Abstract
Pathogenesis-related group 5 (PR5) plant proteins include thaumatin, osmotin, and related proteins, many of which have antimicrobial activity. The recent discovery of PR5-like (PR5-L) sequences in nematodes and insects raises questions about their evolutionary relationships. Using complete plant genome data and discovery of multiple insect PR5-L sequences, phylogenetic comparisons among plants and animals were performed. All PR5/PR5-L protein sequences were mined from genome data of a member of each of two main angiosperm groups—the eudicots (Arabidoposis thaliana) and the monocots (Oryza sativa)—and from the Caenorhabditis nematode (C. elegans and C. briggsase). Insect PR5-L sequences were mined from EST databases and GenBank submissions from four insect orders: Coleoptera (Diaprepes abbreviatus and Biphyllus lunatus), Orthoptera (Schistocerca gregaria), Hymenoptera (Lysiphlebus testaceipes), and Hemiptera (Toxoptera citricida). Parsimony and Bayesian phylogenetic analyses showed that the PR5 family is paraphyletic in plants, likely arising from 10 genes in a common ancestor to monocots and eudicots. After evolutionary divergence of monocots and eudicots, PR5 genes increased asymmetrically among the 10 clades. Insects and nematodes contain multiple sequences (seven PR5-Ls in nematodes and at least three in some insects) all related to the same plant clade, with nematode and insect sequences separating as two clades. Protein structural homology modeling showed strong similarity among animal and plant PR5/PR5-Ls, with divergence only in surface-exposed loops. Sequence and structural conservation among PR5/PR5-Ls suggests an important and conserved role throughout the evolutionary divergence of the diverse organisms from which they reside.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The pathogenesis-related group 5 (PR5) family of plant proteins includes thaumatin and osmotin, closely related proteins. The PR5 proteins are known to accumulate in plants in response to pathogen challenge. Many PR5 proteins have been purified and shown to possess antifungal activity and have been named thaumatin-like, or osmotin-like (Abad et al. 1996; Cheong et al. 1997; Chu and Ng 2003a, b; Hejgaard et al. 1991; Hu and Reddy 1997; Huynh et al. 1992; Ng 2004; Vigers et al. 1991; Vu and Huynh 1994; Wang and Ng 2002; Woloshuk et al. 1991; Ye et al. 1999). The original thaumatin was isolated as a sweet-tasting protein from the fruits of the West African rain forest shrub Thaumatococcus danielli (van der and Loeve 1972) and has been studied in great detail due to its potential use as a noncarbohydrate sweetener. Since that time, the original thaumatin was also found to have antifungal activity (Vigers et al. 1992). Osmotins represent a related set of proteins that were originally isolated by virtue of their regulation by water stress (Singh et al. 1985, 1987). Protein crystallography studies have shown a highly conserved structure for the plant thaumatins and osmotins (Min et al. 2004).
Although the antimicrobial nature of PR5s is not well defined, some have been shown to function by stimulating microbial membrane permeability (Vigers et al. 1991), and as a result the name permatin was coined (Vigers et al. 1991). However, ß-1,3-glucanase activity (Grenier et al. 1999; Trudel et al. 1998) and α-amylase and trypsin inhibition have also been reported as a characteristic of zeamatin (a PR5 from maize) (Svensson et al. 2004; Schimoler-O’Rourke et al. 2001). The amylase/trypsin inhibition activity has been linked to defense against insect feeding (Franco et al. 2002), but these data remain controversial (Gómez-Leyva and Blanco-Labra 2001).
Antimicrobial properties of each PR5 are limited to specific groups of microbes (Abad et al. 1996), and it is hypothesized that an interaction with a membrane glycoprotein is the specificity determining step. Recent work indicates that PR5 binding to the phosphate group of phosphomannoproteins in fungal cell walls may be the initial binding step (Ibeas et al. 2001; Salzman et al. 2004). Since the discovery of the antimicrobial and insect deterrent activity of these enzymes, there has been hope that transgenic expression in plants could be used to produce crops resistant to pathogens and insects. Transgenic expression of PR5s has resulted in increased pathogen resistance in crop plants (Anand et al. 2003; Li et al. 1999; Zhu et al. 1996), however, none has yet reached commercialization. Understanding the mechanism by which these proteins exert their biological activity will aid in developing strategies for commercial use, and advances in genomics now permit comparative sequence and structural studies with the goal of designing or identifying PR5-like (PR5-L) proteins with desired activities.
Comparative studies were initiated through mining of insect expressed sequence tag (EST) libraries to identify cDNA clones encoding PR5-L proteins from representatives of three insect orders: Coleoptera, Hymenoptera, and Hemiptera. Taken with the recent discovery of similar sequences in an insect from the Orthoptera (Brandazza et al. 2004), the sequences provided a broad spectrum of representation within the class Insecta. Also, recent sequencing of complete plant genomes has allowed identification of virtually all PR5 sequences from Arabidopsis thaliana (L.) Heynh. (Thale cress, a eudicot) and Oryza sativa (L.) (rice, a monocot). Comparison of the PR5 protein sequences from these two plants and from the insects and nematodes provided a picture of the evolution of this gene family. These data are presented with protein structural homology modeling showing conservation of core structure among all PR5-L sequences and provide insight into the relationships among members of this conserved multigene family.
Materials and Methods
Bioinformatics
Plants
All annotated PR5-related sequences were obtained from A. thaliana and O. sativa by key word searches of The TIGR Arabidopsis thaliana Database (http://www.tigr.org/tdb/e2k1/ath1) and The TIGR Rice Database (http://www.tigr.org/tdb/e2k1/osa1), respectively, and are listed in Table 1. Since protein signature domains are used for gene annotation, other exclusion criteria were used to avoid the inclusion of PR5 domain containing proteins that have very different structures and most probably very different functions. These criteria required the protein to contain a complete PR5 protein signature domain and not to contain other domains indicative of alternate functions (i.e., protein kinase and membrane spanning domains related to signal transduction functions). Sequences from other plants listed in Table 1 were obtained through GenBank (accession numbers listed in Table 1).
