Abstract
Bacteria belonging to the genus Vibrio are very common to marine and estuarine environments and are found in association with marine plants and animals. Vibrio parahaemolyticus strains that produce thermostable direct hemolysin (TDH) and TDH-related hemolysin (TRH), toxigenic strains of Vibrio cholerae belonging to the serogroups O1 and O139, and Vibrio vulnificus are regarded as important food-borne pathogens, which represent a serious and growing public health hazard. In this study, we established and validated real-time PCR assays for the detection of enteropathogenic Vibrio strains. In a first step, seafood is investigated for the presence of the three Vibrio species. In case of detection of V. cholerae or V. parahaemolyticus, samples are tested for the presence of the cholera toxin gene (ctxA) or tdh/trh genes, respectively, in a second step. All PCR analyses were performed with the same cycling program. Primer/probe sets were thoroughly tested for limit of detection, inclusivity, exclusivity and performance in the matrix. In an interlaboratory study, kits based on these primer/probe sets were successfully tested with cultural and DNA samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vibrionaceae are Gram-negative, halophilic bacteria found worldwide in marine and estuarine environments. The occurrences of Vibrio bacteria rise during warmer months of the year when the seawater temperature increases [15, 16]. Several pathogenic species are known to lead to diseases by ingestion of contaminated seafood or water or through wound infections after contact with contaminated seawater [8, 13]. The majority of food-borne illness is caused by Vibrio cholerae, Vibrio parahaemolyticus or Vibrio vulnificus (http://www.fao.org/food/food-safety-quality/scientific-advice/jemra/risk-assessments/vibrio0/en/). In addition, it is predicted that the number of infections caused by pathogenic Vibrio spp. will increase due to global warming [8, 13, 20].
The most frequent Vibrio species involved in gastrointestinal infections worldwide is V. parahaemolyticus, which is taken up by consumption of raw or undercooked seafood. V. parahaemolyticus strains, can be found in all types of seafood [9]. Strains producing thermostable hemolysin (TDH) or TDH-related hemolysin (TRH), encoded by tdh and trh, respectively, constitute most of the strains with clinical significance [16, 22]. Hence, techniques that are able to detect tdh and trh genes, and therefore enteropathogenic V. parahaemolyticus, would greatly improve quality control of seafood [26].
Vibrio cholerae strains can be divided into two groups. The first group encompasses toxigenic strains of the serogroups O1 and O139, which cause cholera outbreaks and which affect millions of people in developing countries [21, 25]. The second group, designated as non-O1/non-O139, comprises strains belonging to more than 200 different serogroups that are responsible for sporadic diseases [28]. Food safety measures aim to discriminate between toxigenic and harmless strains using PCR assays that detect the cholera toxin gene ctxA. CTX is the key virulence factor of toxigenic strains and is only rarely present in non-O1 and non-O139 strains [25].
Infections due to V. vulnificus are low in number, however, often result in severe outcomes. In the USA, V. vulnificus accounts for 95% of all seafood-related deaths [23]. Numerous virulence factors of V. vulnificus have been identified of which the pilF gene has been suggested to be useful for the detection of human pathogenic strains [27]. In contrast, up to now not a single marker is available for the discrimination between clinical and environmental strains [5]. Thus, PCR detection of V. vulnificus in food control targets only species-specific genes.
In this study, we developed five real-time PCR assays to detect enteropathogenic Vibrio species in a two-step approach. First, the presence of the enteropathogenic species V. vulnificus, V. parahaemolyticus and V. cholerae in seafood is investigated by species-specific PCR. In cases in which V. parahaemolyticus and V. cholerae are detected, an additional PCR is performed to verify the presence of tdh/trh and ctx positive strains, respectively. All PCR conditions are identical and only two fluorescent channels are needed; therefore no specialized PCR machine is required. In addition, the use of only two sets of fluorescent probes per assay increases the sensitivity of the PCR analysis. All PCR assays were characterized with respect to their sensitivity, specificity and performance in seafood matrices. An interlaboratory study was performed to prove the general and reliable application of the developed Vibrio PCR assays.
