Abstract
Introduction
Deep learning–based MRI reconstruction has recently been introduced to improve image quality. This study aimed to evaluate the performance of deep learning reconstruction in pediatric brain MRI.
Methods
A total of 107 consecutive children who underwent 3.0 T brain MRI were included in this study. T2-weighted brain MRI was reconstructed using the three different reconstruction modes: deep learning reconstruction, conventional reconstruction with an intensity filter, and original T2 image without a filter. Two pediatric radiologists independently evaluated the following image quality parameters of three reconstructed images on a 5-point scale: overall image quality, image noisiness, sharpness of gray–white matter differentiation, truncation artifact, motion artifact, cerebrospinal fluid and vascular pulsation artifacts, and lesion conspicuity. The subjective image quality parameters were compared among the three reconstruction modes. Quantitative analysis of the signal uniformity using the coefficient of variation was performed for each reconstruction.
Results
The overall image quality, noisiness, and gray–white matter sharpness were significantly better with deep learning reconstruction than with conventional or original reconstruction (all P < 0.001). Deep learning reconstruction had significantly fewer truncation artifacts than the other two reconstructions (all P < 0.001). Motion and pulsation artifacts showed no significant differences among the three reconstruction modes. For 36 lesions in 107 patients, lesion conspicuity was better with deep learning reconstruction than original reconstruction. Deep learning reconstruction showed lower signal variation compared to conventional and original reconstructions.
Conclusion
Deep learning reconstruction can reduce noise and truncation artifacts and improve lesion conspicuity and overall image quality in pediatric T2-weighted brain MRI.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Magnetic resonance imaging (MRI) is increasingly used in evaluating children’s brain because, different from computed tomography, it entails no radiation exposure and provides superior soft tissue contrast. However, MRI is an inherently slow technique; therefore, parameters should be adjusted to maximize the image quality at a limited scanning time, especially in children who cannot stay still for a long time and frequently need to be sedated for examinations. Many techniques, such as parallel imaging, partial Fourier, and compressed sensing, have been developed to reduce scan time and are used in practice. However, signal-to-noise and/or spatial resolution is often compromised [1, 2].
Recently, deep learning techniques have been applied in various medical imaging fields. In particular, deep learning–based MRI reconstruction has recently been introduced to improve image quality while decreasing computational power and reconstruction time. One of the novel deep learning–based MRI reconstructions (AIR™ Recon DL, GE Healthcare) has become commercially available. Instead of traditional reconstructions using the Fourier transform, this novel deep learning–based MRI reconstruction takes raw k-space data as input and uses a convolutional neural network trained on millions of pairs of low- and high-quality images to directly produce high-fidelity images as output. Recent clinical studies using the prototype of AIR Recon DL have demonstrated improved image quality in brain imaging [3, 4] and orthopedic [5, 6], cardiac [7,8,9], prostate [10, 11], and peripheral nerve MRI [12].
However, to the best of our knowledge, all studies have been conducted in adults, and no study has been conducted regarding the usefulness of DLR in pediatric brain imaging. For children, such image quality improvement techniques are necessary because of their smaller voxel size and relatively tight scan times compared to adults.
Therefore, our study aimed to evaluate the performance of newly developed deep learning reconstruction (DLR) in pediatric brain T2-weighted images by comparing with the conventional filtered and original T2-weighted images.
Materials and methods
Our Institutional Review Board approved this retrospective study and waived the requirement for informed consent.
Study population
From February 2021 to March 2021, 109 consecutive brain MR examinations in the 3.0 T brain MRI (SIGNA Premier, GE Healthcare) with axial T2-weighted sequence were used in this study. Two children underwent brain MRI twice. Therefore, a total of 107 Asian children (51 boys and 56 girls; mean age, 8.5 years [age range, 2 months–18 years]) were included. These patients were imaged for a variety of clinical indications: initial work-up for symptoms (seizure [n = 9], headache [n = 8], developmental delay [n = 2], focal neurologic deficit [n = 2], microcephaly or macrocephaly [n = 4], scalp or skull lesions [n = 3], evaluation of multiple anomalies [n = 4], miscellaneous [n = 5]) or follow-up for tumor (n = 31), or follow-up for Moyamoya disease (n = 12), congenital vascular anomaly (n = 3), hypoxic–ischemic brain injury (n = 8), metabolic disease (n = 4), congenital brain malformation (n = 4), neurocutaneous syndrome (n = 2), and other conditions (n = 6).
