Abstract
Key message
A novel dwarf cucumber mutant, scp-2, displays a typical BR biosynthesis-deficient phenotype, which is due to a mutation in CsDET2 for a steroid 5-alpha-reductase.
Abstract
Brassinosteroids (BRs) are a group of plant hormones that play important roles in the development of plant architecture, and extreme dwarfism is a typical outcome of BR-deficiency. Most cucumber (Cucumis sativus L.) varieties have an indeterminate growth habit, and dwarfism may have its value in manipulation of plant architecture and improve production in certain production systems. In this study, we identified a spontaneous dwarf mutant, super compact-2 (scp-2), that also has dark green, wrinkle leaves. Genetic analyses indicated that scp-2 was different from two previously reported dwarf mutants: compact (cp) and super compact-1 (scp-1). Map-based cloning revealed that the mutant phenotype was due to two single nucleotide polymorphism and a single-base insertion in the CsDET2 gene that resulted in a missense mutation in a conserved amino acid and thus a truncated protein lacking the conserved catalytic domains in the predicted steroid 5α-reductase protein. Measurement of endogenous hormone levels indicated a reduced level of brassinolide (BL, a bioactive BR) in scp-2, and the mutant phenotype could be partially rescued by the application of epibrassinolide (EBR). In addition, scp-2 mutant seedlings exhibited dark-grown de-etiolation, and defects in cell elongation and vascular development. These data support that scp-2 is a BR biosynthesis-deficient mutant, and that the CsDET2 gene plays a key role in BR biosynthesis in cucumber. We also described the systemic BR responses and discussed the specific BR-related phenotypes in cucumber plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant architecture traits related to height, bushy or compact growth habit are important because of their role in lodging resistance and increase of crop productivity (Peng et al. 1999; Clouse and Sasse 1998; Hedden 2003). For example, in wheat, the reduced height genes (Rht) that are involved in the signaling pathway of the plant hormone gibberellic acid (GA) played a critical role in the ‘green revolution’ in crop production (Peng et al. 1999). In cucumber, a typical vining crop, the compact or bushy plant type may be useful to increase planting density and production (Cramer and Wehner 2000).
Several types of dwarf or compact mutants have been described in cucumber. The first dwarf mutant compact (cp) was identified in two plant introduction lines PI 308915 and PI 308916, in which a recessive mutation led to reduced internode length, which has been fine mapped in a region of cucumber chromosome 4, but not yet been cloned (Kauffman and Lower 1976; Li et al. 2011). Kubicki et al. (1986) identified a second compact mutant, compact-2 (cp-2), in which the dwarf phenotype was required to interact with the ‘bushy’ gene. Compared with the previous two mutants, the EMS-induced cucumber mutant ‘super compact’ (scp) was reported to have the most extremely reduced main stem length, no lateral branches, smaller dark green and wrinkled leaves, and deformed pistils (Niemirowicz-Szczytt et al. 1996). Crienen et al. (2009) reported another compact cucumber mutant, which was apparently different from the previous mutants in terms of genomic locations and an intermediate compact phenotype in heterozygous individuals. More recently, two EMS-induced dwarf mutants were reported. The short internode (si) mutant was shown to be associated with a truncated F-box gene in cucumber (Lin et al. 2016). Another mutant, super compact-1 (scp-1), was due to a mutation in the cytochrome P450 gene CsCYP85A1, which encodes the BR-6-oxidase in BR biosynthesis pathway (Wang et al. 2017). Although many plant height mutants have been reported, the relationship between the dwarf phenotype and phytohormones in cucumber needs further study.
The BRs are a group of plant hormones that play important roles in the regulation of plant architecture (Clouse and Sasse 1998). In Arabidopsis, the most characteristic phenotypes of mutants in genes of BR biosynthesis and signaling are compact plant with dark green leaves; the mutant also exhibit a de-etiolation phenotype when grown in the dark (Bishop 2003). BR biosynthesis mutants can generally be restored by exogenous BR application, but BR signaling mutants are not (Clouse et al. 1996; Li et al. 1996). The first BR biosynthesis-deficient mutant, de-etiolated-2 (det2), was identified based on its prominent phenotypes of constitutive photomorphogenesis and extreme dwarfism (Chory et al. 1991). As compared with wild-type dark-grown Arabidopsis seedlings, det2 seedlings are short, have thick hypocotyls, open and expanded cotyledons, and show development of the primary leaf buds (Chory et al. 1991). Under light, det2 plants are smaller and darker green than wild type, show almost complete male sterility, and have delayed leaf and chloroplast senescence (Li et al. 1996). The DET2 gene encodes a steroid 5α-reductase that acts at the early step in brassinolide (BL, the most bioactive BR) biosynthesis; loss-of-function mutants in DET2 result in reduced endogenous BR levels (Fujioka et al. 1997; Noguchi et al. 1999), and subsequently, defects in cell elongation and vascular differentiation (Clouse et al. 1996).