Materials and methods
Culture of Vibrio strains and DNA extraction
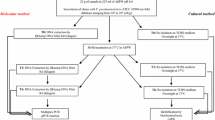
V. cholerae and V. parahaemolyticus were grown on thiosulfate–citrate–bile–sucrose agar (Merck KG, Darmstadt, Germany). V. vulnificus was cultured on salt nutrient agar (SNA) containing 0.5% meat extract, 0.3% peptone and 1% NaCl. LB broth medium (SIFIN Diagnostics GmbH, Berlin, Germany) was inoculated with single colonies and incubated at 37 °C without shaking. Alkaline peptone water (APW; Merck KG, Darmstadt, Germany) was used for enrichment of Vibrio spp. in the matrix. DNA was extracted with the QuickBlue DNA Extraction and Purification Kit (QB-Ex) according to the manufacturer’s instructions (Q-Bioanalytic GmbH, Bremerhaven, Germany).
Primer and probe design
All primers and probes, except primers for V. vulnificus, were developed in this study (Table 1) using the NCBI database and the software PrimerQuest (Integrated DNA Technologies, Inc. Coralville, Iowa, USA). Basic settings included an annealing temperature of 60 °C and an amplicon length of 80 to 240 base pairs. Cross-hybridization of sequences of primers and probes were excluded using nucleotide BLAST [2].
Development of Vibrio-specific real-time PCR assays
All real-time PCR analyses were established based on the TaqMan probe principle [14] using a Roche Light Cycler® 480 equipped with an ABI Prism 7700 Sequence Detector and LightCycler® 480 software (Roche Diagnostics Deutschland GmbH, Mannheim, Germany). Uracil N-glycosylase (UNG) was employed to avoid cross-contamination from previous runs [19]. Plasmid DNA pUC19 was included as an internal amplification control (IAC). A single PCR program was used for all PCR analyses comprising 2 min UNG activation at 50 °C, 10 min initial denaturation at 95 °C, 45 cycles of 20 s for denaturation at 95 °C, 30 s for annealing at 60 °C and 20 s for extension at 72 °C. Reagents were purchased from Life Technologies Corp [TaqMan® Fast Universal PCR Master Mix (2x)], BioRon GmbH (pUC19), Biomers.net GmbH (unlabeled primers), MWG Biotech AG (labeled probes) and PeqLab Biotechnolgie GmbH (dUTP, MgCl2, UNG).
In-/exclusion and limit of detection
Inclusion testing was performed with 46–53 Vibrio target strains with kits for species detection, and with 31 and 13 targets with kits for toxigenic V. parahaemolyticus and V. cholerae detection, respectively (Table S1). The toxigenic V. parahaemolyticus strains tested consisted of nine tdh +/trh + (four trh1 and five trhΨ; Ψ = pseudo gene), 22 tdh −/trh + (four trh1, 17 trh2 and one trhΨ) and no tdh +/trh − genotypes. Exclusion testing was performed with 59 to 108 non-target strains, including 13 non-Vibrio species (Table S2). All strains are listed in tables S1 and S2 and described in table S5. Detection limits were determined by extracting DNA from serial dilutions of fresh overnight cultures of representative Vibrio strains and subsequent PCR analysis. In parallel, each bacterial dilution was grown on agar to determine cell numbers (colony forming units, CFU). Detection limit was defined as the lowest CFU that could be detected by three independently performed PCR assays.
Performance in food matrix
To test the performance of the PCR assays as well as the efficiency of the enrichment procedure, assays were performed with regular trade ware of a seawater fish (Alaska pollock, Theragra chalcogramma), a freshwater fish (striped catfish, Pangasianodon hypophthalmus) and a crustacean (Black tiger prawn, Penaeus monodon). Approximately, 25 g portions of each species were employed as matrix material and diluted tenfold with APW. Each preparation was inoculated with 100 µL aliquots of a series of diluted Vibrio cultures ensuring less than 1 CFU/100 µL in the higher dilutions. After controlled crushing with a paddle blender, samples were incubated overnight at 37 °C without shaking. After DNA extraction, PCR analysis was performed as described above.
