Abstract
N-acetyltransferase 2 (NAT2) is a well-studied phase II xenobiotic metabolizing enzyme relevant in drug metabolism and cancerogenesis. NAT2 activity is largely determined by genetic polymorphisms in the coding region of the corresponding gene. We investigated NAT2 acetylation status in 1556 individuals from Greenland based on four different single nucleotide polymorphism (SNP) panels and the tagging SNP rs1495741. There was good concordance between the NAT2 status inferred by the different SNP combinations. Overall, the fraction of slow acetylators was low with 17.5 % and varied depending on the degree of Inuit ancestry; in individuals with <50 % Inuit ancestry, we observed more than 25 % slow acetylators reflecting European ancestry. Greenland has a high incidence of tuberculosis, and individual dosing of isoniazid according to NAT2 status has been shown to improve treatment and reduce side effects. Our findings could be a first step in pharmacogenetics-based tuberculosis therapy in Greenland.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the first examples of genetic variation in drug metabolism was the N-acetylation of the anti-tubercular agent isoniazid, and it later turned out that NAT2 is mainly driving this variation (Sim et al. 2008). Enzyme activity is divided into three main categories as slow, intermediate and rapid acetylation, with some studies combining intermediate and rapid acetylation. Furthermore, NAT2 plays an important role in N-acetylation of carcinogenic aromatic amines, and slow acetylation status is a risk factor for urinary bladder cancer (Hein 2002).
The coding exon of NAT2 is polymorphic, and the frequencies of the identified mutations are highly variable across the world (Sabbagh et al. 2011). A common genetic variant tagging NAT2 acetylation status, rs1495741, had several findings in genome-wide association studies in recent years; the allele representing slow acetylation status was associated with risk of urinary bladder cancer (Figueroa et al. 2014; Rothman et al. 2010), lower lipid levels (Teslovich et al. 2010; Willer et al. 2013) and increased skin fluorescence (Eny et al. 2014).
The first humans arrived in the North American Arctic around 6000 years ago, and Greenland was populated by Eskimos in several migration waves (Raghavan et al. 2014). Today the Greenlandic population is characterized by this Inuit ancestry and European ancestry introduced by immigration mainly from Denmark and Norway over the last 300 years. Comparing Greenlandic genomes with genomes from the three major global population groups (Africans, Asians and Europeans) showed that they are quite distinct (Pereira et al. 2015).
Greenland has a high incidence of tuberculosis with annually more than 100 new cases per 100,000 individuals (Statistics Greenland 2014), and isoniazid is widely used in both prevention and treatment of tuberculosis. Individual differences in drug metabolism result in substantial variation of isoniazid blood levels (Mitchell and Bell 1957). Slow NAT2 acetylation status is a risk factor for anti-tuberculosis drug induced liver injury (Wang et al. 2012), a severe side effect of treatment. A study on the pharmacokinetics of isoniazid suggested that standard doses are only appropriate for intermediate acetylators, while a 50 % decrease for slow acetylators and a 50 % increase for rapid acetylators could improve both the safety and the efficacy of treatment (Kinzig-Schippers et al. 2005), and a subsequent clinical trial was able to prove these effects (Azuma et al. 2013). We therefore decided to determine the NAT2 acetylation status of 1556 Greenlandic individuals based on genetic data from the NAT2 region to investigate the potential of NAT2 screening in Greenlandic tuberculosis patients.
Materials and methods
Subjects
In 2013, we recruited individuals from seven of the twelve largest towns in Greenland, with populations ranging from 1181 (Upernavik) to 16,454 (Nuuk) (Statistics Greenland 2013). Together, more than 31,000 individuals of the total population of 57,000 were living in these seven towns. Participants had to be born in Greenland and had to be older than 16 years. Individuals were identified through the Greenlandic Civil Registration System and received a letter inviting them to participate. This study was based on 1556 individuals with genotype information available. Basic demographic information of the participants is given in Table 1. The Commission for Scientific Research in Greenland (approval No. 2013-17) and the Danish Data Protection Agency approved the study. Written and informed consent was given by all participants and by parents for participants under 18 years.
