Abstract
Inhibitor cystine knots (ICKs) are a family of structural peptides with a large number of cysteine residues that form intramolecular disulfide bonds, resulting in a knot. These peptides are involved in a variety of biological functions including predation and defense, and are found in various species, such as spiders, scorpions, sea anemones, and plants. The Loxosceles intermedia venom gland transcriptome identified five groups of ICK peptides that represent more than 50 % of toxin-coding transcripts. Here, we describe the molecular cloning of U2-Sicaritoxin-Lit2 (U2-SCRTX-Lit2), bioinformatic characterization, structure prediction, and molecular dynamic analysis. The sequence of U2-SCRTX-Lit2 obtained from the transcriptome is similar to that of μ-Hexatoxin-Mg2, a peptide that inhibits the insect Nav channel. Bioinformatic analysis of sequences classified as ICK family members also showed a conservation of cysteine residues among ICKs from different spiders, with the three dimensional molecular model of U2-SCRTX-Lit2 similar in structure to the hexatoxin from μ-hexatoxin-Mg2a. Molecular docking experiments showed the interaction of U2-SCRTX-Lit2 to its predictable target—the Spodoptera litura voltage-gated sodium channel (SlNaVSC). After 200 ns of molecular dynamic simulation, the final structure of the complex showed stability in agreement with the experimental data. The above analysis corroborates the existence of a peptide toxin with insecticidal activity from a novel ICK family in L. intermedia venom and demonstrates that this peptide targets Nav channels.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Spider venom consists of a complex mixture of toxins used for predation and defense. The heterogeneity of these toxins is directly responsible for ensuring the evolutionary success of spiders, making them the most abundant terrestrial venomous group [1]. Spider venoms are usually colorless liquids enriched with a variety of proteins and chemical compounds, such as enzymes, polypeptides, polyamines, nucleic acids, free amino acids, monoamines, and inorganic salts [2, 3]. Neurotoxins exist among several toxins present in spider venom and act on ion channels in insects and/or mammals, causing paralysis and death. These insecticidal toxins are found not only in spider venoms but also in other venomous species including, scorpions, cone snails, plants, and sea anemones. Most insecticidal toxins are cysteine-rich peptides with a molecular mass between 2 and 12 kDa [4], and these peptides frequently assume a structural scaffold known as an inhibitor cystine knot (ICK). This scaffold is characterized by the presence of two or three antiparallel β-sheets maintained by two disulfide bonds. A third disulfide bond crosses this scaffold resulting in a pseudo-knot arrangement [5, 6]. Due to this knot, these peptides are also known as knottins.
Cystine-knotted peptides are subdivided into three groups—inhibitor cystine knot (ICK), cyclic cystine knot (CCK) and growth factor cystine knot (GFCK), according to their knot structure and amino acid backbone [5]. Cystine-knotted peptides are very stable, as the disulfide bonds in the knot structure make the peptides resistant to protease activity, extreme pH levels, and high temperatures [7, 8]. ICK peptides are structurally similar to CCK peptides; both types of knots are formed by β-sheets that are stabilized by disulfide bonds. Spider venoms are the most abundant source for cystine-knotted peptides from the ICK family, with 775 peptides already deposited in the Knottin Database [9, 10]. It has been suggested that ICK peptides may act upon many different targets, such ion channels, NMDA receptors, and acid-sensing ion channels (ASICs). ICK spider toxins have been widely studied and well described. For example, the biological activity and tertiary structure of atracotoxin (delta-atracotoxin-Ar1), the first NaV channel binding toxin identified, have been extensively studied and identified [11].
Additionally, CCKs are the most stable peptides in the cystine-knotted peptide family and are present in Archaea, Bacteria, and Eukaryotes. CCK peptides, also called cyclotides, have a cyclic backbone in which the N- and C-termini are joined through a conventional amide bond. These peptides are essential in plant defenses against insect predation. Cybase is a database for cyclic peptides containing 818 sequences from 105 species throughout all kingdoms of life [12, 13], with cyclotides representing a large variety of antiretroviral, antimicrobial, hemolytic, and uterotonic properties [14].
Loxosceles intermedia is a brown spider of the Sicariidae family commonly found in Brazil. Loxosceles spiders have a violin-shaped pattern on the dorsal surface of their cephalothorax. Their body length varies from 1 to 5 cm, including the legs, and they have six eyes arranged in non-touching pairs in a U-shaped pattern. Their venom is a complex mixture of proteins and peptides, with phospholipases-D being the most well characterized enzyme [15]. For Loxosceles intermedia venom, three peptides with lepidopteran larvae insecticidal activity have been previously purified and tested. The De Castro laboratory identified and named the peptides as LiTx 1–3 in 2004 [16] (following suggested nomenclature King 2008 [17], U1-sicaritoxin-Li1a, U1-sicaritoxin-Li1b, U2-sicaritoxin-Li1a, respectively). Both LiTx1 and LiTx2 exhibit high amino acid sequence similarity with NaV or CaV channel-targeting toxins. The possible target of LiTx3 is the NaV channel due to its sequence similarity with δ-paluT3 (δ-Amaurobitoxin-Pl1c). δ-paluT3, purified from Pireneitega luctuosa (old Paracoelotes luctuosus) has been demonstrated to have activity against insect Nav channels, but not mammalian Nav channels [18].
All three toxins from L. intermedia are peptides rich in cysteine residues and are putative ICKs. Matsubara and coworkers [19] cloned and expressed a new ICK peptide isoform (U2-SCRTX-Li1b) from L. intermedia. This new recombinant peptide was used as an antigen to produce hyperimmune sera. The recombinant U2-SCRTX-Li1b was also recognized by antisera against L. intermedia, L. laeta, and L. gaucho, suggesting the existence of similar peptides in different Loxosceles species. Transcriptome analysis of the L. intermedia venom gland by Gremski and coworkers [20] revealed additional LiTx toxins with predicted insecticidal activities. They also made note of another toxin group with 2.4 % of toxin-encoding transcripts related to Magi 3 (μ-hexatoxin-Mg2a) from the Macrothele gigas spider, another insect Nav channel inhibitor. μ-hexatoxin-Mg2a binds specifically at site 3 of the NaV channel, causing reversible paralysis in Spodoptera litura [4]. We further characterize these toxins below.
Interest in characterizing knottin peptides stems from their varied activities against different targets and the fact that they have three important characteristics for drug design: excellent stability, high target selectivity, and high specificity [21]. Here, we describe the bioinformatic characterization of a novel ICK peptide from L. intermedia venom, the relationship between U2-SCRTX-Lit2 and other ICK spider toxins, and the predicted peptide and channel structures to infer the molecular target and mode of action for this ICK toxin.