In addition to Arabidopsis, det2 homologous mutants have also been identified in pea (Pisum sativum), Japanese morning glory (Ipomoea nil) and maize (Zea mays); these mutants are named lk, Uzukobito, and na1, respectively (Suzuki et al. 2003; Nomura et al. 2004; Hartwig et al. 2011). These mutants exhibit typical BR-deficient mutant phenotypes such as extreme dwarfism, dark green leaves, and recovery after BR application. However, the pea mutant lk is not de-etiolated, which was the criterion used in identifying det2 and other BR biosynthesis-deficient mutants (Symons et al. 2002). This indicates that the physiological responses to BR can vary in different plants, and exploring BR-related mutants may reveal novel features that would not be revealed in a limited number of plants. In this study, we identified a spontaneous cucumber mutant that exhibited severe dwarfism, smaller dark green and wrinkled leaves, de-etiolation in the dark, and female sterility. All these phenotypes are very similar to those observed in the cucumber scp and scp-1 mutants (Niemirowicz-Szczytt et al. 1996; Wang et al. 2017). Since its allelic relationship with scp is unknown, and the different genomic locations (scp-1 in chromosome 5 and the gene herein in chromosome 3), we designated the mutation super compact-2 (scp-2). Map-based cloning identified the CsDET2 gene (the cucumber homolog of Arabidopsis DET2) as the candidate gene for Scp-2, and two single-base transitions and a 1-bp insertion were found in the coding sequence (CDS) of the mutant allele. The insertion resulted in a predicted truncated protein that was lacking 29 amino acid residues in the C-terminus of the wild-type protein. Examination of the physiological responses and endogenous hormone levels confirmed that scp-2 was a BR biosynthesis-deficient mutant, and that CsDET2 plays a role in BR action in cucumber plants. To the best of our knowledge, this is the first report on an early BR biosynthesis-deficient mutant in cucumber that leads to severe reduction in endogenous BR levels. Therefore, the scp-2 mutant will be valuable for studying the functions of BR in this species.
Materials and methods
Plant materials, mapping populations, and genetic analysis
The super compact-2 (scp-2 hereafter) mutant was discovered during seed multiplication of the cucumber line AM204 by the USDA-ARS Cucumber Breeding Program in 2014. The wild-type AM204W (W = wild type, WT) was originated from PI 618937 (Jin Chun No. 4) through self-pollination, which is a north China fresh market-type (Chinese Long) cucumber from China. Several segregating populations were developed to investigate the mode of inheritance, and for molecular mapping and cloning of the scp-2 mutant, which are listed in Table 1. An F2 population was developed from AM204H, a wild-type plant that was heterozygous at the scp-2 locus (Scp-2scp-2). Since polymorphism of molecular markers in this population was low, two more F2 populations were developed from crosses of AM204M (recessive homozygous mutant) with AM218 (aka, WI7435B) and Gy14. Only mutant plants from the AM204M × AM218 and AM204M × Gy14 F2 populations were used for fine mapping of the scp-2 locus.
The phenotype of the scp-2 mutant was very similar to the ‘compact’ dwarf mutants (Kauffman and Lower 1976; Li et al. 2011). We conducted morphological comparisons between AM204M and the compact mutant WI7201 (PI 308915). All plant materials were grown in the Walnut Street Greenhouses of the University of Wisconsin-Madison, USA.
Phenotyping and analysis of hypocotyl sections
Phenotypes of the cucumber hypocotyls, cotyledons, leaves, and stems were recorded using an optical camera (60D, Canon, Japan). Hypocotyls from WT and scp-2 mutant plants were manually sectioned longitudinally, then observed, measured, and photographed under a light microscope (BX51-32P02, Olympus, Japan).
BR-related physiological analysis
Seventy-two seeds from the F2 (hybrid AM204H) population were germinated and grown in complete darkness at constant temperature (28 °C) and humidity (70%) for 10 days. The de-etiolated phenotype of the scp-2 mutant was identified by short hypocotyls, open cotyledons, and primary leaf bud development.
For responses of hormone treatments, 10 scp-2 mutant plants were grown in a growth chamber with supplemental lights. When the cotyledons were fully expanded, 100 µL EBR solution (0.2 µM, epibrassinolide, a bioactive BR reagent, Sigma-Aldrich, China) was applied to the shoots once a day until the second true leaf was fully expanded, and the physiological responses to BR treatment in mutant plants were recorded.
DNA and RNA extraction, first-strand cDNA synthesis, and quantitative real-time PCR (qRT-PCR) analysis
Genomic DNAs were extracted from cotyledons of cucumber seedlings following Li et al. (2008). To analyze expression of the CsDET2/Csdet2 genes, total RNA was extracted from roots, hypocotyls, cotyledons, leaves, and male buds from WT and mutant plants. First-strand cDNA was synthesized from total RNA as described previously (Li et al. 2012). PCR was performed in a 96-well plate using an ABI 7500 Fast Real-Time PCR System (Applied Biosystems, USA), with SYBR Green Realtime PCR Master Mix (TaKaRa, China). The amplification was initiated by heating to 94 °C for 10 min, followed by 40 cycles of 94 °C for 5 s and 65 °C for 30 s. The amplification specificity was tested by a dissociation curve (65–90 °C). Three biological and three technical replicates were performed for each gene. The cucumber CsACTIN2 gene was used to normalize the gene expression results. The PCR primers used in these experiments are listed in Table S1.
Molecular mapping, cloning, and candidate gene analysis of the scp-2 locus
Genome-wide SSR markers were selected according to Cavagnaro et al. (2010) and Yang et al. (2012). Bulked segregant analysis (BSA; Michelmore et al. 1991) was performed on two genotypic pools consisting of 10 WT and 10 mutant plants that were selected among 190 individuals from the AM218 × AM204M F2 population. Using 244 genome-wide SSR markers, initial mapping placed the scp-2 in cucumber chromosome 3 followed by linkage analysis in a larger AM218 × AM204M F2 population (only the 900 mutant plants were used, the same below), which allowed to identify five SSR markers co-segregating with the scp-2 gene. For further fine mapping of the gene, a new Gy14 × AM204M F2 population was developed, and the 1500 mutant individuals were used for linkage analysis. The 9930 and Gy14 draft genome sequences were then used for scaffold-based chromosome walking to identify the scp-2 gene following Tan et al. (2015).