Of the 107 patients, 36 (33.6%) showed focal abnormalities on MRI. The specific types of neuropathologic abnormalities evaluated in the 36 patients were as follows: brain tumor (n = 11), focal infarction (n = 10), focal parenchymal lesion of metabolic disease (n = 5), focal developmental lesion (n = 4), intracranial hemorrhage (n = 2), and miscellaneous (n = 4).
All of these lesions were annotated on the Picture Archiving and Communication System (PACS) by one radiologist (L.S.B. with 7 years of experience in radiology) prior to subsequent evaluation of lesion conspicuity.
Magnetic resonance imaging acquisition
MRI was performed using a 3-T MRI system (SIGNA Premier, GE Healthcare) with a 48-channel head coil. Axial T2-weighted sequences were obtained using the following parameters: repetition time/echo time, 3386–5294/102–105 ms; flip angle, 142°; field of view, 180–220 mm; matrix sizes, 360 × 260 or 416 × 300; slice thickness, 3–4 mm; slice spacing, 0–1 mm; number of acquisitions, 1; and echo-train length, 13–15. The scanning time was 1 min 50 s, on average.
Axial T2-weighted brain MRI was reconstructed using the three different reconstruction modes: DLR (AIR™ Recon DL), conventional reconstruction with intensity filter A (little sharpening, some smoothing) reconstruction, and original T2 image without a filter. In DLR, users can modify the noise reduction factor. In this study, we employed a high noise reduction factor.
A 50 mg/kg dose of oral chloral hydrate was used to sedate uncooperative children in our institution.
Qualitative analysis
To qualitatively compare image quality among the three reconstruction modes, two pediatric radiologists (C. Y. H. and K. S. H., with 1 and 7 years of experience, respectively) evaluated the following seven image quality parameters of the three reconstructed images on a Likert 5-point scale: overall image quality, image noisiness, sharpness of gray–white matter differentiation, truncation artifact, motion artifact, cerebrospinal fluid (CSF) and vascular pulsation artifacts, and lesion conspicuity. The reviewers evaluated all image series on the PACS database. No restrictions were applied to window level setting adjustments regarding time or ability to scroll through the images. The two reviewers were blinded to patient information, including the patient’s disease and scan parameters that would identify the type of sequence. Reviewers were also asked to report if artifacts that were not present in the conventional or original reconstructions were observed in DLR.
The overall image quality, image noisiness, and sharpness of gray–white matter differentiation were assessed as follows: 1, unacceptable; 2, poor; 3, acceptable; 4, good; and 5, excellent or ideal.
Three parameters related to image artifacts (truncation, motion, and pulsation artifacts) were evaluated as follows: 1, unreadable motion artifact, images of non-diagnostic quality; 2, severe artifact, images degraded but interpretable; 3, moderate artifact with some, but not severe, effect on diagnostic quality; 4, minimal artifact, no effect on diagnostic quality; and 5, no artifact.
Truncation artifact, also known as Gibbs artifact, refers to a series of parallel lines at the interface with abrupt and intense signal changes. Motion artifacts are caused by random motion during the imaging sequence, resulting in blurring and ghosting of the image. Pulsation artifacts indicate ghost images caused by periodic motion, such as pulsating flow of the internal carotid artery or venous sinuses and pulsating CSF flow [13].
For 36 patients with pre-annotated lesions, the lesion conspicuity was scored as follows: 1, unable to see; 2, blurry but visualized; 3, acceptable; 4, good; and 5, excellent.
Quantitative analysis
One radiologist (K.S.H. with 7 years of experience in radiology) performed a quantitative analysis on the PACS database. An axial brain image at the level of the basal ganglia was selected, and 10 regions of interest (ROIs) of the same size (5–100 mm2) were placed, with particular attention paid to avoid inclusion of adjacent non-parenchymal structures (blood vessels, sulci, and cisterns) and patient-to-patient size adjustments. Ten ROIs were placed as follows: four in the deep gray matter (bilateral putamen and thalami), four in the white matter (genu and splenium of the corpus callosum and bilateral centrum semiovale), and two in the CSF (bilateral frontal horns or body of lateral ventricles). To compare the three reconstructions, the mean signal intensity and standard deviations (SDs) were measured for each ROI; subsequently, the measurements were averaged for each tissue type. Signal uniformity was quantified and compared using the coefficient of variation, which is defined as the ratio of the SD to the mean value within the ROIs for each tissue type [14].