Material and methods
Reagents
DNA molecular mass standards, Taq DNA polymerase, Pfu DNA polymerase, T4 DNA ligase, restriction enzymes, and dNTPs from Fermentas (Burlington, Canada). Crude L. intermedia venom was extracted from wild-caught spiders in accordance with the Brazilian Federal System for Authorization and Information on Biodiversity (SISBIO-ICMBIO, N° 29801–1) [22].
U2-SCRTX-Lit2 cDNA cloning
cDNA encoding the putative mature knottin peptide U2-SCRTX-Lit2 was amplified by PCR from the venom library made by Gremski et al. [20]. The forward primer that was used was 5′-GAAGTAGAAGATACACCA-3′, and the reverse primer was 5′-AATCTCGAGTCAGTTGGTGATTCC-3′. The amplified sequence was then subcloned into the pET SUMO vector (Invitrogen, USA). The correct construct was confirmed by sequencing.
Bioinformatics analysis of amino acid sequences
The LIC 327DNA sequence, which was obtained from Gremski et al. 2010 [20], was analyzed using online tools and showed similarity with μ-Hexatoxin-Mg2 (AS000380). The sequence was translated using the EXPASY tool [23], and signal peptide prediction was performed using the SignalP tool [24]. The ProtParam tool was used to calculate different physical chemical peptide parameters [25]. The classification of the cloned sequence, U2-SCRTX-Lit2, in the cystine knot family was determined using the Knotter 1D tool [9, 10]. Others knottin sequences were obtained from the Arachnoserver database, and the hydrophobic signal peptides and propeptides were defined by the SpiderP tool [25]. All analyses were performed using the default parameters. Multiple sequence alignments were generated using ClustalW2 with modified parameters (Gap open penalty = 8; Blosum table cysteine value was set for 100) [26]. The accession numbers of the aligned sequences obtained from the Arachnoserver database are as follows: U11-ctenitoxin-Pn1a (AS000266), U11-ctenitoxin-Pn1b (AS000161), U12-ctenitoxin-Pn1a (AS000267), U18-ctenitoxin-Pn1a (AS000254), U19-ctenitoxin-Pn1a (AS000213), U1-ctenitoxin-Pn1a (AS000282), U1-ctenitoxin-Pr1a (AS000216), U2-ctenitoxin-Pk1a (AS000230), U2-ctenitoxin-Pn1a (AS000234), U2-ctenitoxin-Pn1b (AS000232), U7-ctenitoxin-Pk1a (AS000268), U8-ctenitoxin-Pk1a (AS000238), U8-ctenitoxin-Pr1a (AS000263), U9-ctenitoxin-Pr1a (AS000214), Γ-ctenitoxin-Pn1a (AS000279), δ-ctenitoxin-Asp2e (AS000016), δ-ctenitoxin-Pn1a (AS000280), δ-ctenitoxin-Pn1b (AS000278), δ-ctenitoxin-Pn2a (AS000243), δ-ctenitoxin-Pn2b (AS000242), δ-ctenitoxin-Pn2c (AS000241), δ-ctenitoxin-Pr2d (AS000259), ω-ctenitoxin-Pn4a (AS000269), ω-ctenitoxin-Pr2a (AS000239), U1-hexatoxin-Hi1a (AS000646), U1-hexatoxin-Hv1a (AS000209), U1-hexatoxin-Iw1a (AS000601), U1-hexatoxin-Iw1b (AS000643), U1-hexatoxin-Iw1c (AS000564), U1-hexatoxin-Iw1d (AS000544), U1-hexatoxin-Iw1e (AS000691), μ-hexatoxin-Mg2a (AS000380), ω-oxotoxin-Ol1a (AS000207), ω-oxotoxin-Ol1b (AS000208), ω-oxotoxin-Ot1a (AS000779), U1-plectoxin-Pt1a (AS000405), U1-plectoxin-Pt1b (AS000409), U1-plectoxin-Pt1c (AS000392), U1-plectoxin-Pt1d (AS000393), U1-plectoxin-Pt1f (AS000395), U3-plectoxin-Pt1a (AS000391), ω-plectoxin-Pt1a (AS000412), U1-sicaritoxin-Li1a (AS000225), U1-sicaritoxin-Li1b (AS000251), U1-sicaritoxin-Li1c (AS000283) and U2-sicaritoxin-Li1a (AS000273).
Molecular modeling and structure equilibration
Molecular modeling of Spodoptera litura voltage-gated sodium channel (SlNaVSC – UniProt reference Q3Y5H0) was performed by MODELLER 9.15 [27], using a pore-only sodium channel derived from Alkalilimnicola ehrlichei (PBD 4LTO) as a template [28]. Human agouti related protein (PBD 1HYK) [29] was used as a template for U2-SCRTX-Lit2 modeling. Both templates were chosen after multiple amino acid sequence alignments performed in HHpred webserver [29]. 4LTO structure showed the possibility of 99.96 % for homologous relationship and 1.5 × 10−29 E-value, while for 1HYK it was 97.36 % and 3.6 × 10−4, respectively. Disulfide bonds were explicit in input parameters. PROCHECK webserver [30] was used to evaluate the stereochemical quality of the models. Over 93 % and 70 % of the residues of the modeled SlNaVSC and U2-SCRTX-Lit2, respectively, were assigned on the best allowed regions of the Ramachandran plot, and the remaining residues were located in the marginal regions of the plot; except for Cys25 of U2-SCRTX-Lit2 which occurred in the disallowed region (data not shown).
We performed molecular dynamics to optimize the SlNaVSC structure in a DLPA bilayer and water using Gromacs simulation package, version 5.1 [31]. The system was built with CHARMM-GUI webserver [32–34] with TIP3P water molecules [35] modified for the CHARMM force field. To ensure the successful equilibration of the system, relaxing the structure before production MD, we opted to perform six steps of equilibration dynamics, as described in Jo, Kim, and Im (2007) [33]. Equilibration consisted of applying restraints to the protein, water, ions, and lipid molecules as follows: i) harmonic restraints to ions and heavy atoms of the protein; ii) repulsive planar restraints to prevent water from entering into the membrane hydrophobic region; and iii) planar restraints to hold the position of head groups of membranes along the Z-axis. The restraint forces were gradually decreased step by step and completely removed during the production MD of 200 ns. The reference structure for the normal modes and docking studies in this work was obtained at the lowest energy state during production MD, which was achieved in an RMSD plateau, in other words, when the system was in equilibrium.