The candidate genes in the final genomic interval were analyzed using the Cucumber Genome Database (http://cucumber.genomics.org.cn), and the Arabidopsis homologs were identified by searches of the TAIR database (http://www.arabidopsis.org/). The genomic sequences of the candidate genes from WT and mutant plants, including the approximately 1.7-kb upstream promoter and 1.8-kb downstream sequences, were cloned and sequenced. DNAMAN v6.0 software (http://dnaman.software.informer.com/6.0/) was used to compare the DNA sequences of WT and mutant plants and their deduced protein sequences.
Protein sequence alignment and phylogenetic analysis
Multiple sequence alignment of full-length predicted protein sequences was performed using ClustalW (http://www.ebi.ac.uk/Tools/clustalw2). An unrooted neighbor-joining (NJ) phylogenetic tree was constructed using the MEGA 5.10 software (Tamura et al. 2011) with 1000 bootstrap replications, pair-wise deletion, and a Poisson model.
Measurement of endogenous hormone levels
When the first true leaf was fully expanded, the cotyledons, leaves, and shoots of WT and mutant seedlings grown under the same conditions were harvested to analyze the endogenous IAA, GA (including GA3 and GA4), ABA, Zeatin, and BL (brassinolide) level using HPLC–MS/MS. For the BL measurement, 0.8 g cotyledons, leaves, and shoots (with young leaves and male floral buds) from 24 WT and mutant seedlings were mixed, respectively, and ground to fine powder in liquid nitrogen. The extraction and pretreatment procedures followed Wu et al. (2013). For analysis of other plant hormone, 1.0 g tissues from the same WT and mutant plants were treated as described by Kojima et al. (2009). The HPLC–MS/MS analyses were performed on an Agilent 1290 HPLC (Agilent, USA) coupled with an SCIEX-6500® Qtrap system (A-B, USA). Three technical replicates were conducted for each measurement, and the endogenous levels of plant hormones (ng/g fresh weight) were expressed as the means of three HPLC–MS/MS runs with detectable results.
Results
Origin and phenotypic characterization of the scp-2 mutant
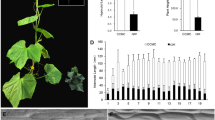
The dwarf mutant was first discovered in the self-pollinated progeny of the cucumber line AM204. The wild-type AM204W was a monoecious, north China type cucumber derived from PI 618937 (Jinchun No. 4). As compared with AM204W, the mutant AM204M plants had a short and inflated hypocotyl, dark green cotyledons, smaller dark green and wrinkled leaves, reduced petioles, and extremely short plant height (Fig. 1a–d). The mutant plants produced normal male flowers (except for the wrinkled corolla) and fertile pollen; the female flowers seemed sterile under greenhouse conditions, although fruits could occasionally be observed on mutant plants under field conditions. These phenotypes were very similar to those reported for the super compact (scp) and super compact-1 mutants (Niemirowicz-Szczytt et al. 1996; Wang et al. 2017). Therefore, before allelism test, the mutant under investigation was designated as super compact-2 (scp-2).
Phenotypic characterization of cucumber AM204W (wild-type plant), scp-2 mutant AM204M, and morphological comparison with cp (compact) mutant (WI7201). a As compared with the AM204W (center), the hypocotyl is severely shortened in scp-2 (left and right), b top view of mutant seedlings showing dark green cotyledons and true leaves with short petioles. c The leaves from five consecutive nodes on the main stem (left to right) of the scp-2 mutant are dark green in color and wrinkled. d Hypocotyl length is reduced in scp-2 (right) as compared to cp (left). e The length of the first two internodes in cp mutant (left) is relatively normal as compared with scp-2 mutant (right). f Top view of mature plants showing wrinkled, dark green leaves in scp-2 mutant (right) as compared with those of cp mutant (left) that appear normal. g–j Longitudinal sections of wild-type (g, h) and scp-2 (i, j) hypocotyls from 10-day old seedlings. Note the disorderly vascular development in scp-2 (i) and the changes in cell shape and size in scp-2 (j). Bar 500 µm in g, i, 200 µm in h, j (color figure online)
The scp-2 mutant was also morphologically similar to the compact mutant (cp) (Li et al. 2011). We compared the cp mutant (WI7201) with the scp-2 mutant (AM204M). We found that both hypocotyl length and seedling height in the scp-2 mutant were shorter than that of the cp mutant (Fig. 1d). They also differed in the internode length. Internodes 1 and 2 of the cp mutant were similar to the WT, but the later (upper) internodes were reduced in length (Li et al. 2011, also observed in this study). In contrast, all internodes in scp-2 mutant plants were severely shortened (Fig. 1d, e). In adult plants, the most obvious difference between the two mutants was the leaf phenotype. The cp mutant was similar to the WT in leaf appearance; however, the scp-2 mutant exhibited dark green and wrinkled leaves throughout the entire growth period (Fig. 1c, f). Genetic mapping results (below) indicated that the scp-2 (in chromosome 3) and cp (in chromosome 4) are two independent loci in the cucumber genome.
The extremely dwarf phenotype is often caused by defective cell elongation. Therefore, we conducted microscopic observation of the hypocotyl of 10-day old seedlings (Fig. 1g–j). The average cell size in the scp-2 mutant was significantly smaller than that of the WT plants (Fig. 1g–j), whereas no obvious difference was observed in the number of cells along the length and cross-sections of hypocotyls in the WT and scp-2 mutant (data not shown). Irregular growth of the vascular system (Fig. 1g, i) and cell shapes (Fig. 1h, j) were also observed in the mutant, which are the typical features of BR-deficient mutants in previous studies.