Nematodes
All the nematode, Caenorhabditis, PR5-L sequences were obtained either from wormbase (http://www.wormbase.org; for C. elegans) or by NCBI blast searches (for C. briggsase).
Insects
The insect sequences were obtained as follows.
Coleoptera
Diaprepes abbreviatus (L.) (Coleoptera: Curculionidae): A set of 8 973 sequences in a cDNA EST dataset derived from a cDNA library of teneral female D. abbreviatus was mined. The sequences were clustered using Sequencher (Gene Codes Inc., Ann Arbor, MI). Three separate PR5-L clones, Dab-PR5-L1, -L2, and -L3, were identified as PR5-Ls in EST clusters by blastx searching of the GenBank nr database. Three full-length or near-full-length clones were chosen from each of three clustered alignments of EST sequences containing 4, 14, and 6 separate clones, respectively. A single clone representing the longest cDNA sequence was isolated and bidirectionally sequenced. The resulting sequence was submitted to GenBank and used in this study.
Biphyllus lunatus (F.) (Coleoptera: Biphyllidae): Putative full-length sequences were constructed from four EST sequences in GenBank (gi:25956508, 25956492, 25956846, 25956831).
Hymenoptera and Hemiptera
Lysiphlebus testaceipes (Cresson) (Hymenoptera: Braconidae) and Toxoptera citricida (Kirkaldy) (Hemiptera: Aphididae): The same procedure as for the D. abbreviatus clone discovery and characterization was followed using a L. testaceipes cDNA library constructed from adults and used to produce 3576 EST sequences, and a Toxoptera citricida library constructed from parthenogenic adult females used to produce 8760 EST sequences.
Orthoptera
Schistocerca gregaria (Forskål 1775) (Othoptera: Acrididae): Two sequences previously described by (Brandazza et al. 2004) were obtained from GenBank.
Amino Acid Sequence Alignments
Amino acid sequences deduced from cDNA clones were aligned using ClustalX (Thompson et al. 1997) with the following parameters: pairwise parameters were 35 for gap opening and 0.75 for gap extension; multiple alignment parameters were 15 for gap opening and 0.30 for gap extension as suggested by Hall (2004). Minor alignment issues were corrected using Se-Al (A. Rambaut; distributed by the author at http://evolve.zoo.ox.ac.uk/software/).
Determination of “Mature Protein”
Most of the PR5 proteins contained a signal sequence that targets the nascent peptide into the secretory pathway for either an extracellular or a vacuolar destination. Since this is a transient sequence that is cleaved from the mature protein, and sequence similarity among signal sequences is typically not conserved, all comparisons were performed with protein sequences minus the signal sequence. Determination of the amino acids comprising the N-terminal signal sequence was performed using the SignalP web server (Bendtsen et al. 2004). It should be noted that an original thaumatin precursor protein contains a C-terminal “pro-domain” of six amino acids that is cleaved in the mature form. Since this domain was determined empirically and cannot be predicted, and since it is relatively small, all comparisons were performed without attempts to identify or remove potential C-terminal pro-regions.
Phylogenetics
Amino acid sequences were evaluated using N-J (neighbor-joining) and Bayesian methods. N-J analyses were performed using PAUP* (Swofford 2003). Bayesian analyses were done using MrBayes version 3.0b4 (Huelsenbeck and Ronquist 2003). Heuristic searches with ten random addition sequence replicates and tree bisection reconstruction (TBR) branch swapping were performed for N-J estimates. The N-J estimates of the thaumatin phylogenies were obtained by estimating distance parameters in PAUP*. The N-J bootstrap analyses of 100 replicates were performed on each data set using a heuristic search with 10 random addition sequence replicates and TBR branch swapping. A Bayesian approach was used because of its easy interpretation of results, its ability to incorporate prior information (uniform in our case) (Huelsenbeck and Ronquist 2003), and its computational advantages (Larget and Simon 1999). MrBayes employs Markov chain Monte Carlo (MCMC) to approximate the posterior probabilities of phylogenies (Green 1995; Metropolis et al. 1953; Hastings 1970). The model of evolution used for the Bayesian analyses was a mixed model of evolution, which allows the MCMC chain to integrate over the 10 fixed amino acid rate matrices implemented in MrBayes. MrBayes was run with four chains from 10 different starting points. All runs were done for 10 million generations and trees were sampled every 100 generations. All runs reached a plateau in likelihood score, which indicated that the MCMC chains converged. One thousand trees were suboptimal at the beginning of the runs and therefore were discarded (burn-in phase). All trees saved from all 10 runs were summarized in PAUP* (see Mr Bayes manual) and the posterior probabilities were recorded (Figs. 1 and 2).
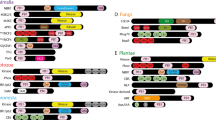
Comparison of PR5-L protein sequences from nematodes and insects with zeamatin from Zea mays. Abbreviations were constructed from the first letter of the genus name and the first and second letters of the species name (Dab, Diaprepes abbreviatus; Blu, Biphyllus lunatus; Sgr, Schistocerca gregaria; Lte, Lysiphlebus testaceipes; Tci, Toxoptera citricida; Cbr, Caenorhabditis briggsae; Cel, C. elegans) followed by the gene designation PR5-L and clone number (see Table 2). A ClustalX alignment of select insect and nematode PR5-L sequences with zeamatin. Pairwise parameters were 35 for gap opening and 0.75 for gap extension. Multiple alignment parameters were 15 for gap opening and 0.30 for gap extension. The shaded labels indicate the insect sequences. Arrows mark the location of the conserved 16 cysteine residues. The G1 through G6 labels and the associated stippled box on the zeamatin sequence indicate the location of gaps formed in the alignment, with a gap being identified as a space greater than one residue inserted in the majority of the sequences. The bracket at the top of the sequence delineates the first 44 residues in the insect sequences that are highly variable. B Most likely (−ln = −6207.89) MrBayes tree generated using MrBayes3.0 and the insect and nematode PR5-L amino acid alignment. Bayesian analyses were performed using a mixed model of evolution for 10 million generations. Numbers above the nodes are N-J parsimony bootstrap/posterior probabilities. Shading is provided to delineate insect orders.