Interlaboratory study
In total, five assays comprising three for species-specific (Kit_Vulnificus, Kit_Cholerae and Kit_Parahaemolyticus) and two for toxin gene detection (Kit_Para_Tox for tdh/trh and Kit_Chol_Tox for ctxA), were evaluated. All participating laboratories (Table S3) received their materials from Q-Bioanalytic, including kits, matrix samples (i.e., portions of Pangasius filets), Vibrio cultures (including a control without any bacteria) and DNA samples.
Vibrio cultures included V. parahaemolyticus CH443 (tdh/trh negative), V. cholerae NCTC 4711 (ctx negative), V. vulnificus VN-0016 and V. alginolyticus ATCC 14582. Analysis of cultured samples was initiated by inoculating 10 g of matrix material with 1 mL of Vibrio culture (100–200CFU/mL) followed by addition of 90 ml of APW and homogenization. After overnight incubation, DNA was extracted as described from 1 ml of homogenate.
DNA samples (2.4–31.0 ng/µl) provided separately were prepared by Q-Bioanalytic from V. parahaemolyticus CH443 (tdh/trh negative), V. parahaemolyticus SZ5367/00 (tdh positive, trh positive), V. cholerae NCTC 4711 (ctx negative), V. cholerae O1 1376 (ctx positive), V. vulnificus VN-0016 and V. alginolyticus ATCC 14582.
PCR analysis was performed by all laboratories from five cultured samples and nine DNA samples resulting in a total of 55 and 99 samples, respectively. Of the cultured samples, 11 were expected to be positive using Kit_Vulnificus, Kit_Cholerae and Kit_Parahaemolyticus, whereas all were expected to be negative for Kit_Para_Tox and Kit-Chol-Tox (Table S4). 33 of the DNA samples were expected to be positive for Kit_Vulnificus, 66 for Kit_Cholerae, 44 for Kit_Parahaemolyticus, 33 for Kit_Para_Tox and 55 for Kit-Chol-Tox (Table S4). Results were statistically evaluated by the National Reference Laboratory (NRL) of the BfR.
Statistical measures
Performance of the assays was assessed by determining the sensitivity and specificity with confidence intervals [10], as well as negative (NPV) and positive (PPV) predictive values, which is a measure of the ratio of the expected negative (and positive) to all results [6] (Table S4). To measure the agreement of results obtained by the laboratories participating in the interlaboratory study with the reference results of Q-Bioanalytic, Cohen’s kappa statistic value κ was applied [1]. κ is a measure between 0 and 1 and indicates the range of no to complete agreement [29]. Corresponding confidence intervals were calculated according to Fleiss et al. [11]. All statistical evaluations were performed with IBM® SPSS Statistics version 21 and Excel 2010.
Statement of human and animal rights
Non-applicable.
Results
Primer and probe design
To develop PCR assays to detect enteropathogenic Vibrio strains, we chose the genes encoding hemolysin/cytolysin as species-specific targets in Kit_Vulnificus (vvhA), Kit_Cholerae (hlyA) and Kit_Parahaemolyticus (tlh). The cholera toxin A gene of V. cholerae and the clinically relevant hemolysin genes of V. parahaemolyticus were chosen as targets for the detection of toxigenic strains with Kit-Chol-Tox (ctxA) and Kit_Para_Tox (tdh, trh1, trh2) (Table 1). For the simultaneous detection of all clinically relevant hemolysin genes of V. parahaemolyticus, we included all primers and probes in the Kit_Para_Tox (Table 2b). To control for PCR inhibition, we adjusted individually the concentration of PCR reagents for all PCR kits and included an IAC (Table 2).