Mutations in the CsDET2 gene correspond to scp-2
We examined segregation of WT and mutant phenotypes in two F2 populations from self-pollinated AM204H (72 plants) and the cross between AM218 × AM204M. The results are presented in Table 1, which suggested that the mutation was controlled by a single recessive locus, scp-2. Initial marker analysis in the AM218 × AM204M F2 population placed the scp-2 locus to a 2.0-cM region flanked by markers UW040671 and UW040452 on cucumber chromosome 3 (Fig. 2a). A third marker, SSR16667, was co-segregating with the gene. The size of the population (F2B AM218 × AM204M) was then increased to 900 mutant plants for fine mapping of scp-2. Six new markers were mapped to the 2.0-cM region, and the scp-2 gene was located between markers UW040598 and UW040503 that were physically 174-kb apart in the Gy14 draft genome scaffold01225 (Fig. 2b). To resolve the order of the co-segregating markers in this region, we conducted marker analysis in the Gy14 × AM204M F2 population, from which 1500 mutant plants were used for fine mapping. With the cucumber reference genome sequences, all putative SSR and SNP loci in this region were predicted, screened, and the polymorphic markers between Gy14 and scp-2 mutant plant were identified (Fig. 2c). Finally, one SSR marker UW040544, and one SNP marker S433755 delimited the scp-2 locus to a genomic interval of 30.75 kb (Fig. 2d).
Map-based cloning of the super compact-2 (scp-2) gene. a Linkage analysis with genome-wide SSR markers placed the scp-2 locus to a 2.0 cM interval flanked by markers UW040671 and UW040452, and marker SSR16667 is co-segregating with the locus in 190 F2 individuals. b In an enlarged segregating population, scp-2 was fine mapped to a genomic region flanked by markers UW040598 and UW040503 in scaffold01225 of the Gy14 draft genome, and five markers are co-segregating with the scp-2 locus. c The local physical map of Scaffold01225 helps to identify 7 SSR markers that are polymorphic between Gy14 and scp-2 mutant (AM204M). Additional SSR and SNP markers delimit the scp-2 locus to a genomic interval (d) with 6 predicted genes including CsDET2 (e). A 1-bp insertion is identified in CsDET2 gene between AM204W and AM204M (e). “RE” in b, d indicates recombinant events
Genome annotation indicated that there were six predicted genes within this 30.75-kb interval (Fig. 2e) including Csa3G732550 that is predicted to encode a steroid 5-alpha-reductase. The deduced protein sequence showed highest similarity with the Arabidopsis AtDET2 (Table S2); and the cucumber gene was subsequently named CsDET2. The amino acid alignment and phylogenetic analysis with well-studied DET2 proteins revealed that, besides the elongated N-terminal region, CsDET2 had a predicted 5α-steroid reductase domain and all of the conserved binding sites for cofactors and steroid substrate (Figs. S1 and S2). These data indicated that CsDET2 should encode a functional steroid 5α-reductase.
Homologous genomic sequences for the six genes in the interval from the WT and scp-2 mutant were subsequently obtained and analyzed, and only the nucleotide mutations presented in CsDET2 were predicted to result in changes to the deduced protein sequence. The open reading frame (ORF) of the WT CsDET2 gene was 876 bp, which possessed a single exon (Fig. 3a, Fig. S3), and was predicted to encode a protein with 291 amino acid residues (aa, Fig. 3b). In the scp-2 mutant, the genomic sequence showed two single nucleotide mutations and a 1-bp insertion (Fig. 3a, Fig. S3). The first mutation, a C to T transition, would result in a missense mutation (R182W) in the deduced amino acid sequence. The amino acid alignment indicated that R182 in CsDET2 is homologous to R152 in AtDET2, R150 in HsSRD5A1, and R145 in HsSRD5A2 (Fig. S2). The conserved R residue, which was originally identified in the human DET2 ortholog, is important for cofactor binding (Russell and Wilson 1994). The second mutation, a T to G transition would produce another missense mutation (C257W). Furthermore, the 1-bp insertion is predicted to cause a frame-shift mutation that would produce a truncated protein of 262 aa with a loss of 29 aa in the C-terminus as compared with the wild-type (Fig. 3).
Sequence alignment between the wild-type CsDET2 and mutant Csdet2 alleles. a Schematic representation of nucleotide variations between the CsDET2 and Csdet2 alleles. The nucleotide sequences were aligned, and changes predicted to cause missense mutations in the amino acid sequence are underlined. The 1-bp insertion in the Csdet2 coding sequence, which leads to early termination of translation, is indicated by triangle. Asterisks indicate stop codons. b Alignment of the deduced amino acid sequence between CsDET2 and Csdet2. Identical amino acids are highlighted in dark blue. The bar above the sequences indicates the predicted 3-oxo-5α-steroid 4-dehydrogenase domain (pfam02544). Conserved residues that are important for sterol binding are indicated by arrows; asterisks indicate residues important for cofactor binding; a diamond marks the glutamic acid residue shown to be important for human DET2 function. Detailed information about the conserved domains and residues is provided in Figure S1 (color figure online)
The amino acid alignment also showed that the C-terminal region in the DET2 protein is highly conserved in all species examined (Fig. S1). The 3-oxo-5α-steroid 4-dehydrogenase domain, which is critical for enzyme activity, contains the whole C-terminal region of the well-studied DET2 proteins. Moreover, in the missing C-terminal sequence of the cucumber mutant protein, the amino acid residue H268 (H239 in AtDET2) is important for sterol binding, and R283 (R254 in AtDET2) is also related to cofactor binding (Fig. 3b, Fig. S1. Hartwig et al. 2011). These results supported the notion that the CsDET2 allele, Csdet2, in the scp-2 mutant encodes a defective protein. In addition, genome-wide analysis indicated that there is only one copy of the DET2 sequence in the cucumber genome (data not shown). Therefore, the Csdet2 mutant should exhibit a BR biosynthesis-deficient phenotype.