Normal modes analysis
Normal modes analysis was used to investigate possible opening motions of the channel, which is important for toxin docking. We calculated the first 87 internal lowest frequency modes of the minimized channel structure. These first modes describe the most representative global conformational changes for the molecule. The opening motions were observed in modes 11 and 23 (Fig. 4). The simulations were performed using CHARMM software [36] and CHARMM36 force field, excluding CMAP [37].
Modes 11 and 23 were used as directional constraints to generate low-energy structures along the normal mode vectors using the VMOD algorithm in CHARMM [38–40]. The VMOD routine restricts only the normal mode coordinate whereas others degrees of freedom remain unaltered [41]. The structures were displaced from −3.0 Å to +3.0 Å using steps of 0.1 Å, resulting in 61 intermediate low-energy structures along each mode. During minimization, the cutoff for non-bonded interactions was set to 10 Å, with a switching function applied between 8 Å and 10 Å [42]. This procedure allowed us 122 different receptor structures to docking calculation.
Molecular docking and evaluation of the complex stability by molecular dynamic simulation
Docking simulations were performed with Hex 8.0.0 [43]. Hex treats proteins as rigid bodies and makes a blind search evaluating the interaction correlation between structures using the fast Fourier transformation algorithm. Each molecule is modeled using an orthogonal three-dimensional expansion to encode its shape, electrostatic charge, and distribution of potential. The effects of solvation and desolvation were treated as surface phenomena, since the algorithm assumes an exclusion model volume and complementarity of form [44]. For each receptor structure 1000 solutions were determined. The best 50 solutions of each round were then clustered into eight final clusters. Further analyses were made using BINANA 1.2.0 [45] to evaluate the specific molecular interactions between channel and toxin. The best solution was used as reference structure to new molecular dynamics analysis to evaluate the behavior of the docked complex over time. Molecular dynamics were performed by inserting the complex in a DLPA bilayer and TIP3P water, as described in the previous section.
Results
Analysis and obtainment of nucleotide and amino acid sequences
The transcriptome analysis of L. intermedia venom, described by Gremski and coworkers, identified a group of toxins that were similar to μ-hexatoxin-Mg2a (Magi 3) [20]. This novel, putative toxin-encoding sequence was registered by 19 expressed sequence tags (ESTs), which were grouped in nine clusters, and coded for two distinct amino acid sequences, both 82 residues in length. These two amino acid sequences have only seven different residues; three residues are in the mature form of the peptide sequence, and one is a conservative substitution. In the present study, we focused on one of these two main sequences (LIC 327), and investigated the peptide, U2-SCRTX-Lit2, using bioinformatics tools available online. The predicted amino acid sequence was obtained using the EXPASY translate tool and showed the presence of 10 cysteine residues. The SignalP tool predicated that U2-SCRTX-Lit2 contains a signal peptide of 19 amino acid residues. Further analysis with the ArachnoServer using the SpiderP tool showed a propeptide of 15 amino acid residues, resulting in a mature toxin of 48 residues. The analysis of the disulfide bond patterns performed using the Knotter 1D tool in the Knottin Database suggested that six cysteine residues formed a C1-C4/C2-C6/C3-C9 structural motif. The acquired disulfide bond pattern agrees with the predicted pattern for the μ-hexatoxin-Mg2a neurotoxin (Fig. 1). Due to the sequence similarity between U2-SCRTX-Lit2 and μ-hexatoxin-Mg2a (52 %) and the presence of 10 cysteine residues, the predicted U2-SCRTX-Lit2 amino acid sequence was submitted to the Knottin Database for further analysis. The U2-SCRTX-Lit2 predicted sequence was defined as a putative ICK family member by the Knotter 1D tool.
Amino acid alignment of ICK sequences
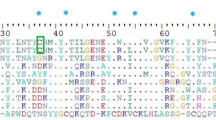
To compare the U2-SCRTX-Lit2 sequence with structurally related peptides, we performed a search in the ArachnoServer database for sequences that were predicted to form five disulfide bonds, equal to the number of predicted disulfide bonds in the U2-SCRTX-Lit2 peptide. The search returned 42 potential sequences containing 10 to 12 cysteine residues. These sequences and four predicted ICK peptides from L. intermedia venom, which also contained 10 cysteine residues, were used to generate multiple sequence alignments. The 47 chosen sequences were from eight genera and 12 species of spiders, with these toxins acting on many distinct molecular targets. Multiple sequence alignments were performed using ClustalW2 with modified parameters: the gap open penalty was 8 and the Blosum table cysteine value was set to 100. Only mature sequences were used for alignment. The ClustalW2 alignment showed that nine cysteine residue positions were highly conserved among all sequences. Additionally, four other cysteine residues (positions 31, 36, 45, and 106) were conserved among some sequences, although the positions were not the same in all sequences (Fig. 2). BLAST analysis showed that U2-SCRTX-Lit2 had more similarity with toxins from the Hexathelidae, Plectreuridae, and Sicariidae spider families. Toxins from these families are able to inhibit sodium and calcium channels, causing paralysis and death in insects. This result in addition with the above bioinformatic findings suggest a putative biological function for U2-SCRTX-Lit2.
Alignment among ICK toxin sequences. The alignment shows nine cysteine residues aligned among all sequences, and four other residues found in some groups of toxins. U2-SCRTX-Lit2 has the same cysteine profile as the plectoxins and μ-hexatoxin-Mg2a, the most similar one by BLASTp analysis. The difference in length and composition of amino acids in the loops and the regions between cysteine residues which are related with broad spectrum of activities from ICK toxins are also highlighted. Alignment performed using ClostalW2 with modified parameters gap open penalty = 8 and Blosum table cysteine value set to 100
Three-dimensional modeling of Spodoptera litura NaVSC and U2-SCRTX-Lit2
The modeled SlNaVSC is built from five α-helices. Helices H1 and H4 span the transmembrane, H2 and H3 are responsible for the control of channel penetrability, and H5 is intracellular (Fig. 3b and c). The tetramer contains a predominantly hydrophilic cavity; the external helices show a highly hydrophobic pattern, as expected for the transmembrane regions (Fig. 3d and e). Each chain interacts with the others by interweaving hydrophilic and hydrophobic interactions with its neighbor. SlNaVSC helices H2 and H3 are more flexible than others regions, consistent with their role in controlling the opening and closing of the channel cavity. Other regions of the channel are quite rigid.
