Abstract
The seed habit represents a remarkable evolutionary advance in plant sexual reproduction. Since the Paleozoic, seeds carry a seed coat that protects, nourishes and facilitates the dispersal of the fertilization product(s). The seed coat architecture evolved to adapt to different environments and reproductive strategies in part by modifying its thickness. Here, we review the great natural diversity observed in seed coat thickness among angiosperms and its molecular regulation in Arabidopsis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The plant sexual reproduction cycle—meiosis, sex differentiation, and fertilization—times the alternation of haploid and diploid generations [1]. Mitosis intercalates the cycle phases and determines the predominance of one generation versus the other. Whereas most non-vascular plants display a predominant gametophytic phase, vascular plants tend to develop large sporophytes [1]. Non-seed plants develop gametophytes physically separated from the sporophyte through the dispersal of spores. By contrast, the evolution of the seed habit in gymnosperms and angiosperms marked the retention of the female gametophyte on the sporophyte and the dispersal of the zygotic embryo (next sporophytic generation) [63]. Seeds greatly contributed to the successful colonization of land by vascular plants. Compared to spores, seeds carry nutrients, rely less on water for germination, convey a higher degree of protection to physical stress, and can disperse in different ways. Such evolutionary advantages were, to a great degree, achieved through the evolution of the seed coat, cell layers that surround, protect, and facilitate the dispersal of the seed fertilization product(s) [52].
The seed coat is a maternal sporophytic tissue that originates from the ovule integument(s). A typical ovule (the seed precursor) comprises four sporophytic tissues: (1) the funiculus, which transports nutrients from the placental tissue, (2) the chalaza, which forms one or more protective (3) integuments, and (4) the nucellus, whose megaspore mother cell undergoes meiosis to originate the gametophyte (Fig. 1). Ovule integuments grow as primordia from the chalaza and are referred to as dermal, if initiated solely by chalazal dermal cells, or sub-dermal, if originated by chalazal dermal and sub-dermal tissue [18, 43]. The developmental patterning of integuments has been thoroughly analyzed in Arabidopsis thaliana [13, 24, 72]. Arabidopsis plants display an outer (oi) and an inner (ii) integument that develop from the chalaza of a finger-like protruding ovule primordium (Fig. 1). The ii primordium initiates as a ring-like structure from dermal chalazal cells (ii initials) at the boundary with the nucellus. ii initials become visible when they undergo a periclinal or oblique cell division followed by cell elongation. The oi primordium grows adjacent to the ii proximal side and extends to the proximal extremity of the chalaza. Whereas the distal part of the oi primordium is of dermal origin and follows ii ontogeny, the basal part is subtended by sub-dermal chalazal tissue undergoing periclinal cell divisions. ii and oi annular primordia grow by anticlinal cell divisions to surround and shape the nucellus and female gametophyte. The distal extremity of the integuments forms the micropyle, a small opening through which the pollen tube penetrates. After fertilization, integument cell layers follow different pathways of cell differentiation [34] and grow in coordination with the fertilization product(s) [40]. Finally, the seed coat undergoes programmed cell death and establishes a protective barrier in between fertilization product(s) and surrounding environment [41].
Seed coat structure and molecular composition deeply influence seed physiology and evolved to adapt to different environmental conditions [43]. Natural diversity has been observed in the thickness of the seed coat, which affects seed germination, dormancy, and mortality [27, 47, 60]. Overall, seed coat thickness is determined by both integument number and cell thickness. Such a character changes along the seed polarity axes and varies among individuals and species. This review explores the evolutionary patterns of seed coat thickness development in angiosperms.
Natural diversity in seed coat thickness
Integument number
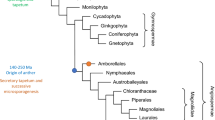
Fossil records of Paleozoic pre-ovules from the earliest seed plants revealed the presence of only one integument (unitegmic, Fig. 2) [28, 80]. The integument structure of such pre-ovules varied among species, but it was typically lobed, vascular bundled, and detached from the nucellus. Leaf-like structures, termed cupules, have been found to partially enclose the ovule and are commonly viewed by paleo-botanists as the precursor of a second integument. Extant gymnosperm ovules are also unitegmic and the integument is probably homologous to that of ancestral seed plants. By contrast, angiosperms evolved two integuments (bitegmic), the so-called oi and ii (Fig. 2). Phylogenetic analyses suggest that bitegmy is the ancestral condition in angiosperms, nevertheless unitegmy arose several times during angiosperm evolution [18].
The transition from bitegmic to unitegmic ovules happened several times during angiosperms evolution along two major ontogenic pathways: (1) developmental suppression or retardation of one integument, (2) congenital integument fusion. Morphological analyses in the Piperaceae family revealed that the single integument of Peperomia shows similarity to the ii of Piper, suggesting that the oi of the former has been lost [88]. Similarly, early arrest of ii has been described in Populus [81]. Congenital fusion of ii and oi probably arose through different mechanisms. Genetic analyses in Arabidopsis (see below) suggest that integumentary fusion can be achieved through lack of boundary formation between oi and ii [14, 44]. This interpretation is confirmed by the analysis of molecular markers of boundary formation in Prunus bitegmic and unitegmic ovules [53] and by morphological studies in Anacardiaceae and Balsaminaceae [43]. By contrast, a mechanism of integumentary shifting has been proposed for the development of the single integument of a number of Ranunculaceae [6]. According to Bouman and Calis, oi sub-dermal tissue subtending ii initials undergoes periclinal cell divisions and, in conjunction with oi, shifts the ii upward. An incision at the tip of the single integument reveals oi and ii remnants. According to the authors, this mechanism would create a bifid integument in between uni- and bitegmic and would only be possible in ovules carrying a sub-dermal oi and dermal ii (Fig. 2). The process of integumentary shifting has been also invoked to explain the development of the unitegmic Gentiana ovule, where no oi and ii remnants are visible [7] (Fig. 2). According to this model the sequence and position of dermal and sub-dermal cell divisions might have led to a unitegmic ovule through integumentary shifting of a sub-dermal oi and dermal ii. Interestingly, species with bitegmic, bifid and unitegmic ovules are grouped in the Impatiens gender. Boesewinkel and Bouman have presented the development and evolution of Impatiens bifid and unitegmic ovules as integumentary shifting [5]. More recently, McAbee et al. have revised this theory and described a change of sub-dermal growth from beneath the oi to the region beneath both oi and ii, without invoking an upward shift of the ii [55]. According to this hypothesis, unitegmy would be the result of ii–oi intercalary growth. All considered, we believe that available data on integument congenital fusion in angiosperms point to two non-exclusive mechanisms: (1) total or partial lack of boundary formation and (2) intercalary growth between ii and oi primordia, regardless of their dermal or sub-dermal origin.