Genotyping, quality control, imputation and haplotype estimation
All 1556 individuals were genotyped on the Illumina Human Omni Express Exome chip. A total of 640,842 SNPs passed quality control; the other SNPs were excluded based on a missing rate >2 %, deviation from Hardy–Weinberg equilibrium (P < 1 × 10−6), minor allele frequency <1 % or discrepancies (P < 1 × 10−6) in allele frequencies between sexes. Subsequently, we imputed unobserved genotypes using phased haplotypes from the integrated phase I release of the 1000 Genomes Project (http://www.1000genomes.org/) with the software packages SHAPEIT (Delaneau et al. 2012) and IMPUTE2 (Howie et al. 2009). This study focused on imputed variants on chromosome 8p22. Accuracy of the estimated allele counts was assessed by SNPTEST (Marchini and Howie 2010), and all reported SNPs had average maximum posterior probabilities of 1 (indicating that there was no uncertainty) and an information measure of 1 (indicating perfect information). Haplotype frequencies were estimated with the expectation–maximization algorithm implemented in PLINK (Purcell et al. 2007).
Assessment of NAT2 acetylation status
We studied acetylation status based on previously described NAT2 SNP panels (Hein and Doll 2012). For the 2-, 3- and 4-SNP panels and the tag-SNP rs1495741, acetylation status was determined by directly counting the number of slow haplotypes/alleles per individual. For the 7-SNP panel, we retrieved acetylation status probabilities for the 21 observed genotype combinations from reference data (Kuznetsov et al. 2009). All slow and intermediate acetylators were assessed with a probability of at least 99.6 %; for rapid acetylation status, all probabilities were >97.9 %.
Determination of Inuit ancestry
The Greenlandic population has become genetically admixed over the last centuries after Northern European visitors settled on the island. Therefore, we investigated admixture with the software tool ADMIXTURE (Alexander et al. 2009) based on a genome-wide set of 43,336 independent SNPs. The results confirmed admixture with two major components referring to Inuit and European ancestry. We estimated the proportions of Inuit and European ancestry of all individuals to investigate differences in NAT2 acetylation status between groups with different levels of Inuit ancestry.
Results
Coverage of NAT2 variants
The current NAT2 nomenclature describes human NAT2 alleles/haplotypes based on combinations of 38 exonic variants (Arylamine N-acetyltransferase Gene Nomenclature Committee 2013). Imputation provided high-accuracy information for 13 of the described SNPs and the tagging SNP rs1495741 (Table 2); seven SNPs were monomorphic in our study group. Genotype frequencies for the 14 SNPs are given in Supplementary Table 1.
Tagging of NAT2 acetylation status and comparison of different SNP panels
We investigated the accuracy of several previously studied NAT2 SNP combinations (Hein and Doll 2012) in assessing NAT2 status, i.e., the tag-SNP rs1495741 and the panels comprised of two SNPs (rs1041983 and rs1801280), three SNPs (rs1799929, rs1799930 and rs1799931), four SNPs (rs1801279, rs1801280, rs1799930 and rs1799931) and seven SNPs (rs1801279, rs1041983, rs1801280, rs1799929, rs1799930, rs1208 and rs1799931). The SNP rs1801279 was monomorphic in our samples, but we kept the naming 4-SNP and 7-SNP panel. Supplementary Table 2 provides the linkage disequilibrium between the seven polymorphic SNPs in all Greenlandic samples.
Table 3 lists inferred slow, intermediate and rapid acetylation status for the 1556 individuals. Concordance between the different panels was high; the 2-SNP and 4-SNP panels agreed perfectly with the reference 7-SNP panel, while slow acetylation status agreed in 98.9 and 97.8 % of individuals for the tag-SNP rs1495741 and the 3-SNP panel, respectively. Supplementary Table 3 additionally provides estimated haplotype frequencies for the six polymorphic SNPs of the 7-SNP panel.
Variation of NAT2 acetylation status in Greenland
Overall, we observed 17.5 % slow acetylators in the study group. The Greenlandic population is admixed with two major components: an Inuit part reaching back to the first migration waves from North America and a Northern European part resulting from interaction with Denmark and Norway in the last 300 years (Table 4). We split the study group in three parts according to the percentage of Inuit ancestry and observed a frequency of 12.2 % slow acetylators among individuals with a high percentage of Inuit ancestry, which is substantially lower than the 25.6 % observed in individuals with <50 % Inuit ancestry.