Statistical analyses
Continuous variables for the study population are summarized as means and SDs. Categorical variables are summarized as counts and percentages.
Qualitative scores were analyzed statistically using the Friedman test, followed by post hoc Dunn’s pairwise comparisons.
The interobserver agreement between the two radiologists was evaluated using the weighted Cohen kappa (κ) test. A κ value ≤ 0.20 indicated slight agreement; 0.21–0.40, fair agreement; 0.41–0.60, moderate agreement; 0.61–0.80, substantial agreement; and 0.81–0.99, almost perfect agreement.
Quantitative signal uniformity values were compared using one-way repeated measures analysis of variance followed by the Bonferroni post hoc test.
All statistical analyses were performed using SPSS Statistics for Windows version 25.0 (IBM Corp., Armonk, NY, USA). A value of P < 0.05 was considered significant.
Results
The overall image quality, noisiness, and gray–white matter sharpness were significantly better with DLR than with conventional or original reconstructions (all P < 0.001, post hoc Dunn’s pairwise comparisons). The conventional reconstruction showed better overall image quality, noisiness, and gray–white matter sharpness scores compared to the original image (all P < 0.05, except for image noise score of radiologist 1, post hoc Dunn’s pairwise comparisons) (Fig. 1).
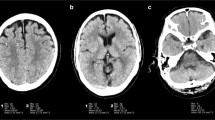
Axial T2-weighted images of a 2-month-old boy with three different reconstructions (a original reconstruction; b conventional reconstruction; c deep learning reconstruction). Deep learning reconstruction shows lower noise and better image quality than the other two reconstructions. Motion artifacts are unaffected with the deep learning reconstruction (arrows)
The DLR had significantly fewer truncation artifacts than the other two reconstructions (all P < 0.001, post hoc Dunn’s pairwise comparisons) for both readers. Truncation artifacts showed no significant difference between the conventional and original reconstructions (Fig. 2).
Magnified views of T2-weighted images of an 8-year-old girl with a history of neonatal meningitis. In the periphery of the brain, truncation artifact (annotated by ovals) is significantly reduced with the deep learning reconstruction (a original reconstruction; b conventional reconstruction; c deep learning reconstruction)
Motion and pulsation artifacts showed no significant difference among three reconstructions (Fig. 1). No identifiable artifacts related to DLR were reported.
Regarding lesion conspicuity, both radiologist reported that DLR scored significantly higher lesion conspicuity than original reconstruction (P = 0.001 and 0.002 for radiologist 1 and 2, respectively, post hoc Dunn’s pairwise comparisons). Lesion conspicuity showed no significant difference between the conventional and original reconstructions for both readers (Figs. 3 and 4).
Magnified views of axial T2-weighted images obtained from a 16-year-old girl with recurred atypical teratoid rhabdoid tumor. There is a recurred mass in the right parietal lobe with extensive peritumoral edema. The margin of the tumor and internal content are more clearly depicted with the deep learning reconstruction (c), compared to the other two images (a original reconstruction; b conventional reconstruction)
Magnified views of axial T2-weighted images obtained from an 18-year-old girl with a history of acute myeloid leukemia. Multifocal hyperintense white matter lesions are more clearly demonstrated with the deep learning reconstruction image (c) than the other two reconstructions (a original reconstruction; b conventional reconstruction)
The weighted Cohen kappa values for assessing interobserver agreement showed almost perfect agreement in overall image quality and moderate agreement in gray–white matter sharpness, noisiness, truncation artifact, and lesion conspicuity scores between the two reviewers.
Table 1 summarizes the results of qualitative analysis and interobserver agreement.
DLR was associated with lower signal variation compared with conventional and original reconstructions, respectively, indicating higher signal inhomogeneity (coefficients of variation were 0.042, 0.055, and 0.060 in white matter; 0.035, 0.045, and 0.050 in gray matter; and 0.020, 0.024, and 0.025 in CSF, for DLR, conventional, and original reconstructions, respectively, all P < 0.001, Bonferroni post hoc test after one-way repeated measures analysis of variance, Table 2).
Discussion
Over the past several years, deep learning technique has been applied to many studies at the cutting edge of MR neuroimaging [15]. The main clinical applications of deep learning in neuroimaging are (1) automated detection or diagnosis [16,17,18], (2) prediction of outcome and disease status [19], (3) improving the image quality [20], and (4) improving the clinical workflow [21]. In particular, MR images often suffer from low signal-to-noise-ratio (SNR) and low contrast-to-noise ratio (CNR) along with image artifacts under the clinical pressure for faster scanning. Therefore, many efforts have been made to improve the image quality of MRI [22,23,24,25,26]. As part of such effort, deep learning reconstructions like AUTOMAP [26] have been proposed to provide a robust image reconstruction of noisy image acquisitions.