Ribbon diagrams of the modeled SlNaVSC. The five α-helices building the 3D-folding of the channel are differently colored and numbered H1–H5 (a). View by side (b) and top (c). Surface hydrophobicity of SlNaVSC (d and e). Orange and blue represent maximum hydrophobicity and hydrophilicity, respectively. N and C correspond to the N- and C-terminus of the polypeptide chains, respectively
Based on molecular modeling, the U2-SCRTX-Lit2 predicted structure exhibits an N-terminal antiparallel β-hairpin that is stabilized by a complex cysteine knot, formed by five putative disulfide bonds between Cys02-Cys17, Cys09-Cys23, Cys16-Cys35, Cys20-Cys33, and Cys-25-Cys42 including the ICK domain (Fig. 4a). This knottin fold is important in all spider toxins targeting vertebrate or insect voltage-gated sodium channels. However, this does not rule out the possibility of other targets, such as calcium, potassium, proton or mechanosensitive channels [46–48], or hemagglutination in erythrocytes [49, 50]. According to King and collaborators, the ICK biological activities are often present or similar in molecules containing a structural scaffold defined by disulfide bond patterns [51]. The alignment of U2-SCRTX-Lit2 with the structural models of the μ-hexatoxin-Mg2a (Magi-3) and δ-hexatoxin-Mg1a (Magi-4) toxins, which were used to experimentally test for Nav channel binding [4, 49], showed that U2-SCRTX-Lit2 shares characteristics with these neurotoxins. The charge distribution is an important feature that can be observed in the U2-SCRTX-Lit2 model. The presence and arrangement of positively charged residues (Lys19 and Arg39) followed by non-polar residues (Leu40, Leu41, Trp5, and Val21) and negatively charged residues (Asp7 and Asp36) strongly suggest an affinity for SlNaVSC channel site 3 [52, 53] (Fig. 4c). Due to the complex cysteine-knot and highly packed structure, U2-SCRTX-Lit2 shows much more rigidity in comparison with SlNaVSC. The surface of U2-SCRTX-Lit2 is mainly hydrophilic (Fig. 4b).
Ribbon diagrams of the modeled U2-SCRTX-Lit2. The five salt bridges are specified in ball and stick representation (a). N and C correspond to the N- and C-terminus of the polypeptide chains, respectively. Surface hydrophobicity of U2-SCRTX-Lit2 (b). Orange and blue represent maximum hydrophobicity and hydrophilicity, respectively. Qualitative surface electrostatic potential of U2-SCRTX-Lit2 (c). Blue represent positive regions, white neutral and red negative regions
Normal modes analysis
We observed that modes 11 and 23 showed different opening motions: mode 11 had alternating opening of two monomers, while mode 23 showed the opening of the four monomers simultaneously (Fig. 5). These motions can be verified by the increase of the solvent accessible surface of the main cavity in the open structures (Table 1). We obtained 61 new structures related to the structure displacement along each mode by using VMOD on CHARMM. The VMOD protocol uses the normal mode as a constraint to successive molecular dynamics and energy minimization, aiming to generate relaxed structures at a displacement range of −3.0 Å to +3.0 Å [40–42]. Using these relaxed structures as receptor models, we performed the docking calculations.
Molecular docking
Docking calculations were performed using 122 channel structures obtained from normal mode analysis. In every docking round, 1000 different solutions were calculated. We picked the best 50 solutions of each complex structure and performed cluster analysis. The results showed eight clusters, from which cluster 1 had the best results. The normal modes approach to generate different conformations of the SlNaVSC channel was effective, since from the top 100 solutions in cluster 1, 73 % were displaced ≥ 1.0 Å. The best solution was found by docking the toxin into the SlNaVSC structure mode 23, at a displacement of −2.2 Å. After BINANA revaluation, the summed electrostatic energy in complex interface contacts was −974.23 kJ mol−1 (Fig. 6a). The interface observed is composed of four pockets of hydrophobic contacts (Fig. 6b), five salt bridges (Fig. 6c), and three H-bonds (Fig. 6d). Details about these interactions can be found in Table 2. Geometric complementarity was not optimized because the docking was performed with two rigid proteins.
Molecular interactions observed in docked SlNaVSC and U2-SCRTX-Lit2 (a). Hydrophobic contacts form four pockets represented by orange mesh (b). In navy blue mesh, five salt bridges form three pockets (c). Three hydrogen bonds were observed and are highlighted in black circles, with the distance between donor and acceptor (d)
Molecular dynamics
To better understand the relationship between channel and toxin, we performed molecular dynamics with SlNaVSC buried in a DLPA bilayer and water as well for the docked complex. The secondary structure during the molecular dynamics was evaluated by using DSSP method. No significant changes were observed for both molecules (data not shown). System total energy was sensibly decreased in the complex, from a mean of −6.5 × 105 kJ mol−1 for SlNaVSC alone to −8.5 × 105 kJ mol−1 for the complex. U2-SCRTX-Lit2 interaction plays a key role in SlNaVSC stabilization, decreasing the channel mobility during dynamics. Figure 7 shows this decrease of the stable plateau of RMSD for SlNaVSC (between 125 and 200 ns). However, the flexibility remained almost unaltered excepting residues 240–246 that showed a decrease in docked complex (Fig. 8). The decrease of mobility but not the flexibility indicates that the toxin acts as a steric blocker, hiding the gate access. As expected, U2-SCRTX-Lit2 shows a very stable motility pattern due to its disulfide bonds spread along the structure, exhibiting a stable plateau between 100 and 200 ns (Fig. 7). The geometric complementarity were modified during MD. Although the toxin was shown less buried at the surface of the channel during MD, the conformation assumed was quite stable, and it also promoted a rearrangement of hydrophobic contacts, salt bridges, and hydrogen bonds (Fig. 9 and Table 3). The BINANA evaluation of the final confirmation of the complex showed a significant increase summed electrostatic energy in the interface contacts, resulting in a total of −2371.15 kJ mol−1.