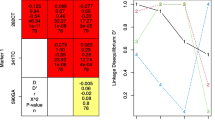
Figure 1 illustrates the distribution of NAT2 acetylation status in Greenland. Slow acetylation status was less frequent in East Greenland, where there was also a larger fraction of Inuit ancestry. This can be explained by the fact that East Greenland was more isolated over the last 300 years, with European visits concentrating on the West Coast; especially, Tasiilaq stood out with an Inuit ancestry of 94 % and only 10 % slow acetylation status.
Pie charts with frequencies of slow, intermediate and rapid acetylation status at the different recruitment sites, and the respective percentage of Inuit ancestry (IA). Figure generated with R (www.r-project.org)
Discussion
Greenland has an admixed population with a dominating Inuit and an additional European component. The frequency of specific genetic variants in Greenland cannot easily be inferred from public databases of the major human populations. The global variation in NAT2 acetylation status has important implications for the metabolism of the anti-tuberculosis drug isoniazid, and the high Greenlandic incidence of tuberculosis motivated us to investigate NAT2 gene variants. The fraction of 17.5 % slow acetylators observed in Greenland is one of the lowest frequencies in the world, driven by the Inuit ancestry. The observed NAT2 acetylation frequencies were close to figures from North-East Asia (Sabbagh et al. 2011), which fits with the current opinion that Greenland was populated from North America and the first humans coming to North America originally came from Siberia (Raghavan et al. 2014).
Even though substantial individual differences in isoniazid metabolism related to NAT2 acetylation status are known for more than 15 years (Parkin et al. 1997) and NAT2 genotypes explain 88 % of the variability in isoniazid clearance (Kinzig-Schippers et al. 2005), assessment of NAT2 status is not routine clinical practice yet. Increased dosages for individuals with rapid acetylation could reduce treatment failure rates. On the other hand, a meta-analysis found a 4.7-fold increased risk of liver injury after anti-tuberculosis treatment in slow acetylators compared with rapid acetylators (Wang et al. 2012), and a lower dosage for these individuals could be indicated. A recent clinical trial investigated a NAT2 genotype-guided treatment of tuberculosis (Azuma et al. 2013) and found that accounting for NAT2 status reduced the rate of early treatment failures in rapid acetylators compared with standard treatment (15 vs. 38 %). In the slow acetylation group, none of the seven patients with genotype-guided lower dosage experienced liver injury during the 6 month of follow-up, whereas there were seven cases of liver injury among the nine patients on standard therapy. The clinical trial was carried out in Japan, and the study group showed a distribution of NAT2 status comparable to Greenland.
Our study relied on a good characterization of NAT2 acetylation status based on the established 7-SNP panel. Additional rare variants associated with slow acetylation are known (Arylamine N-acetyltransferase Gene Nomenclature Committee 2013), and we cannot rule out that these variants and maybe even Inuit-specific variants are present in Greenland. Therefore, the agreement between the inferred NAT2 acetylation status and actual enzyme activity has to be studied before treatment regimens based on genotypes can be introduced. Several methods are available for measuring enzyme activity directly (Hein and Doll 2012), and the results showed good concordance with genetically determined NAT2 acetylation status (Cascorbi et al. 1995; Chen et al. 2006; Grant et al. 1984; Hein and Doll 2012; Kinzig-Schippers et al. 2005; Parkin et al. 1997; Selinski et al. 2011; Smith et al. 1997).