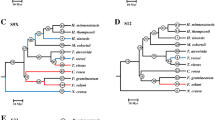
Comparison of PR5/PR5-L protein sequences from plants and animals. Animal sequence abbreviations as described in the legend to Fig. 1. Plant sequence abbreviations for Arabidopsis thaliana and Oryza sativa correspond to the gene locus name in the corresponding TIGR database, with the number following the two-letter genus and species name indicating the chromosome on which the gene is located. The asterisks indicate an O. sativa sequence that is only represented by an older pseudomolecule version nomenclature. Other plant sequences and the fungal sequence used as the outgroup (Magnaporthe griseae gi:38100706) are listed with genus and species followed by the gene designation. NSS is added to label sequences with no predicted signal sequence. A ClustalX alignment of select plant, insect, and nematode PR5/PR5-L amino acid sequences. Pairwise parameters were 35 for gap opening and 0.75 for gap extension. Shading of sequence labels delineates related sequences. Sequence names within stippled area are miscellaneous plant PR5 sequences including the type sequences for osmotin (OSM) and thaumatin (Thau). Multiple alignment parameters were 15 for gap opening and 0.30 for gap extension. The CG1 through CG8 labels and the associated shaded box on the zeamatin sequence indicate the location of gaps formed in the alignment, with a gap being identified as a space greater than one residue inserted in the majority of the sequences. B Majority rule MrBayes consensus tree generated using MrBayes3.0 and the insect, nematode, and plant PR5-L amino acid alignment. The most likely tree found during MrBayes search was −ln = −21,648.866. Bayesian analyses were performed using a mixed model of evolution for 10 million generations. Numbers above the nodes are N-J parsimony bootstrap/posterior probabilities.
Homology Modeling
Homology modeling of the PR5/PR5-L proteins was performed using Swiss-Model (Guex and Peitsch 1997; Schwede et al. 2003). Using DeepView version 3.7 the sequence of interest was opened in Swiss-Model and used as the target in a Blastp search of the protein data bank (pdb; a database of 3-D protein structure data) database. The pdb file for the best match was downloaded and used as a threading template for the query sequence. Finally, the sequence fit was submitted to the Swiss-Model homology server for energy minimization. The quality of the model produced from this alignment was determined from the root mean square (RMS) deviation value of the position of amino acids in the modeled protein to the paired amino acid in the template. The model was also analyzed for violations of main chain Phi/Psi dihedral bond angle ratios and backbone/side chain and side chain/side chain steric conflicts.
Results and Discussion
Identification of Animal PR5-L Sequences
Both in-house and GenBank insect EST databases were mined for the presence of cDNA clones that encoded antimicrobial proteins. Clones producing a protein with significant similarity to known plant PR5s were discovered in in-house databases from cDNA libraries of three insect orders: Coleoptera, Hemiptera, and Hymenoptera (Table 2). Other sequences were obtained from a previously reported PR5-L in the desert locust Schistocerca gregaria (Orthoptera) (Brandazza et al. 2004) and from another Coleopteran, Biphyllus lunatus, for which EST data were deposited in GenBank (Theodorides et al. 2002). Although the PR5-L sequences were found in four insect orders and represent a multigene family in at least two of these (Coleoptera and Orthoptera), no homologues were found in the Diptera (including Aedes and Anopheles [mosquitos] and Drosophila species for which whole-genome sequence data are available and EST projects are extensive). Also, extensive EST databases in honey bee, Apis mellifera L. (representing the most advanced genomics effort in the Hymenoptera), and silkworm (Bombyx mori L., representing the Lepidoptera) were devoid of PR5-L sequences. This was previously observed by Brandazza et al. (2004) and confirmed by our analysis. Caenorhabditis PR5-L sequences were previously reported from genome sequence and from EST data of two separate species: C. elegans and C. briggsae (Brandazza et al. 2004). No PR5-related proteins were identified in searches that included EST and genomic data from fish, mammals, and birds.
Comparison of Animal PR5-Ls
Although nucleic acid identity among all PR5/PR5-L sequences was very low (data not shown), amino acid identity was >35% for all but two sequences and, therefore, suitable for phylogenetic and structural comparisons (Table 2). For this reason all subsequent comparisons were performed using amino acid sequences. A sequence alignment was constructed from the animal PR5-Ls and Zea mays zeamatin, a related plant PR5 for which structural data were available (Fig. 1). Two of the animal sequences, PR5-L1 from D. abbreviatus and Cel_PR5-L7 from C. elegans, were highly divergent from the rest of the sequences (Table 2, showing percentage identity, and Fig. 1, showing poor alignment) and were thus removed from further comparison. In the alignment (Fig. 1), insect sequences were highly divergent in the first 44 amino acids including the gap spaces, whereas the nematode sequences were highly conserved and most similar to the Tci-PR5-L1 (T. citricida) in this region. It is also notable that all sequences from coleopterans lacked the first and last cysteine residues.