Further ovule reduction has been observed in Santalales [74]. Members of this clade do not develop any ovule integument (ategmy). Expression analyses revealed that integument molecular markers are expressed at the distal end of Santalales ategmic ovules [10]. These data have been interpreted as the fusion of an integument remnant to the nucellus. By contrast, some angiosperms evolved a third cell sheath in addition to oi and ii. Annonaceae ovules develop a so-called third integument in between oi and ii [12], whereas Myristicaceae and other groups initiate a third envelop, referred to as aril (not to be confused with arillodes), later on in development in a region proximal to the oi that has been interpreted as the funiculus [17, 66, 77]. Interestingly, arils can also be found in gymnosperms, such as Taxus baccata [54]. Arils either work as attractive organs that facilitate seed dispersal or are reduced to a tiny protuberance, as in a number of basal angiosperms [39].
Finally, the number of integument cell layers can change after ovule fertilization during early seed development. Rice ovules display two-cell-layered oi and ii. Within 3 days after fertilization, oi and ii of most rice cultivars are absorbed by the pericarp leaving only the cuticle layer [50].
Integument cell thickness
Two-cell-layered dermal integuments originate from the chalazal dermal tissue, whereas three-cell-layered sub-dermal integuments display two dermal and one sub-dermal cell layer. Furthermore, integuments can develop more cell layers through periclinal cell division of sub-dermal or dermal cell layers and are referred to as multiplicative [43]. Integument cell thickness is considered a stable character worth using in macrosystematic analyses [18]. Nevertheless, 3D morphological analyses of Arabidopsis ovules suggest the contrary. Arabidopsis ovules were thought to display a regular two- and three-cell-layered oi and ii, respectively [72]. The ii grows a third cell layer (ii1′) by periclinal cell divisions of the innermost ii cell layer (ii1). Coen et al. have described the formation of a fourth cell layer (ii1″) by periclinal cell division of the ii1′ in 70 and 35% of Wassilewskija and Columbia Arabidopsis ovules, respectively [13]. Furthermore, the oi has been recently shown to initiate sub-dermal cell stripes (oi′) in 10% of the Columbia Arabidopsis ovules tested [24]. These data highlight the importance of 3D imaging for the analysis of integument cell patterning and suggest that cell thickness is a less stable character than previously thought.
Finally, integument thickness is affected by cell size. During ovule development, cell size does not vary considerably among integument cell layers [72]. Nevertheless, integument cell layers follow different fates and expand dishomogeneously after fertilization. In Arabidopsis, ii1′ and ii1″ undergo a more dramatic cell expansion process compared to the other integument cell layers. Coen et al. have revealed that ii1′ and ii1″ growth is not tightly coordinated with the rest of the integument cell layers and have suggested a role in cushioning seed coat development and offsetting perturbations in its developmental program [13].
Integument thickness along the ovule polarity axes
Both integument number and cell thickness can vary considerably along the ovule polarity axes (Fig. 1). Integument growth leads to the formation of a micropyle at the distal side of the ovule. The majority of bitegmic ovules are amphistomic, as both oi and ii contribute to micropyle formation. More rarely, angiosperm ovules can be endostomic or exostomic if only the ii or the oi, respectively, generate the micropyle [18]. Endostomic ovules show distal thickening of the ii at the micropyle where the ii is not covered by the oi, suggesting that oi limits ii growth [38]. Thickening of the distal region of the integument might be instrumental to enclose the nucellus apex and form a micropylar pore. Proximal ii thickening is instead visible in Arabidopsis where ii1′ and ii1″ do not develop all the way to the micropyle. Ectopic ii1′ growth in the micropylar region interferes with embryo development by limiting its growth space and is repressed by a tight regulatory mechanism (see below) [13].
In several basal angiosperms, oi and/or ii radial patterning shows discontinuous growth and leads to lobation. Lobed iis have been considered remnants of ancestral integuments or a sign of atavism as paleozoic ovules displayed a lobbed integument [83]. Nevertheless, it has been argued that lobation might have evolved as a way to restrict the circumference of the integument and allow the formation of a micropyle, as an alternative mechanism to distal integument thickening [38].
Finally, integument number and cell thickness deeply affect ovule curvature along the adaxial–abaxial polarity axis. The micropyle of anatropous (curved) ovules is placed side by side to the chalaza, whereas orthotropous (non-curved) ovules develop micropyle and chalaza along a straight line. Orthotropous ovules are either unitegmic or display a thin or partially developed oi suggesting that the oi plays a major role in ovule curvature [18]. This hypothesis is supported by a number of Arabidopsis mutants that revert from anatropous to orthotropous when the oi is missing or partially developed (see below) [14, 44]. Nevertheless, gymnosperm Podocarpaceae develop unitegmic anatropous ovules. In such ovules, the function of the oi is taken over by the epimatium, an ovuliferous scale that resembles the oi and is involved in ovule curvature [82]. Angiosperm anatropous ovules have either annular or semi-annular oi primordia that grow into cup-shaped or hooded ois [92]. Ovules with a hooded oi display a difference in thickness along the ovule adaxial–abaxial polarity axis: the abaxial side shows a fully developed oi whereas the adaxial side misses partially or entirely the oi. The development of a hooded or cup-shaped oi morphology has been correlated to the speed of developmental curvature: pronounced semi-annular oi primordia tend to curve faster than annular primordia [18].