Overall, we provide a first overview of NAT2 acetylation status in Greenland. The frequency of rapid acetylation is particularly high in East Greenland, where the incidence of tuberculosis is highest. Both clinical trial simulations (Gumbo et al. 2007) and an actual clinical trial provided evidence that increased doses of isoniazid can reduce treatment failure. At the advent of personalized medicine, the optimizing of isoniazid dosage based on NAT2 genotypes in Greenland might be a practicable example.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Arylamine N-acetyltransferase Gene Nomenclature Committee (2013) Human NAT2 alleles (haplotypes), Update 19/11/2013. http://nat.mbg.duth.gr/Human%20NAT2%20alleles_2013.htm. Assessed 10 Oct 2014
Azuma J, Ohno M, Kubota R, Yokota S, Nagai T, Tsuyuguchi K, Okuda Y, Takashima T, Kamimura S, Fujio Y, Kawase I (2013) NAT2 genotype guided regimen reduces isoniazid-induced liver injury and early treatment failure in the 6-month four-drug standard treatment of tuberculosis: a randomized controlled trial for pharmacogenetics-based therapy. Eur J Clin Pharmacol 69:1091–1101
Cascorbi I, Drakoulis N, Brockmoller J, Maurer A, Sperling K, Roots I (1995) Arylamine N-acetyltransferase (NAT2) mutations and their allelic linkage in unrelated Caucasian individuals: correlation with phenotypic activity. Am J Hum Genet 57:581–592
Chen B, Zhang WX, Cai WM (2006) The influence of various genotypes on the metabolic activity of NAT2 in a Chinese population. Eur J Clin Pharmacol 62:355–359
Delaneau O, Marchini J, Zagury JF (2012) A linear complexity phasing method for thousands of genomes. Nat Methods 9:179–181
Eny KM, Lutgers HL, Maynard J, Klein BE, Lee KE, Atzmon G, Monnier VM, van Vliet-Ostaptchouk JV, Graaff R, van der Harst P, Snieder H, van der Klauw MM, Sell DR, Hosseini SM, Cleary PA, Braffett BH, Orchard TJ, Lyons TJ, Howard K, Klein R, Crandall JP, Barzilai N, Milman S, Ben-Avraham D, Wolffenbuttel BH, Paterson AD (2014) GWAS identifies an NAT2 acetylator status tag single nucleotide polymorphism to be a major locus for skin fluorescence. Diabetologia 57:1623–1634
Figueroa JD, Ye Y, Siddiq A, Garcia-Closas M, Chatterjee N, Prokunina-Olsson L, Cortessis VK, Kooperberg C, Cussenot O, Benhamou S, Prescott J, Porru S, Dinney CP, Malats N, Baris D, Purdue M, Jacobs EJ, Albanes D, Wang Z, Deng X, Chung CC, Tang W, Bas Bueno-de-Mesquita H, Trichopoulos D, Ljungberg B, Clavel-Chapelon F, Weiderpass E, Krogh V, Dorronsoro M, Travis R, Tjonneland A, Brenan P, Chang-Claude J, Riboli E, Conti D, Gago-Dominguez M, Stern MC, Pike MC, Van Den Berg D, Yuan JM, Hohensee C, Rodabough R, Cancel-Tassin G, Roupret M, Comperat E, Chen C, De Vivo I, Giovannucci E, Hunter DJ, Kraft P, Lindstrom S, Carta A, Pavanello S, Arici C, Mastrangelo G, Kamat AM, Lerner SP, Barton GH, Lin J, Gu J, Pu X, Hutchinson A, Burdette L, Wheeler W, Kogevinas M, Tardon A, Serra C, Carrato A, Garcia-Closas R, Lloreta J, Schwenn M, Karagas MR, Johnson A, Schned A, Armenti KR, Hosain GM, Andriole G Jr., Grubb R III, Black A, Ryan DW, Gapstur SM, Weinstein SJ, Virtamo J, Haiman CA, Landi MT, Caporaso N, Fraumeni JF Jr., Vineis P, Wu X, Silverman DT, Chanock S, Rothman N (2014) Genome-wide association study identifies multiple loci associated with bladder cancer risk. Hum Mol Genet 23:1387–1398