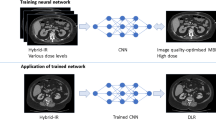
In the conventional MR image reconstruction pipeline, the image is mathematically reconstructed from data in k-space using Fourier transform. Vendor-provided DLR (AIR™ Recon DL, GE Healthcare) of this study uses a convolutional neural network (CNN) to reconstruct the image directly from k-space data. This CNN reconstruction algorithm was trained with a supervised learning approach, using these pairs of images to generate a high-quality image from a low-quality image with truncation artifacts and noise. The DL reconstruction pipeline uses raw k-space data as its input and generates high-fidelity images with higher signal-to-noise ratio, reduced truncation artifacts, and higher spatial resolution as its output [27]. Currently, this algorithm is applied to two-dimensional sequences and offers tunable noise reduction factors to accommodate the user’s preference.
To date, a few studies using this DLR technique have been conducted for evaluating small structures, such as the pituitary gland, prostate, and peripheral nerves, in adult patients [3, 4, 10,11,12]. For example, Kim and Lee et al. reported that thin section images using DLR show enhanced diagnostic accuracy in evaluating the pituitary lesions and surrounding small structures in adult patients [3, 4]. Therefore, they suggested that DLR could be a promising technique for improving the visualization of small structures with increased image quality. Similarly, other groups applied the DLR to prostate and peripheral nerve imaging, where improved visualization of small fine structures is critical. Additionally, several preliminary studies using DLR have been conducted to improve image quality of cardiac MRI of adults [7,8,9]. This is because the image quality of cardiac MRI is often impaired due to narrow time window for capturing moving hearts. Those preliminary studies of cardiac MRI in adults have shown that the image quality of cardiac MRI can be improved with DLR. In a similar context, we believed that DLR could be ideally applied in pediatric brain imaging to improve image quality and diagnostic accuracy. This is because brain structures in children are generally smaller than in adults and MR image quality in children is often impaired due to tighter time window for scanning. As a result, our study demonstrated decreased noise, truncation artifacts, and improved overall image quality of DLR over conventional filtered reconstruction and original images, resulting in improved lesion conspicuity in pediatric T2-weighted brain imaging.
The DLR pipeline was designed to suppress truncation artifacts by estimating truncated high-frequency k-space data [27]. Truncation artifacts are more pronounced at a high-contrast interface. In case of T2-weighted images, truncation artifacts are often observed in the peripheral brain due to the high contrast between hyperintense CSF and relatively hypointense cortex. In conventional reconstruction, software filters are applied to mitigate noise and truncation artifacts, but they result in reduced effective spatial resolution and blurred images. By omitting the software filters, DLR can greatly remove truncation artifacts while decreasing image noise without compromising the image sharpness and effective spatial resolution. Our study demonstrated a significant reduction in truncation artifacts with DLR without impairment of image sharpness.
Regarding the motion and pulsation artifacts, DLR was initially designed to reduce image noise, truncation artifacts, and improve edge sharpness and was not designed to remove other types of artifacts, such as motion, flow, banding, and ghosting. Therefore, there were no significant differences in the motion and pulsation artifact scores in our study. To date, several studies have reported that deep learning techniques could show promising performance in reducing motion artifacts [28,29,30]. Therefore, we hope that the motion and pulsation artifact reduction function will be incorporated into the vendor-provided DLR in the near future.
For practical point of view, one of the promising applications of DLR would be to reduce the scanning time while maintaining MRI image quality in children. The time required to perform an MRI is often limited in children. In that sense, obtaining quality images in a short time is a very necessary task in pediatric MRI. Thus, we believe that further studies are needed to validate the benefit of reducing scan time with DRL.
Our study has some limitations. First, we had a relatively small number of children with lesions covering a wide range of pediatric brain diseases. Nevertheless, our data suggest that DLR could provide a higher overall image quality and lesion conspicuity for pediatric brain diseases. Another limitation was the inability to completely blind the two radiologists to sequence types. During image evaluation, we performed a blind analysis. Despite this, there were noticeable differences between DLR and other reconstructions, which often allowed the blinded readers to distinguish between them. Third, we did not evaluate the whole pediatric brain sequences in our study. Instead, we evaluated only T2-weighted sequences, because T2-weighted sequences are fundamental and representative sequences in brain imaging. We decided so, because the results would have been redundant, if other 2d sequences had been included. In our clinical experience, the same effects of DLR have been confirmed in other applicable 2d brain sequences (e.g., T1-weighted and FLAIR sequences).