Molecular regulation of seed coat thickness in arabidopsis
Establishment of the ovule proximal–distal polarity axis
In Arabidopsis, the ovule proximal–distal polarity axis is morphologically evident at stage 2 of ovule development when the distal nucellus develops a megaspore mother cell, the central chalaza initiates integument primordia, and the proximal funiculus differentiates a vascular strand (Fig. 1) [72]. Genetic and molecular evidences suggest that ovule proximal–distal patterning occurs sequentially starting from the distal nucellus toward the funiculus [73]. The establishment of chalazal identity, which has obvious repercussions on integument initiation, is therefore deeply influenced by nucellus development. More than 25 years of genetic studies on Arabidopsis ovule development have revealed a handful of genes involved in ovule proximal–distal patterning.
Mutations in the AINTEGUMENTA (ANT) gene, encoding for an APETALA2 domain transcription factor, lead to bare ovules with no or reduced integument primordia [16, 49]. Starting at stage 2, ANT is expressed in the chalaza and integuments primordia where it is speculated to regulate cell proliferation underlying integument outgrowth [16, 49]. Furthermore, ANT and the mitochondrial ribosomal protein HUELLENLOS (HLL) have redundant functions in establishing the ovule proximal–distal axis [73, 79]. Ipomorph hll alleles display early arrest of integument development and, in combination with ant mutations, lead to reduced chalaza and funiculus regions [73]. Similarly, HLL works synergistically with the SHORT INTEGUMENTS 2 (SIN2) mitochondrial DAR GTPase [9, 36]. sin2 ovules show reduced integument growth, whereas sin2;hll double mutants undergo early arrest of ovule primordia [9]. The nature of HLL and SIN2 protein indicates that metabolic defects can affect developmental patterning through mechanisms yet to be discovered.
Another early marker of the chalazal region, the BEL1 homeodomain transcription factor, has been implicated in the acquisition of chalazal identity [59, 67, 69]. bel1 ovules fail to develop an ii and the oi turns into a swollen collar-like tissue or secondary ectopic carpeloid structure, depending on the bel1 mutation. A similar phenotype has been observed in ovules over-expressing the AGAMOUS (AG) MADS-box gene [67] and in the triple MADS-Box genes mutant seedstick;shatterproof1;shatterproof2 (stk;shp1;shp2) [65]. Interestingly, the carpeloid outgrowths of bel1 ovules are converted into green leaf-like structures in the double apetala2;bel1 and quadruple stk;shp1;shp2;bel1 mutants [8, 59]. Brambilla and coworkers have proposed a model for cell identity acquisition in ovule integuments based on genetic analyses and in vitro MADS-box proteins interactions [8]. According to it, BEL1 interacts in the chalaza with AG and SEPALLATA3 MADS-box proteins to repress carpel identity in the integuments; this protein complex is then stabilized by an ovule identity complex comprising STK, SHP1, and SHP2 proteins [8].
Whereas BEL1 and ANT work in a cell-autonomous fashion, the WUSCHEL (WUS) homeodomain transcription factor promotes integument initiation from the nucellus, non-cell autonomously [33]. wus ovules fail to develop integuments and show defects in megagametogenesis. WUS and ANT do not regulate each other’s expression suggesting that they establish independent pathways from the nucellus and chalaza, respectively. These data indicate that ANT determines responsiveness of chalazal cells to a WUS-mediated signal from the nucellus that ultimately induces integument formation. WUS expression in the nucellus is tightly regulated by a plethora of transcription factors. Defects in integument development of bel1, sporocyteless/nozzle (spl/nzz, see below), stk;shp1;shp2, and phabulosa;phavoluta;corona (phb;phv;cna, see below) mutants have been linked to ectopic WUS mRNA accumulation in the chalaza [8, 75, 93]. Furthermore, exogenous application of cytokinin on ovules leads to integument arrest and concomitant expression of WUS in the integuments [4]. Finally, the expression of WUS in the chalaza under the control of the ANT promoter region expands the chalazal domain and leads to the initiation of an ii and two or more proximal integuments, some of which express oi markers [33, 75]. Only the ii of proANT:WUS ovules grows around the nucellus, whereas more proximal integuments fail to develop further.
The adaptor-like transcriptional repressor SPL/NZZ, another master regulator of the ovule proximal–distal polarity axis, is expressed throughout the ovule at stage 2, but more strongly in the dermal cells of the chalaza and in integument primordia [2, 71, 90, 94]. spl/nzz ovules display reduced nucellus and integuments growth as well as impaired sporogenesis, a phenotype that is partially suppressed by the bel1 mutation [2, 71, 94]. SPL/NZZ negatively regulates the expression of ANT and BEL1 in the nucellus while positively regulating WUS expression [2, 75]. By contrast, SPL/NZZ expression in the chalaza is activated by ANT and BEL1 and it is necessary for the establishment of the adaxial–abaxial polarity axis, thus working as a link between proximal–distal and adaxial–abaxial patterning [2, 3].
Establishment of ovule and integuments adaxial–abaxial polarity axes
In Arabidopsis, ovule primordia acquire adaxial–abaxial polarity and transition from radial to bilateral symmetry at stage 2-III (Fig. 1) [72]. Whereas the ii primordium arises simultaneously on the adaxial and abaxial side of the ovule primordium, the oi primordium initiates on the abaxial face and its predominant abaxial growth is responsible for the Arabidopsis ovule curvature [18]. Adaxial–abaxial patterning characterizes then the laminar growth of each integument (Fig. 1). Here again, genetic studies have helped better understand regulatory pathways underlying such processes.