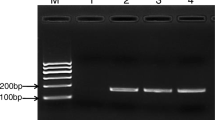
The expected length of 80–240 base pairs for the PCR amplicons was validated using high-percentage agarose gel electrophoresis (data not shown). Real-time PCR with serial dilutions of DNA from selected Vibrio strains consistently resulted in the amplification of the expected PCR products. Standard curves depicted a strong linear correlation between C T values and the DNA concentrations, with a correlation coefficient R 2 of at least 0.98 (Fig. S1).
In-/exclusion and limit of detection
The five different real-time PCR systems detected all but two target strains (Table 3). The two exceptions were strains V. parahaemolyticus VN-0070, which contains a pseudo-trh gene that was not detected by Kit_Para_Tox, and V. cholerae V166/12 (non-O1/non-O139), which does not contain a hylA gene, and therefore could not be detected by Kit_Cholerae. On the other hand, two non-target strains, Grimontia hollisae M106 and M107, which contain a tdh gene, were detected with Kit_Para_Tox (Table 3).
Limits of detection were 0.6 ± 0.3 CFU for V. vulnificus and V. cholerae and 0.7 ± 0.4 CFU for V. parahaemolyticus (Fig. 1). Depending on the presence of either the tdh or a trh gene, 4.5 ± 2.2 or 45.0 ± 22.1 CFU of the toxigenic strains of V. parahaemolyticus was detected (Fig. 1). We did not determine the detection limits for the toxigenic V. cholerae strains.
Internal amplification control (IAC) and performance in food matrix
An IAC allows the identification of false-negative results due to inhibiting substances impairing Taq-polymerase activity. Different labels used for Vibrio (6-FAM) and IAC-specific (HEX) probes enabled parallel monitoring of co-amplified PCR products by employing different filter sets (Fig. S2). Analysis of fish and shellfish obtained from customary trade ware inoculated with serial dilutions of Vibrio cultures, revealed that, in principle, one CFU was sufficient for the detection of Vibrio using enrichment in APW followed by real-time PCR (data not shown). No inhibitory effects, as indicated by complete PCR failure (drop out of the IAC) or shifts in CT values, were observed.
Interlaboratory study
Evaluation of the results from all participating laboratories by the NRL showed a success rate between 89 and 98% for the different Vibrio PCR kits (Fig. 2). Sensitivity and specificity were above 85% in all but one case. NPV and PPV were above 90% with cultural samples generally exhibiting a lower PPV, presumably because of the low (20%) prevalence (Table 4 and Table S4). All κ coefficients calculated for the combined experimental results obtained with cultured and DNA samples were in the range of 0.858–0.898 (Fig. 3), showing an almost perfect concordance with the reference results of Q-Bioanalytic.
Degree of agreement between the expected results (reference results of Q-Bioanalytic GmbH) and the observed results of the laboratories. The degree and strength of agreement is quantified by the κ coefficient [1, 29]. Error bars indicate a confidence interval of 95% [11]. The results of the cultural and of the DNA samples were combined for calculation of the κ coefficient
Discussion
The aim of this study was to develop a system of real-time PCR assays that enables the detection of enteropathogenic V. cholerae, V. parahaemolyticus and V. vulnificus in seafood in a two-step process using the same cycling conditions for all assays. The first step detects strains of the three species with a limit of detection of approximately one CFU per reaction. If V. cholerae or V. parahaemolyticus is present, putative pathogenic strains are identified by performing a second PCR targeting the ctxA gene and tdh/trh genes, respectively. The detection limits for V. parahaemolyticus strains harboring the trh gene were slightly higher. However, CFU numbers should be sufficient as PCR is performed from enrichment cultures.
The selection of genes to identify different species was based on previously published PCR assays and targeted hemolysin/cytolysin genes of all three species. Application of the tlh gene for the identification of V. parahaemolyticus, vvhA for V.vulnificus and hlyA for V. cholerae had been described by Panicker et al. [24] and the same target genes were used for species identification in several other published approaches [12, 17, 26]. We used the V. vulnificus primers and probe described by Campbell and Wright [7] without any further modifications in our protocol as they work with the same cycling profile. Inclusivity and exclusivity confirmed the suitability of the selected target genes for species identification. In the case of V. cholerae, we detected one environmental strain lacking the hemolysin gene; however, all toxigenic strains possess this gene [24, 25] and thus will be detected.