The alignment resulted in six variable domains due to insertions/deletions (labeled G1–G6 in Fig. 1). Five of these six domains were from insertions in at least two of the insect sequences. One of these, G2, represented a four-amino acid insertion in both the zeamatin and the orthopteran sequences (Glu-Arg-Glu-Ser in zeamatin and Sgr-PR5-L1 and Glu-Arg-Glu-Arg in Sgr-PR5-L2). The sixth gap, G4, resulted from a large insertion of ∼39 amino acids in the nematode Cel-PR5-L3 sequence but was variable in length among the other sequences (Fig. 1A). The most likely tree generated using MrBayes was generated from the amino acid alignment using zeamatin as the outgroup (Fig. 1B). Figure 1B contains two distinct animal clades stemming from the node separating them from a common animal ancestor: insects and nematodes. The gene phylogeny was similar to the organism phylogeny (i.e., all coleopteran sequences were monophyletic and all orthopteran sequences were monophyletic) with the exception of the grouping of the hemipteran, T. citricida PR5-L sequence (Tci-PR5-L1) with the hymenopteran aphid-parasitoid, L. testaceipes, (Lte-PR5-L1). Current insect phylogeny groups Hymenoptera with the Coleoptera in the Endopterygota superorder and Hemiptera in the related but distinct Hemipteroid assemblage (Paraneoptera group) (information taken from the Tree of Life web project; http://tolweb.org/tree?group=Neoptera&contgroup=Pterygota). Despite this organism relationship, the PR5-L protein from L. testaceipes (a hymenopteran) was more closely related to the T. citricida (Hemipteran) sequence than to any of the coleopteran PR5-Ls. If these are indeed pathogen defensive proteins (noting that activity of these proteins in animals has yet to be determined), the nature of the pathogens that threaten the species could drive evolution of the PR5-L sequences to a greater degree than phylogeny. It is likely that T. citricida and L. testaceipes would be exposed to similar pathogens since they have such an intimate association (L. testaceipes parasitizes T. citricida). Therefore, it is possible that the need to defend against similar pathogens could provide the evolutionary pressure to keep the sequences of these PR5-Ls more similar than to those of more evolutionarily related organisms that exist in different environments (the coleopterans listed have a subterranean larval stage, whereas the aphid and aphid parasitoid do not).
The paraphyletic relationship of the insect sequences with the nematode sequences, as well as among the insect groups, suggests a single-gene inheritance of PR5-Ls within the insects and within each insect order followed by gene duplication within these groups. However, this relationship may not hold for all PR5-L gene family members since they are not necessarily all represented in the limited insect EST sequence data. Complete genome data have allowed identification of potentially all PR5-L sequences from C. elegans but only partial representation of the C. briggseae PR5-L gene family. In one case, two C. briggseae PR5-Ls (Cbr-PR5-L4 and Cbr-PR5-L5) group with a single C. elegans sequence (Cel-PR5-L6), suggesting continued duplication of these genes after separation of these two species. Since hyperdivergence of antimicrobial peptides (generally fewer than 50 amino acids) is well documented (Nicolas et al. 2003; Vanhoye et al. 2003; Duda et al. 2002), evolutionary pressure for the divergence of other antimicrobials such as PR5-Ls could also be expected for the same reason: adaptation to diverse and continually evolving pathogens.
Comparison of Plant and Animal PR5s/PR5-Ls
A more complete phylogenetic comparison of all related PR5-L proteins among plants and animals was performed by aligning every PR5-L identified in the sequence data from entire genomes of one eudicot (A. thaliana) and one monocot (O. sativa) with those from the nematode C. elegans (includes those discovered from a related nematode C. briggsase) and the insect sequences (Fig. 2). The comparison also included two PR5-L sequences from Pinus taeda (loblolly pine, a gymnosperm), the type sequences for thaumatin (from Thaumatococcus danielli) and osmotin (from Nicotiana tabaci), and one sequence from Triticum aestivum (wheat, a monocot) that represents a described structural variant of the PR5-Ls (Rebmann et al. 1991). The sequence alignment was similar to the animal sequence alignment (compare Fig. 1A with Fig. 2A), with conserved regions separated by variable length gaps. A total of eight variable gap regions were identified, six of which matched those identified in the animal sequence alignment (Fig. 1). The new gaps were CG3 and CG5. The CG3 gap represents a relatively large region (∼15 residues) that was variable only among the plant sequences. The CG5 gap resulted from a proline and an arginine present in only five monocot sequences.
The phylogeny from this alignment is provided with the bootstrap values and posterior probabilities for the branching points as determined both by N-J bootstrap and Bayesian analyses (Fig. 2). This phylogeny shows a paraphyletic grouping of plant PR5s with at least 10 distinct clades, with each clade containing at least one A. thaliensis and one O. sativa sequence (Fig. 2B). In this comparison, the animal proteins grouped into a single clade stemming from the node at which they separate from plant sequences, with the closest relationship to plant sequences in clades 6 and 7. A phylogenetic interpretation would be that the animal PR5-L genes diverged from a single ancestor to the plant sequences that was the ancestor to clades 6 and 7.
The paraphyletic distribution of the plant PR5-L sequences supports a model where a common ancestor to the monocots and the eudicots had 10 PR5-L genes (with the divergence of monocots and dicots corresponding to 130 million to 240 million years ago [Crane et al. 1995; Wolfe et al. 1989]), each of which gave rise to one of the distinct clades observed in the plant PR5-L phylogeny where the clades are defined as separate groups containing at least one member from O. sativa and A. thaliana. Then in the case of clades containing multiple members from a single species, gene duplication events continued to occur throughout eudicot and moncot evolution. Chromosomal locations of each A. thaliana and O. sativa gene were indicated in the gene designations on the cladogram and show that PR5 genes were on 10 of the 12 O. sativa chromosomes and on 4 of the 5 A. thaliana chromosomes. Chromosome locations suggest that subsequent duplication of members within these clades resulted from interchromosomal and intrachromosomal duplications. In O. sativa, clades 3, 5, 6, 7, and 8 have members from multiple chromosomes, whereas A. thaliana has members from multiple chromosomes in only clades 1, 6, 7, and 9. Tandem duplications were observed for each species, with O. sativa chromosome 12 containing six PR5 genes within a cluster (clade 5) that contained only two other small (<50-amino acid) hypothetical proteins (mapping location obtained from The TIGR Rice Genome Database; http://www.tigr.org/tdb/e2k1/osa1). Arabidopsis also had tandemly duplicated PR5 genes on chromosomes 1 (At1g75030, At1g75040, and At1g75050) and 4 (At4g36000 with At4g36010 and At4g 38660 with At4g38670) as determined using mapping data obtained from the TIGR Arabidopsis thaliana database (http://www.tigr.org/tdb/e2k1/ath1). Gene duplication within clades was an asymmetric occurrence in comparisons both within and between A. thaliana and O. sativa (Fig. 2B). In O. sativa, there were nine members of clade 5, while A. thaliana only had a single representative. In A. thaliana, however, there were five members of clade 9, while O. sativa only had one. In each of these cases the increase in gene number was only partly the result of tandem gene duplication.