Grant DM, Tang BK, Kalow W (1984) A simple test for acetylator phenotype using caffeine. Br J Clin Pharmacol 17:459–464
Gumbo T, Louie A, Liu W, Brown D, Ambrose PG, Bhavnani SM, Drusano GL (2007) Isoniazid bactericidal activity and resistance emergence: integrating pharmacodynamics and pharmacogenomics to predict efficacy in different ethnic populations. Antimicrob Agents Chemother 51:2329–2336
Hein DW (2002) Molecular genetics and function of NAT1 and NAT2: role in aromatic amine metabolism and carcinogenesis. Mutat Res 506–507:65–77
Hein DW, Doll MA (2012) Accuracy of various human NAT2 SNP genotyping panels to infer rapid, intermediate and slow acetylator phenotypes. Pharmacogenomics 13:31–41
Howie BN, Donnelly P, Marchini J (2009) A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5:e1000529
Kinzig-Schippers M, Tomalik-Scharte D, Jetter A, Scheidel B, Jakob V, Rodamer M, Cascorbi I, Doroshyenko O, Sorgel F, Fuhr U (2005) Should we use N-acetyltransferase type 2 genotyping to personalize isoniazid doses? Antimicrob Agents Chemother 49:1733–1738
Kuznetsov IB, McDuffie M, Moslehi R (2009) A web server for inferring the human N-acetyltransferase-2 (NAT2) enzymatic phenotype from NAT2 genotype. Bioinformatics 25:1185–1186
Marchini J, Howie B (2010) Genotype imputation for genome-wide association studies. Nat Rev Genet 11:499–511
Mitchell RS, Bell JC (1957) Clinical implications of isoniazid blood levels in pulmonary tuberculosis. N Engl J Med 257:1066–1070
Parkin DP, Vandenplas S, Botha FJ, Vandenplas ML, Seifart HI, van Helden PD, van der Walt BJ, Donald PR, van Jaarsveld PP (1997) Trimodality of isoniazid elimination: phenotype and genotype in patients with tuberculosis. Am J Respir Crit Care Med 155:1717–1722
Pereira V, Tomas C, Sanchez JJ, Syndercombe-Court D, Amorim A, Gusmao L, Prata MJ, Morling N (2015) The peopling of Greenland: further insights from the analysis of genetic diversity using autosomal and X-chromosomal markers. Eur J Hum Genet 23:245–251
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Raghavan M, DeGiorgio M, Albrechtsen A, Moltke I, Skoglund P, Korneliussen TS, Gronnow B, Appelt M, Gullov HC, Friesen TM, Fitzhugh W, Malmstrom H, Rasmussen S, Olsen J, Melchior L, Fuller BT, Fahrni SM, Stafford T Jr, Grimes V, Renouf MA, Cybulski J, Lynnerup N, Lahr MM, Britton K, Knecht R, Arneborg J, Metspalu M, Cornejo OE, Malaspinas AS, Wang Y, Rasmussen M, Raghavan V, Hansen TV, Khusnutdinova E, Pierre T, Dneprovsky K, Andreasen C, Lange H, Hayes MG, Coltrain J, Spitsyn VA, Gotherstrom A, Orlando L, Kivisild T, Villems R, Crawford MH, Nielsen FC, Dissing J, Heinemeier J, Meldgaard M, Bustamante C, O’Rourke DH, Jakobsson M, Gilbert MT, Nielsen R, Willerslev E (2014) The genetic prehistory of the New World Arctic. Science 345:1255832