KANADI (KAN) and YABBY (YAB) transcription factors are known to promote abaxial identity in vegetative organs and ovules [48, 76]. kan1;kan2 mutant ovules display reduced oi growth [21]. Similarly, mutations in the YAB gene INNER NO OUTER (INO) lead to the development of an amorphous protuberance at the place of the oi [85]. INO expression in the proximal–abaxial side of the chalaza (stage 2-I) is tightly regulated in a spatial–temporal fashion. Whereas BEL1 is a pre-requisite for INO expression, ANT and SPL/NZZ act antagonistically to time the onset of INO expression which in turn establishes a positive auto-regulatory loop and feedbacks on ANT expression [3, 58]. The SUPERMAN (SUP) zinc-finger protein interferes with INO positive regulatory loop in the adaxial domain of ovule primordia, promoting oi abaxial development and ovule bilateral symmetry [26, 58, 85]. Similarly, WUS, SPL/NZZ, HLL, and KAN4/ABERANT TESTA SHAPE (ATS) work redundantly in delimitating INO expression to the abaxial side of the oi [3, 75, 85]. Finally, INO has been shown to directly interact with the co-repressors LEUNIG and SEUSS and the co-activator ADA2b/PROPORZ1 (PRZ1) to promote oi growth [78]. Acquisition of ovule adaxial identity is instead promoted by the class III homeodomain-leucine zipper transcription factor PHB. PHB mRNA initially localizes in the adaxial side of ovule primordia (stage 1-I) [75] and phb-1d/PHB gain-of-function mutants, insensitive to miRNA negative regulation, exhibit orthotropous ovules with reduced oi growth [45, 57].
Abaxial–adaxial polarity appears also within integuments, with both oi and ii displaying an adaxial and abaxial side, necessary to establish the oi–ii boundary (Fig. 1). KAN4/ATS is expressed in the abaxial side of ii primordia and loss of KAN4/ATS function leads to the fusion of ii and oi into a single thicker integument [56]. KAN4/ATS is post-translationally repressed by the UNICORN (UCN) active AGC VIII kinase [19, 20]. In ucn mutants as in sk21-D, a kan4/ats dominant allele, integuments form protrusions that originate from periclinal or oblique divisions of dermal cells [19]. Moreover, KAN4/ATS interacts with DELLA proteins, repressors of gibberellin-dependent pathways, to regulate the expression of genes involved in integument growth [32]. In situ hybridization analyses of the class III homeodomain-leucine zipper transcription factors PHB, PHV, and CNA have revealed their ii adaxial pattern of expression [45, 75]. phb;phv;cna triple mutant displays a variety of phenotypes such as reduced ii or oi growth and the formation of amorphous integuments [45]. PHB expression in the ii is regulated by WUS, ANT, and SPLL/NZZ, whereas it is independent of INO and KAN4/ATS [45, 75]. Overall, these genetic analyses suggest that a balanced expression of adaxial and abaxial polarity determinants is necessary for both ii and oi growth, similar to what observed in leaves [45, 56].
Regulation of multiplicative integuments
The growth of Arabidopsis sub-dermal integument cell layers has been found to be tightly regulated. The development of ii sub-dermal cell layers (ii1′ and ii1″) by periclinal cell divisions of the ii1 is promoted by STK and TRANSPARENT TESTA 16 (TT16) MADS-Box transcription factors [13]. tt16;stk ovules lack ii sub-dermal cell layers almost completely. Furthermore, TT16 is a master regulator of ii1′ and ii1″ cell patterning [13, 15]. Wild-type ii1′ arrests before the micropylar region, thus creating a proximal–distal polarity in seed coat thickness. tt16 ovules exhibit a distal shift of ii1′ and ii1″ development, thus creating a thicker micropylar and thinner chalazal region, compared to the wild type. TT16 expression pattern in the ii1 marks in advance the development of the ii1′ and is responsible for the correct positioning and progression of ii1 periclinal cell divisions. In tt16 seeds, embryo development is impaired by the mechanical action of the seed ii1′ invading the micropylar pole, highlighting the importance of tightly regulating sub-dermal integument growth. Finally, TT16 has been involved in the formation of oi sub-dermal cell stripes (oi′), which arise by periclinal cell divisions of sub-dermal chalazal cells [24]. The chalaza of tt16 ovules appears more extended along the proximal distal axis, compared to the wild type, thus favoring the growth of sub-dermal cells in between oi1 and oi2.
After fertilization, ii1′ and ii1″ undergo dramatic cell expansion, compared to all other integument cells, thus affecting considerably seed coat thickness. The differentiation of all integument cell layers was thought to be repressed by FERTILIZATION INDEPENDENT SEED (FIS) Polycomb group (PcG) proteins (see below) and induced by a signaling pathway initiated by the endosperm [22, 70]. Nevertheless, genetic analyses indicate that ii1′ and ii1″ cell expansion is not repressed by FIS PcG proteins and requires a signal from both embryo and endosperm [13, 23]. ii1′ and ii1″ sub-dermal position is not responsible for such a unique regulatory mechanism as oi′ cell expansion responds to the same molecular pathways underlying the development of integument dermal cell layers [24].
Hormonal signaling
As described above, cytokinins and gibberellins regulate ovule proximal–distal [4] and adaxial/abaxial patterning [32], respectively. Brassinosteroids induce ANT and HLL while repressing AP2 expression during early ovule development [37]. Finally, auxin has been shown to play a fundamental role in Arabidopsis integument initiation. Mutations in the auxin efflux facilitator PIN-FORMED 1 (PIN1) as well as treatments with the auxin flux inhibitor NPA lead to finger-like ovule structures [4]. SPL/NZZ and BEL1 regulate PIN1 expression by acting through the cytokinin signaling pathway [4]. Furthermore, SPL/NZZ has a role in auxin homeostasis by repressing the expression of YUCCA genes, which are involved in the biosynthesis of auxin [51]. Likewise, NAC family transcription factors CUP-SHAPED COTYLEDON 1 (CUC1) and 2 promote integument formation by modulating auxin flux through a PIN1-dependent mechanism [42]. Auxin affects not only ovule proximal–distal, but also adaxial–abaxial patterning. KAN4/ATS interacts with ETTIN (ETT), also known as AUXIN RESPONSIVE FACTOR 3 (ARF3), on the abaxial side of the ii [46]. KAN4/ATS-ETT/ARF3 protein complex promotes ii laminar growth and oi–ii separation by regulating PIN1 expression [46]. Finally, microRNA167 has been shown to promote integument growth by preventing ARF6 and ARF8 expression in the ovule [91].