Antimicrobial activity comparisons of members from these groups may provide interesting insight into the relationship of plant-pathogen interactions and duplication of specific PR5 genes. One hypothesis is that the greatest amplification of a gene clade occurred because that clade contained PR5-Ls most active against pathogens impacting that plant group. Interestingly, the cogrouping of the two type species from both osmotin (Ntabacum OSM) and thaumatin (Tdaniellii Thau) into a single clade that contains both osmotin-like (At4g11650) and thaumatin-like sequences indicates that the phylogenetic relationship of the PR5-L sequences does not support the separate nomenclature for these proteins and argues for a single name such as PR5.
Predicted Structural Similarities of Plant and Animal PR5s/PR5-Ls
Further similarities among the PR5/PR5-L proteins from plants and animals were observed by comparative homology modeling of protein structures. Comparative modeling was possible because the three-dimensional structures for both the original thaumatin, osmotin, and an osmotin-related antimicrobial peptide, zeamatin, are available (de Vos et al. 1985; McPherson and Weickmann 1990; Batalia et al. 1996; Koiwa et al. 1999; Lay et al. 2003; Min et al. 2004). Of those plant PR5 proteins for which structural data were available, blast search analysis showed that insect PR5-Ls were most similar to either zeamatin or osmotin (Table 2). Comparative modeling was performed using the Swiss-Model server for initial structural alignments. The animal PR5-Ls produced structures similar to those of their plant counterparts, with minor steric and bond angle conflicts that did not invalidate the potential models as close approximations (Table 3). The few conflicts in side chain-side chain or side chain-backbone and Psi:Phi dihedral angle ratios mapped to either loop or loop/β-sheet intersections (Fig. 3A), where such conflicts are often found in acceptable structural models since these are the most variable regions among comparisons of homologous proteins (Lesk 2003). Our data indicate that the core structure of insect sequences is highly conserved with that for the plant proteins, as are the spatial relationships of the three domains that make up the PR5 structure (Fig 3B). These are domain I, a core 10–11 stranded ß-sheet core; domain II, a set of disulfide-rich loops exposed on the surface of the protein juxtaposed with domain I to form a surface cleft in the protein; and domain III, a second, smaller loop region containing two disulfide bonds.
Three-dimensional structure of zeamatin with labeled domains and regions of divergence from homology model of animal PR5-Ls. A Ribbon structure for zeamatin protein. Red indicates the regions were the structural predictions for the animal sequences cannot be reliably determined. The quality of the model was considered unreliable in regions where violations in any of the Phi:Psi dihedral angle ratios or steric hindrances between backbone/side groups and side-groups/side groups were detected (see Table 3). B Three-dimensional ribbon structure of zeamatin showing structural domains: blue—domain I, a core 10–11 stranded ß-sheet core; red—domain II, a set of disulfide-rich loops exposed on the surface of the protein juxtapositioned with domain I to form a surface cleft (labeled as acidic cleft) in the protein; and green—domain III, a second smaller loop region containing two disulfide bonds. C Zeamatin ribbon structure with the eight gap regions identified in the PR5/PR5-L alignments (Fig. 2A) mapped to the structure. Red identifies the variable regions labeled CG1, CG2, CG4, and CG6 that are variable among all phylogenetic groups. Blue delineates the variable regions that have a consistent insert in at least two of the insect sequences (CG7 and CG8). Orange signifies a region that is variable among the plant sequences (CG3) and pink identifies a region that has a two-base insert (proline and arginine) only in monocot sequences (CG5).
The finding that the core structure (domain I) of all PR5/PR5-Ls remained conserved among plant and insects allowed mapping of the conserved and variable regions identified in the ClustalX alignment (Fig. 2) to a structural model for a plant PR5. Similar mapping was observed for all plant sequences and zeamatin was used for visualization (Fig. 3). The variable regions CG1, GC2, CG4, and CG6, variable among all of the phylogenetic groups (Fig. 2), map specifically to loop regions on the structure. The variable regions, which have a consistent insert in at least two of the insect sequences, CG7 and CG8, map to a loop-α-helix border in domain II and a loop region in domain I, respectively. Finally, two plant-variable regions, CG3 and CG5, both map to domain I, with CG3 comprising a loop region and CG5, the monocot proline and arginine insert, mapping to a ß-strand exposed to the surface of PR5 on the opposite side of the molecule from a well-characterized cleft. The side group of this arginine protrudes from the protein surface and is only present in zeamatin, thaumatin, and four O. sativa proteins (Os03g45960, Os03g46060, and Os03g46070), all of which are members of clade 5 (Fig. 2B).
Mapping of all the variable regions to predicted loop structures, with the exception of CG5, further supports the validity of the predicted core structure for the animal PR5-L sequences. The conservation of sequence similarity across diverse taxa and the continued increase in gene copy number during more recent evolution suggest an ancient (prior to the separation of plants and animals) and current important role for this gene family. If antimicrobial properties are observed for the animal PR5-Ls, the diversity of sequence will provide an excellent source of data for comparative genomics studies linking sequence, structure, and function.