Molecular interactions observed in docked SlNaVSC and U2-SCRTX-Lit2 after 200 ns of production molecular dynamics (a). Hydrophobic contacts form four pockets represented by orange mesh (b). In navy blue mesh, four salt bridges form three pockets (c). Twelve hydrogen bonds were observed and are highlighted in black circles, with the distance between donor and acceptor (d)
Discussion
L. intermedia venom is a complex mixture of many proteins and peptides, with the most characterized proteins belonging to the phospholipase-D family [15, 54, 55]. Nevertheless, transcriptome analyses of L. intermedia venom glands showed that more than half of the toxin transcripts encode for peptides with insecticidal potential. Due to their abundance, interesting biological activity and promising biotechnological applications, we characterized a novel toxin peptide from L. intermedia venom, named U2-SCRTX-Lit2. Analysis of the U2-SCRTX-Lit2 sequence using the Knotter 1D tool confirmed that this peptide is a novel member of the ICK peptide family. The ICK family contains a structural motif in which peptides have three to seven disulfide bonds [5, 56, 57]. ICK family members have many molecular targets and are derived from various organisms, including animals, plants, and fungi. The cysteine connectivity framework is extremely diverse, with the best-described pattern being C1-C4, C2-C5, and C3-C6 [9, 10]. The connectivity framework may vary in ICK sequences that contain more than six cysteine residues; cysteine residues involved in the formation of the knot motif do not follow the conventional connective framework of C1-C4, C2-C5, C3-C6. Our peptide of interest, U2-SCRTX-Lit2, is composed of ten cysteine residues. Sequence similarity to μ-hexatoxin-Mg2a, data from Knotter 1D and modeling predict a connectivity pattern of C1-C4, C2-C6, C3-C9, C5-C8, C7-C10. This difference in the connectivity pattern of U2-SCRTX-Lit2 and disposition of cysteine residues is in agreement with Vassilevski and coworkers (2009) [2]. Different patterns of connectivity associated with the amino acid composition of loops between cysteine residues are responsible for the broad activity spectrum of cystine-knotted peptides, while the amino acids between the cysteines are involved in loop organization. The most well studied loops are found in the classical ICK peptides and contain six cysteine residues. Cysteine residues III and IV are adjacent, a characteristic frequently found in spider ICKs, including U2-SCRTX-Lit2.
The SignalP and SpiderP bioinformatic analysis of the predicted amino acid sequence of U2-SCRTX-Lit2 revealed that it is composed of a signal peptide containing 19 amino acid residues, which is also present in other ICK peptides [58–60]. The SpiderP tool analysis also identified a 15 amino acid propeptide in U2-SCRTX-Lit2, resulting in a mature toxin of 48 residues. Propeptides are often found in spider toxins and are usually cleaved after an arginine residue by specific peptidases as found in U2-SCRTX-Lit2 [61]. The U2-SCRTX-Lit2 molecular structure containing the signal peptide, propeptide, and mature toxin is in agreement with the structural organization described for several spider toxins [62, 63].
To highlight the importance of the cysteine positions, we performed multiple alignment of peptides with five disulfide bonds, which included peptides in L. intermedia venom. The alignment showed nine cysteine residues at the same amino acid position in all sequences analyzed. The other cysteine residues were found in four patterns and were associated with different biological activities. U2-SCRTX-Lit2 has vicinal cysteines III and IV, common for spider toxins [2, 9, 10]. The fifth cysteine residue was only found in U2-SCRTX-Lit2, μ-hexatoxin-Mg2a and plectoxins, but not in other ICK toxins from the Loxosceles family (sicaritoxins) (Fig. 2). In sicaritoxins the additional cysteine is found at position IX, the same position that is described in two other groups of ICKs: the hexatoxins that lack biological activity and ctenitoxin, which has 12 cysteine residues and confirmed biological activity. These results strengthened the idea that amino acid composition and disulfide bond patterns are important for ICK activities.
U2-SCRTX-Lit2 shows similarity by BLASTp, with toxins with activity on Nav and Cav channels and are able to cause paralysis and death in insects. The most similar toxin to U2-SCRTX-Lit2 is μ-hexatoxin-Mg2a, (52 % identity) which has a paralytic effect on Spodoptera litura, due to its binding at site 3 of Nav channels, but does not exhibit activity in mice [4]. Beyond this cited toxin, it also has similarity with previous reported L. intermedia toxins that exhibit insecticidal activity. U2-SCTX-Li1a most likely interacts with Na+ channels, causing flaccid paralysis in lepidopteran larvae [16]. Other sicaritoxins are toxic to insects, but their molecular targets are not defined. For plectoxins, only ω-plectoxin-Pt1a had confirmed activity, and is related to the inhibition of presynaptic Cav channels in Drosophila nerve terminals [64]. The alignment suggested that U2-SCRTX-Lit2 was more closely related to plectoxins than sicaritoxins. This was expected because the cysteine framework of plectoxins and U2-SCRTX-Lit2 are the same, and these peptides share some additional amino acid conservation, which is not observed with sicaritoxins. Another difference is that sicaritoxins contain a C-terminal peptide, the biological function of which is not well understood [16]. This C-terminal peptide was found in U1-SCTX-Li1a, U1-SCTX-Li1b, and U1-SCTX-Li1c, but not in U2-SCTX-Li1a and U2-SCRTX-Lit2. This C-terminal peptide is probably cleaved as a prost-translational modification as described for other ICK peptides. These peptides frequently show post-translational modifications, including amidation at the C-terminus, which often increases toxin activity and potency [65, 66]. De Castro and collaborators 2004 [16] showned that the lysine at U1-SCRTX-Lit1a, and arginine residues at U1-SCRTX-Lit1b and U1-SCRTX-Lit1c are amidated after the last cysteine, corroborating the idea of C-terminal propeptide processing.
Next, we used the structural model for μ-hexatoxin-Mg2a available in the Knottin Database to compare the predicted structure of U2-SCRTX-Lit2. U2-SCRTX-Lit2 apparently contains the inhibitory cystine-knot structural motif and shows sequence homology with neurotoxin peptides. Indeed, the predicted disulfide bond arrangement of U2-SCRTX-Lit2 resembles spider neurotoxin peptides [46–48]. The obtained ICK motif consists of double-antiparallel β-sheets connected by three disulfide bonds, which form a small stable globular domain, often observed in a variety of scorpion, spider, cone snail, and snake toxins [53, 67, 68]. Because U2-SCRTX-Lit2 is structurally similar to the fold found in spider toxins that is capable of modulating NaV channels and contains the key residues for binding to the NaV channel target, it is expected that this peptide also has the same structure-function relationship. The molecular model and the structural alignment showed residues that are conserved in spider toxins targeting Nav channel site-3 [49, 50, 69]. Sequence and structural homology suggest that a number of residues in these spider toxins are oriented in a similar manner. U2-SCRTX-Lit2 contains a cluster of positively charged (Lys18 and Arg38) and aromatic (Trp5 and Phe28) residues that appear in similar positions on the surface of almost all spider active neurotoxins [4, 52, 53, 70, 71]. Although not yet confirmed, these conserved regions are most likely responsible for the affinity for Nav channels. Interestingly, structure prediction analysis revealed a structural similarity between U2-SCRTX-Lit2 and human ASIP, a peptide with an ICK scaffold. Human ASIP and AgRP are described as unique peptides with an ICK fold that modulate the activity of G protein-coupled receptors [72]. The biological activity of ASIP is associated with antagonism of melanocortin receptors, which could be due to the way the peptide interacts with its target. Similar to what occurs with insecticidal ICK peptides, this weaponization of hormones has already been described for centipedes and spiders [73].