Kinase signaling
In addition to UCN (see above), a number of other Arabidopsis kinase proteins, acting in different signaling pathways, have been involved in ovule integument outgrowth.
The ERECTA (ER) gene family, encoding for LRR-receptor-like kinases (LRR-RLKs), is expressed in the ovule integuments and is important for proper integument growth [64]. When the expression level of ER and ER-like genes was dramatically reduced, the integuments failed to completely surround the nucellus. The ER gene family genetically interacts with the PRETTY FEW SEEDS 2 (PFS2) homeo-box gene, which also promotes integument outgrowth [62, 64]. Similarly, mutations in the MITOGEN-ACTIVATED PROTEIN KINASES 3 (MPK3) and 6 (MPK6) lead to premature arrest of ii and oi growth and have been speculated to work downstream of the ERECTA pathway [86].
The LRR-RLK strubbelig (sub) mutant leads to incomplete and irregular growth of the oi [11]. SUB trafficking is mediated by HAPLESS13 (HAP13), the μ subunit of adaptor protein 1 that regulates protein sorting at the trans-Golgi network/early endosome. hap13 ovules display reduced oi development probably due to SUB mistargeting [87]. A sub-like ovule phenotype was observed in pfs2;ino and pfs2;nzz ovules and was compared to integument lobes of primitive ovules [61]. A forward genetic approach, aimed at finding mutations causing sub-like ovule phenotypes, allowed Fulton and coworkers to identify the quirky (qry), detorqueo and zerzaust mutants [25]. QKY encodes a predicted membrane-anchored C2-domain protein that co-localizes with SUB at the plasmodesmata, thus raising the possibility that both proteins influence symplastic trafficking of molecules [84].
Finally, ARABIDOPSIS CRINKLY 4 encodes another RLK protein localized at the plasma membrane of L1 cell layers [29]. In acr4 mutants, ovule integuments display a wide range of phenotypes, including arrest of integument growth and lack of integument cell layer organization [29, 30, 89].
Epigenetic determinants
dsRNA processing and epigenetic control of transcriptional activation have been shown to regulate ovule integument development in Arabidopsis. Mutations in SHORT INTEGUMENTS 1 (SIN1) (also known as SUSPENSOR1 or CARPEL FACTORY), which encodes a Dicer protein, result in premature integument arrest [31]. Nevertheless, the RNA targets of SIN1 are yet to be identified.
As mentioned above, PcG proteins have also been involved in integument development. Two FIS components of the Polycomb repressive complex 2, MULTICOPY SUPPRESOR OF IRA1 (MSI1) and FERTILIZATION INDEPENDENT ENDOSPERM (FIE), act sporophytically to repress integument cell expansion and differentiation before fertilization [70]. The study of FIE and MSI1 is complicated by the seed lethal phenotype of loss of function mutants. Nevertheless, FIE and MSI1 are haploinsufficient and emasculated fie/+ and msi1/+ flowers produce both wild-type-looking ovules and enlarged autonomous seeds having a partially differentiated seed coat. Furthermore, Hennig and coworkers discovered a positive role for MSI1 in integument outgrowth [35]. MSI1 co-suppression lines exhibited orthotropous ovules with limited oi development, resembling ino mutants.
Conclusive remarks
Genetic analyses of Arabidopsis ovule development revealed a number of master regulators of integument number and cell thickness that might well be responsible for the natural diversity observed in seed coat thickness (Fig. 3). Genes, such as ANT, ATS, BEL1, INO, and ETT, have been used as markers to better understand the evolution of unitegmy and ategmy in angiosperms [10, 53, 55]. Furthermore, the expression pattern of AGL6-like genes suggests that the angiosperm ii is homologous to the gymnosperm single integument [68]. By contrast, the evolution of the oi remains more controversial. It has been hypothesized that cupules might be ancestral ois. In line with this model, the Arabidopsis oi expresses INO, member of a gene family implicated in leaf development [85]. Moreover, Arabidopsis ap2;bel1 and stk;shp1;shp2;bel1 mutant ovules show a conversion of the oi into a leaf-like structure. Alternatively, the oi might have evolved de novo through the WUS or UCN pathway. Less attention has been given to arils while addressing the evolution of the oi. The study of gymnosperms B-sister MADS-box genes, orthologous to Arabidopsis TT16 and GORDITA, revealed their high expression in the integument of Ginkgo biloba ovules and low expression in integument and aril of T. baccata ovules [54]. These data suggest, despite being interpreted differently by the authors of such study, that T. baccata arils express an integument molecular marker and might be seen as integuments. Expression analyses of other integument master regulator genes in angiosperms and gymnosperms arils would help better address nature and evolution of these integument-like structures.