Rothman N, Garcia-Closas M, Chatterjee N, Malats N, Wu X, Figueroa JD, Real FX, Van Den Berg D, Matullo G, Baris D, Thun M, Kiemeney LA, Vineis P, De Vivo I, Albanes D, Purdue MP, Rafnar T, Hildebrandt MA, Kiltie AE, Cussenot O, Golka K, Kumar R, Taylor JA, Mayordomo JI, Jacobs KB, Kogevinas M, Hutchinson A, Wang Z, Fu YP, Prokunina-Olsson L, Burdett L, Yeager M, Wheeler W, Tardon A, Serra C, Carrato A, Garcia-Closas R, Lloreta J, Johnson A, Schwenn M, Karagas MR, Schned A, Andriole G Jr., Grubb R III, Black A, Jacobs EJ, Diver WR, Gapstur SM, Weinstein SJ, Virtamo J, Cortessis VK, Gago-Dominguez M, Pike MC, Stern MC, Yuan JM, Hunter DJ, McGrath M, Dinney CP, Czerniak B, Chen M, Yang H, Vermeulen SH, Aben KK, Witjes JA, Makkinje RR, Sulem P, Besenbacher S, Stefansson K, Riboli E, Brennan P, Panico S, Navarro C, Allen NE, Bueno-de-Mesquita HB, Trichopoulos D, Caporaso N, Landi MT, Canzian F, Ljungberg B, Tjonneland A, Clavel-Chapelon F, Bishop DT, Teo MT, Knowles MA, Guarrera S, Polidoro S, Ricceri F, Sacerdote C, Allione A, Cancel-Tassin G, Selinski S, Hengstler JG, Dietrich H, Fletcher T, Rudnai P, Gurzau E, Koppova K, Bolick SC, Godfrey A, Xu Z, Sanz-Velez JI, Garcia-Prats D, Sanchez M, Valdivia G, Porru S, Benhamou S, Hoover RN, Fraumeni JF Jr., Silverman DT, Chanock SJ (2010) A multi-stage genome-wide association study of bladder cancer identifies multiple susceptibility loci. Nat Genet 42:978–984
Sabbagh A, Darlu P, Crouau-Roy B, Poloni ES (2011) Arylamine N-acetyltransferase 2 (NAT2) genetic diversity and traditional subsistence: a worldwide population survey. PLoS One 6:e18507
Selinski S, Blaszkewicz M, Lehmann ML, Ovsiannikov D, Moormann O, Guballa C, Kress A, Truss MC, Gerullis H, Otto T, Barski D, Niegisch G, Albers P, Frees S, Brenner W, Thuroff JW, Angeli-Greaves M, Seidel T, Roth G, Dietrich H, Ebbinghaus R, Prager HM, Bolt HM, Falkenstein M, Zimmermann A, Klein T, Reckwitz T, Roemer HC, Lohlein D, Weistenhofer W, Schops W, Hassan Rizvi SA, Aslam M, Banfi G, Romics I, Steffens M, Ekici AB, Winterpacht A, Ickstadt K, Schwender H, Hengstler JG, Golka K (2011) Genotyping NAT2 with only two SNPs (rs1041983 and rs1801280) outperforms the tagging SNP rs1495741 and is equivalent to the conventional 7-SNP NAT2 genotype. Pharmacogenet Genomics 21:673–678
Sim E, Lack N, Wang CJ, Long H, Westwood I, Fullam E, Kawamura A (2008) Arylamine N-acetyltransferases: structural and functional implications of polymorphisms. Toxicology 254:170–183
Smith CA, Wadelius M, Gough AC, Harrison DJ, Wolf CR, Rane A (1997) A simplified assay for the arylamine N-acetyltransferase 2 polymorphism validated by phenotyping with isoniazid. J Med Genet 34:758–760
Statistics Greenland (2013) Greenland in Figures 2013. http://www.stat.gl/publ/kl/GF/2013/pdf/Greenland%20in%20Figures%202013.pdf
Statistics Greenland (2014) Greenland in Figures 2014. http://www.stat.gl/publ/en/GF/2014/pdf/Greenland%20in%20Figures%202014.pdf