Nucleotide BLAST [2] analysis revealed sequence similarities of the V. cholerae ctxA gene and the V. parahaemolyticus tdh/trh genes to related virulence genes of other bacterial species. The primer and probe sequences targeting the tdh toxin gene of V. parahaemolyticus also hybridize to tdh genes of strains of V. mimicus and V. hollisae. Similarly, the trh primer and probe sequences of V. parahaemolyticus were detected in sequences of the trh genes of strains of Aeromonas veronii, V. anguillarum (formerly Listonella anguillarum) and V. alginolyticus. The trh2 primer and probe sequences, however, are present only in a strain of V. alginolyticus. Few strains of V. alginolyticus can also harbor the ctxA gene, and hence are likely to be detected with the primer–probe system developed for the detection of toxigenic V. cholerae. However, the detection of other bacterial strains harboring tdh/trh or ctxA genes should lead to the same measures concerning contaminated foods as if enteropathogenic Vibrio were present. Of all tested target strains of V. cholerae and V. parahaemolyticus harbouring ctxA and tdh/trh genes, respectively, only V. parahaemolyticus strain VN-0070 was negative, most likely caused by internal nucleotide deletions within the trh pseudogene [4].
All five kits developed on the basis of the primer/probe systems performed well in the interlaboratory study. V. alginolyticus was included as a non-target Vibrio control in this study, as this species is found frequently in seafood. The κ coefficients, which were calculated from the reports of the laboratories, demonstrated that the participants’ results were in strong agreement with the expected results.
In conclusion, we developed a two-step real-time PCR approach that is suitable for the detection of enteropathogenic V. cholerae, V. parahaemolyticus and V. vulnificus, with the advantage that all PCR assays are optimized for the use of the same cycling settings. The PCR assays were validated and tested for their sensitivity, specificity and performance in the food matrix. In addition, the interlaboratory study confirmed the general applicability of the Vibrio PCR kits.
References
Altman DG (1999) Practical statistics for medical research. Chapman & Hall/CRC Press, New York
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Baker-Austin C, Trinanes JA, Taylor NGH, Hartnell R, Siitonen A, Martinez-Urtaza J (2012) Emerging Vibrio risk at high latitudes in response to ocean warming. Nature Clim Change 3:73–77
Bechlars S, Jäckel C, Diescher S, Wüstenhagen DA, Kubick S, Dieckmann R, Strauch E (2015) Characterization of trh2 harbouring Vibrio parahaemolyticus strains isolated in Germany. PLoS One. doi:10.1371/journal.pone.0118559
Bier N, Bechlars S, Diescher S, Klein F, Hauk G, Duty O, Strauch E, Dieckmann R (2013) Genotypic diversity and virulence characteristics of clinical and environmental Vibrio vulnificus isolates from the Baltic Sea region. Appl Environ Microbiol 79:3570–3581
Bier N, Diescher S, Strauch E (2015) Multiplex PCR for detection of virulence markers of Vibrio vulnificus. Lett Appl Microbiol 60:414–420
Campbell MS, Wright AC (2003) Real-time PCR analysis of Vibrio vulnificus from oysters. Appl Environ Microbiol 69:7137–7144
Chuang YC, Yuan CY, Liu CY Lan CK Huang AH (1992) Vibrio vulnificus infection in Taiwan: report of 28 cases and review of clinical manifestations and treatment. Clin Infect Dis 15:271–276