References
Bai SN (2015) The concept of the sexual reproduction cycle and its evolutionary significance. Front Plant Sci 6:11
Balasubramanian S, Schneitz K (2000) NOZZLE regulates proximal–distal pattern formation, cell proliferation and early sporogenesis during ovule development in Arabidopsis thaliana. Dev Suppl 127:4227–4238
Balasubramanian S, Schneitz K (2002) NOZZLE links proximal–distal and adaxial–abaxial pattern formation during ovule development in Arabidopsis thaliana. Dev Suppl 129:4291–4300
Bencivenga S, Simonini S, Benkova E, Colombo L (2012) The transcription factors BEL1 and SPL are required for cytokinin and auxin signaling during ovule development in Arabidopsis. Plant Cell 24:2886–2897
Boesewinkel FD, Bouman F (1991) The development of bi- and unitegmic ovules and seeds in Impatiens (Balsaminaceae). Bot Jahrb Syst Pflanzengesch Pflanzengeogr 113:87–104
Bouman F, Calis JIM (1977) Integumentary shifting—a third way to unitegmy. Ber Deutsch Bot Ges 90:15–28
Bouman F, Schier S (1979) Ovule ontogeny and seed coat development in Gentiana, with a discussion on the evolutionary origin of the single integument. Acta Bot Neerl 28:467–478
Brambilla V, Battaglia R, Colombo M, Masiero S, Bencivenga S, Kater MM, Colombo L (2007) Genetic and molecular interactions between BELL1 and MADS box factors support ovule development in Arabidopsis. Plant Cell 19:2544–2556
Broadhvest J, Baker SC, Gasser CS (2000) SHORT INTEGUMENTS 2 promotes growth during Arabidopsis reproductive development. Genetics 155:899–907
Brown RH, Nickrent DL, Gasser CS (2010) Expression of ovule and integument-associated genes in reduced ovules of Santalales. Evol Dev 12:231–240
Chevalier D, Batoux M, Fulton L, Pfister K, Yadav RK, Schellenberg M, Schneitz K (2005) STRUBBELIG defines a receptor kinase-mediated signaling pathway regulating organ development in Arabidopsis. Proc Natl Acad Sci USA 102:9074–9079
Christmann M (1989) Die tritegmischen Annonaceen-Samen. Botanische Jahrbücher fur Systematik 110:433–439
Coen O, Fiume E, Xu W, De Vos D, Lu J, Pechoux C, Lepiniec L, Magnani E (2017) Developmental patterning of the sub-epidermal integument cell layer in Arabidopsis seeds. Dev Suppl 144:1490–1497
Colombo L, Battaglia R, Kater MM (2008) Arabidopsis ovule development and its evolutionary conservation. Trends Plant Sci 13:444–450
Ehlers K, Bhide AS, Tekleyohans DG, Wittkop B, Snowdon RJ, Becker A (2016) The MADS box genes ABS, SHP1, and SHP2 are essential for the coordination of cell divisions in ovule and seed coat development and for endosperm formation in Arabidopsis thaliana. PLoS One 11:e0165075
Elliott RC, Betzner AS, Huttner E, Oakes MP, Tucker WQ, Gerentes D, Perez P, Smyth DR (1996) AINTEGUMENTA, an APETALA2-like gene of Arabidopsis with pleiotropic roles in ovule development and floral organ growth. Plant Cell 8:155–168
Endress PK (1973) Arils and aril-like structures in woody Ranales. New Phytol 72:1159–1171
Endress PK (2011) Angiosperm ovules: diversity, development, evolution. Ann Bot 107:1465–1489
Enugutti B, Schneitz K (2013) Genetic analysis of ectopic growth suppression during planar growth of integuments mediated by the Arabidopsis AGC protein kinase UNICORN. BMC Plant Biol 13:2
Enugutti B, Kirchhelle C, Oelschner M, Torres Ruiz RA, Schliebner I, Leister D, Schneitz K (2012) Regulation of planar growth by the Arabidopsis AGC protein kinase UNICORN. Proc Natl Acad Sci USA 109:15060–15065
Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11:1251–1260
Figueiredo DD, Batista RA, Roszak PJ, Hennig L, Kohler C (2016) Auxin production in the endosperm drives seed coat development in Arabidopsis. Elife 5:e20542. https://doi.org/10.7554/eLife.20542
Fiume E, Coen O, Xu W, Lepiniec L, Magnani E (2017) Growth of the Arabidopsis sub-epidermal integument cell layers might require an endosperm signal. Plant Signal Behav 12:e1339000
Fiume E, Coen O, Xu W, Lepiniec L, Magnani E (2017) Developmental patterning of sub-epidermal cells in the outer integument of Arabidopsis seeds. PLoS One 12:e0188148
Fulton L, Batoux M, Vaddepalli P, Yadav RK, Busch W, Andersen SU, Jeong S, Lohmann JU, Schneitz K (2009) DETORQUEO, QUIRKY, and ZERZAUST represent novel components involved in organ development mediated by the receptor-like kinase STRUBBELIG in Arabidopsis thaliana. PLoS Genet 5:e1000355
Gaiser JC, Robinson-Beers K, Gasser CS (1995) The Arabidopsis SUPERMAN gene mediates asymmetric growth of the outer integument of ovules. Plant Cell 7:333–345
Gardarin A, Durr C, Mannino MR, Busset H, Colbach N (2010) Seed mortality in the soil is related to seed coat thickness. Seed Sci Res 20:243–256
Gerrienne P, Meyer-Berthaud B, Fairon-Demaret M, Streel M, Steemans P (2004) Runcaria, a middle devonian seed plant precursor. Science 306:856–858
Gifford ML, Dean S, Ingram GC (2003) The Arabidopsis ACR4 gene plays a role in cell layer organisation during ovule integument and sepal margin development. Dev Suppl 130:4249–4258
Gifford ML, Robertson FC, Soares DC, Ingram GC (2005) ARABIDOPSIS CRINKLY4 function, internalization, and turnover are dependent on the extracellular crinkly repeat domain. Plant Cell 17:1154–1166
Golden TA, Schauer SE, Lang JD, Pien S, Mushegian AR, Grossniklaus U, Meinke DW, Ray A (2002) SHORT INTEGUMENTS1/SUSPENSOR1/CARPEL FACTORY, a Dicer homolog, is a maternal effect gene required for embryo development in Arabidopsis. Plant Physiol 130:808–822