Our models showed very good results in quality analysis, indicating the power of threading modeling to perform in silico calculations from structures that do not have any experimental resolution available. Both channel and toxin show hydrophobic patterns consistent with their putative characteristics: SlNaVSC shows high hydrophobic regions in the transmembrane region while the gate region is mainly hydrophilic as is most of the U2-SCRTX-Lit2 surface (Figs. 3 and 4) [74]. Futhermore, surface charge complementarity was observed in these regions. Protein-protein docking studies can predict the interaction pose and energy between two proteins by using their geometric and charge complementarity [43, 75]. To predict the inhibitory action of U2-SCRTX-Lit2 on SlNaVSC with reasonable confidence, the channel is required to be in an open-state conformation. In this way, NMA calculations were performed to investigate these opening and closing motions, since they are essential to Na+ ion intake. We verified that normal modes 11 and 23 presented two different kinds of open/close motions: alternating AC/BD opening/closing and simultaneous opening/closing.
Docking and MD analysis indicate that the toxin acts by not only blocking the channel, but also preventing its opening motion. Interactions occur in three chains at least, preventing the movement of helices H2 and H3 of each chain. Our hypothesis is that water molecules guide the interface formation, bridging hydrogen bond donors and acceptors, aided by hydrophobic contacts and salt bridges holding the channel and toxin together. Notwithstanding the weaker forces of salt bridges and hydrogen bonds in protein-protein interfaces, water competition in protein-protein binding can stabilize protein association [76]. Polar and charged residues at the protein surface may contribute to these interactions. Xu and coworkers (1997) [76] found an average of 10.7 hydrogen bonds and 2.0 salt bridges per interface in a study with 319 non-redundant protein–protein interfaces, previously assembled from protein X-ray structures.
In summary, the results suggest the existence of a novel class of peptides from the ICK family in Loxosceles intermedia venom. Bioinformatic analysis reveals U2-SCRTX-Lit2 contains characteristics that are similar to other ICK peptides. Multiple sequence alignments grouped U2-SCRTX-Lit2 with peptides that act on Nav or Cav channels. Molecular docking and protein modeling suggest that the toxin is able to bind and modulate insect Nav channels.
References
King GF, Hardy MC (2013) Spider-venom peptides: structure, pharmacology, and potential for control of insect pests. Annu Rev Entomol 58:475–496
Vassilevski AA, Kozlov SA, Grishin EV (2009) Molecular diversity of spider venom. Biochem Biokhimiia 74:1505–1534
Jackson H, Parks TN (1989) Spider toxins: recent applications in neurobiology. Annu Rev Neurosci 12:405–414
Corzo G, Gilles N, Satake H, Villegas E, Dai L, Nakajima T, Haupt J (2003) Distinct primary structures of the major peptide toxins from the venom of the spider Macrothele gigas that bind to sites 3 and 4 in the sodium channel. FEBS Lett 547:43–50
Iyer S, Acharya KR (2011) Tying the knot: The cystine signature and molecular-recognition processes of the vascular endothelial growth factor family of angiogenic cytokines. FEBS J 278:4304–4322
Craik DJ, Daly NL, Waine C (2001) The cystine knot motif in toxins and implications for drug design. Toxicon 39:43–60
Werle M, Loretz B, Entstrasser D, Foger F (2007) Design and evaluation of a chitosan-aprotinin conjugate for the peroral delivery of therapeutic peptides and proteins susceptible to enzymatic degradation. J Drug Target 15:327–333
Werle M, Kolmar H, Albrecht R, Bernkop-Schnurch A (2008) Characterisation of the barrier caused by luminally secreted gastro-intestinal proteolytic enzymes for two novel cystine-knot microproteins. Amino Acids 35:195–200
Gracy JRM, Le-Nguyen D, Gelly J-C, Kaas Q, Heitz A, Chiche L (2007) The knottin or inhibitor cystine knot scaffold in. Nucleic Acids Res 36(2008):D314–D319
Gelly JC, Gracy JRM, Kaas Q, Le†Nguyen D, Heitz A, Chiche L (2004) The KNOTTIN website and database: a new information system dedicated to the knottin scaffold. Nucleic Acids Res 32:D156–D159
Pallaghy PK, Alewood D, Alewood PF, Norton RS (1997) Solution structure of robustoxin, the lethal neurotoxin from the funnel-web spider Atrax robustus. FEBS Lett 419:191–196
Wang CK, Kaas Q, Chiche L, Craik DJ (2008) CyBase: a database of cyclic protein sequences and structures, with applications in protein discovery and engineering. Nucleic Acids Res 36:D206–210
Mulvenna JP, Wang C, Craik DJ (2006) CyBase: a database of cyclic protein sequence and structure. Nucleic Acids Res 34:D192–194
Craik DJ, Swedberg JE, Mylne JS, Cemazar M (2012) Cyclotides as a basis for drug design. Expert Opin Drug Discovery 7:179–194
Chaim OM, Trevisan-Silva D, Chaves-Moreira D, Carolina AM, Valéria Pereira Ferrer W, Hitomi Matsubara F, Mangili OC, da Silveira RB, Luiza Helena G, Waldemiro G, Andrea S-R, Silvio Sanches V (2011) Brown spider (Loxosceles genus) venom toxins: tools for biological purposes. Toxins 3:309–344
de Castro CS, Silvestre FG, Araujo SC, de Gabriel MY, Mangili OC, Cruz I, Chavez-Olortegui C, Kalapothakis E (2004) Identification and molecular cloning of insecticidal toxins from the venom of the brown spider Loxosceles intermedia. Toxicon 44:273–280
King GF, Gentz MC, Escoubas P, Nicholson GM (2008) A rational nomenclature for naming peptide toxins from spiders and other venomous animals. Toxicon 52:264–276
Corzo G, Escoubas P, Stankiewicz M, Pelhate M, Kristensen CP, Nakajima T (2000) Isolation, synthesis and pharmacological characterization of δ-palutoxins IT, novel insecticidal toxins from the spider Paracoelotes luctuosus (Amaurobiidae). Eur J Biochem 267:5783–5795