Limited research on the antifungal activity of PR5s suggests membrane permeability as the mode of action (Abad et al. 1996; Roberts and Selitrennikoff 1990). However, the mechanism by which this occurs is unknown. Direct pore formation by PR5 proteins (especially zeamatin) has been dismissed for three primary reasons (Koiwa et al. 1997): (1) membrane permeabalization is not inhibited at 4°C, (2) PR5 proteins share none of the features that are normally associated with pore-forming proteins that breach the membrane (Osmond et al. 2001; Batalia et al. 1996; Koiwa et al. 1999), and (3) osmotically stabilizing mannitol does not protect cells from the osmotic imbalance caused by one member of the PR5 family (PR5d). Although there is strong evidence that at least one PR5, osmotin, interacts with phosophomannoprotein components of the cell wall (Ibeas et al.2000) and that this interaction is necessary for osmotin activity, this does not preclude direct pore formation as a subsequent step in permeabilization. A two-step process of first binding receptors and subsequent pore formation is a documented mechanism for numerous pore-forming protein toxins (Hurley and Misra 2000; Hong et al. 2002). Interestingly, the core β-sheet structure of zeamatin (and present in all PR5/PR5-L models) shares structural similarity to the cytolysin sticholysin II, a pore-forming toxin from the sea anemone Stichodactyla helianthus. This similarity was first mentioned by Hong et al. (2002). Sticholysin II exists as a free monomer in solution but forms a tetrameric membrane-permeating pore upon association specifically with sphingomyelin-containing membranes (De et al. 1998). The predicted mass of mature sticholysin II is 19,284 D and is slightly smaller than most PR5s, with MW ∼22,000 Da. However, it is larger than a group of antifungal plant PR5s belonging to clade five (Fig. 1B) and typified by a Triticum aestivum protein, PWIR2, that has a MW of 15,673 Da (Rebmann et al. 1991). This clade also contained osmotin and zeamatin. Furthermore, when structural homology modeling was performed with PWIR2, acceptable structural homology models were obtained when this protein was modeled to either zeamatin or the sticholysin II (Fig. 4). It is unclear whether this represents a possible functional relationship between sticholysin II and PWIR2 (and similar clade 7 proteins) or whether it demonstrates artifactual similarities that may arise when modeling relatively small proteins that share a common motif, i.e., multiple acceptable structural models that could be formed through homology modeling that do not have biological significance. In any case, the second of the three reasons for excluding direct pore formation by PR5s is no longer valid: some PR5 proteins share similar structure (albeit not sequence similarity) to known pore-forming protein toxins. Whatever the cause, the structural relationship of PWIR2 with the stycholysin structure offers enticing comparative data that could be the basis for functional studies of PR5/PR5-L-membrane interactions. The first reason for dismissing direct pore formation by PR5s is also in question since biological membranes are not necessarily crystalline at 4°C. Evidence from pore-forming proteins indicates that temperature sensitivity may be a function of pore-forming protein stability, not membrane fluidity (Zitzer et al. 2000). The stabilization of the PR5/PR5-L structure with multiple disulfide bonds, a characteristic not shared by many other pore-forming peptides, may impair greater stability at 4°C. These insights suggest the need to consider pore formation by at least some PR5s as a testable hypothesis.
Structural homology modeling of the T. aestivum PWIR2 protein to both the stycholysin II (left) and the zeamatin (right) structures. PWIR2 modeled and color-coded to show the three main structural domains of PR5s. Yellow: core domain I; blue, domain II; green, domain III. Red zeamatin) indicates the location of poor model structure predictions due to RMS (root mean square data for spatial deviation of aligned amino acid pairs) values for residue relationships between zeamatin and animal PR5-Ls.
The phylogenetic analysis of the relationship of the PR5-L sequences from insects, nematodes, and plants indicates a strong evolutionary pressure to conserve sequence and structure of the protein. Associated with this pressure is the continued increase in copy number of these genes during evolution in both plants and animals. Sequence conservation between plants and insects was also noted for antimicrobial plant defensins and an antimicrobial drosomycin peptide from Drosophila that share 40% identity (Fehlbaum et al. 1994). Antimicrobial peptides are considered part of an organism’s innate immunity and there is convincing evidence that signal transduction pathways that trigger innate immunity can be traced back to a common genetic ancestor in plants and animals (Baker et al. 1997). This similarity led to the statement that “...it is a provocative thought that innate immunity in both plants and animals may have evolved from common ancestral modules that have been used to protect against infection for more than 1 billion years of evolution. (Hoffmann et al. 1999). Our analyses show that the phylogenetic and structural relationship of insect, nematode, and plant PR5-L proteins support this ancient relationship and provide insight into the continued nature of the evolutionary adaptation of these proteins.