Gomez MD, Ventimilla D, Sacristan R, Perez-Amador MA (2016) Gibberellins regulate ovule integument development by interfering with the transcription factor ATS. Plant Physiol 172:2403–2415
Gross-Hardt R, Lenhard M, Laux T (2002) WUSCHEL signaling functions in interregional communication during Arabidopsis ovule development. Genes Dev 16:1129–1138
Haughn G, Chaudhury A (2005) Genetic analysis of seed coat development in Arabidopsis. Trends Plant Sci 10:472–477
Hennig L, Taranto P, Walser M, Schonrock N, Gruissem W (2003) Arabidopsis MSI1 is required for epigenetic maintenance of reproductive development. Dev Suppl 130:2555–2565
Hill TA, Broadhvest J, Kuzoff RK, Gasser CS (2006) Arabidopsis SHORT INTEGUMENTS 2 is a mitochondrial DAR GTPase. Genetics 174:707–718
Huang HY, Jiang WB, Hu YW, Wu P, Zhu JY, Liang WQ, Wang ZY, Lin WH (2013) BR signal influences Arabidopsis ovule and seed number through regulating related genes expression by BZR1. Mol Plant 6:456–469
Igersheim A, Endress PK (1997) Gynoecium diversity and systematics of the Magnoliales and winteroids. Bot J Linn Soc 124:213–271
Igersheim A, Endress PK (1998) Gynoecium diversity and systematics of the paleoherbs. Bot J Linn Soc 127:289–370
Ingram GC (2010) Family life at close quarters: communication and constraint in angiosperm seed development. Protoplasma 247:195–214
Ingram GC (2016) Dying to live: cell elimination as a developmental strategy in angiosperm seeds. J Exp Bot 68:785–796
Ishida T, Aida M, Takada S, Tasaka M (2000) Involvement of CUP-SHAPED COTYLEDON genes in gynoecium and ovule development in Arabidopsis thaliana. Plant Cell Physiol 41:60–67
Johri BM, Ambegaokar KB, Srivastava PS (2013) Comparative embryology of angiosperms. Springer, Berlin
Kelley DR, Gasser CS (2009) Ovule development: genetic trends and evolutionary considerations. Sex Plant Reprod 22:229–234
Kelley DR, Skinner DJ, Gasser CS (2009) Roles of polarity determinants in ovule development. Plant J 57:1054–1064
Kelley DR, Arreola A, Gallagher TL, Gasser CS (2012) ETTIN (ARF3) physically interacts with KANADI proteins to form a functional complex essential for integument development and polarity determination in Arabidopsis. Dev Suppl 139:1105–1109
Kelly KM, Van Staden J, Bell WE (1992) Seed coat structure and dormancy. Plant Growth Regul 11:201–209
Kerstetter RA, Bollman K, Taylor RA, Bomblies K, Poethig RS (2001) KANADI regulates organ polarity in Arabidopsis. Nature 411:706–709
Klucher KM, Chow H, Reiser L, Fischer RL (1996) The AINTEGUMENTA gene of Arabidopsis required for ovule and female gametophyte development is related to the floral homeotic gene APETALA2. Plant Cell 8:137–153
Krishnan S, Dayanandan P (2003) Structural and histochemical studies on grain-filling in the caryopsis of rice (Oryza sativa L.). J Biosci 28:455–469
Li LC, Qin GJ, Tsuge T, Hou XH, Ding MY, Aoyama T, Oka A, Chen Z, Gu H, Zhao Y, Qu LJ (2008) SPOROCYTELESS modulates YUCCA expression to regulate the development of lateral organs in Arabidopsis. New Phytol 179:751–764
Linkies A, Graeber K, Knight C, Leubner-Metzger G (2010) The evolution of seeds. New Phytol 186:817–831
Lora J, Hormaza JI, Herrero M (2015) Transition from two to one integument in Prunus species: expression pattern of INNER NO OUTER (INO), ABERRANT TESTA SHAPE (ATS) and ETTIN (ETT). New Phytol 208:584–595
Lovisetto A, Guzzo F, Busatto N, Casadoro G (2013) Gymnosperm B-sister genes may be involved in ovule/seed development and in some species, in the growth of fleshy fruit-like structures. Ann Bot 112:535–544
McAbee JM, Kuzoff RK, Gasser CS (2005) Mechanisms of derived unitegmy among Impatiens species. Plant Cell 17:1674–1684
McAbee JM, Hill TA, Skinner DJ, Izhaki A, Hauser BA, Meister RJ, Venugopala Reddy G, Meyerowitz EM, Bowman JL, Gasser CS (2006) ABERRANT TESTA SHAPE encodes a KANADI family member, linking polarity determination to separation and growth of Arabidopsis ovule integuments. Plant J 46:522–531
McConnell JR, Barton MK (1998) Leaf polarity and meristem formation in Arabidopsis. Dev Suppl 125:2935–2942
Meister RJ, Kotow LM, Gasser CS (2002) SUPERMAN attenuates positive INNER NO OUTER autoregulation to maintain polar development of Arabidopsis ovule outer integuments. Dev Suppl 129:4281–4289
Modrusan Z, Reiser L, Feldmann KA, Fischer RL, Haughn GW (1994) Homeotic transformation of ovules into carpel-like structures in Arabidopsis. Plant Cell 6:333–349
Nooden LD, Blakley KA, Grzybowski JM (1985) Control of seed coat thickness and permeability in soybean: a possible adaptation to stress. Plant Physiol 79:543–545
Park SO, Hwang S, Hauser BA (2004) The phenotype of Arabidopsis ovule mutants mimics the morphology of primitive seed plants. Proc Biol Sci 271:311–316
Park SO, Zheng Z, Oppenheimer DG, Hauser BA (2005) The PRETTY FEW SEEDS2 gene encodes an Arabidopsis homeodomain protein that regulates ovule development. Dev Suppl 132:841–849
Pettitt J (1970) Heterospory and the origin of the seed habit. Biol Rev 45:401–415
Pillitteri LJ, Bemis SM, Shpak ED, Torii KU (2007) Haploinsufficiency after successive loss of signaling reveals a role for ERECTA-family genes in Arabidopsis ovule development. Dev Suppl 134:3099–3109
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, Wisman E, Yanofsky MF (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88
Povilus RA, Losada JM, Friedman WE (2015) Floral biology and ovule and seed ontogeny of Nymphaea thermarum, a water lily at the brink of extinction with potential as a model system for basal angiosperms. Ann Bot 115:211–226