Conclusion
The vendor-provided deep learning reconstruction could reduce noise and truncation artifact and improve lesion conspicuity and overall image quality in pediatric T2-weighted brain MRI.
References
Choi KS, Choi YH, Cheon J-E, Kim WS, Kim IO (2020) Application of T1-weighted BLADE sequence to abdominal magnetic resonance imaging of young children: a comparison with turbo spin echo sequence. Acta Radiol 61:1406–1413
Lee JH, Choi YH, Cheon JE, Lee SM, Cho HH, Shin SM, Kim WS, Kim IO (2015) Improved abdominal MRI in non-breath-holding children using a radial k-space sampling technique. Pediatr Radiol 45:840–846
Kim M, Kim HS, Kim HJ, Park JE, Park SY, Kim YH, Kim SJ, Lee J, Lebel MR (2021) Thin-slice pituitary MRI with deep learning-based reconstruction: diagnostic performance in a postoperative setting. Radiology 298:114–122
Lee DH, Park JE, Nam YK, Lee J, Kim S, Kim YH, Kim HS (2021) Deep learning-based thin-section MRI reconstruction improves tumour detection and delineation in pre- and post-treatment pituitary adenoma. Sci Rep 11:21302
Hahn S, Yi J, Lee HJ, Lee Y, Lim YJ, Bang JY, Kim H, Lee J (2021) Image quality and diagnostic performance of accelerated shoulder MRI with deep learning-based reconstruction. AJR Am J Roentgenol.
Koch KM, Sherafati M, Arpinar VE, Bhave S, Ausman R, Nencka AS, Lebel RM, McKinnon G, Kaushik SS, Vierck D, Stetz MR, Fernando S, Mannem R (2021) Analysis and evaluation of a deep learning reconstruction approach with denoising for orthopedic MRI. Radiol Artif Intell 3:e200278
Ogawa R, Kido T, Nakamura M, Nozaki A, Lebel RM, Mochizuki T, Kido T (2021) Reconstruction of cardiovascular black-blood T2-weighted image by deep learning algorithm: a comparison with intensity filter. Acta Radiol Open 10.
van der Velde N, Hassing HC, Bakker BJ, Wielopolski PA, Lebel RM, Janich MA, Kardys I, Budde RPJ, Hirsch A (2021) Improvement of late gadolinium enhancement image quality using a deep learning-based reconstruction algorithm and its influence on myocardial scar quantification. Eur Radiol 31:3846–3855
Muscogiuri G, Martini C, Gatti M, Dell’Aversana S, Ricci F, Guglielmo M, Baggiano A, Fusini L, Bracciani A, Scafuri S, Andreini D, Mushtaq S, Conte E, Gripari P, Annoni AD, Formenti A, Mancini ME, Bonfanti L, Guaricci AI, Janich MA, Rabbat MG, Pompilio G, Pepi M, Pontone G (2021) Feasibility of late gadolinium enhancement (LGE) in ischemic cardiomyopathy using 2D-multisegment LGE combined with artificial intelligence reconstruction deep learning noise reduction algorithm. Int J Cardiol 343:164–170
Wang X, Ma J, Bhosale P, Ibarra Rovira JJ, Qayyum A, Sun J, Bayram E, Szklaruk J (2021) Novel deep learning-based noise reduction technique for prostate magnetic resonance imaging. Abdom Radiol (NY) 46:3378–3386
Park JC, Park KJ, Park MY, Kim MH, Kim JK (2021) Fast T2-weighted imaging with deep learning-based reconstruction: evaluation of image quality and diagnostic performance in patients undergoing radical prostatectomy. J Magn Reson Imaging.
Zochowski KC, Tan ET, Argentieri EC, Lin B, Burge AJ, Queler SC, Lebel RM, Sneag DB (2022) Improvement of peripheral nerve visualization using a deep learning-based MR reconstruction algorithm. Magn Reson Imaging 85:186–192
Park JE, Choi YH, Cheon J-E, Kim WS, Kim I-O, Ryu YJ, Paek M (2018) Three-dimensional radial VIBE sequence for contrast-enhanced brain imaging: an alternative for reducing motion artifacts in restless children. Am J Roentgenol 210:876–882
Kim T-H, Baek M-Y, Park JE, Ryu YJ, Cheon J-E, Kim I-O, Choi YH (2018) Comparison of DWI methods in the pediatric brain: PROPELLER turbo spin-echo imaging versus readout-segmented echo-planar imaging versus single-shot echo-planar imaging. Am J Roentgenol:1352–1358.