References
Abad LR, D’Urzo MP, Liu D, Narasimhan ML, Reuveni M, Zhu JK, Niu X, Singh NK, Hasegawa PM, Bressan RA (1996) Antifungal activity of tobacco osmotin has specificity and involves plasma membrane permeabilization. Plant Sci 118:11–23
Anand A, Zhou T, Trick HN, Gill BS, Bockus WW, Muthukrishnan S (2003) Greenhouse and field testing of transgenic wheat plants stably expressing genes for thaumatin-like protein, chitinase and glucanase against Fusarium graminearum. J Exp Bot 54:1101–1111
Baker B, Zambryski P, Staskawicz B, Nesh-Kumar SP (1997) Signaling in plant–microbe interactions. Science 276:726–733
Batalia MA, Monzingo AF, Ernst S, Roberts W, Robertus JD (1996) The crystal structure of the antifungal protein zeamatin, a member of the thaumatin-like, PR-5 protein family. Nat Struct Biol 3:19–23
Bendtsen JD, Nielsen H, von HG, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Brandazza A, Angeli S, Tegoni M, Cambillau C, Pelosi P (2004) Plant stress proteins of the thaumatin-like family discovered in animals. FEBS Lett 572:3–7
Cheong NE, Choi YO, Kim WY, Bae IS, Cho MJ, Hwang I, Kim JW, Lee SY (1997) Purification and characterization of an antifungal PR-5 protein from pumpkin leaves. Mol Cells 7:214–219
Chu KT, Ng TB (2003a) Isolation of a large thaumatin-like antifungal protein from seeds of the Kweilin chestnut Castanopsis chinensis. Biochem Biophys Res Commun 301:364–370
Chu KT, Ng TB (2003b) Mollisin, an antifungal protein from the chestnut Castanea mollissima. Planta Med 69:809–813
Crane PR, Friis EM, Raunsgaard-Pedersen K (1995) The origin and early diversificaiton of angiosperms. Nature 374:27–33
de Vos AM, Hatada M, van der WH, Krabbendam H, Peerdeman AF, Kim SH (1985) Three-dimensional structure of thaumatin I, an intensely sweet protein. Proc Natl Acad Sci USA 82:1406–1409
De LR, V, Mancheno JM, Lanio ME, Onaderra M, Gavilanes JG (1998) Mechanism of the leakage induced on lipid model membranes by the hemolytic protein sticholysin II from the sea anemone Stichodactyla helianthus. Eur J Biochem 252:284–289
Duda TF Jr, Vanhoye D, Nicolas P (2002) Roles of diversifying selection and coordinated evolution in the evolution of amphibian antimicrobial peptides. Mol Biol Evol 19:858–864
Fehlbaum P, Bulet P, Michaut L, Lagueux M, Broekaert WF, Hetru C, Hoffmann JA (1994) Insect immunity. Septic injury of Drosophila induces the synthesis of a potent antifungal peptide with sequence homology to plant antifungal peptides. J Biol Chem 269:33159–33163
Franco OL, Rigden DJ, Melo FR, Grossi-De-Sa MF (2002) Plant alpha-amylase inhibitors and their interaction with insect alpha-amylases. Eur J Biochem 269:397–412
Gómez-Leyva JF, Blanco-Labra A: (2001) Bifunctional a–amylase/trypsin inhibitor activity previously ascibed to the 22 KDa TL protein, resided in a contaminant protein of 14 KDa. J Plant Physiol 158(2):177–183
Green PJ (1995) Reversible jump Markov chain Monte Carlo compuation and Bayesian model determinations. Biometrika 82:711–732
Grenier J, Potvin C, Trudel J, Asselin A (1999) Some thaumatin-like proteins hydrolyse polymeric beta-1,3-glucans. Plant J 19:473–480
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Hastings MK (1970) Monte Carlo sampling methods using Markov chains and their applications. Biometrika 57:97–109
Hejgaard J, Jacobsen S, Svendsen I (1991) Two antifungal thaumatin–like proteins from barley grain. FEBS Lett 291:127–131
Hoffmann JA, Kafatos FC, Janeway CA, Ezekowitz RA (1999) Phylogenetic perspectives in innate immunity. Science 284:1313–1318
Hong Q, Gutierrez-Aguirre I, Barlic A, Malovrh P, Kristan K, Podlesek Z, Macek P, Turk D, Gonzalez-Manas JM, Lakey JH, Anderluh G (2002) Two-step membrane binding by Equinatoxin II, a pore-forming toxin from the sea anemone, involves an exposed aromatic cluster and a flexible helix. J Biol Chem 277:41916–41924
Hu X, Reddy AS (1997) Cloning and expression of a PR5-like protein from Arabidopsis: inhibition of fungal growth by bacterially expressed protein. Plant Mol Biol 34:949–959
Huelsenbeck JP, Ronquist F (2003) MrBayes 3: Baysian inference under mixed models. Bioinformatics 19:1572–1574
Hurley JH, Misra S (2000) Signaling and subcellular targeting by membrane-binding domains. Annu Rev Biophys Biomol Struct 29:49–79
Huynh QK, Borgmeyer JR, Zobel JF (1992) Isolation and characterization of a 22 kDa protein with antifungal properties from maize seeds. Biochem Biophys Res Commun 182:1–5
Ibeas JI, Lee H, Damsz B, Prasad DT, Pardo JM, Hasegawa PM, Bressan RA, Narasimhan ML (2000) Fungal cell wall phosphomannans facilitate the toxic activity of a plant PR-5 protein. Plant J 23:375–383
Ibeas JI, Yun DJ, Damsz B, Narasimhan ML, Uesono Y, Ribas JC, Lee H, Hasegawa PM, Bressan RA, Pardo JM (2001) Resistance to the plant PR-5 protein osmotin in the model fungus Saccharomyces cerevisiae is mediated by the regulatory effects of SSD1 on cell wall composition. Plant J 25:271–280
Koiwa H, Kato H, Nakatsu T, Oda J, Yamada Y, Sato F (1997) Purification and characterization of tobacco pathogenesis-related protein PR-5d, an antifungal thaumatin-like protein. Plant Cell Physiol 38:783–791
Koiwa H, Kato H, Nakatsu T, Oda J, Yamada Y, Sato F (1999) Crystal structure of tobacco PR-5d protein at 1.8 A resolution reveals a conserved acidic cleft structure in antifungal thaumatin-like proteins. J Mol Biol 286:1137–1145
Larget B, Simon D (1999) Markov chain Mente Carlo algorithms for the baysian analysis of phylogenetic trees. Mol Biol Evol 16:111–120
Lay FT, Schirra HJ, Scanlon MJ, Anderson MA, Craik DJ (2003) The three-dimensional solution structure of NaD1, a new floral defensin from Nicotiana alata and its application to a homology model of the crop defense protein alfAFP. J Mol Biol 325:175–188