The scp-2 mutant shows reduced levels of endogenous BR
Because DET2 is a rate-limiting enzyme in early steps of BR biosynthesis, the endogenous BR level in scp-2 mutant plants expressing the Csdet2 gene was expected to be altered. Due to their extremely low concentrations, detection of endogenous BRs in plants has always been a challenge (Bajguz 2011). Therefore, we first examined the transcript abundance of CsDET2, and the tissues with relatively high expression levels of the gene were sampled for analysis. We found that, as compared with the root, the expression of CsDET2 was up-regulated in the cotyledons, leaves, and male buds of WT. The highest accumulation of CsDET2 mRNA was in the cotyledon implying that the BR level might be high in this vegetative organ (Fig. 4). Meanwhile, expression of the mutant allele (Csdet2) was up-regulated in the hypocotyl, leaf, and male buds in the mutant line (Fig. 4). As a result, the cotyledons, leaves, and shoots (with young leaves and male floral buds) of WT and scp-2 mutant seedlings were harvested and used to measure the endogenous BR levels.
Expression analysis of the CsDET2 and Csdet2 alleles in cucumber plants. Conserved sequence regions of the two alleles, which exclude the conserved domains, were used to design for the qPCR assay. After sequencing the gene coding sequence, plants with homozygous genotypes were selected. Tissues from roots, hypocotyls, cotyledons, leaves, and male floral flower buds from homozygous SCP-2SCP-2 and scp-2scp-2 plants were used to assay the expression of the CsDET2/Cdet2 alleles. The reference gene, CsACTIN2, was used to normalize the gene expression data. All experiments were repeated in triplicate with independent samples, error bars represent the SE, and asterisks indicate significant difference between the scp-2 and wild-type plants (t test, P < 0.05)
The HPLC–MS/MS analysis indicated that, as compared with the WT, the endogenous BL level in tissues (mixed from cotyledons, leaves, and shoots) of the scp-2 mutant was significantly reduced, and it was below the level of detection in two technical replicates (Table 2). Levels of five other endogenous phytohormones, IAA, ABA, GA3, GA4, and Zeatin, were also assayed in samples from the same plants used for the BL measurements. Interestingly, the levels of both GA3 and GA4 were significantly increased in the scp-2 mutant suggesting a possible antagonistic relationship with BL (Table 2). Therefore, the relationship between BR and GA in the cucumber dwarf mutant needs to be investigated further.
The scp-2 mutant shows systemic changes in BR-related features
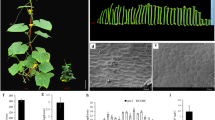
In the scp-2 mutant, the extreme dwarfing, along with the dark green and wrinkled leaves, are typical symptoms of BR biosynthesis deficiency. We tested the etiolation response with the WT and scp-2 mutant seedlings. After 10 days of growth in the dark, the WT seedlings, as expected, had a typical etiolated appearance, with a highly elongated hypocotyl (Fig. 5a) and closed, unexpanded cotyledons (Fig. 5b). However, hypocotyls on the scp-2 seedlings failed to elongate (Fig. 5a), and the mutant seedlings displayed partially open cotyledons and primary leaf bud (Fig. 5c). These results show that BR biosynthesis-deficient mutants in Arabidopsis and cucumber share similar de-etiolation features.
BR physiological responses in cucumber plants. a–c Phenotypes of dark-grown scp-2 and wild-type cucumber seedlings. a Ten-day-old dark-grown (etiolated) wild-type (right) and scp-2 (left) seedlings. b A close-up view of the top of a wild-type seedling shows the two closed cotyledons. c A close-up view of an scp-2 seedling shows the partially open cotyledons and primary leaf bud. d–i Recovery of the scp-2 mutant by exogenous application of EBR. d Wild-type seedling at the two-leaf stage and (e) its first leaf. f scp-2 mutant seedling at the two-leaf stage after EBR application (see “Materials and methods”) and g its first leaf. Note the elongated first leaf petiole, and the shape and color of the first leaf. h scp-2 mutant seedling at the two-leaf stage and i its first leaf (color figure online)
BR-treatment recovery is another feature observed in BR biosynthesis-deficient mutants, so we treated scp-2 seedlings with exogenous EBR (epibrassinolide). The results clearly showed that the color, shape, and petiole length of leaves treated with EBR were restored to wild-type appearance (Fig. 5d–i). Exogenous GA was also used to treat the mutant seedlings, but no obvious responses (plant height or color) were observed (data not shown). Although the continuous application of EBR could not restore the mutant plant to wild-type height (in detail, the EBR treatment had no significant impact on the internode length in the mutant plant), the exogenous BR could, at least partially, rescue the scp-2 mutant in cucumber. It has been reported that BRs appear to be synthesized and function in the same tissue or even within the same cell (Bishop et al. 1996; Shimada et al. 2003; Symons and Reid 2004). This explains why exogenous BR cannot completely complement the mutant with severely reduced levels of endogenous BR. This also implies that scp-2 is deficient in the early step of BR biosynthesis, which blocks endogenous BR production.