Choi KS, Sunwoo L (2022) Artificial intelligence in neuroimaging: clinical applications. Investig Magn Reson Imaging 26:1–9
Di Ieva A, Russo C, Liu S, Jian A, Bai MY, Qian Y, Magnussen JS (2021) Application of deep learning for automatic segmentation of brain tumors on magnetic resonance imaging: a heuristic approach in the clinical scenario. Neuroradiology 63:1253–1262
Tanaka KW, Russo C, Liu S, Stoodley MA, Di Ieva A (2022) Use of deep learning in the MRI diagnosis of Chiari malformation type I. Neuroradiology 64:1585–1592
Zhang Y, Duan Y, Wang X, Zhuo Z, Haller S, Barkhof F, Liu Y (2022) A deep learning algorithm for white matter hyperintensity lesion detection and segmentation. Neuroradiology 64:727–734
Ugga L, Cuocolo R, Solari D, Guadagno E, D’Amico A, Somma T, Cappabianca P, Basso D, de Caro ML, Cavallo LM, Brunetti A (2019) Prediction of high proliferative index in pituitary macroadenomas using MRI-based radiomics and machine learning. Neuroradiology 61:1365–1373
Uetani H, Nakaura T, Kitajima M, Yamashita Y, Hamasaki T, Tateishi M, Morita K, Sasao A, Oda S, Ikeda O, Yamashita Y (2021) A preliminary study of deep learning-based reconstruction specialized for denoising in high-frequency domain: usefulness in high-resolution three-dimensional magnetic resonance cisternography of the cerebellopontine angle. Neuroradiology 63:63–71
Ginat DT (2020) Analysis of head CT scans flagged by deep learning software for acute intracranial hemorrhage. Neuroradiology 62:335–340
Choi KS, You SH, Han Y, Ye JC, Jeong B, Choi SH (2020) Improving the reliability of pharmacokinetic parameters at dynamic contrast-enhanced MRI in astrocytomas: a deep learning Approach. Radiology 297:178–188
Kim KH, Choi SH, Park SH (2018) Improving arterial spin labeling by using deep learning. Radiology 287:658–666
Chung H, Cha E, Sunwoo L, Ye JC (2021) Two-stage deep learning for accelerated 3D time-of-flight MRA without matched training data. Med Image Anal 71:102047
Han Y, Sunwoo L, Ye JC (2020) k-space deep learning for accelerated MRI. IEEE Trans Med Imaging 39:377–386
Zhu B, Liu JZ, Cauley SF, Rosen BR, Rosen MS (2018) Image reconstruction by domain-transform manifold learning. Nature 555:487–492
Lebel RM (2020) Performance characterization of a novel deep learning-based MR image reconstruction pipeline. arXiv preprint arXiv:200806559.
Lee S, Jung S, Jung K-J, Kim D-H (2020) Deep learning in MR motion correction: a brief review and a new motion simulation tool (view2Dmotion). Investig Magn Reson Imaging 24:196–206
Duffy BA, Zhao L, Sepehrband F, Min J, Wang DJ, Shi Y, Toga AW, Kim H, Alzheimer’s Disease Neuroimaging I (2021) Retrospective motion artifact correction of structural MRI images using deep learning improves the quality of cortical surface reconstructions. Neuroimage 230:117756
Kromrey ML, Tamada D, Johno H, Funayama S, Nagata N, Ichikawa S, Kuhn JP, Onishi H, Motosugi U (2020) Reduction of respiratory motion artifacts in gadoxetate-enhanced MR with a deep learning-based filter using convolutional neural network. Eur Radiol 30:5923–5932
Acknowledgements
The authors appreciate Jaeseung Kim Jae for technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and informed consent
Our Institutional Review Board approved this retrospective study and waived the requirement for informed consent.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, SH., Choi, Y.H., Lee, J.S. et al. Deep learning reconstruction in pediatric brain MRI: comparison of image quality with conventional T2-weighted MRI. Neuroradiology 65, 207–214 (2023). https://doi.org/10.1007/s00234-022-03053-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-022-03053-1