Lesk AM (2003) Introduction to protein architecture. Oxford University Press, New York
Li R, Wu N, Fan Y, Song B (1999) Transgenic potato plants expressing osmotin gene inhibits fungal development in inoculated leaves. Chin J Biotechnol 15:71–75
McPherson A, Weickmann J (1990) X-ray analysis of new crystal forms of the sweet protein thaumatin. J Biomol Struct Dyn 7:1053–1060
Metropolis N, Rosenbluth AW, Rosenbluth MN, Teller AH, Teller E (1953) Equations of state calculations by fastr computing machines. J Chem Phys 21:1087–1091
Min K, Ha SC, Hasegawa PM, Bressan RA, Yun DJ, Kim KK (2004) Crystal structure of osmotin, a plant antifungal protein. Proteins 54:170–173
Ng TB (2004) Antifungal proteins and peptides of leguminous and non-leguminous origins. Peptides 25:1215–1222
Nicolas P, Vanhoye D, Amiche M (2003) Molecular strategies in biological evolution of antimicrobial peptides. Peptides 24:1669–1680
Osmond RI, Hrmova M, Fontaine F, Imberty A, Fincher GB (2001) Binding interactions between barley thaumatin-like proteins and (1,3)-beta-D–glucans. Kinetics, specificity, structural analysis and biological implications. Eur J Biochem 268:4190–4199
Rebmann G, Mauch F, Dudler R (1991) Sequence of a wheat cDNA encoding a pathogen–induced thaumatin-like protein. Plant Mol Biol 17:283–285
Roberts W, Selitrennikoff CP (1990) Zeamatin, an antifungal protein from maize with membrane-permeabilizing activity. J Gen Microbiol 136:1771–1778
Salzman RA, Koiwa H, Ibeas JI, Pardo JM, Hasegawa PM, Bressan RA (2004) Inorganic cations mediate plant PR5 protein antifungal activity through fungal Mnn1- and Mnn4-regulated cell surface glycans. Mol Plant Microbe Interact 17:780–788
Schimoler-O’Rourke R, Richardson M, Selitrennikoff CP (2001) Zeamatin inhibits trypsin and alpha-amylase activities. Appl Environ Microbiol 67:2365–2366
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Singh NK. Handa AK, Hasegawa PM, Bressan RA (1985) Protein associated with adaptation of cultured tobacco cells to NaCl. Plant Physiol 79:126–137
Singh NK, Bracker CA, Hasegawa PM, Handa AK, Buckel S, Hermodson MA, Pfankoch E, Regnier FA, Bressan RA (1987) Characterization of osmotin. A thaumatin-like protein associated with osmotic adaptation in plant cells. Plant Physiol 85:529–536
Svensson B, Fukuda K, Nielsen PK, Bonsager BC (2004) Proteinaceous alpha-amylase inhibitors. Biochim Biophys Acta 1696:145–156
Swofford DL (2003) PAUP*: Phylogenetic analysis using parsiomony (*and other methods). Sinauer Associates, Sunderland, MA
Theodorides K, De RA, Gomez-Zurita J, Foster PG, Vogler AP (2002) Comparison of EST libraries from seven beetle species: towards a framework for phylogenomics of the Coleoptera. Insect Mol Biol 11:467–475
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Trudel J, Grenier J, Potvin C, Asselin A (1998) Several thaumatin-like proteins bind to beta-1,3-glucans. Plant Physiol 118:1431–1438
van der WH, Loeve K (1972) Isolation and characterization of thaumatin I and II, the sweet-tasting proteins from Thaumatococcus daniellii Benth. Eur J Biochem 31:221–225
Vanhoye D, Bruston F, Nicolas P, Amiche M (2003) Antimicrobial peptides from hylid and ranin frogs originated from a 150-million-year-old ancestral precursor with a conserved signal peptide but a hypermutable antimicrobial domain. Eur J Biochem 270:2068–2081
Vigers AJ, Roberts WK, Selitrennikoff CP (1991) A new family of plant antifungal proteins. Mol Plant Microbe Interact 4:315–323
Vigers AJ, Wiedemann S, Roberts WK, Legrand M, Selitrennikoff CP, Fritig B (1992) Thaumatin-like pathogenesis-related proteins are antifungal. Plant Sci 83:155–161
Vu L, Huynh QK (1994) Isolation and characterization of a 27-kDa antifungal protein from the fruits of Diospyros texana. Biochem Biophys Res Commun 202:666–672
Wang H, Ng TB (2002) Isolation of an antifungal thaumatin-like protein from kiwi fruits. Phytochemistry 61:1–6
Wolfe KH, Gouy Y-W, Sharp PM, Li W-H (1989) Date of the monocot-dicot divergence estimated from chloropast DNA seqeunce data. Proc Natl Acad Sci USA 86:6201–6205
Woloshuk CP, Meulenhoff JS, Sela-Buurlage M, van den Elzen PJ, Cornelissen BJ (1991) Pathogen-induced proteins with inhibitory activity toward Phytophthora infestans. Plant Cell 3:619–628
Ye XY, Wang HX, Ng TB (1999) First chromatographic isolation of an antifungal thaumatin-like protein from French bean legumes and demonstration of its antifungal activity. Biochem Biophys Res Commun 263:130–134
Zhu B, Chen TH, Li PH (1996) Analysis of late-blight disease resistance and freezing tolerance in transgenic potato plants expressing sense and antisense genes for an osmotin-like protein. Planta 198:70–77
Zitzer A, Harris JR, Kemminer SE, Zitzer O, Bhakdi S, Muething J, Palmer M (2000) Vibrio cholerae cytolysin: assembly and membrane insertion of the oligomeric pore are tightly linked and are not detectably restricted by membrane fluidity. Biochim Biophys Acta 1509:264–274
Acknowledgments
The authors would like to acknowledge Dr. Phat Dang, Bryan Backlaski, and Kimberly Poole for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
[Reviewing Editor: Dr. Rafael Zardoya]
Rights and permissions
About this article
Cite this article
Shatters, R.G., Boykin, L.M., Lapointe, S.L. et al. Phylogenetic and Structural Relationships of the PR5 Gene Family Reveal an Ancient Multigene Family Conserved in Plants and Select Animal Taxa. J Mol Evol 63, 12–29 (2006). https://doi.org/10.1007/s00239-005-0053-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-005-0053-z