothrops jararaca venom. Insights into the disintegrin gene family. J Biol Chem 267:22869–22876
Sanz L, Bazaa A, Marrakchi N, Pérez A, Chenik M, Bel Lasfer Z, El Ayeb M, Calvete JJ (2006) Molecular cloning of disintegrins from Cerastes vipera and Macrovipera lebetina transmediterranea venom gland cDNA libraries. Insight into the evolution of the snake venom’s integrin inhibition system. Biochem J 395:385–392
Smith JB, Theakston RDG, Coelho ALJ, Barja-Fidalgo C, Calvete JJ, Marcinkiewicz C (2002) Characterization of a monomeric disintegrin, ocellatusin, present in the venom of the Nigerian carpet viper, Echis ocellatus. FEBS Lett 512:111–115
Scarborough RM, Rose JW, Hsu MA, Phillips DR, Fried VA, Campbell AM, Nannizzi L, Charo IF (2001) Barbourin, a GPIIb-IIIa-specific integrin antagonist from the venom of Sistrurus m. barbouri. J Biol Chem 266:9359–9362
Tani A, Ogawa T, Nose T, Nikandrov NN, Deshimaru M, Chijiwa T, Chang CC, Fukumaki Y, Ohno M (2002) Characterization, primary structure and molecular evolution of anticoagulant protein from Agkistrodon actus venom. Toxicon 40:803–813
Vidal N (2002) Colubroid systematics: evidence for an early appearance of the venom apparatus followed by extensive evolutionary tinkering. J Toxicol Toxin Rev 21:21–41
Acknowledgments
This work was financed by Grant BFU2004-01432 from the Ministerio de Educación y Ciencia, Madrid, Spain (to J.J.C.). P.J. and L.S. are recipients of a predoctoral fellowship (FPI; formación de personal investigador) from the Spanish Ministerio de Educación y Ciencia and a postdoctoral I3P contract, respectively. R.A.H. and S.C.W. were funded by the Wellcome Trust.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Bryan Grieg Fry]
Rights and permissions
About this article
Cite this article
Juárez, P., Wagstaff, S.C., Sanz, L. et al. Molecular Cloning of Echis ocellatus Disintegrins Reveals Non-Venom-Secreted Proteins and a Pathway for the Evolution of Ocellatusin. J Mol Evol 63, 183–193 (2006). https://doi.org/10.1007/s00239-005-0269-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0269-y