Guidelines on the Application of General Principles of Food Hygiene to the Control of Pathogenic Vibrio Species in Seafood (2010) Codex Alimentarius CCFH, Document CAC/GL 73-2010, Rome, Italy. http://www.fao.org/fao-who-codexalimentarius/standards/list-of-standards/en/. Accessed 26 Oct 2016
Collett D (1999) Modelling binary data. Chapman & Hall/CRC Press, New York
Fleiss JL, Bruce L, Myunghee CP (2003) Statistical Methods for Rates & Proportions. Wiley, New York
Fykse EM, Skogan G, Davies W, Olsen JS, Blatny JM (2007) Detection of Vibrio cholerae by real-time nucleic acid sequence-based amplification. Appl Environ Microbiol 73:1457–1466
Hlady WG, Klontz KC (1996) The epidemiology of Vibrio infections in Florida, 1981–1993. J Infect Dis 173:1176–1183
Holland PM, Abramson RD, Watson R, Gelfand DH (1991) Detection of specific polymerase chain reaction product by utilizing the 5´→3´ exonuclease activity of Thermus aquaticus DNA polymerase PNAS USA 88:7276–7280
Huehn S, Eichhorn C, Urmersbach S, Breidenbach J, Bechlars S, Bier N, Alter T, Bartelt E, Frank C, Oberheitmann B, Gunzer F, Brennholt N, Böer S, Appel B, Dieckmann R, Strauch E (2014) Pathogenic Vibrios in environmental, seafood and clinical sources in Germany. Int J Med Microbiol 304:843–850
Johnson C, Bowers J, Griffitt K, Molina V, Clostio R, Pei S, Laws E, Paranjpye RN, Strom MS, Chen A, Hasan NA, Huq A, Noriea NF 3rd, Grimes DJ, Colwell RR (2012) Ecology of Vibrio parahaemolyticus and Vibrio vulnificus in the coastal and estuarine waters of Louisiana, Maryland, Mississippi, and Washington (United States). Appl Environ Microbiol 78:7249–7257
Jones JL, Lüdeke CH, Bowers JC, DeRosia-Banick K, Carey DH, Hastback W (2014) Abundance of Vibrio cholerae, V. vulnificus, and V. parahaemolyticus in oysters (Crassostrea virginica) and clams (Mercenaria mercenaria) from Long Island sound. Appl Environ Microbiol 80:7667–7672
Lindgren E, Andersson Y, Suk JE, Sudre B, Semenza JC (2012) Monitoring EU emerging infectious disease risk due to climate change. Science 336:418–419
Longo MC, Berninger MS, Harley JL (1990) Use of uracil DNA glycosylase to control carry-over contamination in polymerase chain reactions. Gene 93:125–128
Martinez-Urtaza J, Bowers JC, Trinanes J, DePaola A (2010) Climate anomalies and the increasing risk of Vibrio parahaemolyticus and Vibrio vulnificus illnesses. Food Res Int 43:1780–1790
Mutreja A, Kim DW, Thomson NR, Connor TR, Lee JH, Kariuki S, Croucher NJ, Choi SY, Harris SR, Lebens M, Niyogi SK, Kim EJ, Ramamurthy T, Chun J, Wood JLN, Clemens JD, Czerkinsky C, Nair GB, Holmgren J, Parkhill J, Dougan G (2011) Evidence for several waves of global transmission in the seventh cholera pandemic. Nature 477:462–465
Nishibuchi M, Taniguchi T, Misawa T, Khaeomanee-Iam V, Honda T, Miwatani T (1989) Cloning and nucleotide sequence of the gene (trh) encoding the hemolysin related to the thermostable direct hemolysin of Vibrio parahaemolyticus. Infect Immun 57:2691–2697
Oliver JD (2015) The biology of Vibrio vulnificus. Microbiol Spectr. doi:10.1128/microbiolspec.VE-0001-2014
Panicker G, Call D, Krug M, Bej A (2004) Detection of pathogenic Vibrio spp. in shellfish by using multiplex PCR and DNA microarrays. Appl Environ Microbiol 70:7436–7444
Rivera IN, Chun J, Huq A, Sack RB, Colwell RR (2001) Genotypes associated with virulence in environmental isolates of Vibrio cholerae. Appl Environ Microbiol 67:2421–2429
Robert-Pillot A, Copin S, Himber C, Gay M, Quilici ML (2014) Occurrence of the three major Vibrio species pathogenic for human in seafood products consumed in France using real-time PCR. Int J Food Microbiol 189:75–81