Discussion
Cucumber dwarf/compact mutants
Cucumber is an annual vining plant, and indeterminate growth of the shoot produces a single long main stem. The longer vine and growth period mean higher production in the European and Asian cucumber plants. Therefore, plant height in cucumber is not only a plant architecture trait, but also is closely related to the yield. There are many reports describing plant height in cucumber, including determinate habit (de) (Fazio et al. 2003) and dwarf/compact mutants (cp, cp-2, scp, scp-1, and si) (Kubicki et al. 1986; Niemirowicz-Szczytt et al. 1996; Li et al. 2011; Lin et al. 2016; Wang et al. 2017). Among these, the scp and scp-1 mutants was reported to have pronounced dwarfism and typical features related to BR-deficiency. In this study, we identified a mutant similar to the two scp mutants, which shared the dwarf/compact plant architecture, dark green and wrinkled leaves, and female sterility. Unfortunately, the scp mutant was not available, and the relationship between these two mutations could not be confirmed without a test of allelism. The scp-1 locus (CsCYP85A1) has been recently cloned, which was located in cucumber chromosome 5 and encodes a BR-C6-oxidase in the BR biosynthesis pathway. Our work presented herein suggested that scp-2 is a lesion in the CsDET2 gene that results in a defect in BR synthesis. In addition to the appearance of the plants, the BR-related responses in scp-2, such as low endogenous BR level, de-etiolation when grown in the dark, and recovery after exogenous BR application, confirmed that scp-2 is a BR biosynthesis-deficient mutant. Together with the map-based cloning results, the wild-type allele of the scp-2 locus should be CsDET2.
One feature of the scp-2 mutant was a 1-bp insertion in the CsDET2 gene, and the wild-type nucleotide sequence was changed from ‘AAA’ to ‘AAAA’. Since the original source, AM204W, was an inbred line, and scp-2 was discovered in the natural self-pollinated progeny, the mutation can be explained by slipped-strand mispairing (Levinson and Gutman 1987). Both the scp and scp-1 mutations were induced by EMS treatment (Niemirowicz-Szczytt et al. 1996; Wang et al. 2017). Although the scp mutant was not recoverable, based on its similar phenotypes with the scp-1 and scp-2 mutants, both of which are deficiency in same BR biosynthesis pathway (showed in Fig. S4), it is reasonable to speculate that scp could also be a mutation in the BR-related pathway.
The BR response in cucumber
Brassinosteroids (BRs) are a widely distributed class of steroid hormones in plants that play roles in many biological processes including cell expansion, vascular differentiation, photomorphogenesis, male fertility, flowering, senescence, seed germination, and the stress response (Clouse and Sasse 1998; Gudesblat and Russinova 2011). However, the BR responses in cucumber have not been well established. Many studies focused on the BR-induced stress tolerance (Xia et al. 2011; Wang et al. 2012; Li et al. 2013; Wei et al. 2015; An et al. 2016). Fu et al. (2008) indicated that application of EBR could induce parthenocarpic fruit growth, and suggested that BRs play an important role during early fruit development in cucumber. However, a lack of characterized mutants limits the study of the systemic BR response in this species. In this study, the scp-2 mutant exhibited the typical BR-deficient phenotype that included severe dwarfing, dark green and wrinkled cotyledons and leaves, dark-grown de-etiolation, cell elongation and vascular development defects, and recovery after exogenous BR application. All these features, along with reduced endogenous BR levels, confirmed that cucumber shares major conserved BR-related features with Arabidopsis.
However, we found that the endogenous GA level was increased in this BR biosynthesis-deficient mutant. The hormone measurements indicated that the endogenous levels of GA3 and GA4 increased 4.2- and 4.5-fold in the scp-2 mutant, respectively, as compared with WT plants (Table 2). Similarly, the OsGSR1 RNAi transgenic rice line showed a reduced level of endogenous BR and an elevated level of endogenous GA (Wang et al. 2009). GA and BR deficiencies can often result in similar phenotypes, such as dwarfism, reduced seed germination, and delayed flowering, and the GA-deficient mutants also show de-etiolation phenotypes in the dark (Alabadí et al. 2004; Wang et al. 2009). Here, we found that exogenous GA application could not restore the dwarf hypocotyl in scp-2 seedlings. The same result was also shown for Arabidopsis det2, in which hypocotyl elongation was insensitive to GA (Steber and McCourt 2001). In contrast, the GA-deficient mutants show a normal BR response and are partly rescued by BR (Bai et al. 2012). All these results indicated that GA may be not required for the BR response, and BR antagonistically regulates GA levels in cucumber.
GA can induce male flowers and arrest the development of female flowers in cucumber (Atsmon 1968), and this could explain the rare female flowers observed on the scp-2 mutant plant. Interestingly, the male flowers appeared to be normal (except the wrinkled corolla), and the pollen grains are fertile and can be used in backcrosses to heterozygous individuals to maintain the mutant genotype. This is very different from what is known about the relationship between BR and pollen. Pollen are thought to be a rich source of endogenous BRs, and the first BR, brassinolide (BL) was isolated from pollen of Brassica napus (Grove et al. 1979). Moreover, the Arabidopsis BR biosynthetic and signaling mutants show varying degrees of male sterility with reduced pollen number, viability, and release efficiency (Ye et al. 2010). In this study, we used brassinazole (Brz, a BR biosynthesis inhibitor) to treat the wild-type male floral buds, and no obvious changes were detected in the mature stamens and pollen grains (data not shown). Therefore, based on our observations, we propose that the development of male flowers and pollen in cucumber may act in a BR-independent mode, and there could be a different regulation pathway in this organ. We have identified the WUS-AG-SPL/NZZ pathway functions in the development of the cucumber male flower, and down-regulated expression of these genes could explain the sterile male flowers in another cucumber mutant (mango fruit, unpublished data). Since BRs are considered to control male fertility by regulating the expression of genes in the WUS-AG-SPL/NZZ pathway in Arabidopsis (Ye et al. 2010), the different regulation mechanism between BR and male flowers in cucumber should be studied in the future.
Author contribution statement
ZL, YW, and ZG conceived the research and designed the experiments. SL, YW, and ZL identified the mutant and developed the mapping populations and initial mapping of the mutant gene. SH and HN performed fine mapping and cloning of the candidate gene. QT, SW, and ZL participated in genotyping and phenotyping. YW and ZL supervised the experiments and wrote the manuscript. All authors reviewed and approved the final version of the manuscript before submission.