Ray A, Robinson-Beers K, Ray S, Baker SC, Lang JD, Preuss D, Milligan SB, Gasser CS (1994) Arabidopsis floral homeotic gene BELL (BEL1) controls ovule development through negative regulation of AGAMOUS gene (AG). Proc Natl Acad Sci USA 91:5761–5765
Reinheimer R, Kellogg EA (2009) Evolution of AGL6-like MADS box genes in grasses (Poaceae): ovule expression is ancient and palea expression is new. Plant Cell 21:2591–2605
Reiser L, Modrusan Z, Margossian L, Samach A, Ohad N, Haughn GW, Fischer RL (1995) The BELL1 gene encodes a homeodomain protein involved in pattern formation in the Arabidopsis ovule primordium. Cell 83:735–742
Roszak P, Kohler C (2011) Polycomb group proteins are required to couple seed coat initiation to fertilization. Proc Natl Acad Sci USA 108:20826–20831
Schiefthaler U, Balasubramanian S, Sieber P, Chevalier D, Wisman E, Schneitz K (1999) Molecular analysis of NOZZLE, a gene involved in pattern formation and early sporogenesis during sex organ development in Arabidopsis thaliana. Proc Natl Acad Sci USA 96:11664–11669
Schneitz K, Hulskamp M, Pruitt RE (1995) Wild-type ovule development in Arabidopsis thaliana: a light microscope study of cleared whole-mount tissue. Plant J 7:731–749
Schneitz K, Baker SC, Gasser CS, Redweik A (1998) Pattern formation and growth during floral organogenesis: HUELLENLOS and AINTEGUMENTA are required for the formation of the proximal region of the ovule primordium in Arabidopsis thaliana. Dev Suppl 125:2555–2563
Shamrov II, Anisimova GM, Batygina TB, Sita GL (2001) The types and morphological evolution of the ovule in the order Santalales. Botanicheskij Zhurnal (St. Petersburg) 86:1–14
Sieber P, Gheyselinck J, Gross-Hardt R, Laux T, Grossniklaus U, Schneitz K (2004) Pattern formation during early ovule development in Arabidopsis thaliana. Dev Biol 273:321–334
Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL (1999) Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Dev Suppl 126:4117–4128
Silveira SR, Dornelas MC, Martinelli AP (2016) Perspectives for a framework to understand aril initiation and development. Front Plant Sci 7:1919
Simon MK, Skinner DJ, Gallagher TL, Gasser CS (2017) Integument development in Arabidopsis depends on interaction of YABBY protein INNER NO OUTER with co-activators and co-repressors. Genetics 207:1489–1500
Skinner DJ, Baker SC, Meister RJ, Broadhvest J, Schneitz K, Gasser CS (2001) The Arabidopsis HUELLENLOS gene, which is essential for normal ovule development, encodes a mitochondrial ribosomal protein. Plant Cell 13:2719–2730
Smith DL (1964) The evolution of the ovule. Biol Rev 39:137–159
Tieghem PV (1900) Sur la structure de l’ovule et de la graine et sur les affinités des Salicacées. Bull Mus Natl Hist Nat 6:197–201
Tomlinson PB (1992) Aspects of cone morphology and development in Podocarpaceae (Coniferales). Int J Plant Sci 153:572–588
Umeda A, Imaichi R, Kato R (1994) Ovular development and morphology of the outer integument of Magnolia grandiflora (Magnoliaceae). Am J Bot 81:361–367
Vaddepalli P, Herrmann A, Fulton L, Oelschner M, Hillmer S, Stratil TF, Fastner A, Hammes UZ, Ott T, Robinson DG, Schneitz K (2014) The C2-domain protein QUIRKY and the receptor-like kinase STRUBBELIG localize to plasmodesmata and mediate tissue morphogenesis in Arabidopsis thaliana. Dev Suppl 141:4139–4148
Villanueva JM, Broadhvest J, Hauser BA, Meister RJ, Schneitz K, Gasser CS (1999) INNER NO OUTER regulates abaxial–adaxial patterning in Arabidopsis ovules. Genes Dev 13:3160–3169
Wang H, Liu Y, Bruffett K, Lee J, Hause G, Walker JC, Zhang S (2008) Haplo-insufficiency of MPK3 in MPK6 mutant background uncovers a novel function of these two MAPKs in Arabidopsis ovule development. Plant Cell 20:602–613
Wang JG, Feng C, Liu HH, Ge FR, Li S, Li HJ, Zhang Y (2016) HAPLESS13-mediated trafficking of STRUBBELIG is critical for ovule development in Arabidopsis. PLoS Genet 12:e1006269
Warming E (1878) De l’ovule. Ann Sci Nat Bot 6:177–266
Watanabe M, Tanaka H, Watanabe D, Machida C, Machida Y (2004) The ACR4 receptor-like kinase is required for surface formation of epidermis-related tissues in Arabidopsis thaliana. Plant J 39:298–308
Wei B, Zhang J, Pang C, Yu H, Guo D, Jiang H, Ding M, Chen Z, Tao Q, Gu H, Qu LJ, Qin G (2015) The molecular mechanism of sporocyteless/nozzle in controlling Arabidopsis ovule development. Cell Res 25:121–134
Wu MF, Tian Q, Reed JW (2006) Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Dev Suppl 133:4211–4218
Yamada T, Imaichi R, Kato M (2001) Developmental morphology of ovules and seeds of Nymphaeales. Am J Bot 88:963–974
Yamada T, Sasaki Y, Hashimoto K, Nakajima K, Gasser CS (2016) CORONA, PHABULOSA and PHAVOLUTA collaborate with BELL1 to confine WUSCHEL expression to the nucellus in Arabidopsis ovules. Dev Suppl 143:422–426
Yang WC, Ye D, Xu J, Sundaresan V (1999) The SPOROCYTELESS gene of Arabidopsis is required for initiation of sporogenesis and encodes a novel nuclear protein. Genes Dev 13:2108–2117
Acknowledgements
Funding was provided by Labex Saclay Plant Sciences-SPS (Grant no. ANR-10-LABX-0040-SPS) and FP7-PEOPLE-2012-CIG Marie Skłodowska (Grant no. SEEDNET EU-CIG no. 321710).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Coen, O., Magnani, E. Seed coat thickness in the evolution of angiosperms. Cell. Mol. Life Sci. 75, 2509–2518 (2018). https://doi.org/10.1007/s00018-018-2816-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-018-2816-x