Matsubara FH, Gremski LH, Meissner GO, Constantino Lopes ES, Gremski W, Senff-Ribeiro A, Chaim OM, Veiga SS (2013) A novel ICK peptide from the Loxosceles intermedia (brown spider) venom gland: Cloning, heterologous expression and immunological cross-reactivity approaches. Toxicon 71:147–158
Gremski LH, da Silveira RB, Chaim OM, Probst CM, Ferrer VP, Nowatzki J, Weinschutz HC, Madeira HM, Gremski W, Nader HB, Senff-Ribeiro A, Veiga SS (2010) A novel expression profile of the Loxosceles intermedia spider venomous gland revealed by transcriptome analysis. Mol Biosyst 6:2403–2416
Reinwarth M, Nasu D, Kolmar H, Avrutina O (2012) Chemical synthesis, backbone cyclization and oxidative folding of cystine-knot peptides: promising scaffolds for applications in drug design. Molecules 17:12533–12552
Feitosa L, Gremski W, Veiga SS, Elias MC, Graner E, Mangili OC, Brentani RR (1998) Detection and characterization of metalloproteinases with gelatinolytic, fibronectinolytic and fibrinogenolytic activities in brown spider (Loxosceles intermedia) venom. Toxicon 36:1039–1051
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E, Grosdidier A, Hernandez C, Ioannidis V, Kuznetsov D, Liechti R, Moretti S, Mostaguir K, Redaschi N, Rossier G, Xenarios I, Stockinger H (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597–603
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Wilkins MR, Gasteiger E, Bairoch A, Sanchez JC, Williams KL, Appel RD, Hochstrasser DF (1999) Protein identification and analysis tools in the ExPASy server. Methods Mol Biol 112:531–552
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
N. Eswar, B Webb, MA Marti-Renom, MS. Madhusudhan, D. Eramian, MY. Shen, U Pieper, A Sali (2007) Comparative protein structure modeling using MODELLER. Curr Protoc Protein Sci Chapt 2:Unit 2.9. doi:10.1002/0471140864.ps0209s50
Shaya D, Findeisen F, Abderemane-Ali F, Arrigoni C, Wong S, Nurva SR, Loussouarn G, Minor DL Jr (2014) Structure of a prokaryotic sodium channel pore reveals essential gating elements and an outer ion binding site common to eukaryotic channels. J Mol Biol 426:467–483
Soding J, Biegert A, Lupas AN (2005) The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res 33:W244–248
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Jo S, Lim JB, Klauda JB, Im W (2009) CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys J 97:50–58
Jo S, Kim T, Im W (2007) Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS ONE 2, e880
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Brooks BR, Brooks CL 3rd, Mackerell AD Jr, Nilsson L, Petrella RJ, Roux B, Won Y, Archontis G, Bartels C, Boresch S, Caflisch A, Caves L, Cui Q, Dinner AR, Feig M, Fischer S, Gao J, Hodoscek M, Im W, Kuczera K, Lazaridis T, Ma J, Ovchinnikov V, Paci E, Pastor RW, Post CB, Pu JZ, Schaefer M, Tidor B, Venable RM, Woodcock HL, Wu X, Yang W, York DM, Karplus M (2009) CHARMM: the biomolecular simulation program. J Comput Chem 30:1545–1614
Buck M, Bouguet-Bonnet S, Pastor RW, MacKerell AD Jr (2006) Importance of the CMAP correction to the CHARMM22 protein force field: dynamics of hen lysozyme. Biophys J 90:L36–38
Floquet N, Durand P, Maigret B, Badet B, Badet-Denisot MA, Perahia D (2009) Collective motions in glucosamine-6-phosphate synthase: influence of ligand binding and role in ammonia channelling and opening of the fructose-6-phosphate binding site. J Mol Biol 385:653–664
Louet M, Perahia D, Martinez J, Floquet N (2011) A concerted mechanism for opening the GDP binding pocket and release of the nucleotide in hetero-trimeric G-proteins. J Mol Biol 411:298–312
Philot EA, Perahia D, Braz AS, Costa MG, Scott LP (2013) Binding sites and hydrophobic pockets in Human Thioredoxin 1 determined by normal mode analysis. J Struct Biol 184:293–300
Batista PR, Pandey G, Pascutti PG, Bisch PM, Perahia D, Robert CH (2011) Free energy profiles along consensus normal modes provide insight into HIV-1 protease flap opening. J Chem Theory Comput 7:2348–2352
Louet M, Karakas E, Perret A, Perahia D, Martinez J, Floquet N (2013) Conformational restriction of G-proteins Coupled Receptors (GPCRs) upon complexation to G-proteins: a putative activation mode of GPCRs? FEBS Lett 587:2656–2661
Ritchie DW, Venkatraman V (2010) Ultra-fast FFT protein docking on graphics processors. Bioinformatics 26:2398–2405
Ritchie DW (2008) Recent progress and future directions in protein-protein docking. Curr Protein Pept Sci 9:1–15
Durrant JD, McCammon JA (2011) BINANA: a novel algorithm for ligand-binding characterization. J Mol Graphics Model 29:888–893
Nicholson GM (2007) Insect-selective spider toxins targeting voltage-gated sodium channels. Toxicon 49:490–512
Nicholson GM, Little MJ, Birinyi-Strachan LC (2004) Structure and function of d-atracotoxins: lethal neurotoxins targeting the voltage-gated sodium channel. Toxicon 43:587–599
Nicholson GM, Blanche T, Mansfield K, Tran Y (2002) Differential blockade of neuronal voltage-gated Na(+) and K(+) channels by antidepressant drugs. Eur J Pharmacol 452:35–48
Liang S (2004) An overview of peptide toxins from the venom of the Chinese bird spider Selenocosmia huwena Wang [=Ornithoctonus huwena (Wang)]. Toxicon 43:575–585
Liang SP, Pan X (1995) A lectin-like peptide isolated from the venom of the Chinese bird spider Selenocosmia huwena. Toxicon 33:875–882
King GF, Tedford HW, Maggio F (2002) Structure and function of insecticidal neurotoxins from Australian funnel-web spiders. Toxin Rev 21:361–389
Li D, Xiao Y, Xu X, Xiong X, Lu S, Liu Z, Zhu Q, Wang M, Gu X, Liang S (2004) Structure--activity relationships of hainantoxin-IV and structure determination of active and inactive sodium channel blockers. J Biol Chem 279:37734–37740