References
Alabadí D, Gil J, Blázquez MA, García-Martínez JL (2004) Gibberellins repress photomorphogenesis in darkness. Plant Physiol 134:1050–1057
An Y, Zhou H, Zhong M, Sun J, Shu S, Shao Q, Guo S (2016) Root proteomics reveals cucumber 24-epibrassinolide responses under Ca(NO3)2 stress. Plant Cell Rep 35:1081–1101
Atsmon D (1968) The interaction of genetic, environmental, and hormonal factors in stem elongation and floral development of cucumber plants. Ann Bot 32:877–882
Bai M, Shang J, Oh E, Fan M, Bai Y, Zentella R, Sun T, Wang Z (2012) Brassinosteroid, gibberellin and phytochrome impinge on a common transcription module in Arabidopsis. Nat Cell Biol 14:810–817
Bajguz A (2011) Brassinosteroids: a class of plant hormone: brassinosteroids—occurrence and chemical structures in plants. Wiley-Interscience, New York, pp 1–27
Bishop GJ (2003) Brassinosteroid mutants of crops. J Plant Growth Regul 22:325–335
Bishop GJ, Harrison K, Jones JDG (1996) The tomato Dwarf gene isolated by heterologous transposon tagging encodes the first member of a new cytochrome P450 family. Plant Cell 8:959–969
Cavagnaro PF, Senalik DA, Yang L, Simon PW, Harkins TT, Kodira CD, Huang S, Weng Y (2010) Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.). BMC Genom 11:569
Chory J, Nagpal P, Peto CA (1991) Phenotypic and genetic analysis of det2, a new mutant that affects light-regulated seedling development in Arabidopsis. Plant Cell 3:445–459
Clouse SD, Sasse JM (1998) Brassinosteroids: essential regulators of plant growth and development. Annu Rev Plant Physiol Plant Mol Biol 49:427–451
Clouse SD, Langford M, McMorris TC (1996) A brassinosteroid-insensitive mutant in Arabidopsis thaliana exhibits multiple defects in growth and development. Plant Physiol 111:671–678
Cramer CS, Wehner TC (2000) Path analysis of the correlation between fruit number and plant traits of cucumber populations. HortScience 35:708–711
Crienen J, Reuling G, Segers B, van de Wal M (2009) New cucumber plants with a compact growth habit. Patent, International publication number WO 2009/059777 A1
Fazio G, Staub JE, Stevens MR (2003) Genetic mapping and QTL analysis of horticultural traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Theor Appl Genet 107:864–874
Fu F, Mao W, Shi K, Zhou Y, Asami T, Yu J (2008) A role of brassinosteroids in early fruit development in cucumber. J Exp Bot 59:2299–2308
Fujioka S, Li J, Choi YH, Seto H, Takatsuto S, Watanabe T, Kuriyama H, Yokota T et al (1997) The Arabidopsis de-etiolated2 mutant is blocked early in brassinosteroid biosynthesis. Plant Cell 9:1951–1962
Grove MD, Spencer GF, Rohwedder WK, Mandava NB, Worley JF, Warthen JD, Steffens GL, Flippen-Anderson JL et al (1979) Brassinolide, a plant growth-promoting steroid isolated from Brassica napus pollen. Nature 281:216–217
Gudesblat GE, Russinova E (2011) Plants grow on brassinosteroids. Curr Opin Plant Biol 14:530–537
Hartwig T, Chuck GS, Fujioka S, Klempien A, Weizbauer R, Potluri DP, Choe S, Johal GS et al (2011) Brassinosteroid control of sex determination in maize. Proc Natl Acad Sci USA 108:19814–19819
Hedden P (2003) The genes of the green revolution. Trends Genet 19:5–9
Kauffman CS, Lower RL (1976) Inheritance of an extreme dwarf plant type in the cucumber. J Am Sci Hortic Sci 101:150–151
Kojima M, Kamada-Nobusada T, Komatsu H, Takei K, Kuroha T, Mizutani M, Ashikari M, Ueguchi- Tanaka M et al (2009) Highly sensitive and high-throughput analysis of plant hormones using MS-probe modification and liquid chromatography-tandem mass spectrometry: an application for hormone profiling in Oryza sativa. Plant Cell Physiol 50:1201–1214
Kubicki B, Soltysiak U, Korzeniewska A (1986) Induced mutations in cucumber (Cucumis sativus L.) V. Compact type of growth. Genet Pol 27:289–298
Levinson G, Gutman GA (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Bio Evol 4:203–221
Li J, Nagpal P, Vitart V, McMorris TC, Chory J (1996) A role for brassinosteroids in light-dependent development of Arabidopsis. Science 272:398–401
Li Z, Pan J, Guan Y, Tao Q, He H, Si L, Cai R (2008) Development and fine mapping of three co-dominant SCAR markers linked to the M/m gene in the cucumber plant (Cucumis sativus L.). Theor Appl Genet 117:1253–1260
Li Y, Yang L, Pathak M, Li D, He X, Weng Y (2011) Fine genetic mapping of cp: a recessive gene for compact (dwarf) plant architecture in cucumber, Cucumis sativus L. Theor Appl Genet 123:973–983
Li Z, Wang S, Tao Q, Pan J, Si L, Gong Z, Cai R (2012) A putative positive feedback regulation mechanism in CsACS2 expression suggests a modified model for sex determination in cucumber (Cucumis sativus L.). J Exp Bot 63:4475–4484
Li P, Chen L, Zhou Y, Xia X, Shi K, Chen Z, Yu J (2013) Brassinosteroids-induced systemic stress tolerance was associated with increased transcripts of several defence-related genes in the phloem in Cucumis sativus. PLoS One 8:e66582
Lin T, Wang S, Zhong Y, Gao D, Cui Q, Chen H, Zhang Z, Shen H et al (2016) A truncated F-box protein confers the dwarfism in cucumber. J Genet Genom 43:223–226