Teslovich TM, Musunuru K, Smith AV, Edmondson AC, Stylianou IM, Koseki M, Pirruccello JP, Ripatti S, Chasman DI, Willer CJ, Johansen CT, Fouchier SW, Isaacs A, Peloso GM, Barbalic M, Ricketts SL, Bis JC, Aulchenko YS, Thorleifsson G, Feitosa MF, Chambers J, Orho-Melander M, Melander O, Johnson T, Li X, Guo X, Li M, Shin CY, Jin GM, Jin KY, Lee JY, Park T, Kim K, Sim X, Twee-Hee OR, Croteau-Chonka DC, Lange LA, Smith JD, Song K, Hua ZJ, Yuan X, Luan J, Lamina C, Ziegler A, Zhang W, Zee RY, Wright AF, Witteman JC, Wilson JF, Willemsen G, Wichmann HE, Whitfield JB, Waterworth DM, Wareham NJ, Waeber G, Vollenweider P, Voight BF, Vitart V, Uitterlinden AG, Uda M, Tuomilehto J, Thompson JR, Tanaka T, Surakka I, Stringham HM, Spector TD, Soranzo N, Smit JH, Sinisalo J, Silander K, Sijbrands EJ, Scuteri A, Scott J, Schlessinger D, Sanna S, Salomaa V, Saharinen J, Sabatti C, Ruokonen A, Rudan I, Rose LM, Roberts R, Rieder M, Psaty BM, Pramstaller PP, Pichler I, Perola M, Penninx BW, Pedersen NL, Pattaro C, Parker AN, Pare G, Oostra BA, O’Donnell CJ, Nieminen MS, Nickerson DA, Montgomery GW, Meitinger T, McPherson R, McCarthy MI, McArdle W, Masson D, Martin NG, Marroni F, Mangino M, Magnusson PK, Lucas G, Luben R, Loos RJ, Lokki ML, Lettre G, Langenberg C, Launer LJ, Lakatta EG, Laaksonen R, Kyvik KO, Kronenberg F, Konig IR, Khaw KT, Kaprio J, Kaplan LM, Johansson A, Jarvelin MR, Janssens AC, Ingelsson E, Igl W, Kees HG, Hottenga JJ, Hofman A, Hicks AA, Hengstenberg C, Heid IM, Hayward C, Havulinna AS, Hastie ND, Harris TB, Haritunians T, Hall AS, Gyllensten U, Guiducci C, Groop LC, Gonzalez E, Gieger C, Freimer NB, Ferrucci L, Erdmann J, Elliott P, Ejebe KG, Doring A, Dominiczak AF, Demissie S, Deloukas P, de Geus EJ, de FU, Crawford G, Collins FS, Chen YD, Caulfield MJ, Campbell H, Burtt NP, Bonnycastle LL, Boomsma DI, Boekholdt SM, Bergman RN, Barroso I, Bandinelli S, Ballantyne CM, Assimes TL, Quertermous T, Altshuler D, Seielstad M, Wong TY, Tai ES, Feranil AB, Kuzawa CW, Adair LS, Taylor HA Jr., Borecki IB, Gabriel SB, Wilson JG, Holm H, Thorsteinsdottir U, Gudnason V, Krauss RM, Mohlke KL, Ordovas JM, Munroe PB, Kooner JS, Tall AR, Hegele RA, Kastelein JJ, Schadt EE, Rotter JI, Boerwinkle E, Strachan DP, Mooser V, Stefansson K, Reilly MP, Samani NJ, Schunkert H, Cupples LA, Sandhu MS, Ridker PM, Rader DJ, van Duijn CM, Peltonen L, Abecasis GR, Boehnke M, Kathiresan S (2010) Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466:707–713
Wang PY, Xie SY, Hao Q, Zhang C, Jiang BF (2012) NAT2 polymorphisms and susceptibility to anti-tuberculosis drug-induced liver injury: a meta-analysis. Int J Tuberc Lung Dis 16:589–595
Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S, Kanoni S, Ganna A, Chen J, Buchkovich ML, Mora S, Beckmann JS, Bragg-Gresham JL, Chang HY, Demirkan A, Den Hertog HM, Do R, Donnelly LA, Ehret GB, Esko T, Feitosa MF, Ferreira T, Fischer K, Fontanillas P, Fraser RM, Freitag DF, Gurdasani D, Heikkila K, Hypponen E, Isaacs A, Jackson AU, Johansson A, Johnson T, Kaakinen M, Kettunen J, Kleber ME, Li X, Luan J, Lyytikainen LP, Magnusson PK, Mangino M, Mihailov E, Montasser ME, Muller-Nurasyid M, Nolte IM, O’Connell JR, Palmer CD, Perola M, Petersen AK, Sanna S, Saxena R, Service SK, Shah S, Shungin D, Sidore C, Song C, Strawbridge RJ, Surakka I, Tanaka T, Teslovich TM, Thorleifsson G, Van den Herik EG, Voight BF, Volcik KA, Waite LL, Wong A, Wu Y, Zhang W, Absher D, Asiki G, Barroso I, Been LF, Bolton JL, Bonnycastle LL, Brambilla P, Burnett MS, Cesana G, Dimitriou M, Doney AS, Doring A, Elliott P, Epstein SE, Eyjolfsson GI, Gigante B, Goodarzi