Yamaji N, Little MJ, Nishio H, Billen B, Villegas E, Nishiuchi Y, Tytgat J, Nicholson GM, Corzo G (2009) Synthesis, solution structure, and phylum selectivity of a spider delta-toxin that slows inactivation of specific voltage-gated sodium channel subtypes. J Biol Chem 284:24568–24582
O.M. Chaim (2005) Estudo da atividade citotóxica da proteína dermonecrótica do veneno de aranha-marrom (Loxosceles intermedia) com enfase no efeito nefrotóxico
Appel MH, da Silveira RB, Chaim OM, Paludo KS, Silva DT, Chaves DM, da Silva PH, Mangili OC, Senff-Ribeiro A, Gremski W, Nader HB, Veiga SS (2008) Identification, cloning and functional characterization of a novel dermonecrotic toxin (phospholipase D) from brown spider (Loxosceles intermedia) venom. Biochim Biophys Acta 1780:167–178
Moore SR, Cochran JR, Wittrup KD, Gregory LV (2012) Engineering knottins as novel binding agents. . Methods Enzymol 503:223–251
Richardson M, Pimenta AMC, Bemquerer MP, Santoro MM, Beirao PSL, Lima ME, Figueiredo SG, Bloch C Jr, Vasconcelos EAR, Campos FAP, Gomes PC, Cordeiro MN (2006) Comparison of the partial proteomes of the venoms of Brazilian spiders of the genus Phoneutria. Comp Biochem Physiol C: Toxicol Pharmacol 142:173–187
Liu Z, Yu Z, Liu N, Zhao C, Hu J, Dai Q (2010) cDNA cloning of conotoxins with framework XII from several Conus species. Acta Biochim Biophys Sin 42:656–661
Liu Z,Li H, Liu N, Wu C, Jiang J, Yue J, Jing Y, Dai Q (2012) Diversity and evolution of conotoxins in Conus virgo, Conus eburneus, Conus imperialis and Conus marmoreus from the South China Sea. Toxicon 60:982–989
Kalapothakis E, Penaforte CL, Leao RM, Cruz JS, Prado VF, Cordeiro MN, Diniz CR, Romano-Silva MA, Prado MA, Gomez MV, Beirao PS (1998) Cloning, cDNA sequence analysis and patch clamp studies of a toxin from the venom of the armed spider (Phoneutria nigriventer). Toxicon 36:1971–1980
Kozlov SA, Grishin EV (2007) The universal algorithm of maturation for secretory and excretory protein precursors. Toxicon 49:721–726
Sollod BL, Wilson D, Zhaxybayeva O, Gogarten JP, Drinkwater R, King GF (2005) Were arachnids the first to use combinatorial peptide libraries? Peptides 26:131–139
Escoubas P, Sollod B, King GF (2006) Venom landscapes: mining the complexity of spider venoms via a combined cDNA and mass spectrometric approach. Toxicon 47:650–663
Branton WD, Kolton L, Jan YN, Jan LY (1987) Neurotoxins from Plectreurys spider venom are potent presynaptic blockers in Drosophila. J Neurosci : Off J Soc Neurosci 7:4195–4200
Sermadiras I, Revell J, Linley JE, Sandercock A, Ravn P (2013) Recombinant expression and in vitro characterisation of active Huwentoxin-IV. PLoS ONE 8: e83202
Windley MJ, Herzig V, Dziemborowicz SAA, Hardy MC, King GF, Nicholson GM (2012) Spider-venom peptides as bioinsecticides. Toxins 4:191–227
Wang X-h, Connor M, Smith R, Maciejewski MW, Howden MEH, Nicholson GM, Christie MJ, King GF (2000) Discovery and characterization of a family of insecticidal neurotoxins with a rare vicinal disulfide bridge. Nat Struct Mol Biol 7:505–513
Wen S, Wilson DT, Kuruppu S, Korsinczky ML, Hedrick J, Pang L, Szeto T, Hodgson WC, Alewood PF, Nicholson GM (2005) Discovery of an MIT-like atracotoxin family: spider venom peptides that share sequence homology but not pharmacological properties with AVIT family proteins. Peptides 26:2412–2426
Liang PH, Ko TP, Wang AH (2002) Structure, mechanism and function of prenyltransferases. Eur J Biochem 269:3339–3354
Villegas E, Adachi-Akahane S, Bosmans F, Tytgat J, Nakajima T, Corzo G (2008) Biochemical characterization of cysteine-rich peptides from Oxyopes sp. venom that block calcium ion channels. Toxicon 52:228–236
Wang P, Liao Z, Guo L, Li W, Chen M, Pi Y, Gong Y, Sun X, Tang K (2004) Cloning and functional analysis of a cDNA encoding Ginkgo biloba farnesyl diphosphate synthase. Mol Cells 18:150–156
McNulty JC, Jackson PJ, Thompson DA, Chai B, Gantz I, Barsh GS, Dawson PE, Millhauser GL (2005) Structures of the agouti signaling protein. J Mol Biol 346:1059–1070
Undheim EAB, Grimm LL, Low C-F, Morgenstern D, Herzig V, Zobel-Thropp P, Pineda SS, Habib R, Dziemborowicz S, Fry BG, Nicholson GM, Binford GJ, Mobli M, King GF (2015) Weaponization of a hormone: convergent recruitment of hyperglycemic hormone into the venom of arthropod predators. Structure 23:1283–1292
Rinkevich FD, Du Y, Dong K (2013) Diversity and convergence of sodium channel mutations involved in resistance to pyrethroids. Pestic Biochem Physiol 106:93–100
Tedford HW, Steinbaugh BA, Bao L, Tait BD, Tempczyk-Russell A, Smith W, Benzon GL, Finkenbinder CA, Kennedy RM (2013) In silico screening for compounds that match the pharmacophore of omega-hexatoxin-Hv1a leads to discovery and optimization of a novel class of insecticides. Pestic Biochem Physiol 106:124–140
Xu D, Tsai CJ, Nussinov R (1997) Hydrogen bonds and salt bridges across protein-protein interfaces. Protein Eng 10:999–1012
Acknowledgments
This work is supported by grants from CNPq, CAPES, Fundação Araucária-Paraná, UFABC, FAPESP. And Secretaria de Tecnologia e Ensino Superior do Paraná, SETI-PR, Brasil.
Author information
Authors and Affiliations
Corresponding author
Additional information
Highlights
• The importance of cysteine residue conservation among knottin peptide.
• Bioinformatic characterization of an ICK peptide from L. intermedia venom.
• Toxin and channel structure predictions and dynamic interactions.
Rights and permissions
About this article
Cite this article
Meissner, G.O., de Resende Lara, P.T., Scott, L.P.B. et al. Molecular cloning and in silico characterization of knottin peptide, U2-SCRTX-Lit2, from brown spider (Loxosceles intermedia) venom glands. J Mol Model 22, 196 (2016). https://doi.org/10.1007/s00894-016-3067-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-016-3067-0