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Niemirowicz-Szczytt K, Rucinska M, Korzeniewsia A (1996) An induced mutation in cucumber (Cucumis sativus L.): super compact. Cucurbit Genet Coop Rep 19:1–3 (article 1)
Noguchi T, Fujioka S, Takatsuto S, Sakurai A, Yoshida S, Li J, Chory J (1999) Arabidopsis det2 is defective in the conversion of (24R)-24-methylcholest-4-en-3-one to (24R)-24-methy-5α-cholestan-3-one in brassinosteroid biosynthesis. Plant Physiol 120:833–839
Nomura T, Jager CE, Kitasaka Y, Takeuchi K, Fukami M, Yoneyama K, Matsushita Y, Nyunoya H et al (2004) Brassinosteroid deficiency due to truncated steroid 5α-reductase causes dwarfism in the lk mutant of pea. Plant Physiol 135:2220–2229
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ et al (1999) ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400:256–261
Russell DE, Wilson JD (1994) Steroid 5 α-reductase: two genes/two enzymes. Annu Rev Biochem 63:25–61
Shimada Y, Goda H, Nakamura A, Takatsuto S, Fujioka S, Yoshida S (2003) Organ-specific expression of brassinosteroid-biosynthetic gene and distribution of endogenous brassinosteroids in Arabidopsis. Plant Physiol 131:287–297
Steber CM, McCourt P (2001) A role for brassinosteroids in germination in Arabidopsis. Plant Physiol 125:763–769
Suzuki Y, Saso K, Fujioka S, Yoshida S, Nitasaka E, Nagata S, Nagasawa H, Takatsuto S et al (2003) A dwarf mutant strain of Pharbitis nil, Uzukobito (kobito), has defective brassinosteroid biosynthesis. Plant J 36:401–410
Symons GM, Reid JB (2004) Brassinosteroids do not undergo long-distance transport in pea. Implications for the regulation of endogenous brassinosteroid levels. Plant Physiol 135:2196–2206
Symons GM, Schultz L, Kerckhoffs LH, Davies NW, Gregory D, Reid JB (2002) Uncoupling brassinosteroid levels and de-etiolation in pea. Physiol Plant 115:311–319
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tan J, Tao Q, Niu H, Zhang Z, Li D, Gong Z, Weng Y, Li Z (2015) A novel allele of monoecious (m) locus is responsible for elongated fruit shape and perfect flowers in cucumber (Cucumis sativus L.). Theor Appl Genet 128:2483–2493
Wang L, Wang Z, Xu Y, Joo SH, Lim SK, Xue Z, Xu Z, Wang Z et al (2009) OsGSR1 is involved in crosstalk between gibberellins and brassinosteroids in rice. Plant J 57:498–510
Wang B, Li Y, Zhang W (2012) Brassinosteroids are involved in response of cucumber (Cucumis sativus) to iron deficiency. Ann Bot 110:681–688
Wang H, Li W, Qin Y, Pan Y, Wang X, Weng Y, Chen P, Li Y (2017) The cytochrome P450 gene CsCYP85A1 is a putative candidate for super compact-1 (scp-1) plant architecture mutation in cucumber (Cucumis sativus L.). Front Plant Sci 8:266
Wei L, Deng X, Zhu T, Zheng T, Li P, Wu J, Zhang D, Lin H (2015) Ethylene involved in brassinosteroids induced alternative respiratory pathway in cucumber (Cucumis sativus L.) seedlings response to abiotic stress. Front Plant Sci 6:982
Wu Q, Wu D, Shen Z, Duan C, Guan Y (2013) Quantification of endogenous brassinosteroids in plant by on-line two-dimensional microscale solid phase extraction-on column derivatization coupled with high performance liquid chromatography-tandem mass spectrometry. J Chromatogr A 1297:56–63
Xia X, Zhou Y, Ding J, Shi K, Asami T, Chen Z, Yu J (2011) Induction of systemic stress tolerance by brassinosteroid in Cucumis sativus. New Phytol 191:706–720
Yang L, Koo D-H, Li Y, Zhang X, Luan F, Havey MJ, Jiang J, Weng Y (2012) Chromosome rearrangements during domestication of cucumber as revealed by high-density genetic mapping and draft genome assembly. Plant J 71:895–906
Ye Q, Zhu W, Li L, Zhang S, Yin Y, Ma H, Wang X (2010) Brassinosteroids control male fertility by regulating the expression of key genes involved in Arabidopsis anther and pollen development. Proc Natl Acad Sci USA 107:6100–6105
Acknowledgements
The authors thank Kristin Haider (USDA-ARS) for technical help and Professor Xiaofeng Wang (Northwest A&F University) for valuable suggestions on BR-related analysis. ZL and SL’s work in the University of Wisconsin at Madison visit was partially supported by the China Scholarship Council. This work was supported by the National Natural Science Foundation of China (Nos. 31471879, 31672150) (to ZL), the Innovation of Agricultural Science and Technology in Shaanxi Province (No. 2015NY081) (to ZL), the Young Talent Cultivation Project (Northwest A&F University) (to ZL) and the Agriculture and Food Research Initiative Competitive Grant 2013-67013-21105 from the U.S. Department of Agriculture National Institute of Food (to YW).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Sanwen Huang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hou, S., Niu, H., Tao, Q. et al. A mutant in the CsDET2 gene leads to a systemic brassinosteriod deficiency and super compact phenotype in cucumber (Cucumis sativus L.). Theor Appl Genet 130, 1693–1703 (2017). https://doi.org/10.1007/s00122-017-2919-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2919-z