MO, Grallert H, Gravito ML, Groves CJ, Hallmans G, Hartikainen AL, Hayward C, Hernandez D, Hicks AA, Holm H, Hung YJ, Illig T, Jones MR, Kaleebu P, Kastelein JJ, Khaw KT, Kim E, Klopp N, Komulainen P, Kumari M, Langenberg C, Lehtimaki T, Lin SY, Lindstrom J, Loos RJ, Mach F, McArdle WL, Meisinger C, Mitchell BD, Muller G, Nagaraja R, Narisu N, Nieminen TV, Nsubuga RN, Olafsson I, Ong KK, Palotie A, Papamarkou T, Pomilla C, Pouta A, Rader DJ, Reilly MP, Ridker PM, Rivadeneira F, Rudan I, Ruokonen A, Samani N, Scharnagl H, Seeley J, Silander K, Stancakova A, Stirrups K, Swift AJ, Tiret L, Uitterlinden AG, van Pelt LJ, Vedantam S, Wainwright N, Wijmenga C, Wild SH, Willemsen G, Wilsgaard T, Wilson JF, Young EH, Zhao JH, Adair LS, Arveiler D, Assimes TL, Bandinelli S, Bennett F, Bochud M, Boehm BO, Boomsma DI, Borecki IB, Bornstein SR, Bovet P, Burnier M, Campbell H, Chakravarti A, Chambers JC, Chen YD, Collins FS, Cooper RS, Danesh J, Dedoussis G, de FU, Feranil AB, Ferrieres J, Ferrucci L, Freimer NB, Gieger C, Groop LC, Gudnason V, Gyllensten U, Hamsten A, Harris TB, Hingorani A, Hirschhorn JN, Hofman A, Hovingh GK, Hsiung CA, Humphries SE, Hunt SC, Hveem K, Iribarren C, Jarvelin MR, Jula A, Kahonen M, Kaprio J, Kesaniemi A, Kivimaki M, Kooner JS, Koudstaal PJ, Krauss RM, Kuh D, Kuusisto J, Kyvik KO, Laakso M, Lakka TA, Lind L, Lindgren CM, Martin NG, Marz W, McCarthy MI, McKenzie CA, Meneton P, Metspalu A, Moilanen L, Morris AD, Munroe PB, Njolstad I, Pedersen NL, Power C, Pramstaller PP, Price JF, Psaty BM, Quertermous T, Rauramaa R, Saleheen D, Salomaa V, Sanghera DK, Saramies J, Schwarz PE, Sheu WH, Shuldiner AR, Siegbahn A, Spector TD, Stefansson K, Strachan DP, Tayo BO, Tremoli E, Tuomilehto J, Uusitupa M, van Duijn CM, Vollenweider P, Wallentin L, Wareham NJ, Whitfield JB, Wolffenbuttel BH, Ordovas JM, Boerwinkle E, Palmer CN, Thorsteinsdottir U, Chasman DI, Rotter JI, Franks PW, Ripatti S, Cupples LA, Sandhu MS, Rich SS, Boehnke M, Deloukas P (2013) Discovery and refinement of loci associated with lipid levels. Nat Genet 45:1274–1283
Acknowledgments
We thank all study participants for their cooperation in this study, as well as all individuals involved in collecting and handling of data and samples. We thank Mikael Andersson (data management), Poul Berger, Henrik Hjalgrim, Mikkel Koch, Jacqueline M. Mistry, Helle Møller and Johan E. Navne (all field-work). Sascha Wilk Michelsen and Karen Bjorn-Mortensen are both supported by Commission for Scientific Research in Greenland/Danish Research Council scholarships, and Bjarke Feenstra is supported by an Oak Foundation fellowship. The study was conducted with support from: The Danish Research Council, The Greenlandic Ministry of Education, Church, Culture and Gender Equality, the Maersk foundation (Fonden til lægevidenskabens Fremme), and the Aase and Ejnar Danielsens Foundation.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Geller, F., Soborg, B., Koch, A. et al. Determination of NAT2 acetylation status in the Greenlandic population. Arch Toxicol 90, 883–889 (2016). https://doi.org/10.1007/s00204-015-1501-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-015-1501-1