Roig FJ, Sanjuán E, Llorens A, Amaro C (2010) pilF polymorphism-based PCR to distinguish Vibrio vulnificus strains potentially dangerous to public health. Appl Environ Microb 76:1328–1333
Schirmeister F, Dieckmann R, Bechlars S, Bier N, Faruque SM, Strauch E (2014) Genetic and phenotypic analysis of Vibrio cholerae non-O1, non-O139 isolated from German and Austrian patients. Eur J Clin. Microbiol Infect Dis 33:767–778
Sim J, Wright CC (2005) The Kappa statistic in reliability studies: use, interpretation, and sample size requirements. Phys Ther 85:257–268
Acknowledgements
The project VibrioNet was funded by the Federal Ministry of Education and Research (BMBF), Germany, grant no. 01KI1015H. The study is based on the bachelor theses of R. Heuer, J. Weber and J.-H. Schumacher performed at Hochschule Bremerhaven and Q-Bioanalytic GmbH (Germany). For participation in the interlaboratory study, we thank E. Bartelt (Niedersächsisches Landesamt für Verbraucherschutz und Lebensmittelsicherheit, LAVES, Cuxhaven), C. Eichhorn and F. Gunzer (Medizinische Mikrobiologie, Technische Universität Dresden), G. Gerdts (Alfred-Wegener-Institut Helmholtz-Zentrum für Polar- und Meeresforschung, Helgoland), T. Hühn and A. Alter (Institut für Lebensmittelhygiene, Freie Universität Berlin), U. Messelhäusser (Bayerisches Landesamt für Gesundheit und Lebensmittelsicherheit, Oberschleißheim), C. Wind and L. Böhme (Chemisches und Veterinäruntersuchungsamt, Freiburg), N. Matthes, P. Becker and C. Wittmann (Landesamt für Landwirtschaft, Lebensmittelsicherheit und Fischerei Mecklenburg-Vorpommern, Rostock), A.-C. Geuthner and D. Mäde (Landesamt für Verbraucherschutz Sachsen-Anhalt, Halle), H. Monse and M. Müller (Chemisches und Veterinäruntersuchungsamt Rhein-Ruhr-Wupper, Krefeld), O. Sindt and T. Trumpf (Landeslabor Schleswig–Holstein, Neumünster).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
BO, J-OA, EE, JH, CS, and SU are employed with the Company Q-Bioanalytic GmbH (Bremerhaven, Germany). RH and J-HS are former employees of Q-Bioanalytic GmbH.
Electronic supplementary material
Below is the link to the electronic supplementary material.
217_2017_2844_MOESM1_ESM.tif
Standard curves with logarithmic amounts of DNA (0 = 1 ng) plotted against cycle threshold (ct) values: A = Kit_Parahaemolyticus tested with strain CH 445, B = Kit_Para_Tox tested with strains VN-0077 (tdh+), VN-0028 (trh1+) and VN-0084 (trh2+), C = Kit_Cholerae tested with strain VN-0014, D = Kit_Chol_Tox tested with strain 1360 (R4), E = Kit_Vulnificus tested with strain VN-3457 (TIF 483 KB)
217_2017_2844_MOESM2_ESM.tif
Real-time PCR amplification of a serial dilution (120 ng to 1.2 pg) of genomic DNA of the toxic Vibrio cholerae strain 1360. Curves in (A) were obtained with a 6-FAM and TAMRA-labeled probe directed against the ctxA gene. Curves in (B) were obtained with a 5´-HEX and 3´-TAMRA-labeled probe directed against plasmid pUC19 employed as an internal amplification control (TIF 284 KB)
Rights and permissions
About this article
Cite this article
Eschbach, E., Martin, A., Huhn, J. et al. Detection of enteropathogenic Vibrio parahaemolyticus, Vibrio cholerae and Vibrio vulnificus: performance of real-time PCR kits in an interlaboratory study. Eur Food Res Technol 243, 1335–1342 (2017). https://doi.org/10.1007/s00217-017-2844-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-017-2844-z